Abstract
Previously, we found that the quorum sensing transcription factor SdiA up-regulates AcrAB. Others found that a 4-quinolone was a quorum-sensing signal in Pseudomonas aeruginosa. In Escherichia coli, there are at least three multidrug transporters (AcrAB∕TolC, MdfA, and NorE) that exude fluoroquinolones. Here, we show that ΔacrAB, tolC210, or ΔnorE mutants have the same growth rate as WT cells in exponential phase but grow to higher cell density in stationary phase. Overproduction of either pump caused cells to reach lower density. mdfA had no effect. Conditioned medium (CM) from cells overexpressing acrAB represses cell growth more than CM from WT cells. CM from pump mutant cells represses cell growth less than CM from WT cells. These results were not affected by the deletion of luxS, which synthesizes the quorum-sensing signal autoinducer 2 (AI-2). Expression of the rpoS gene encoding the stationary phase σ factor is induced earlier in cells overexpressing acrAB and later in acrAB mutant cells. These results support a model in which a natural function of AcrAB∕TolC and NorE is to export signals for cell–cell communication. Drugs exported by pumps may resemble communication molecules normally exuded.
Bacteria communicate using small chemical molecules, and this process is termed quorum sensing. Accumulation of quorum-sensing signals (QSS) in the growth medium mirrors cell density. Once a threshold concentration is reached, the QSS activate transcription factors that, in turn, up-regulate the signal synthase and multiple other genes. QSS have been shown to control bioluminescence, virulence factor expression, biofilm formation, entry into stationary phase, conjugal transfer of plasmid DNA, spore formation, and transformation competence (1). Distinct classes of QSS have been discovered in Gram-negative bacteria: the N-acyl homoserine lactones (AHL), the LuxS-derived family of furanone signals (2), the diketopiperazines (3), and 2-heptyl-3-hydroxy-4-quinolone (PQS) (4) and additional signals are likely to exist. PQS activates quorum sensing-regulated virulence factors (5, 6) and, when overproduced, results in cell autolysis (7). It is surprising that bacteria produce a quinolone to communicate because quinolones are among the most potent, broad-spectrum antimicrobial agents used to treat bacterial infections (8). However, PQS does not seem to be bactericidal against Escherichia coli (4).
QSS identified thus far from Gram-negative bacteria regulate biofilm formation or the expression of bioluminescence or virulence genes (1). QSS thought to control cell density have been shown to block DNA replication in E. coli. However, despite intensive efforts, neither this QSS nor any other involved in stationary phase regulation has been purified (9). Known QSS vary widely in their ability to diffuse across bacterial membranes. The Vibrio fischeri AHL signal, 3OC6-HSL, freely diffuses (10). C4-HSL, a Pseudomonas aeruginosa AHL signal, is also freely diffusible (11). However, a larger P. aeruginosa AHL signal, 3OC12-HSL, exits the cell inefficiently (11). The multidrug resistance (MDR) efflux pump MexAB and, perhaps, MexCD, MexEF, and MexGHI assist the diffusion of the 3OC12-HSL signal from P. aeruginosa (11, 12, 13–14). MexEF may also emit PQS from the cell (12).
Gram-negative bacteria contain four families of MDR transporters: RND, MFS, MATE, and SMR (15). RND pumps, through a connection to an outer membrane channel, are able to emit substrate (from the cytoplasm or periplasm) directly to the outside of the cell (16). The other types are located in the inner membrane and emit substrate into the periplasmic space.
An integration of transporters and quorum sensing is suggested by the following. (i) MDR pumps can efflux otherwise non-freely diffusible QSS (11). Overproduction of MexAB, MexCD, and MexEF in P. aeruginosa decreased production of virulence factors positively regulated by quorum sensing (12, 14, 17), and the overproduction of MexAB decreased extracellular concentrations of 3OC12-HSL (17). Although suggestive of a link between MDR pumps and quorum sensing, the process is not clear. (ii) The MexGHI pump was identified in a screen for gene products that affect the transcriptional expression of a quorum sensing-regulated virulence factor (18); deletion of MexGHI decreased virulence factor production (13). (iii) Microarray analyses in P. aeruginosa show that the expression of at least seven MDR pumps is regulated positively by C4-HSL and 3OC12-HSL (19, 20–21).
In E. coli, SdiA, which is homologous to the LuxR family of quorum-sensing transcription factors, is regulated in a density-dependent manner (22). SdiA overproduction leads, perhaps indirectly (23), to increased resistance to quinolones and other antibiotics through up-regulation of AcrAB (24, 25). Lack of SdiA led to slightly lower AcrB levels and decreased drug resistance (25). Like its P. aeruginosa Mex efflux pump homologs, AcrAB is an RND transporter. These data suggest that AcrAB may play a role in quorum sensing through the active efflux of QSS.
Results
acrAB Null Mutants Overgrow in Stationary Phase.
In E. coli, adaptation to growth in stationary phase is regulated by quorum sensing (26). Thus, it seems logical that entry of cells into stationary phase may be defective in the absence of a gene product that is important in quorum sensing. Therefore, we monitored the growth of ΔacrAB and its isogenic WT strain (Fig. 1). Growth rates of WT and ΔacrAB cells during exponential growth phase were indistinguishable, as has been shown (27). However, as the cultures approached stationary phase, ΔacrAB deviated from WT and grew to a statistically significant increased density of 17 ± 4% (Table 1). This overgrowth has not been reported previously, and was the same in LB (Fig. 1A) or M9 (Fig. 1B) medium. The stationary phase section of the growth curve was done nine times with the same results.
Fig. 1.
Growth of WT and MDR transporter mutant cells. (A) Strains W4573 (WT, ○) and LZ2184 (ΔacrAB, •) were inoculated to an A600 = 0.02 (time = 0) into LB medium and grown at 37°C. A600 readings were taken at the time points shown. (B) Growth curve as in A except strains were grown in M9 minimal medium throughout the experiment. (C) WT, tolC, ΔacrAB, and ΔacrAB tolC strains were grown as in A. A600 readings taken 5–12 h after inoculation were averaged for each experiment. The ratio of mutant∕WT is shown from four separate experiments done in triplicate. Error bars represent SEM. (D) Growth curve in LB medium as in A, using strains W4573 (WT, ○), LZ2184 (ΔacrAB, •); LZ2096 (ΔnorE, ▵), LZ2301 (ΔmdfA, □), and LZ2306 (ΔacrAB ΔnorE, ◇). Cultures were inoculated in duplicate, and the values shown are the mean with SEM. Where no error bars, the deviation is smaller than the symbol. Three-way factorial analysis of data from D and two additional trials is shown. The three factors were trial number, strain, and time. Time was <10 h and ≥10 h. The strain data (for time >10 h) were statistically significant (WT was different from ΔacrAB and ΔacrAB ΔnorE) with an overall probability of type I error = 0.05. No other factors were significant. (E) Effect of CM from WT or ΔacrAB ΔnorE on growth of WT cells. The ratio of A600 readings from WT cells grown in CM from ΔacrAB ΔnorE to that of CM from WT is plotted with time. The experiment was performed twice in triplicate; SEM is shown.
Table 1.
Summary of stationary phase cell measurements
| Strain | A600 (12 h)n = 1 | A600* (12–20 h)n = 9 | Protein (8 h) mg∕mln = 1 | Protein* (8–14 h)n = 4 | No. of cells × 109 per ml (10 h)n = 1 | No. of cells (10–14 h)n = 2 |
|---|---|---|---|---|---|---|
| W4573 (WT) | 5.16 ± 0.06 | 1 ± 0.03 | 1.81 ± 0.02 | 1 ± 0.03 | 7.28 ± 0.73 | 1 ± 0.1 |
| LZ2184 (ΔacrAB) | 5.83 ± 0.18 | 1.17 ± 0.04 | 2.04 ± 0.06 | 1.15 ± 0.04 | 8.85 ± 0.8 | 1.22 ± 0.09 |
Strains were grown in LB medium at 37°C with shaking for the times indicated. n, Number of separate experiments used to calculate the normalized values. Because of day-to-day variation, the WT was normalized to 1. The ΔacrAB was calculated relative to that. Within each experiment, strains were analyzed in duplicate. The standard deviation is shown.
*Normalized.
Averaged from four experiments done in duplicate, the stationary phase ΔacrAB cultures contained 15 ± 4% more protein than WT (Table 1) (protein levels were identical in logarithmic phase). In exponential growth, where the cell densities were identical, the ΔacrAB cells were, on average, ≈15% longer than WT cells (data not shown), as determined by using light microscopy. In stationary phase, the WT and ΔacrAB cells were all short and approximately equal in size (data not shown). Therefore, cell length does not seem to account for the increased density and protein levels seen in ΔacrAB. Using a Petroff-Hausser counter, we found that the ΔacrAB cultures contained 22 ± 9% more cells in stationary phase (Table 1). Therefore, the overgrowth phenotype of the ΔacrAB strain seems to be caused by an increase in cell number. A fraction of the cultures used for cell counting was spread onto LB-agar and incubated overnight. At 6–8 h (late logarithmic phase), the ratio of cells able to form colonies to total cell number was 54 ± 5.5% in WT cells and 58.4 ± 12.6% in ΔacrAB. After 14 h (stationary phase), the WT ratio was 45 ± 4.8%, and the mutant ratio was 42 ± 8.8%. Thus, there is no difference in the viability of the two strains.
The outer membrane channel TolC associates with AcrAB to transport substrates out into the environment. tolC210::Tn10 was transduced into WT and ΔacrAB strains, and growth of the resultant strains was measured. In four experiments (in triplicate), tolC210, ΔacrAB, and ΔacrAB tolC210 each overgrew by an average of ≈10% (Fig. 1C) compared with WT (P < 0.01 by Student’s t test).
ΔnorE, but Not ΔmdfA, Mutants Overgrow in Stationary Phase.
No difference was found in logarithmic phase, but ΔnorE cells overgrew in stationary phase (Fig. 1C). The ΔacrAB ΔnorE strain grew identically to WT in logarithmic phase and overgrew in stationary phase to the same extent as ΔacrAB or ΔnorE (Fig. 1C). The removal of mdfA had no effect on growth of WT (Fig. 1C), ΔacrAB, or ΔnorE strains (data not shown).
Effect of Conditioned Medium (CM) on Repression of Cell Growth.
The overgrowth of ΔacrAB, ΔnorE, and ΔacrAB ΔnorE would be expected if these cells were defective in emitting a growth cessation QSS. If so, CM from the mutant should not be as efficient as CM from WT cells at inhibiting cell growth. We measured the effects of CM harvested from WT, ΔacrAB, ΔnorE, or ΔacrAB ΔnorE on WT cell growth. CM was harvested when the strains reached a density of A600 = 3.0, at a point before the overgrowth of the mutant strains when cell density and time in the culture medium are identical between the donor strains. Exponentially growing (A600 ≈ 0.4) WT cells were then inoculated into CM from the WT or mutant strains to an A600 = 0.02, and growth was followed spectrophotometrically. CM from ΔacrAB or ΔnorE inhibited growth identically to CM from WT (data not shown). However, CM from the ΔacrAB ΔnorE cells allowed more growth than the CM from WT cells, with the maximal difference (20%) observed at 1.5 h after inoculation (Fig. 1E). This finding suggests that there was more QSS exuded by the WT strain compared with the double mutant. A synergistic effect between NorE and AcrAB (28), combined with NorE emitting its substrate only into the periplasm, may explain why either single mutant had no differential effect. After 3 h of growth, there was no difference in the density of the cells grown in any of the CM. This finding suggests that, over time, QSS produced by the growing cells overwhelms QSS present in the CM.
Overexpression of MDR Transporters Represses Growth.
We determined the effect of acrAB overexpression on growth by using plasmid pAB (29), which contains the acrAB genes controlled by an arabinose inducible promoter (Fig. 2A–C). WT cells transformed with either pAB or the parent vector, pACYC184, were grown in LB medium with or without arabinose. Without arabinose, cells with pAB grew similarly to cells with pACYC184. However, cells that contained pAB, in arabinose, had a decrease (≈50%) in growth compared with cells with pACYC184 (Fig. 2 A–C). Cells kept below A600 = 0.4 by serial dilution (preincubation for 3, 5, or 7 h in arabinose-containing medium before inoculation into fresh arabinose containing medium at A600 = .02) grew identically with pAB or pACYC18 (data not shown). Thus, acrAB deletion or overexpression has no effect on logarithmic growth, even over many cell doublings. Arabinose levels of 0.2% and 0.02% led to a 10-fold increase in AcrA levels at 3.5 h (28). We found that AcrA levels present at A600 ≈ 0.2, in cells grown for 4, 6, or 8 h in arabinose, were also 10-fold higher (data not shown). Thus, cells overproducing AcrAB exhibit decreased growth at high cell densities and not at low cell densities, despite being overproduced in both.
Fig. 2.
Growth of cells overexpressing MDR transporters. LZ24 (WT) cells vector pACYC184 (○) or the acrAB-encoding plasmid pAB (□) were grown in LB medium containing chloramphenicol and either no (A), 0.02% (B), or 0.2% (C) arabinose for 3 h and then inoculated into fresh medium to A600 = 0.02 (time = 0). Cultures were grown at 37°C. A600 readings were taken at the times shown. (D) Cell growth as above except in LB medium with ampicillin, and the strain (LZ24) with vector pC108 (○), pSUP4 (norE; □), pSUP5 (mdfA; ▵), or pSY2 (norE1; ◇). Cultures were inoculated in duplicate, and the values shown are the mean with SEM. Where there are no error bars, the SEM is smaller than the symbol. The entire experiment was repeated three times with the same results. (E) Growth rates of WT (W4573) cells in CM from cells containing either pACYC184 or pAB. Exponentially growing WT cells containing pACYC184 were inoculated at A600 = 0.02 (time = 0) into either LB medium with chloramphenicol and 0.2% arabinose (circles); CM harvested at A600 = 1.1 (squares) from cells with pACYC184 (open symbols) or pAB (closed symbols); or CM harvested at A600 = 1.8 (triangles) from cells with pACYC184 or pAB and grown at 37°C. A600 readings were taken as shown. Cultures were inoculated in duplicate into the specified CM; SEM was smaller than the symbols. The experiment was repeated twice with the same results.
pSUP4 and pSUP5 contain, respectively, the norE and mdfA genes under the control of their natural promoters inserted within the tetA gene in pBR322. pBR322-based pSY2 contains norE1, an inactive allele of norE (28). Cells with pSUP4 were the same as control cells in logarithmic growth, but grew to a final stationary phase density ≈65% that of control cells (Fig. 2D). pSUP5 and pSY2 had little or no effect.
CM from Cells Overexpressing acrAB Represses Cell Growth.
If overexpression of acrAB leads to decreased stationary phase growth because of increased efflux of a growth cessation signal, than CM derived from cells overexpressing acrAB should repress growth more than CM from WT cells. CM was prepared from cells containing pAB or pACYC184. CM was harvested at A600 = 1.1 or 1.8. Exponentially growing (A600 ≈ 0.4) WT cells with pACYC184 grown in LB broth with arabinose were inoculated into fresh LB medium with arabinose, or into supplemented CM to an A600 = 0.02, and growth was monitored. Vector CM harvested at either A600 = 1.1 or 1.8 repressed cell growth compared with fresh LB medium. The CM harvested at A600 = 1.8 repressed growth more than CM harvested at A600 = 1.1 (Fig. 2E). The acrAB overexpression CM harvested at A600 = 1.1 had the same effect on growth as the vector CM. However, the acrAB overexpression CM harvested at A600 = 1.8 repressed cell growth more than the vector CM. The repression was quick (≈35% reduction, ≈30 min) and increased to ≈60% reduction at 3.5 h (Fig. 2E). At 24 h, all cultures reached similar cell density (data not shown), suggesting that the growth inhibition signal becomes ineffective. That the pAB and vector CM harvested at A600 = 1.1 had the same effect on cell growth suggests that the signal needs to accumulate to higher density outside of the cell to exert an affect.
Overexpression of acrAB or norE, but not mdfA results in decreased stationary phase growth (Fig. 2A–D). We tested whether CM derived from cells overexpressing norEor mdfA affected cell growth. No difference between these CM (harvested at A600 = 1.8) was observed (data not shown). The difference between AcrAB and NorE CM effects may result from AcrAB emitting its substrate outside of the cell and NorE emitting into the periplasm.
Effect of Mixing ΔacrAB and WT Cells.
WT and mutant cells were grown together at a 1:1 ratio in LB medium. The replacement of the acrAB genes with the kan gene in the mutant allowed us to distinguish it from WT cells. Antibiotic markers do not affect viability in the absence of antibiotic selection (30).
The mixed cells reached an intermediate density between that of ΔacrAB and WT (Fig. 3A), which could have indicated that each strain grew as before and that the signals were not provided to the mutant from the WT strain. Indeed, the viability of the mixed population was the same as with either mutant alone (Fig. 3B and data not shown). However, whereas the WT and mutant cells are represented equally initially, at the same cell density where the mutant strain deviated from WT (Fig. 1), the mutant strains undergrew in the mixed population, to finally represent ≈30% of the total cells (Fig. 3C). The ratio of mutant to WT cells was unchanged for an additional 24 h. For the mixed population to reach the intermediate cell density shown in Fig. 3A, the WT cells must overgrow. We interpret this finding to mean that, when provided by WT cells, QSS are able to penetrate the ΔacrAB cells to regulate entry into stationary phase and prevent overgrowth, but that these signals may not readily escape in the absence of the AcrAB pump.
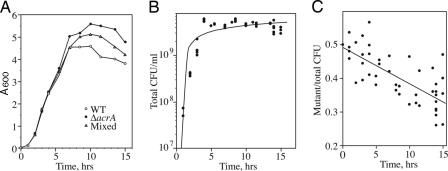
Fig. 3.
Effect of mixing ΔacrAB and WT cells. Exponentially growing WT alone (W4573), ΔacrAB alone (LZ2184), or WT and mutant mixed at a 1:1 ratio were inoculated at A600 = 0.02 (time = 0) and grown at 37°C. At the times shown, aliquots were taken. (A) The optical density of WT alone (○), ΔacrAB alone (•), and mixed cells (▵) are plotted over time. (B) From the mixed cells experiment, the total cell viability was measured by cfu on LB-agar plates. (C) WT and ΔacrAB were distinguished by spreading the mixed cells onto LB-agar plates with or without kanamycin. The ratio of the cfu with over without kanamycin is plotted against time. The combined data are from four experiments. The best fit line (y = −0.011x + 0.491) has an r-value = 0.705.
Effect of LuxS on acrAB Overexpression-Mediated Growth Repression.
If overproduction of the MDR transporters represses growth because of increased efflux of a growth-cessation signal, then cells that are defective in generating this signal should not be affected by overproduction of the MDR pumps. Only one QSS, synthesized by LuxS, has thus far been identified in E. coli. This QSS is in the autoinducer 2 (AI-2) family, which may form altered chemical forms depending upon environment (31, 32). QSS made by LuxS has been proposed to regulate stationary phase in E. coli (33). We transformed a ΔluxS strain with pAB or pBAD to test whether AI-2 is emitted by AcrAB. Removal of luxS did not prevent the growth repression caused by acrAB overexpression when the cells were grown with arabinose (Fig. 6, which is published as supporting information on the PNAS web site). Similar results were also observed for norE overexpression. Thus, the acrAB and norEoverexpression phenotype is not dependent on LuxS, indicating that AI-2 is not the putative QSS emitted by these transporters.
Effect of acrAB Overexpression on rpoS Expression.
RpoS is a σ factor that controls expression of multiple genes involved in adaptation and survival in stationary phase. It is present at low levels during logarithmic growth but increases dramatically as cells transition into stationary phase (34, 35). RpoS also is involved in how cells react to starvation, osmotic stress, and the stringent response (36). Studies have linked HSL (37, 38) and PQS (39) with RpoS expression. Thus, RpoS could be a downstream target of a possible growth cessation QSS emitted by AcrAB and NorE.
We measured rpoS expression in cells overexpressing acrAB (Fig. 4 A and B). Strain HS143 (40) contains rpoS::lacZ. Cells were transformed with either pAB or pACYC184 and grown in LB medium with 0.02% arabinose for >4 h before the start of the assay. In cells containing the vector plasmid, rpoS transcript levels at low cell density were low but were up-regulated dramatically at high cell density, consistent with previous results (Fig. 4A). At low cell density (A600 ≤ 0.4), overexpression of acrAB did not affect rpoS, but overexpression of acrAB stimulated rpoS expression at a lower cell density than in cells with normal levels of acrAB. Cells overexpressing acrAB had the same rpoS transcript levels at a density of A600 = 1.2 as that of cells with vector at a density of A600 = 1.8 (Fig. 4B).
Fig. 4.
Effect of MDR transporters on rpoS expression. (A and B) Effect of acrAB overexpression on rpoS expression. (A) Strain HS143 contains, on the chromosome, the lacZ gene controlled by the rpoS promoter. HS143, containing pACYC184 (□) or pAB (■), was grown in LB broth with chloramphenicol. At various times, cell densities and β-galactosidase activities were measured. (B) The experiment was performed as in A, except cells were grown with 0.02% arabinose for 4 h before the first data point and during the time course. (C) Effect of acrAB or norE mutations on rpoS expression. Isogenic strains containing lacZ under control of the rpoS promoter (LZ2755, WT, □), and acrA1 (LZ2756, ○), ΔnorE (LZ2757, ▵), or acrA1 ΔnorE (LZ2758, ◇) were grown in LB broth, and β-galactosidase activity was measured as in A. Experiments were repeated four times with the same results.
Cells overexpressing acrAB grow slower once they reach mid-logarithmic phase (A600 < 0.4). If rpoS expression is sensitive to growth rate, then one reason for the changed rpoS transcript level might be a difference in growth rates. We determined the rpoS expression profile for WT cells grown at 30°C and 37°C. WT cells grown at 30°C double ≈50% slower as compared with cells grown at 37°C (data not shown). The rpoS promoter activity, at various cell densities, was identical for cells grown at either temperature (data not shown) indicating that growth rate, alone, does not affect rpoS expression. Cells overexpressing either mdfA or norE exhibited the same rpoS transcript levels as WT cells (data not shown).
Effect of Decreased acrAB on rpoS Expression.
If cells overexpressing acrAB cause an increase in rpoS transcript levels because a QSS is accumulating faster, then cells lacking acrAB should exhibit the opposite effect. The rpoS::lacZ allele was transduced into an acrAB mutant (N43) and its isogenic WT strain (W4573). At low cell densities, rpoS promoter activity in the acrAB mutant cells was ≈85% that of WT cells (Fig. 4C). WT cells had increased rpoS transcript levels with increased cell density, as expected. However, rpoS transcripts were reduced ≈50% in acrAB mutant cells (Fig. 4C). This finding suggests that acrAB is required for the full increase in rpoS transcription levels during the transition to stationary phase. The removal of norE had no effect on rpoS transcript levels, in either WT or acrAB cells (Fig. 4C).
Discussion
E. coli exhibits several quorum sensing-regulated phenotypes: inhibition of DNA replication initiation (9), regulation of the sdiA promoter (22), regulation of virulence genes in pathogenic E. coli (41, 42), regulation of genes involved in energy production in stationary phase (33, 43), and regulation of RpoS (26, 34).
AcrAB is positively regulated by the quorum sensing-regulated transcription factor SdiA. We have shown here that ΔacrAB cells grew to higher cell density than WT as measured by optical density, total protein levels, and visible and viable cell counts. The ΔnorE and tolC mutant strains grew to similar cell density as the ΔacrAB strain. Previously, a ΔluxS mutant was reported to grow faster than WT cells (42). However, closer examination of the data reveals that the logarithmic growth was the same. Only when cells exited logarithmic growth did ΔluxS overgrow. Thus, ΔluxS is another potential example of an overgrowth phenotype.
A possible explanation for the increased density of the ΔacrAB or ΔnorE strains is that they emit less QSS into the medium. Consistent with this finding, WT cells cultured in CM derived from ΔacrAB ΔnorE were less efficient at suppressing growth than CM from WT cells. This effect was temporary, peaking at 90 min after inoculation. An unidentified E. coli QSS found in CM that inhibits DNA replication also exerted only a temporary inhibitory effect (9).
We propose that AcrAB, NorE, and perhaps other MDR pumps promote cell–cell communication by extruding QSS more efficiently than the signals can diffuse on their own (Fig. 5). Although AcrAB and NorE might be extruding multiple QSS, we consider the simplest case, one QSS (Fig. 5). As cells grow and synthesize QSS, the extracellular concentration increases. Once the extracellular QSS concentration is high enough (quorum is reached), the entire population of cells enters stationary phase. The effects of AcrAB, MdfA, and NorE upon entry into stationary phase is illustrated in Fig. 5B. Strains lacking the acrAB or norE gene have decreased, and strains overexpressing acrAB or norE have increased, extracellular QSS relative to WT cells. Thus, those cells enter stationary phase accordingly. MdfA had no effect on this process. This transporter may function instead to allow alkali tolerance (44).
Fig. 5.
Model for a role of MDR transporters in quorum sensing. (A) The outer membrane of Gram-negative bacteria is more permeable than the inner membrane. Although some QSS diffuse freely across cell membranes, efflux pumps assist other QSS (denoted as squares). The AcrAB∕TolC transporter expels QSS from the cytoplasm and periplasm to outside the cell. The NorE pump is located in the inner membrane and expels its substrate into the periplasm. When QSS reach threshold density, a signal transduction cascade is triggered, perhaps through a receptor, R, as shown. RpoS increases, causing entry into stationary phase. (B) Effect of acrAB, norE, or mdfA efflux pump levels on entry into stationary phase. At low cell density, intra- and extracellular QSS concentrations are low. As cell density increases, extracellular QSS concentration increases until the probability that the QSS encounters its periplasmic receptor is high. Increasing efflux pump levels by overexpression from plasmids pAB (acrAB), pSUP4 (norE), or pSUP5 (mdfA) are denoted as arrows pointed up. Overexpression of either acrAB or norE hastens the increase in the extracellular concentration of QSS and, therefore, entry into stationary phase. Cells lacking acrAB or norE have a delayed increase in extracellular QSS concentration that delays entry into stationary phase.
In Vibrio harveyi, the AI-2 signal binds to a periplasmic LuxP receptor that transduces the signal through sensor kinases and a phosphotransferase to a response regulator (1). Our model proposes a periplasmic receptor for QSS because cells lacking efflux pumps accumulate QSS inside the cell, but this does not trigger entry into stationary phase. In addition, when WT and acrAB mutant cells are mixed, the acrAB cells seem to act as a sink for the QSS. This finding may result from QSS entering the periplasm of both WT and mutant cells equally well, but WT cells can efficiently efflux the QSS from the periplasm unlike the acrAB cells.
RpoS is up-regulated dramatically as cells exit logarithmic growth. A reasonable model is that QSS may trigger rpoS expression. rpoS transcripts are reduced in acrAB mutant cells. In addition, overproduction of AcrAB resulted in earlier induction of rpoS, which may indicate that AcrAB emits QSS that trigger rpoS. The general stress caused by protein overexpression indirectly may trigger RpoS expression (45). If this is the case, then even WT levels of AcrAB cause such stress to the cells. Also, overproduction of MdfA or NorE has no effect on rpoS promoter activity.
We found that the luxS null mutant still shows repression of growth by acrAB or norE overexpression. Thus, it is unlikely that LuxS synthesizes a signal emitted by these pumps. The only molecules that AcrAB and NorE are both known to efflux are a subset of the fluoroquinolone class of antibiotics (28). Perhaps the QSS affecting stationary phase resembles these fluoroquinolones.
Bacterial genomes contain tens to hundreds of transporters (46). Different efflux pumps, or families of pumps, might have different signal specificities, leading to a wide range of regulation and communication possibilities within a microbial community. QSS can act as agonists or antagonists with each other (47). The multiple signals and their hierarchies may provide a resilient and adaptable “intelligence network” that permits bacteria to survey and react quickly to the surrounding environment.
Xenobiotics may be expelled by MDR transporters, not because they are recognized as toxins, but because their structure resembles communication signals normally exported. Sublethal concentrations of several antibiotics, including fluoroquinolones, induce quorum-sensing regulator lux genes (ref. 48 and J. Davies, personal communication). In addition, PQS and 3OC12-HSL signals exert immunomodulatory effects on host organisms (49), further blurring the line between communication and drug molecules. It is imperative that the functions of MDR pumps be delineated before we can hope to understand the complex nature of infection and drug resistance or use pump inhibitors in chemotherapy.
Murine hematopoietic stem cells have very high levels of efflux activity (50, 51), which inversely correlates with stem cell differentiation (52, 53). Overexpression of P-glycoprotein seems to increase hematopoietic stem cell activity (54), and the ABC transporter Bcrp1∕ABCG2 is expressed in stem cells (55). In the tissue proportioning that dictates prestalk and prespore cells in Dictyolstelium, pharmacological evidence suggests that prespore-specific transporters preclude the response of those cells to a stalk cell inducer (56). Perhaps a general function of these transporters is to efflux signals that would otherwise promote differentiation, as part of a cell–cell communication mechanism similar to that proposed here. The natural signals emitted by both prokaryotic and eukaryotic pumps will be of enormous future interest.
Materials and Methods
Strains, Plasmids, and Reagents.
E. coli strains, plasmids, and buffer conditions used are listed in Table 2, which is published as supporting information on the PNAS web site. For all comparative analyses, isogenic strains were used. For simplicity, we refer to the strains by their relevant genotype. Unless specified otherwise, all chemicals were reagent grade obtained from Fisher Scientific or Sigma.
Cell Density Measurements.
Logarithmically growing cells were inoculated to A600 = 0.02 into LB medium (time = 0), and spectrophotometric readings were taken over time. Aliquots were plated on LB-agar and incubated 12–15 h at 37°C to obtain colony-forming units∕ml (cfu∕ml). To determine total cell protein, 1 ml of cells grown overnight was harvested, washed with ddH20, resuspended in 200 μl of lysis buffer (1% SDS∕350 mM 2-mercaptoethanol∕62.5 mM Tris·Cl, pH 6.8), incubated at 100°C for 5 min, and subjected to Bradford assays (Bio-Rad). Cell number was determined from cultures fixed with formaldehyde (5% final concentration) by counting, using a Petroff-Hausser slide counter and a Zeiss phase contrast microscope.
For growth in medium with arabinose, exponentially growing cells were inoculated into LB medium with arabinose and grown for 3 h for induction to A600 = 0.2–0.4. These arabinose-induced cells were then diluted to A600 = 0.02 in fresh medium with arabinose (time = 0). For the mixed growth experiments, exponentially growing cells were inoculated to a cell density of A600 = 0.02. At various times, A600 was measured, a dilution was spread onto LB-agar (for total cfu∕ml) or onto LB-agar with 50 μg∕ml kanamycin (for cfu∕ml of ΔacrAB::kan). Replica plating from LB-agar plates onto LB-agar plates with kanamycin was also used.
CM Assays.
Exponentially growing WT (W4573) or ΔacrAB ΔnorE (LZ2186) cells were inoculated to A600 = 0.005 into LB medium, grown until A600 = 3.0 (≈5 h), and pelleted by centrifugation (Beckman) at 3,000 × g for 10 min. The supernatant was decanted and sterilized through 0.22-μM filters (Millipore) to generate CM, which was stored in single-use aliquots at −80°C. Before use, CM was supplemented with 0.5× LB medium (from 20× stock, 50 g∕liter Difco yeast extract, 100 g∕liter Difco tryptone peptone). Recipient cells were grown with shaking at 37°C to a cell density of A600 ≈ 0.4 in LB medium, and then diluted into CM (A600 = 0.02) and grown, with shaking, for the duration of the experiment. Aliquots were taken over time, and cell densities were measured.
For CM from WT cells (LZ24) overexpressing acrAB or not, exponentially growing arabinose-induced WT cells containing parent plasmid, pACYC184, or pAB were inoculated to A600 = 0.01 into LB medium with arabinose and grown until either A600 = 1.1 (≈3 h) or A600 = 1.8 (≈4 h) before being processed as above, except that the CM was supplemented with 0.2% glycerol, 0.25% casamino acids, and 2.5 μg∕liter thiamine.
β-Galactosidase Assays.
Exponentially growing arabinose-induced HS143 cells containing pAB or pACYC184 were inoculated into LB medium with arabinose to A600 = 0.02 (time = 0). Over time, aliquots were taken to measure A600 and β-galactosidase activity (57). To determine rpoS promoter activity in cells with inactive efflux pumps, we used strains LZ2755, LZ2756, LZ2757, or LZ2758.
Supplementary Material
Acknowledgments
We thank Drs. H. Nikaido (University of California, Berkeley), L. I. Rothfield (University of Connecticut, Farmington), B. Wanner (Purdue Univeristy, West Lafayette, IN), and H. I. Zgorskaya (University of Oklahoma, Norman) for strains and plasmids; Dr. J. Rojo for statistical analyses; Drs. G. Bennett, J. S. Butel, C. F. Earhart, M. A. Goodell, A. Kuspa, L. I. Rothfield, and D. A. Siegele for advice; M. J. Maynard for technical assistance; and Dr. M. J. Lombardo for reading the manuscript. This work was funded by National Science Foundation Grant MCB-0090880, National Institutes of Health Grant R01 AI054830, a Burroughs Wellcome Fund New Investigator Award in the Toxicological Sciences (to E.L.Z.), and National Research Service Award CA09197 from the National Cancer Institute (to S.Y.).
Glossary
ABBREVIATIONS:
- CM
conditioned medium
- MDR
multidrug resistant
- QSS
quorum-sensing signal
- AHL
N-acyl homoserine lactone
- AI-2
autoinducer 2
- PQS
2-heptyl-3-hydroxy-4-quinolone
- cfu
colony-forming unit.
Footnotes
Conflict of interest statement: No conflicts declared.
This paper was submitted directly (Track II) to the PNAS office.
References
- 1.Miller M. B., Bassler B. L. Annu. Rev. Microbiol. 2001;55:165–199. doi: 10.1146/annurev.micro.55.1.165. [DOI] [PubMed] [Google Scholar]
- 2.Xavier K. B., Bassler B. L. Curr. Opin. Microbiol. 2003;6:191–197. doi: 10.1016/s1369-5274(03)00028-6. [DOI] [PubMed] [Google Scholar]
- 3.Holden M. T., Ram Chhabra S., de Nys R., Stead P., Bainton N. J., Hill P. J., Manefield M., Kumar N., Labatte M., England D., et al. Mol. Microbiol. 1999;33:1254–1266. doi: 10.1046/j.1365-2958.1999.01577.x. [DOI] [PubMed] [Google Scholar]
- 4.Pesci E. C., Milbank J. B. J., Pearson J. P., McKnight S., Kende A. S., Greenberg E. P., Iglewski B. H. Proc. Natl. Acad. Sci. USA. 1999;96:11229–11234. doi: 10.1073/pnas.96.20.11229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Calfee M. W., Coleman J. P., Pesci E. C. Proc. Natl. Acad. Sci. USA. 2001;98:11633–11637. doi: 10.1073/pnas.201328498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Gallagher L. A., McKnight S. L., Kuznetsova M. S., Pesci E. C., Manoil C. J. Bacteriol. 2002;184:6472–6480. doi: 10.1128/JB.184.23.6472-6480.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.D’Argenio D. A., Calfee M. W., Rainey P. B., Pesci E. C. J. Bacteriol. 2002;184:648–6489. doi: 10.1128/JB.184.23.6481-6489.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cunha B. A. Adv. Ther. 1998;15:277–287. [PubMed] [Google Scholar]
- 9.Withers H. L., Nordstrom K. Proc. Natl. Acad. Sci. USA. 1998;95:15694–15699. doi: 10.1073/pnas.95.26.15694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kaplan H. B., Greenberg E. P. J. Bacteriol. 1985;163:1210–1214. doi: 10.1128/jb.163.3.1210-1214.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Pearson J. P., van Delden C., Iglewski B. H. J. Bacteriol. 1999;181:1203–1210. doi: 10.1128/jb.181.4.1203-1210.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kohler T., van Delden C., Curty L. K., Hamzehpour M. M., Pechere J. C. J. Bacteriol. 2001;183:5213–5222. doi: 10.1128/JB.183.18.5213-5222.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Aendekerk S., Ghysels B., Cornelis P., Baysse C. Microbiology. 2002;148:2371–2381. doi: 10.1099/00221287-148-8-2371. [DOI] [PubMed] [Google Scholar]
- 14.Sanchez P., Linares J. F., Ruiz-Diez B., Campanario E., Navas A., Baquero F., Martinez J. L. J. Antimicrob. Chemother. 2002;50:657–664. doi: 10.1093/jac/dkf185. [DOI] [PubMed] [Google Scholar]
- 15.Saier M. H., Paulsen I. T. Semin. Cell Dev. Biol. 2001;12:205–213. doi: 10.1006/scdb.2000.0246. [DOI] [PubMed] [Google Scholar]
- 16.Murakami S. N. R., Yamashita E., Yamaguchi A. Nature. 2002;419:587–593. doi: 10.1038/nature01050. [DOI] [PubMed] [Google Scholar]
- 17.Evans K., Passador L., Srikumar R., Tsang E., Nezezon J., Poole K. J. Bacteriol. 1998;180:5443–5447. doi: 10.1128/jb.180.20.5443-5447.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Diggle S. P., Winzer K., Lazdunski A., Williams P., Camara M. J. Bacteriol. 2002;184:2576–2586. doi: 10.1128/JB.184.10.2576-2586.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Whiteley M., Lee K. M., Greenberg E. P. Proc. Natl. Acad. Sci. USA. 1999;96:13904–13909. doi: 10.1073/pnas.96.24.13904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Schuster M., Lostroh C. P., Ogi T., Greenberg E. P. J. Bacteriol. 2003;185:2066–2079. doi: 10.1128/JB.185.7.2066-2079.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wagner V. E., Bushnell D., Passador L., Brooks A. I., Iglewski B. H. J. Bacteriol. 2003;185:2080–2095. doi: 10.1128/JB.185.7.2080-2095.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Garcia-Lara J., Shang L. H., Rothfield L. I. J. Bacteriol. 1996;178:2742–2748. doi: 10.1128/jb.178.10.2742-2748.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ahmer B. M. Mol. Microbiol. 2004;52:933–945. doi: 10.1111/j.1365-2958.2004.04054.x. [DOI] [PubMed] [Google Scholar]
- 24.Wei Y., Lee J. M., Smulski D. R., LaRossa R. J. Bacteriol. 2001;183:2265–2272. doi: 10.1128/JB.183.7.2265-2272.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rahmati S., Yang S., Davidson A. L., Zechiedrich E. L. Mol. Microbiol. 2002;43:677–685. doi: 10.1046/j.1365-2958.2002.02773.x. [DOI] [PubMed] [Google Scholar]
- 26.Huisman G. W., Kolter R. Science. 1997;265:537–539. doi: 10.1126/science.7545940. [DOI] [PubMed] [Google Scholar]
- 27.Ma D., Cook D. N., Alberti M., Pon N. G., Nikaido H., Hearst J. E. Mol. Microbiol. 1995;16:45–55. doi: 10.1111/j.1365-2958.1995.tb02390.x. [DOI] [PubMed] [Google Scholar]
- 28.Yang S., Clayton Rahmati S., Zechiedrich E. L. J. Antimicrob. Chemother. 2003;51:545–556. doi: 10.1093/jac/dkg126. [DOI] [PubMed] [Google Scholar]
- 29.Zgurskaya H. I., Nikaido H. J. Bacteriol. 2000;182:4264–4267. doi: 10.1128/jb.182.15.4264-4267.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Zambrano M. M., Siegele D. A., Almiron M., Tormo A., Kolter R. Science. 1993;259:1757–1760. doi: 10.1126/science.7681219. [DOI] [PubMed] [Google Scholar]
- 31.Chen X., Schauder S., Potier N., Van Dorsselaer A., Pelczer I., Bassler B. L., Hughson F. M. Nature. 2002;415:545–549. doi: 10.1038/415545a. [DOI] [PubMed] [Google Scholar]
- 32.Miller S. T., Xavier K. B., Campagna S. R., Taga M. E., Semmelhack M. F., Bassler B. L., Hughson F. M. Mol. Cell. 2004;15:677–687. doi: 10.1016/j.molcel.2004.07.020. [DOI] [PubMed] [Google Scholar]
- 33.DeLisa M. P., Wu C. F., Wang L., Valdes J. J., Bentley W. E. J. Bacteriol. 2001;183:5239–5247. doi: 10.1128/JB.183.18.5239-5247.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Mulvey M. R., Switala J., Borys A., Loewen P. C. J. Bacteriol. 1990;172:6713–6720. doi: 10.1128/jb.172.12.6713-6720.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Loewen P. C., Hengge-Aronis R. Annu. Rev. Microbiol. 1994;48:53–80. doi: 10.1146/annurev.mi.48.100194.000413. [DOI] [PubMed] [Google Scholar]
- 36.Hengge-Aronis R. Microbiol. Mol. Biol. Rev. 2002;66:373–395. doi: 10.1128/MMBR.66.3.373-395.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Latifi A., Foglino M., Tanaka K., Williams P., Lazdunski A. Mol. Microbiol. 1996;21:1137–1146. doi: 10.1046/j.1365-2958.1996.00063.x. [DOI] [PubMed] [Google Scholar]
- 38.Winzer K., Falconer C., Garber N.C., Diggle S.P., Camara M., Williams P. J. Bacteriol. 2000;182:6401–6411. doi: 10.1128/jb.182.22.6401-6411.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Diggle S. P., Winzer K., Chhabra S. R., Worrall K. E., Camara M., Williams P. Mol. Microbiol. 2003;50:29–43. doi: 10.1046/j.1365-2958.2003.03672.x. [DOI] [PubMed] [Google Scholar]
- 40.Schellhorn H. E., Stones V. L. J. Bacteriol. 1992;174:4769–4776. doi: 10.1128/jb.174.14.4769-4776.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sperandio V., Mellies J. L., Nguyen W., Shin S., Kaper J. B. Proc. Natl. Acad. Sci. USA. 1999;96:15196–15201. doi: 10.1073/pnas.96.26.15196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Sperandio V., Torres A. G., Giron J. A., Kaper J. B. J. Bacteriol. 2001;183:5187–5197. doi: 10.1128/JB.183.17.5187-5197.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Baca-DeLancey R. R., South M. M. T., Ding X., Rather P. N. Proc. Natl. Acad. Sci. USA. 1999;96:4610–4614. doi: 10.1073/pnas.96.8.4610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lewinson O., Padan E., Bibi E. Proc. Natl. Acad. Sci. USA. 2004;101:14073–14078. doi: 10.1073/pnas.0405375101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Gill R. T., Valdes J. J., Bentley W. E. Metab. Eng. 2000;2:178–189. doi: 10.1006/mben.2000.0148. [DOI] [PubMed] [Google Scholar]
- 46.Paulsen I. T., Chen J., Nelson K. E., Saier M. H. J. Mol. Microbiol. Biotechnol. 2001;3:145–150. [PubMed] [Google Scholar]
- 47.Bassler B. L. Curr. Opin. Microbiol. 1999;2:582–587. doi: 10.1016/s1369-5274(99)00025-9. [DOI] [PubMed] [Google Scholar]
- 48.Goh E. B., Yim G., Tsui W., McClure J., Surette M. G., Davies J. Proc. Natl. Acad. Sci. USA. 2002;99:17025–17030. doi: 10.1073/pnas.252607699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Hooi D. S., Bycroft B. W., Chhabra S. R., Williams P., Pritchard D. I. Infect. Immun. 2004;72:6463–6470. doi: 10.1128/IAI.72.11.6463-6470.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Goodell M. A., Rosenzweig M., Kim H., Marks D. F., DeMaria M., Paradis G., Grupp S. A., Sieff C. A., Mulligan R. C., Johnson R. P. Nat. Med. 1997;3:1337–1345. doi: 10.1038/nm1297-1337. [DOI] [PubMed] [Google Scholar]
- 51.Chaudhary P. M., Roninson I. B. Cell. 1991;66:85–94. doi: 10.1016/0092-8674(91)90141-k. [DOI] [PubMed] [Google Scholar]
- 52.Gussoni E., Soneoka Y., Strickland C. D., Buzney E. A., Khan M. K., Flint A. F., Kunkel L. M., Mulligan R. C. Nature. 1999;401:390–394. doi: 10.1038/43919. [DOI] [PubMed] [Google Scholar]
- 53.Jackson K. A., Mi T., Goodell M. A. Proc. Natl. Acad. Sci. USA. 1999;96:14482–14486. doi: 10.1073/pnas.96.25.14482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Bunting K. D., Zhou S., Lu T., Sorrentino B. P. Blood. 2000;96:902–909. [PubMed] [Google Scholar]
- 55.Zhou S., Schuetz J. D., Bunting K. D., Colapietro A. M., Sampath J., Morris J. J., Lagutina I., Grosveld G. C., Osawa M., Nakauchi H., Sorrentino B. P. Nat. Med. 2001;7:1028–1034. doi: 10.1038/nm0901-1028. [DOI] [PubMed] [Google Scholar]
- 56.Good J. R., Kuspa A. Dev. Biol. 2000;220:53–61. doi: 10.1006/dbio.2000.9611. [DOI] [PubMed] [Google Scholar]
- 57.Miller J. H. Experiments in Genetics. Woodbury, NY: Cold Spring Harbor Lab. Press; 1972. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.