Abstract
Infections caused by the obligate intracellular pathogen Chlamydia trachomatis have a marked impact on human health. C. trachomatis serovariants are the leading cause of bacterial sexually transmitted disease and infectious preventable blindness. Despite decades of effort, there is no practical vaccine against C. trachomatis diseases. Here we report that all C. trachomatis reference serotypes responsible for sexually transmitted disease and blinding trachoma synthesize a highly conserved surface-exposed antigen termed polymorphic membrane protein D (PmpD). We show that Ab specific to PmpD are neutralizing in vitro. We also present evidence that Ab against serovariable-neutralizing targets, such as the major outer membrane protein, block PmpD neutralization. This finding suggests that a decoy-like immune evasion strategy may be active in vivo whereby immunodominant type-specific surface antigens block the neutralizing ability of species-common PmpD Ab. Collectively, these results show that PmpD is a previously uncharacterized C. trachomatis species-common pan-neutralizing target. Moreover, a vaccine protocol using recombinant PmpD to elicit neutralizing Ab in the absence of immunodominant type-specific Ab might be highly efficacious and surpass the level of protection achieved through natural immunity.
Keywords: Chlamydia, autotransporter, neutralizing target, surface antigen, vaccine
Chlamydia trachomatis is an obligate intracellular bacterial pathogen that colonizes and infects oculogenital mucosal surfaces. The organism exists as multiple serovariants that infect millions of people worldwide. Ocular infections cause trachoma, a chronic follicular conjunctivitis that results in scarring and blindness. The World Health Organization estimates that 300–500 million people are afflicted by trachoma (1), making it the most prevalent form of infectious preventable blindness (2). Urogenital infections are the leading cause of bacterial sexually transmitted disease (3) in both industrialized and developing nations (4). Moreover, sexually transmitted diseases are risk factors for infertility (5), the transmission of HIV (6), and human papilloma virus-induced cervical neoplasia (7). Control of C. trachomatis infections is an important public health goal. Unexpectedly, however, aggressive infection control measures based on early detection and antibiotic treatment have resulted in an increase in infection rates (8), most likely by interfering with natural immunity, a concept suggested by studies performed in experimental infection models (9). Effective management of chlamydial disease will likely require the development of an efficacious vaccine (8).
Attempts to develop a chlamydial vaccine have met with only limited success (10–15). This lack of progress has been due, in part, to the unavailability of a relevant preclinical model of oculourogenital infection that is amenable to both characterizing functional aspects of host protective immunity and testing vaccine candidates. The development of a murine female genital tract infection and disease model was instrumental in circumventing these limitations (16) and has provided important insights into the mechanism of protective immunity. Protective immunity, as defined by clearance of a primary infection, depends on an IL-12 CD4+ Th1 immune response (17). The obligatory intracellular lifestyle of chlamydiae suggests cellular immunity is more important than Ab in protection from infection. However, there is evidence supporting an important role for Ab in protective immunity. In humans (18) and nonhuman primates (14), resistance to natural or experimental infection correlates with the presence of local chlamydial-specific IgG and IgA Ab. Preclinical models of infection using B cell- (19) or Fc receptor- (20) deficient mice support a role for B cells and/or Ab in protective immunity. In these models, the effect on the protective immune response was most pronounced in rechallenged mice. Additionally, immune serum was recently shown (21) to confer a marked level of resistance against murine chlamydial genital tract reinfection but not against primary infection. Ab-mediated protection depended on a CD4+ T cell immune response that apparently regulate(s) adaptive changes in the local genital tract tissues, which facilitates the protective role of Ab. These studies implicate a previously unrecognized role for B cells in recall immunity, an aspect of adaptive immunity that is relevant for vaccine design.
Surface-exposed antigens (Ag) on the infectious elementary body (EB) are targets of protective Ab. The most extensively studied chlamydial Ag is the immunodominant antigenically variable major outer membrane protein (MOMP) (22, 23), which, to date, is the only known target of neutralizing Ab (24). Ab specific to MOMP neutralize the in vitro infectivity of C. trachomatis by blocking EB attachment (25). Vaccination of rodents with MOMP proteins (26) or passive transfer of MOMP-specific mAb (10) provides only partial immunity. Recently, Pal et al. (27) reported that immunization of mice with native MOMP results in protection against genital tract challenge at levels equal to that generated by infection. However, MOMP might not be the most compelling vaccine candidate for use in humans because protection has been shown to be homotypic and short-lived (28). Therefore, alternative targets of neutralizing Ab may be required to develop an efficacious human vaccine.
Caldwell et al. (29–31) described an immunogenic C. trachomatis species-specific Ag with a mass of ≈155 kDa that is recognized by convalescent sera from individuals with lymphogranuloma venereum (LGV), cervicitis, urethritis, and trachoma. Although purified by immunoaffinity chromatography nearly three decades ago, the molecular nature of the 155-kDa species-common Ag has remained elusive. Here we show that the 155-kDa C. trachomatis species-common Ag is polymorphic membrane protein D (PmpD) and that Ab specific to PmpD are pan-neutralizing. We also present evidence that Ab specific to surface-accessible MOMP and lipopolysaccharide (LPS) block the neutralizing activity of PmpD Ab. These results imply that chlamydiae have evolved a decoy-like immune evasive strategy by which Ab against abundant immunodominant surface Ag block anti-PmpD-mediated neutralization, suggesting an important and common functional role for PmpD in the pathogenesis of infection. Our findings could have important implications for the future design of a univalent vaccine effective against the multiple serovariants that cause C. trachomatis sexually transmitted disease and blinding trachoma.
Results
Rabbit 155-kDa Antiserum Is Specific for PmpD.
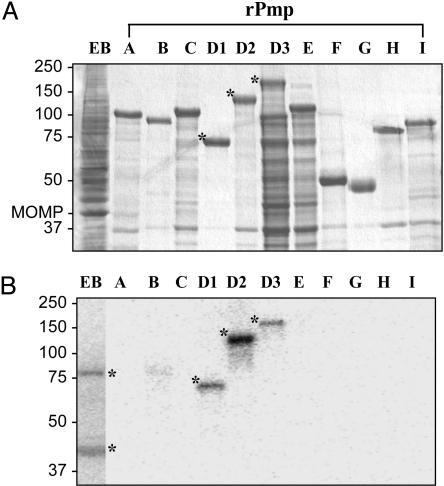
As part of a screen for Ab reactive with C. trachomatis rPmp (r, recombinant), we included rabbit antiserum generated against the 155-kDa Ag. The rPmp and EB were immunoblotted with the 155-kDa antiserum (Fig. 1). The predominant rPmp present in each lysate is shown by gel electrophoresis in Fig. 1A. By immunoblot, 155-kDa antiserum reacted with only EB and rPmpD polypeptides (Fig. 1B). Preimmune rabbit serum was not reactive with EB or rPmpD polypeptides (data not shown). The most intensely reacting rPmpD fragment was D2 (amino acid 45–1079). These findings indicate that antiserum to the 155-kDa Ag is specific for PmpD, a conclusion further supported by the predicted mass of C. trachomatis PmpD, which is ≈161 kDa (32, 33). Interestingly, the antiserum specifically recognized two polypeptides of ≈80 and 42 kDa in EB lysates, suggesting that C. trachomatis PmpD is processed similarly to PmpD of C. pneumoniae (34). With these findings, the 155-kDa antiserum is now referred to as PmpD antiserum.
Fig. 1.
Rabbit 155-kDa antiserum is specific for PmpD. (A) Protein gel of EB and rPmp E. coli lysates used in immunoblotting. (B) Immunoblot with absorbed 155-kDa antiserum. Lanes from left to right are E EB, partially purified rPmp polypeptides A, B, C, D1, D2, D3, E, F, G, H, and I. ∗, the 155-kDa antiserum recognized two polypeptides (≈80 and 42 kDa) in the EB lysate and all three rPmpD polypeptides. The marked polypeptides in A correspond to the polypeptides recognized by the 155-kDa antiserum in B.
PmpD Is a C. trachomatis Species-Common Ag.
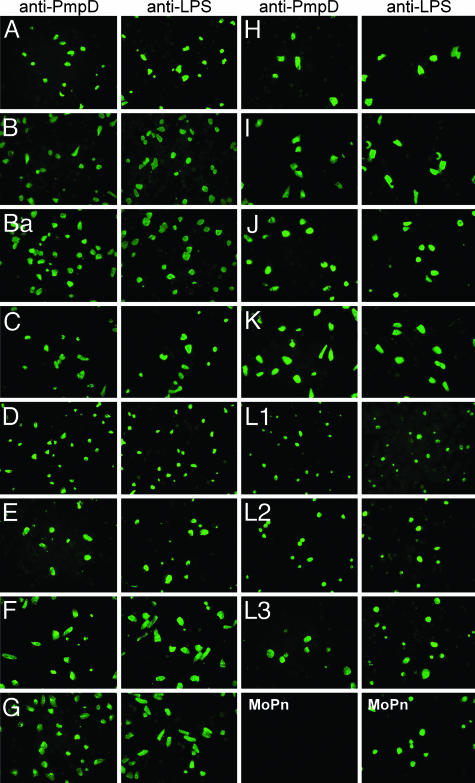
We tested which chlamydial strains reacted with PmpD antiserum by inclusion staining of infected HeLa cells. To confirm that PmpD is a C. trachomatis species-common Ag, we included all 15 C. trachomatis serovars and C. muridarum strain mouse pneumonitis (MoPn). PmpD sequence alignments are 99.15% identical between C. trachomatis serovars (32, 33). In contrast, PmpD of C. trachomatis is 71.46% identical to PmpD of C. muridarum (MoPn), 34.74% identical to PmpD of C. pneumoniae, and 36.50% identical to PmpD of C. caviae (35–37). Inclusions of all 15 C. trachomatis serovars, but not C. muridarum, stained strongly with PmpD antiserum (Fig. 2). In contrast, mAb to genus-specific LPS stained both C. trachomatis and C. muridarum inclusions. To determine whether PmpD antiserum reacted with equal intensity with all C. trachomatis serovars, we conducted endpoint titrations on HeLa cells infected with all 15 C. trachomatis serovars, C. muridarum, C. pneumoniae, and C. caviae. The endpoint titer, by indirect fluorescent Ab, was 1:3,200 for each C. trachomatis serovar, suggesting that the density and exposure of the PmpD protein were similar among C. trachomatis serovars. C. muridarum, C. caviae, and C. pneumoniae did not react with PmpD antiserum at any dilution tested (Table 1).
Fig. 2.
PmpD is a species-common C. trachomatis Ag. HeLa cells infected with the 15 C. trachomatis (A–L3) serovars and C. muridarum (MoPn) were stained by indirect fluorescent Ab with anti-PmpD serum and anti-LPS. Inclusions for each of the C. trachomatis serovars (A–L3) reacted strongly with anti-PmpD. In contrast, inclusions of C. muridarum (MoPn) failed to react with anti-PmpD. Inclusions of all 16 strains reacted with anti-LPS.
Table 1.
Endpoint titration of anti-PmpD to Chlamydial serovars/strains
| C. trachomatis serotype/strain | anti-PmpD (reciprocal dilution) |
||||||
|---|---|---|---|---|---|---|---|
| 100 | 200 | 400 | 800 | 1,600 | 3,200 | 6,400 | |
| B complex serotype | |||||||
| B/TW-5 | +++* | +++ | ++ | ++ | + | + | − |
| Ba/AP-2 | +++ | +++ | ++ | ++ | + | + | − |
| D/UW-3 | +++ | +++ | ++ | ++ | + | + | − |
| E/Bour | +++ | +++ | ++ | ++ | + | + | − |
| L1/440 | +++ | +++ | ++ | ++ | + | + | − |
| L2/434 | +++ | +++ | ++ | ++ | + | + | − |
| C complex serotype | |||||||
| A/HAR-13 | +++ | +++ | ++ | ++ | + | + | − |
| C/TW-3 | +++ | +++ | ++ | ++ | + | + | − |
| H/UW-4 | +++ | +++ | ++ | ++ | + | + | − |
| I/UW-12 | +++ | +++ | ++ | ++ | + | + | − |
| J/UW-36 | +++ | +++ | ++ | ++ | + | + | − |
| Intermediate serotype | |||||||
| F/IC-Cal | +++ | +++ | ++ | ++ | + | + | − |
| G/UW-524 | +++ | +++ | ++ | ++ | + | + | − |
| K/UW-31 | +++ | +++ | ++ | ++ | + | + | − |
| L3/404 | +++ | +++ | ++ | ++ | + | + | − |
| C. muridarium MoPn | − | − | − | − | − | − | − |
| C. pneumoniae | − | − | − | − | − | − | − |
| C. caviae GPIC | − | − | − | − | − | − | − |
*Fluorescence intensity: +++, strong; ++, moderate; +, weak; −, negative.
PmpD Is Surface-Exposed.
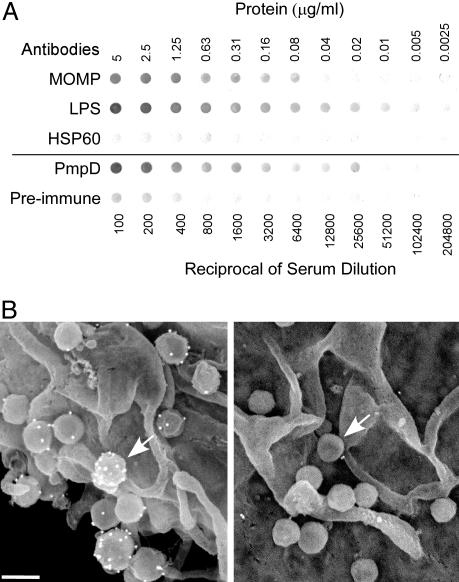
To determine whether PmpD is surface-exposed and a potential target of neutralizing Ab, EB were blotted onto a nitrocellulose membrane and reacted with PmpD antiserum or mAb against MOMP, LPS (which are both surface-exposed), or HSP60 (a cytoplasmic nonsurface-exposed Ag) (Fig. 3A). Consistent with previous reports (23, 38) mAb specific to MOMP and LPS, but not HSP60, were reactive with EB. PmpD antiserum, but not preimmune serum, was also reactive with EB, demonstrating that PmpD is surface-exposed. All mAb reacted with serovar D-infected HeLa cells by indirect fluorescent Ab (data not shown). EM studies were also performed to visually demonstrate the surface exposure of PmpD (Fig. 3B). EB attached to HeLa cells were incubated with either anti-PmpD or preimmune sera, followed by anti-rabbit IgG conjugated to gold particles. Gold labeling of EB bound to cells was seen with anti-PmpD (Fig. 3B Left) but not with preimmune serum (Fig. 3B Right).
Fig. 3.
PmpD is immunoaccessible by immunodot blot and scanning immunoelectron microscopy. (A) D EB were blotted onto nitrocellulose membrane and incubated with different mAb (Upper) and anti-PmpD and preimmune sera (Lower). MAb against MOMP and LPS reacted with EB, whereas mAb specific to HSP60 was nonreactive. Anti-PmpD reacted with EB, but preimmune serum was nonreactive. Protein concentrations are shown for mAb and reciprocal dilutions for anti-PmpD and preimmune sera. (B) L2 EB bound to paraformaldehyde-fixed HeLa cells were reacted with anti-PmpD (Left) or preimmune sera (Right), then probed with gold-conjugated anti-rabbit IgG. Immunogold labeling of PmpD is shown as bright spheres on the EB surface (Left). In contrast there was no association of gold with bound EB incubated with preimmune sera (Right). Left arrow indicates gold-labeled EB; right arrow indicates unlabeled EB. (Scale bar: 0.25 μm.)
PmpD Is a Species-Common Pan-Neutralizing Target.
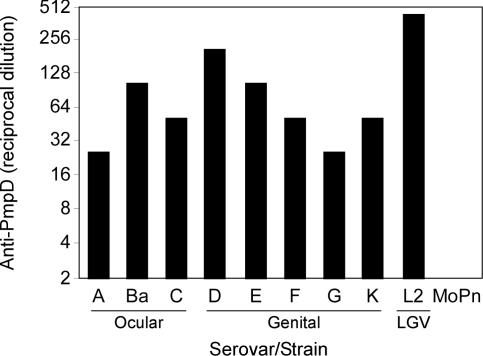
Surface exposure of PmpD suggested it might be a neutralizing target. MOMP is the only known target of neutralizing Ab (24). Chlamydial LPS, although surface-accessible, is not a neutralizing target (39); therefore, it was critical to determine whether PmpD antiserum was neutralizing or nonneutralizing. Neutralizing 50% endpoint determinations were performed in hamster kidney (HaK) cells on representative serovars from the three major C. trachomatis serogroups (B, C, and intermediate complex) representing ocular (A, Ba, and C), genital noninvasive (D, E, F, G, and K), and genital invasive (L2) serovars. The PmpD antiserum titer that resulted in a 50% reduction in inclusion forming units (IFU) for each serovar and for the negative control strain C. muridarum (MoPn) is shown in Fig. 4. The titers resulting in a 50% reduction in IFU varied for each serovar. B complex serovars (Ba, D, E, and L2) were more efficiently neutralized than C complex (A and C) and intermediate complex (F, G, and K) serovars. PmpD antiserum failed to neutralize C. muridarium. These findings show that C. trachomatis PmpD is a species-common pan-neutralizing target.
Fig. 4.
PmpD is a target of neutralizing Ab. The neutralizing activity of anti-PmpD was assayed against C. trachomatis serovars A, Ba, C, D, E, F, G, K, L2, and C. muridarum (MoPn). Results are expressed as the reciprocal of the serum dilution resulting in 50% reduction in IFU (50% endpoints) in HaK cells. The experiment was repeated twice in triplicate; representative data from a single experiment are depicted. Anti-PmpD neutralized C. trachomatis serovars but failed to neutralize the infectivity of C. muridarum (MoPn).
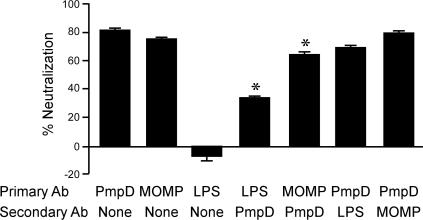
Ab to MOMP and LPS Block the Neutralizing Ability of PmpD Antiserum.
We hypothesized that MOMP immunodominant serovar-specific Ab might mask PmpD species-common neutralizing Ag and, hence, block the neutralizing activity of PmpD antiserum. To test this hypothesis, EB were preincubated with serovar-specific mAb to MOMP or genus-specific mAb to LPS, followed by incubation with PmpD antiserum and assayed for infectivity for HaK cells (Fig. 5). Both PmpD antiserum and mAb to MOMP were neutralizing, whereas mAb to LPS was nonneutralizing. Anti-LPS mAb was very effective in blocking the neutralizing activity of PmpD antiserum compared to PmpD antiserum alone (P < 0.05). Anti-MOMP mAb also significantly blocked the neutralizing activity of PmpD antiserum compared to PmpD antiserum alone (P < 0.05); however, this effect was less marked than that observed for anti-LPS mAb because both anti-MOMP mAb and PmpD antiserum are neutralizing. When PmpD antiserum was incubated with EB before anti-LPS or anti-MOMP mAb, the blocking effect of both mAb was negated.
Fig. 5.
Ab to MOMP and LPS inhibit anti-PmpD-mediated neutralization of chlamydial infectivity. D EB were incubated with different primary and secondary Ab, then plated onto HaK monolayers and assayed for IFU to determine percent neutralization. Both anti-PmpD and mAb to MOMP were neutralizing, whereas mAb to LPS was not neutralizing. Preincubation with mAb to LPS or MOMP significantly blocked the ability of anti-PmpD to neutralize infectivity when compared to anti-PmpD alone (∗, P < 0.05 by using two-tailed unpaired t test). Preincubation with anti-PmpD negated the blocking effect of mAb to LPS or MOMP.
Discussion
We previously described a highly immunogenic C. trachomatis species-specific Ag (155 kDa) that was recognized by sera from patients diagnosed with LGV, cervicitis, urethritis, and trachoma (29–31). In this study, we demonstrate that antiserum to the 155-kDa Ag reacts specifically with PmpD of C. trachomatis. We also show that this PmpD antiserum recognizes all 15 C. trachomatis serovars but not C. muridarum, C. pneumoniae, or C. caviae. Additionally, we show that PmpD is surface-exposed and a C. trachomatis pan-neutralizing target. Finally, we show that Ab to immunodominant surface Ag such as MOMP and LPS can block the ability of PmpD antiserum to neutralize the infectivity of EB.
Chlamydial Pmp resemble autotransporters of Gram-negative bacteria (40, 41). Despite functional diversity, autotransporters possess common characteristics that include translocation across the inner membrane in a Sec-dependent manner, secretion of the amino-terminal portion (the passenger domain) of the protein to the bacterial cell surface, and a carboxyl-terminal translocator domain (42). The translocator domain forms a β-barrel structure through which the passenger domain transverses, resulting in a secreted product capable of interacting with the extracellular environment. The passenger domain may or may not be proteolytically cleaved upon translocation across the outer membrane (42). C. trachomatis possesses nine different pmp genes, all of which are expressed (43). In C. pneumoniae the pmp orthologs include 21 genes that are divided into six gene families: A, B/C, D, E/F, G, and H (40). Among the pmp pmpD is the least variable between species at both the nucleotide and amino acid level. Genes exhibiting a low degree of sequence variation between strains, and between species, often carry out essential functions.
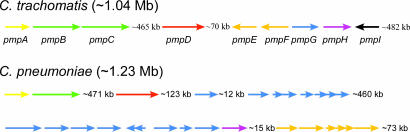
Recently, Wehrl et al. (34) reported that C. pneumoniae PmpD is surface-exposed and a target of neutralizing Ab. These authors also showed that PmpD is proteolytically cleaved and the amino-terminal portion translocates to the cell surface. The C. trachomatis pmpD gene, encoding a predicted polypeptide of 150 kDa, is virtually identical (99.15%) among C. trachomatis serovars (32, 33) but not among other chlamydial species (35–37). Fig. 6 shows the general chromosomal arrangement of the pmp in both C. trachomatis and C. pneumoniae. Although the pmp gene family is highly polymorphic, pmpD is unique. Organizationally, the pmpD genes are physically isolated and unaltered in relative gene organization between the two species. Conservation of pmpD in C. trachomatis serovars suggests that if PmpD is surface-accessible, similar to C. pneumoniae PmpD, it could function as a species-common pan-neutralizing target for C. trachomatis isolates. Our results, demonstrating that C. trachomatis PmpD is surface-exposed and a target of neutralizing Ab, are consistent with those of Wehrl et al. (34) for C. pneumoniae PmpD. Moreover, the C. trachomatis PmpD also appears to be proteolytically processed in a fashion similar to that of C. pneumoniae PmpD. Based on the similarity of PmpD to autotransporters, the majority of the PmpD protein in the mature EB would be predicted to be in the processed rather than the full-length form. Because PmpD is expressed by all C. trachomatis serovars, it likely serves a common and, perhaps, critical function in pathogenesis.
Fig. 6.
Gene arrangement of the pmp gene family in the C. trachomatis and C. pneumoniae genomes. The sequenced C. trachomatis genomes are ≈1.04 Mb, whereas the C. pneumoniae genomes are ≈1.23 Mb in size. The gene families are identified via differing colors, as originally assigned by Grimwood and Stephens (40). pmpA, yellow; pmpB/C, green; pmpD, red; pmpE/F, orange; pmpG, blue; pmpH, purple; pmpI, black. The chromosomal distance between each pmp gene/gene cluster is indicated in kilobases (kb).
Preexisting Ab to MOMP or LPS effectively block anti-PmpD neutralization in vitro. Conversely, if PmpD Ab are preexisting, Ab to MOMP or LPS do not block anti-PmpD neutralization. These in vitro findings may have important implications to in vivo immunity. MOMP and LPS are immunodominant Ag during natural infection of humans (44) and nonhuman primates (14, 28). Primary anti-MOMP responses are serovar-specific (23), and protection is homotypic (28). MOMP and LPS are abundant on the EB surface (38), whereas PmpD appears to be a less abundant protein. In vivo abundant immunodominant surface Ag may function as decoys for the immune system by blocking the binding of more broadly protective species-common pan-neutralizing Ab. This decoy function could explain serovar-specific immunity to reinfection because Ab to PmpD would be prevented from binding its cognate neutralizing target(s). We propose that preexisting Ab to MOMP and LPS might prevent Ab specific to PmpD from functioning in protective immunity; suggesting a vaccine that generates heterotypic PmpD neutralizing Ab in the absence of preexisting MOMP or LPS Ab may be effective.
Previous vaccine studies have used the C. trachomatis MOMP because it is surface-exposed, neutralizing, immunodominant, and possesses antigenic properties that elicit serotype-specific immune responses (14, 28). MOMP subunit vaccines have mostly either failed to evoke protective immunity (11, 13) or provided only partial immunity (15). Pal et al. (27) recently demonstrated that vaccination with native C. muridarum MOMP in the mouse model confers a level of protective immunity that is equivalent to infection itself. These findings are important in not only showing that MOMP is a viable vaccine candidate, but also that an acellular univalent vaccine can protect against chlamydial urogenital infection. However, this approach has inherent shortcomings. First, purification of large amounts of native MOMP is impractical because of the difficulty and expense of in vitro propagation of large quantities of chlamydiae. Second, MOMP is extensively disulfide cross-linked on the EB surface, making it difficult to mimic its native immunogenic conformation by recombinant DNA technology. Third, a MOMP-based vaccine capable of evoking broad coverage against multiple serovars would need to be multivalent, because MOMP is antigenically variable (22) and the serovar-specific domains of the protein are immunodominant (22, 23). Our results indicate that PmpD may be a superior Ag to MOMP as a C. trachomatis vaccine candidate. PmpD is an antigenically stable pan-neutralizing target that, in theory, would provide protection against all human strains, thus allowing the development of a univalent vaccine that is efficacious against both blinding trachoma and sexually transmitted disease.
Materials and Methods
Chlamydial and Cell Cultures.
C. trachomatis serovars A/HAR-13, B/TW-5/OT, Ba/Ap-2, C/TW-3/OT, D/UW-3/Cx, E/Bour, F/IC-Cal-3, G/UW-524/Cx, H/UW-4/Cx, I/UW-12/Ur, J/UW-36/Cx, K/UW-31/Cx, L1/LGV-440, L2/LGV-434, L3/LGV-404, C. muridarum (MoPn), C. pneumoniae (AR-39), and C. caviae strain guinea pig inclusion conjunctivitis were grown in HeLa 229 (ATCC CCL-2.1) and HaK (ATCC CCL-15) cells as previously described (45).
Antibodies.
Rabbit polyclonal antiserum (Pmp-D antiserum) raised against L2 Ag-0.65 (155-kDa Ag) and the mouse mAb EVI-HI (genus-specific anti-LPS), Bb5 (anti-D MOMP), L21–45 (anti-L2 MOMP), and A57-B9 (anti-heat shock protein [HSP60]) were used in these studies.
Indirect Immunofluorescence.
Chlamydial strains were grown on HeLa monolayers until mature inclusion formation, then methanol fixed and stained with either Pmp-D antiserum or mAb to LPS, followed by Alexa Fluor 488-labeled secondary Ab (Invitrogen).
Immunoblot Analysis.
The pmp genes encoding Pmp A-I, including three different clones expressing overlapping polypeptides of the pmpD gene, were expressed in Escherichia coli as His-Tag fusion polypeptides (data not shown). Insoluble inclusion bodies were differentially extracted by using Triton X-100 and sonication. The partially purified rPmp polypeptides corresponding to amino acid residues 47–979 (rPmpA), 949–1747 (rPmpB), 994–1766 (rPmpC), 921–1528 (rPmpD1), 45–1079 (rPmpD2), 45–1528 (rPmpD3), 27–962 (rPmpE), 28–449 (rPmpF), 139–502 (rPmpG), 337–1009 (rPmpH), 30–879 (rPmpI), and E EB were loaded on 10% SDS/PAGE gels and transferred to poly(vinylidene difluoride) membranes (Amersham Pharmacia Biosciences). Each lane was visually standardized by Coomassie brilliant blue staining to contain ≈1–2 μg of protein of full-length rPmp or 10 μg of EB. Preimmune and anti-PmpD sera were preabsorbed with inclusion bodies of the His-tagged recombinant capsid protein from chlamydiaphage phiCPG1, an irrelevant antigen, to absorb nonspecific antibody (data not shown). The PmpD antiserum was diluted 1:1,000 against rPmp and 1:500 against E EB. Membranes were blotted with the preabsorbed serum, followed by a horseradish peroxidase-conjugated secondary Ab (Kirkegaard and Perry Laboratories). The blots were visualized with ECL (SuperSignal West Dura Extended Duration Substrate; Pierce) and read by using imagequant 5.2 software.
Immunodot Blot.
D EB were blotted onto a nitrocellulose membrane in a BIO-DOT microfiltration apparatus (Bio-Rad) and reacted with mAb specific to D MOMP, LPS, and HSP60, anti-PmpD, and preimmune sera as previously described (23). Detection of primary Ab reactivity with EB surface Ag was modified from previous work by incubating nitrocellulose membrane with alkaline phosphatase-conjugated secondary Ab (Zymed, Invitrogen). Blots were developed with solutions of 5-bromo-4-chloro-3-indolyl phosphate plus nitroblue tetrazolium salt (Zymed) as described by the manufacturer.
Immunolocalization by Electron Microscopy.
HeLa cells grown on silicon chips were 2% paraformaldehyde fixed, washed with PBS, blocked with 2% FBS, and infected with L2. Cells were washed, incubated with anti-PmpD or preimmune sera, washed, incubated with goat anti-rabbit IgG 20 nm colloidal gold (BB International, Cardiff, U.K.), washed, and fixed with 2.5% glutaraldehyde/4% paraformaldehyde in 0.1 M sodium cacodylate buffer. Samples were postfixed, alternating two rounds of 15 min in OsO4 and 30 min in 1% tannic acid, then ethanol dehydrated and dried in a Bal-Tec cpd 030 drier (Balzers). Chips were mounted on aluminum studs (Ted Pella, Inc., Redding, CA), coated with 25 Å of chromium in an IBS/TM200S ion beam sputterer (South Bay Technology, Inc., San Clemente, CA) and viewed at 5 kV with an S-5200 in-lens electron microscope in mixed backscatter mode (Hitachi, Tokyo). Digital images were processed with photoshop 7.0 (Adobe Systems, Mountain View, CA).
Neutralization Assays.
In vitro neutralization of chlamydial infectivity for HaK cells was performed by adding 1 × 106 EB/ml to twofold dilutions of anti-PmpD and preimmune sera and assaying for IFU as previously described (38). MAb to MOMP, LPS, or isotype matched irrelevant mAb were used as positive and negative controls, respectively. Percent specific neutralization was calculated as [(preimmune IFU – immune IFU)/preimmune IFU] × 100 for each dilution. Primary (blocking) and secondary Ab concentrations that individually resulted in ≈75% neutralization of D EB were use for blocking assays. Serum was diluted 1:50 and 10 μg/ml of mAb were used. EB were incubated with primary (blocking) Ab for 30 min, secondary Ab was added, and incubation continued for 30 min. EB-Ab mixtures were plated onto monolayers and assayed for IFU as described above.
Acknowledgments
We thank Robert Heinzen, David Nelson, Nathalie Reveneau, and Kena Swanson for their critical reading of the manuscript and helpful suggestions; Anita Mora for assistance in graphic art; and Kelly Matteson for manuscript formatting. This research was supported by the Intramural Research Program of the National Institutes of Health, National Institute of Allergy and Infectious Diseases (NIAID). R.-c.H. and P.B. were supported by NIAID Grant AI051417.
Abbreviations
- Ag
antigen
- EB
elementary body
- HaK
hamster kidney
- IFU
inclusion forming unit
- LGV
lymphogranuloma venereum
- LPS
lipopolysaccharide
- MOMP
major outer membrane protein
- MoPn
mouse pneumonitis
- PmpD
polymorphic membrane protein D
- r
recombinant.
Footnotes
Conflict of interest statement: No conflicts declared.
This paper was submitted directly (Track II) to the PNAS office.
Freely available online through the PNAS open access option.
References
- 1.Resnikoff S., Pascolini D., Etya’ale D., Kocur I., Pararajasegaram R., Pokharel G. P., Mariotti S. P. Bull. W.H.O. 2004;82:844–851. [PMC free article] [PubMed] [Google Scholar]
- 2.Whitcher J. P., Srinivasan M., Upadhyay M. P. Bull. W.H.O. 2001;79:214–221. [PMC free article] [PubMed] [Google Scholar]
- 3.Division of STD Prevention. Sexually Transmitted Disease Surveillance 1997 (Centers Dis. Control Prev., Atlanta) 1998.
- 4.World Health Organization. Global Prevalence and Incidence of Selected Curable Sexually Transmitted Infections: Overview and Estimates. Geneva: World Health Org.; 2001. pp. 1–43. [Google Scholar]
- 5.Westrom L., Joesoef R., Reynolds G., Hagdu A., Thompson S. E. Sex. Transm. Dis. 1992;19:185–192. [PubMed] [Google Scholar]
- 6.Plummer F., Simonsen J. N., Cameron D. W., Ndinya-Achola J., Kreiss J. K., Gakinya M. N., Waiyaki P., Cheang M., Piot P., Ronald A. R. J. Infect. Dis. 1991;163:233–239. doi: 10.1093/infdis/163.2.233. [DOI] [PubMed] [Google Scholar]
- 7.Anttila T., Saikku P., Koskela P., Bloigu A., Dillner J., Ikaheimo I., Jellum E., Lehtinen M., Lenner P., Hakulinen T., et al. J. Am. Med. Assoc. 2001;285:47–51. doi: 10.1001/jama.285.1.47. [DOI] [PubMed] [Google Scholar]
- 8.Brunham R. C., Pourbohloul B., Mak S., White R., Rekart M. L. J. Infect. Dis. 2005;192:1836–1844. doi: 10.1086/497341. [DOI] [PubMed] [Google Scholar]
- 9.Su H., Morrison R. P., Messer R., Whitmire W. M., Hughes S., Caldwell H. D. J. Infect. Dis. 1999;180:1252–1258. doi: 10.1086/315046. [DOI] [PubMed] [Google Scholar]
- 10.Cotter T. W., Meng Q., Shen Z. L., Zhang Y. X., Su H., Caldwell H. D. Infect. Immun. 1995;63:4704–4714. doi: 10.1128/iai.63.12.4704-4714.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Pal S., Barnhart K. M., Wei Q., Abai A. M., Peterson E. M., de la Maza L. M. Vaccine. 1999;17:459–465. doi: 10.1016/s0264-410x(98)00219-9. [DOI] [PubMed] [Google Scholar]
- 12.Pal S., Theodor I., Peterson E. M., de la Maza L. M. Infect. Immun. 1997;65:3361–3369. doi: 10.1128/iai.65.8.3361-3369.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Su H., Parnell M., Caldwell H. D. Vaccine. 1995;13:1023–1032. doi: 10.1016/0264-410x(95)00017-u. [DOI] [PubMed] [Google Scholar]
- 14.Taylor H. R., Whittum-Hudson J., Schachter J., Caldwell H. D., Prendergast R. A. Invest. Ophthalmol. Vis. Sci. 1988;29:1847–1853. [PubMed] [Google Scholar]
- 15.Zhang D., Yang X., Berry J., Shen C., McClarty G., Brunham R. C. J. Infect. Dis. 1997;176:1035–1040. doi: 10.1086/516545. [DOI] [PubMed] [Google Scholar]
- 16.Barron A. L., White H. J., Rank R. G., Soloff B. L., Moses E. B. J. Infect. Dis. 1981;143:63–66. doi: 10.1093/infdis/143.1.63. [DOI] [PubMed] [Google Scholar]
- 17.Morrison R. P., Caldwell H. D. Infect. Immun. 2002;70:2741–2751. doi: 10.1128/IAI.70.6.2741-2751.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Brunham R. C., Kuo C. C., Cles L., Holmes K. K. Infect. Immun. 1983;39:1491–1494. doi: 10.1128/iai.39.3.1491-1494.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Su H., Feilzer K., Caldwell H. D., Morrison R. P. Infect. Immun. 1997;65:1993–1999. doi: 10.1128/iai.65.6.1993-1999.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Moore T., Ananaba G. A., Bolier J., Bowers S., Belay T., Eko F. O., Igietseme J. U. Immunology. 2002;105:213–221. doi: 10.1046/j.0019-2805.2001.01354.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Morrison R. P., Morrison S. G. J. Immunol. 2005;175:7536–7542. doi: 10.4049/jimmunol.175.11.7536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Caldwell H. D., Schachter J. Infect. Immun. 1982;35:1024–1031. doi: 10.1128/iai.35.3.1024-1031.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Zhang Y. X., Stewart S., Joseph T., Taylor H. R., Caldwell H. D. J. Immunol. 1987;138:575–581. [PubMed] [Google Scholar]
- 24.Caldwell H. D., Perry L. J. Infect. Immun. 1982;38:745–754. doi: 10.1128/iai.38.2.745-754.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Su H., Caldwell H. D. Infect. Immun. 1991;59:2843–2845. doi: 10.1128/iai.59.8.2843-2845.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Batteiger B. E., Rank R. G., Bavoil P. M., Soderberg L. S. J. Gen. Microbiol. 1993;139:2965–2972. doi: 10.1099/00221287-139-12-2965. [DOI] [PubMed] [Google Scholar]
- 27.Pal S., Peterson E. M., de la Maza L. M. Infect. Immun. 2005;73:8153–8160. doi: 10.1128/IAI.73.12.8153-8160.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Grayston J. T., Kim K. S. W., Alexander E. R., Wang S.-P. In: Trachoma and Related Disorders Caused by Chlamydial Agents. Nichols R. L., editor. Boston: Excerpta Medica; 1971. pp. 377–385. [Google Scholar]
- 29.Caldwell H. D., Kuo C. C., Kenny G. E. J. Immunol. 1975;115:963–968. [PubMed] [Google Scholar]
- 30.Caldwell H. D., Kuo C. C., Kenny G. E. J. Immunol. 1975;115:969–975. [PubMed] [Google Scholar]
- 31.Caldwell H. D., Kuo C. C. J. Immunol. 1977;118:437–441. [PubMed] [Google Scholar]
- 32.Carlson J. H., Porcella S. F., McClarty G., Caldwell H. D. Infect. Immun. 2005;73:6407–6418. doi: 10.1128/IAI.73.10.6407-6418.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Stephens R. S., Kalman S., Lammel C., Fan J., Marathe R., Aravind L., Mitchell W., Olinger L., Tatusov R. L., Zhao Q., et al. Science. 1998;282:754–759. doi: 10.1126/science.282.5389.754. [DOI] [PubMed] [Google Scholar]
- 34.Wehrl W., Brinkmann V., Jungblut P. R., Meyer T. F., Szczepek A. J. Mol. Microbiol. 2004;51:319–334. doi: 10.1046/j.1365-2958.2003.03838.x. [DOI] [PubMed] [Google Scholar]
- 35.Kalman S., Mitchell W., Marathe R., Lammel C., Fan J., Hyman R. W., Olinger L., Grimwood J., Davis R. W., Stephens R. S. Nat. Genet. 1999;21:385–389. doi: 10.1038/7716. [DOI] [PubMed] [Google Scholar]
- 36.Read T. D., Brunham R. C., Shen C., Gill S. R., Heidelberg J. F., White O., Hickey E. K., Peterson J., Utterback T., Berry K., et al. Nucleic Acids Res. 2000;28:1397–1406. doi: 10.1093/nar/28.6.1397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Read T. D., Myers G. S., Brunham R. C., Nelson W. C., Paulsen I. T., Heidelberg J. F., Holtzapple E., Khouri H., Federova N. B., Carty H. A., et al. Nucleic Acids Res. 2003;31:2134–2147. doi: 10.1093/nar/gkg321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Su H., Watkins N. G., Zhang Y. X., Caldwell H. D. Infect. Immun. 1990;58:1017–1025. doi: 10.1128/iai.58.4.1017-1025.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Byrne G. I., Stephens R. S., Ada G., Caldwell H. D., Su H., Morrison R. P., Van der Pol B., Bavoil B., Bobo L., Everson S. J. Infect. Dis. 1993;168:415–420. doi: 10.1093/infdis/168.2.415. [DOI] [PubMed] [Google Scholar]
- 40.Grimwood J., Stephens R. S. Microb. Comp. Genomics. 1999;4:187–201. doi: 10.1089/omi.1.1999.4.187. [DOI] [PubMed] [Google Scholar]
- 41.Henderson I. R., Lam A. C. Trends Microbiol. 2001;9:573–578. doi: 10.1016/s0966-842x(01)02234-x. [DOI] [PubMed] [Google Scholar]
- 42.Henderson I. R., Navarro-Garcia F., Nataro J. P. Trends Microbiol. 1998;6:370–378. doi: 10.1016/s0966-842x(98)01318-3. [DOI] [PubMed] [Google Scholar]
- 43.Lindquist E., Stephens R. In: Stephens R., Byrne G., Christiansen G., Clarke I., Grayston J., Rank R., Ridgway G., Saikku P., Schachter J., Stamm W., editors. Proceedings of the Ninth International Symposium on Human Chlamydial Infection; San Francisco: Int. Chlamydia Symp.; 1998. pp. 259–262. [Google Scholar]
- 44.Ward M. E., Treharne J. D., Murray A. J. Gen. Microbiol. 1986;132:1599–1610. doi: 10.1099/00221287-132-6-1599. [DOI] [PubMed] [Google Scholar]
- 45.Caldwell H. D., Kromhout J., Schachter J. Infect. Immun. 1981;31:1161–1176. doi: 10.1128/iai.31.3.1161-1176.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]