Abstract
We report here the isolation and analysis of novel repA host range mutants of pPS10, a plasmid originally found in Pseudomonas savastanoi. Upon hydroxylamine treatment, five plasmid mutants were selected for their establishment in Escherichia coli at 37°C, a temperature at which the wild-type form cannot be established. The mutations were located in different functional regions of the plasmid RepA initiation protein, and the mutants differ in their stable maintenance, copy number, and ability to interact with sequences of the basic replicon. Four of them have broadened their host range, and one of them, unable to replicate in Pseudomonas, has therefore changed its host range. Moreover, the mutants also have increased their replication efficiency in strains other than E. coli such as Pseudomonas putida and Alcaligenes faecalis. None of these mutations drastically changed the structure or thermal stability of the wild-type RepA protein, but in all cases an enhanced interaction with host-encoded DnaA protein was detected by gel filtration chromatography. The effects of the mutations on the functionality of RepA protein are discussed in the framework of a three-dimensional model of the protein. We propose possible explanations for the host range effect of the different repA mutants, including the enhancement of limiting interactions of RepA with specific host replication factors such as DnaA.
Many plasmids are able to replicate in their original host and related species, and some of them have an extended host range which allows their establishment in several different bacterial species. At least two essential processes contribute to the host range phenotype: replication and partition (42). However, replication has received more attention in the effort to understand the host range phenotype.
The study of plasmid maintenance in different hosts has been addressed by focusing on broad-host-range replicons (see references 4 and 45 for reviews), mainly RSF1010 and RK2. Plasmid RSF1010 codes for its own replication initiator proteins, allowing a host-independent initiation replication mechanism (7, 39). RK2 uses a different strategy: it synthesizes two variants of the TrfA initiation protein in the oriV region, and the initiation of replication needs the participation of one or the other of these TrfA proteins, depending on the host (11, 12, 41). Moreover, the number, sequence, and spacing of its several DnaA boxes also influence the replication in RK2 (8, 9, 40). Genetic analysis suggests that interactions between the C-terminal region of TrfA and host replication factor(s) can specifically affect plasmid host range (3). Broad-host-range plasmids replicating by rolling-circle mechanisms have also been identified, a classic example of which is pLS1 (4, 5).
Narrow-host range-plasmids can also be used to identify host range determinants. A genetic approach consists in the isolation of plasmid mutants with mutations that allow their establishment in a new host. This approach has been followed with pPS10, a plasmid that can be efficiently established in Pseudomonas species at 30 and 37°C but not in Escherichia coli (33). Transformation of E. coli with pPS10 at 37°C does not yield visible colonies on solid medium, whereas at 30°C minute colonies that are not viable in subsequent cultures are seen (14). These characteristics make pPS10 a borderline plasmid in terms of efficient replication in E. coli and open up the possibility of transferring it from Pseudomonas savastanoi, a bacterium pathogenic to the olive tree, to a human enteric bacterium such as E. coli.
The minimal replicon of pPS10 contains the origin of replication (oriV), including four 22-bp iterons, followed by two 8-bp inverse repeats overlapping the −35 and −10 sequences of the repA promoter, and the repA gene coding for the initiation replication protein RepA, of 230 amino acids (see scheme in reference 30). Analysis of the primary structure of this protein, together with other biochemical and biophysical studies (20), indicates that RepA is built up from a leucine zipper motif, involved in dimerization (17, 22), followed by two winged-helix domains (amino acids 42 to 113 and 135 to 221) connected by a flexible linker. The second domain contains a putative helix-turn-helix motif for DNA binding (19). The RepA protein of pPS10 interacts as a monomer with the iterons of oriV and, jointly with the DnaA protein of the host, promotes the initiation of pPS10 replication (14, 17, 21, 34). On the other hand, the RepA protein, as a dimer, binds to the inverted repeats and represses its own transcription (17, 18).
A previous work (14) reported the isolation of a pPS10 mutant, pMM141, that was readily established as an autonomous replicon in E. coli and in the original host. The mutant contains a single-amino-acid change (A31V) in the leucine zipper region of RepA (14, 21). The mutation does not alter the dimerization state of RepA nor its capability to interact with the iterons of the origin or the repA operator (17), suggesting that it could have affected interactions with a critical host replication factor(s). In this sense, there was recently reported one DnaA mutant that allows the efficient establishment of wild-type pPS10 in E. coli (30). The genetic study of this mutant suggests that concerted interactions between RepA and DnaA play a relevant role in this establishment.
In this work we report the isolation and characterization of new host range mutations in RepA obtained by in vitro mutagenesis of the pPS10 replicon. The study of the pPS10 derivatives shows a variety of phenotypic effects and points to the improvement of interactions between RepA and host factors such as DnaA as a key determinant of its host range.
MATERIALS AND METHODS
Bacterial strains and plasmids.
E. coli K-12 strains used were CC118 (24), C600EL (hsdR) (kindly supplied by K. Nordström), and BT1000 (polA1) (47). Other strains were Pseudomonas aeruginosa PAO1024 (r− m+) (33), Pseudomonas putida KT2440 (r− m+) (1, 32), Agrobacterium tumefaciens DSM30510 (6), and Alcaligenes faecalis (German Research Center for Biotechnology).
Plasmids used were pRG9B, a pPS10-pBR322-derived shuttle vector (14); pMAL-c2 (New England BioLabs), an expression vector to fuse proteins to maltose binding protein (MBP); and pUCProm and pUCIt, pUC18Not derivatives that include the repA operator and the oriV iteron sequences of pPS10, respectively (19).
Media and growth conditions.
Cultures were usually grown in Luria-Bertani (LB) medium according to the method of Sambrook et al. (38) at 30 or 37°C (E. coli) or 30°C (other strains). When required, media were supplemented with antibiotics to select for plasmids conferring resistance to them: ampicillin (100 μg ml−1) and kanamycin (KAN; 50 μg ml−1 for E. coli, A. tumefaciens, and A. faecalis; 75 μg ml−1 for Pseudomonas). A. tumefaciens was grown in TY medium, according to the method of Díaz et al. (6), supplemented with rifampin at 50 μg ml−1. Growth of cells for MBPRepA expression was carried out in 2× YT medium (38) supplemented with 0.2% glucose.
General methods.
Plasmid DNA was purified with the Qiagen Miniprep kit. Restriction enzymes were from New England BioLabs, Roche Diagnostics, and Amersham Biosciences. Cloning procedures, gel electrophoresis of DNA and proteins, and radioactive labeling of DNA were performed as described by Sambrook et al. (38). Primer oligonucleotides were synthesized in a Beckman 1000 M DNA synthesizer, desalted, and used without further purification. DNA sequencing was carried out in a Perkin-Elmer ABI-Prism 377 DNA sequencer with fluorescent chain terminators.
Bacterial transformation and electroporation.
E. coli transformation was carried out as described by Lederberg and Cohen (29). In the case of Pseudomonas, we used the method described by Bagdasarian and Timmis (1). Electroporation-competent cells of E. coli, A. faecalis, and A. tumefaciens were prepared according to the method of Dower et al. (10). Competent cells, with the exception of A. tumefaciens, were kept overnight at 4°C in order to increase their competence. Electroporation-competent cells of Pseudomonas were prepared as described by Smith and Iglewski (43). In all cases one pulse of 25 μF, 200 Ω, 2.5 kV, was delivered with the Gene-Pulser instrument (Bio-Rad). Immediately after electroporation, cells were suspended in 2 ml of SOC medium (38) and incubated for 1 h at 30°C. Cells were then plated at appropriate dilutions, and the number of transformants, grown at 30°C, was calculated with respect to viable cells.
DNA mutagenesis.
Hydroxylamine mutagenesis was carried out by the protocol of Humphreys et al. (25). Twenty microliters of plasmid DNA (0.2 mg ml−1) was mixed with 100 μl of 0.1 M sodium phosphate (pH 6.0) containing 1 mM EDTA and 80 μl of 1 M hydroxylamine at pH 6.0 (Sigma). Upon incubation at 75°C for 30 min, the sample was extensively dialyzed against TE buffer (10 mM Tris, 1 mM EDTA, pH 8.0).
Determination of plasmid copy number.
Cell lysates were obtained from exponential-phase cultures as described by Projan et al. (37). DNA was transferred onto nitrocellulose membranes according to the method of Southern (44) by the protocol described in the work of Sambrook et al. (38). The 964-bp EcoRI fragment from plasmid pRG9B was labeled with [α-32P]dCTP and used as the hybridization probe. Total amounts of plasmid and chromosome DNA were evaluated by densitometry with the Molecular Dynamics densitometer fitted with the analysis program ImageQuant.
Resistance to gentamicin was used as an alternative way to determine copy number. One hundred microliters of a 10−6 dilution of overnight cultures was spread on LB plates containing different concentrations of gentamicin, and the MIC was determined for each mutant plasmid after counting of the colonies grown in 24 h.
Plasmid stability.
In order to check the segregational stability of pPS10 derivatives, a culture was grown at 30°C in KAN-containing LB medium to an optical density at 600 nm of 0.4. Then, 100 μl of a 10−5 dilution was plated on medium without antibiotic. After overnight incubation at 30°C, 100 independent colonies were patched on LB and LB-KAN plates to determine the number of cells that retained the plasmid after the incubation. Colonies growing on nonselective medium were pooled, and individual colonies were again isolated on LB plates. One hundred of these colonies were patched on LB and LB-KAN plates to determine the percentage of plasmid-containing cells after a second propagation on nonselective medium at 30°C. The procedure was repeated a third and fourth time when needed. These values correspond to the stability of the plasmid after approximately 30, 60, 90, and 120 generations without selection, as 30 generations is roughly the time needed to form a clearly visible colony (14).
Construction of pMalRepA plasmids.
The ATG-EcoRI fragment of pSBM plasmids (nucleotides 577 to 1500 according to the work of Nieto et al. [34]) was amplified by PCR with Pfu polymerase (Stratagene) and the oligonucleotides 5′-ATGGTCGAGAACAAAGTCACG-3′ (ATG initiation site underlined) and 5′-CCGGAATTCTCTGTGCCATAGC-3′ (EcoRI site underlined). The former oligonucleotide was phosphorylated by T4 polynucleotide kinase so that the PCR product, once digested with EcoRI, was cloned into the XmnI-EcoRI sites of pMAL-c2. The integrity of the constructs was verified by DNA sequencing.
Purification of MBPRepA proteins.
E. coli CC118 cells harboring the pMalRepA plasmids were grown to mid-exponential phase. MBPRepA expression was then induced with 10 μM isopropyl-β-d-galactopyranoside (IPTG), and incubation proceeded overnight. Cell extracts were applied to an amylose column equilibrated in a buffer containing 10 mM Tris (pH 7.4), 500 mM NaCl, 1 mM EDTA, and 1 mM dithiothreitol, and eluted with the same buffer plus 20 mM maltose. The purity of the samples was checked by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (28). The protein concentration was evaluated by absorption spectroscopy with an extinction coefficient ɛ280 of 78,678 M−1 cm−1, calculated with the parameters of Fasman (13) and making use of the known aromatic amino acid content of the fusion protein.
Circular dichroism spectroscopy.
Circular dichroism experiments were carried out in a Jasco J-720 spectropolarimeter fitted with a thermostatted cell holder and interfaced with a Neslab RTE-110 water bath. Isothermal wavelength spectra were acquired at a scan speed of 20 nm min−1 with a response time of 2 s and averaged over at least six scans at 5°C. The protein was previously dialyzed against a buffer containing 20 mM sodium phosphate (pH 6.5), 1 mM dithiothreitol, 1 mM EDTA, and 500 mM NaCl. The protein concentration was 2.5 μM, and the cuvette path length was 1 or 2 mm. Thermal denaturation experiments were performed with a heating rate of 50°C h−1 and a response time of 2 s. Scans were collected in 1-mm-diameter cuvettes and with a protein concentration of 2.5 μM. To avoid evaporation, samples were overlaid with mineral oil (Sigma). Data were analyzed according to a sequential three-state denaturation mechanism involving sequential unfolding of MBP and RepA moieties.
Electrophoretic mobility shift assays (EMSAs).
EMSA experiments were carried out based on the procedure of Fried and Crothers (15). For the operator binding assays, the radioactive probe used was the 90-bp NotI fragment of pUCProm, whereas for the iteron binding assays, the probe was the 440-bp NotI fragment of pUCIt (19). Both fragments were labeled with [α-32P]dCTP and 1 U of Klenow enzyme. Probe quantitation was carried out by agarose gel electrophoresis, measuring the fluorescence emitted by the sample stained with ethidium bromide, with the Bio-Rad Gel Doc system and the Molecular Analyst program. The RepA-DNA binding reaction took place in a final volume of 20 μl containing 25 mM HEPES-KOH buffer (pH 8.0), plus 1 mM EDTA, 4% saccharose, 50 mg of bovine serum albumin ml−1, 4 mM dithiothreitol, 50 mM KCl, 100 ng of phenolized and sonicated calf thymus DNA, either 24.5 fmol of iteron probe or 8.6 fmol of promoter probe, and increasing concentrations of MBPRepA protein. Reaction mixtures were assembled on ice and then transferred to room temperature for 30 min. Then 5 μl of loading buffer was added to the samples and loaded into a prerun nondenaturing 5% polyacrylamide gel in TBE buffer (89 mM Tris-borate, 89 mM boric acid, 2 mM EDTA, pH 8.0). Electrophoresis was carried out in 0.5× TBE at 200 V, keeping the temperature at 4°C for 7 h (iteron binding assays) or 4 h (operator binding assays). Finally, gels were dried at 80°C and autoradiographed.
Modeling of RepA structure.
RepA structure was modeled with the Swiss-Model threading utility (http://www.expasy.ch/spdbv/) (23) with the published structure of RepE as a template (27). The rough model was subsequently refined by steepest descent energy minimization.
Gel filtration analysis of the interaction between E. coli DnaA and MBPRepA variants.
Seventy-five microliters of a 25-fold-concentrated extract of a stationary-phase E. coli LE392(pUC392) culture (optical density at 600 nm = 3.5) (30) grown at 30°C was mixed with either 25 μl of an extract of a 2.5-fold-concentrated extract of a stationary-phase culture of E. coli CC118 harboring the different pMALRepA constructs grown at 37°C to an optical density at 600 nm of 2.2 or 25 μl of 20 mM sodium phosphate buffer (pH 7.0) plus 150 mM NaCl (control experiment). The mixtures were kept on ice for 30 min and subsequently loaded onto a Sephadex G-75 column (26 by 0.9 cm) equilibrated in 20 mM sodium phosphate buffer (pH 7.0) plus 150 mM NaCl and run with the same buffer at a flow rate of 1 ml min−1 and at 20°C. Exclusion (6.0-ml) and total (21.0-ml) volumes were determined with dextran blue and potassium dichromate, respectively. Two-hundred-microliter fractions were collected, and 100 μl of each fraction was subjected to a dot blot analysis with the ECL kit (Amersham Biosciences) and rabbit anti-DnaA antibodies in a 1:25,000 dilution (kindly supplied by W. Messer and H. Seitz, Max-Planck-Institut für molekulare Genetik, Berlin-Dahlem, Germany).
RESULTS
Isolation of new pPS10 derivative mutants able to replicate in E. coli at 37°C.
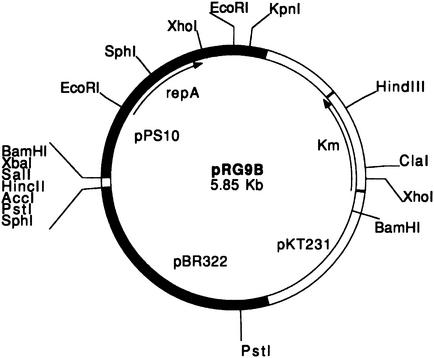
The plasmid chosen for this work was pRG9B (14), a cointegrate containing the pMB9 (from plasmid pBR322) and pPS10 replicons (Fig. 1). Using pRG9B has two major advantages: (i) it allows purification of a high amount of plasmid DNA in polA+ E. coli strains, where the pMB9 replicon directs the replication of the cointegrate, and (ii) pPS10-based replication can be analyzed in E. coli polA mutant strains (such as BT1000), where the pMB9 replicon is inactive (this strain, deficient in DNA polymerase I, cannot sustain replication of ColE1-type replicons). A solution of pRG9B DNA was treated with hydroxylamine, a mutagenic agent that fundamentally induces C→T transitions (36), and the reaction mixture was used to transform E. coli BT1000. Since the wild-type pPS10 replicon cannot be established in E. coli at 37°C, suitable plasmid mutants were directly selected by plating at this temperature.
FIG. 1.
Circular map of plasmid pRG9B containing the basic replicon of plasmids pPS10 (upper left) and pBR322 (lower left) and the Kmr gene from pKT231 (right). The most relevant restriction endonuclease sites are also shown.
In order to map the mutations induced by hydroxylamine, and to eliminate other transitions induced by the reagent outside the repA gene, we replaced the 964-bp EcoRI fragment of pRG9B with the corresponding fragments of the mutants, yielding the pSBM plasmid series (Table 1). All the recombinant plasmids conserved the ability to replicate in E. coli BT1000 at 37°C. Moreover, pSBM plasmids were also able to transform P. aeruginosa, with the exception of pSBM135 (data not shown). Therefore, the latter is a case of host range modification instead of host range broadening.
TABLE 1.
Mutations in the RepA protein that amplify or modify the host range of the pPS10 plasmid
| Plasmid | Amino acid change | Protein |
|---|---|---|
| pSBM31 | Ala31→Val (A31V) | RepA(A31V) |
| pSBM44 | Gly44→Ser (G44S) | RepA(G44S) |
| pSBM47 | Thr47→Ile (T47I) | RepA(T47I) |
| pSBM93 | Arg93→Cys (R93C) | RepA(R93C) |
| pSBM135 | Gly135→Ser (G135S) | RepA(G135S) |
The 964-bp EcoRI fragments in which the mutations were located were sequenced. Five different single-base mutations were found, leading to single-amino-acid changes in the RepA proteins (Table 1). pSBM31 possesses the same mutation as does the host range plasmid pMM141, previously described by our laboratory (14), i.e., the A31V change in the leucine zipper region of the RepA protein. pSBM44, pSBM47, and pSBM93 present their mutations (G44S, T47I, and R93C, respectively) within the first of the two proposed winged-helix domains of the protein (20). Finally, pSBM135 contains a G135S change at the beginning of the second winged-helix domain (20). It should be noted that no mutations in the Shine-Dalgarno region of the repA gene were found.
In order to check the establishment of the pPS10 derivatives in E. coli polA+ strains (such as C600EL), as well as in other gram-negative bacteria, pSBM plasmids with the pMB9 replicon deleted were constructed (pBM series). This was carried out by digestion with PstI (Fig. 1) followed by intramolecular religation of the plasmid and transformation in the P. aeruginosa PAO1024 strain. Plasmids pBM31, pBM44, pBM47, and pBM93 were thus obtained from the corresponding pSBM plasmids. The pBM9B plasmid was obtained from nonmutated pRG9B. All pBM plasmids except pBM9B were also able to efficiently transform BT1000.
Since pSBM135 cannot replicate in Pseudomonas, transformation to obtain the pBM135 construct was carried out directly in the E. coli BT1000 strain at 30°C. Kmr transformants formed larger colonies than those obtained with pRG9B and were viable in subsequent cultures in the presence of antibiotic selection, but it was not possible to detect plasmid DNA from them by standard purification procedures. For this reason, we did not carry out any studies on pBM135, using pSBM135 instead. This should be taken into account, especially when comparisons are carried out with the pBM derivatives of the other repA mutants (see below).
Stability of the host range mutants.
Although wild-type pPS10 can replicate in E. coli at 30°C, establishment in this host is very inefficient, and the plasmid is rapidly lost in the absence of selection (14). Table 2 shows the segregational stability of both the parental plasmid and the pBM mutants in P. aeruginosa and E. coli. In all cases the stability approaches 100% in P. aeruginosa at 30°C after 120 generations, whereas in E. coli C600EL the stability is different among the mutants: pBM44 and pBM47 are slightly more stable than the wild type, whereas pBM31 and pBM93 turned out to be the most stable ones. Plasmid pSBM135 is very unstable in E. coli BT1000 and, as described above, has lost its capacity to replicate in P. aeruginosa. Whereas pBM plasmids represent a host range broadening case, pSBM135 seems to be an example of a shift in host range, somehow defective, as the establishment in E. coli is only marginal.
TABLE 2.
Stability of mini-pPS10 host range mutants in E. coli and P. aeruginosa
| Plasmid | % Plasmid-containing cells after cyclea
|
|||||||
|---|---|---|---|---|---|---|---|---|
|
E. coli C600EL
|
P. aeruginosa PAO1024
|
|||||||
| 1 | 2 | 3 | 4 | 1 | 2 | 3 | 4 | |
| pBM9B | <1 | >99 | >99 | >99 | >99 | |||
| pBM31 | 69 | 34 | 7 | 3 | >99 | >99 | >99 | >99 |
| pBM44 | 23 | <1 | >99 | >99 | >99 | >99 | ||
| pBM47 | 19 | <1 | >99 | >99 | >99 | >99 | ||
| pBM93 | 78 | 53 | 37 | 21 | >99 | >99 | >99 | >99 |
| pSBM135b | <1 | NDc | ND | ND | ND | |||
Cycles of propagation in the absence of selection. One cycle corresponds to the number of generations (about 30) needed to form a colony from a single cell (14).
The stability was determined in the polA1 E. coli strain BT1000.
ND, not determined, as pSBM135 cannot be established in P. aeruginosa.
It is noteworthy that the pMM141 host range mutant of pPS10 shows better stability in E. coli than does pBM31, in spite of sharing the same repA mutation (14). This can be attributed to a second mutation detected outside the repA gene in pMM141, which is currently under investigation in our laboratory.
Copy number of the host range mutants.
The copy number of pPS10 in E. coli must be very low even at 30°C. Kmr colonies obtained at 30°C after transformation are very small and grow poorly either on plates or in liquid media. Moreover, we were unable to detect pPS10 in such transformants by standard DNA isolation procedures. However, its residual presence in E. coli is demonstrated by the ability of such preparations to transform the Pseudomonas strain PAO1024 (14). Taking this into account, it was important to test whether an increase in copy number could be a factor contributing to the establishment of pPS10 in E. coli.
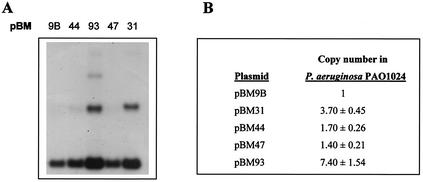
Densitometry of Southern hybridization gels with the pRG9B 964-bp EcoRI fragment as a probe (Fig. 2) clearly shows an increase in copy number for pBM31 and pBM93 in P. aeruginosa (four- and eightfold, respectively), relative to the wild-type pBM9B plasmid. pBM44 and pBM47 are found in roughly the same copy number as the wild-type derivative pBM9B. pSBM135, as described above, is unable to be established in P. aeruginosa. The presence of multimers in the case of pBM31 and pBM93 is noteworthy (Fig. 2).
FIG. 2.
Copy number of the host range mutants in P. aeruginosa. (A) Autoradiograph from total lysates of P. aeruginosa containing wild-type pBM9B and mutated pBM plasmids hybridized to a repA DNA probe. (B) Copy number of pBM mutants relative to that of pBM9B. Values are the averages obtained from five different experiments and were obtained by autoradiograph densitometry corrected to the same amount of chromosomal DNA in all the samples. pSBM135 is not included, as it cannot be established in P. aeruginosa.
Copy number in E. coli was studied in the polA+ and polA strains C600EL and BT1000, respectively (data not shown). We could not calculate the copy number of the mutants relative to the wild-type plasmid due to the marginal establishment of the latter in E. coli. As in P. aeruginosa, pBM93 displayed the highest copy number in both strains, followed by pBM31, pBM44, and pBM47. In BT1000, pBM47 and pSBM135 must be present in a very low copy number because a long exposure of the autoradiogram was necessary to detect the signal. In addition, replication efficiency was always higher in C600EL than in BT1000, suggesting that the polA1 mutation may influence the efficiency of pPS10 plasmid replication even though it should be, in principle, independent of DNA polymerase I.
An alternative method to measure copy number is to determine levels of resistance to antibiotics (46). Since the kanamycin resistance determinant confers resistance to gentamicin also (2), we used the latter antibiotic because Pseudomonas species are more sensitive to it than to KAN. Levels of resistance to gentamicin of P. aeruginosa PAO and E. coli C600 harboring the different plasmid variants are shown in Table 3. It can be seen that the MICs for the two sets of strains are comparable for the two bacterial species and are qualitatively similar to the data obtained by Southern blotting (Fig. 2). We did not attempt to quantify these values in terms of copy number since a possible nonlinear correlation can occur in cases of high values of this parameter (35).
TABLE 3.
MICs of gentamicin that abolish growth of bacterial strains harboring the pBM variants
| Plasmid | MIC (μg/ml) for strain:
|
|
|---|---|---|
| P. aeruginosa | E. coli C600EL | |
| None | 0.78 | 1.03 |
| pBM9B | 1.24 | NDa |
| pBM31 | 1.63 | 1.80 |
| pBM44 | 1.30 | 1.45 |
| pBM47 | 1.28 | 1.45 |
| pBM93 | 2.18 | 2.22 |
ND, not determined.
Mutant establishment in other bacterial strains.
It was of interest to study the establishment of the mutated plasmids in different bacterial strains other than P. aeruginosa and E. coli and belonging to different Proteobacteria subdivisions (48). Accordingly we determined the transformation efficiency and the stability of the pBM plasmids in P. putida (γ subdivision), A. faecalis (β subdivision), and A. tumefaciens (α subdivision), at 30°C.
Results are shown in Tables 4 and 5. Transformation efficiencies after electroporation of the mutant pBM plasmids in P. aeruginosa, E. coli, and A. faecalis were not significantly different from that of the wild type (Table 4). In P. putida, 100 to 1,000 times more transformants were obtained with the mutants than with pBM9B. Neither the wild type nor the mutants are able to transform A. tumefaciens. Stability of the wild-type and mutant plasmids is high in both species of Pseudomonas (>90%) (Table 5). On the other hand, the mutant plasmids are more stable than the wild type in E. coli and A. faecalis, in accordance with their increased host range phenotype. In both organisms, pBM31 and pBM93 show the highest stabilities (Table 5).
TABLE 4.
Electroporation efficiencies at 30°C of different gram-negative bacteria with the pBM variants
| Plasmid | No. of transformants/viable cell and ng of DNA
|
||||
|---|---|---|---|---|---|
| P. aeruginosa | E. coli | P. putida | A. faecalis | A. tumefaciens | |
| pBM9B | 0.6 | 9.7 | 0.04 | 16 | 10−4 |
| pBM31 | 2.9 | 0.9 | 3.0 | 5.3 | 10−4 |
| pBM44 | 3.8 | 1.6 | 18 | 24 | <10−4 |
| pBM47 | 0.8 | 2.6 | 12 | 7.8 | <10−4 |
| pBM93 | 3.1 | 6.3 | 6.0 | 21 | <10−4 |
TABLE 5.
Stability of pBM variants in different bacterial strains at 30°C
| Plasmid | % of cells containing plasmida
|
|||
|---|---|---|---|---|
| P. aeruginosa | E. coli | P. putida | A. faecalis | |
| pBM9B | >99 | <1 | 92 | <1 |
| pBM31 | >99 | 69 | >99 | 25 |
| pBM44 | >99 | 23 | >99 | 6 |
| pBM47 | >99 | 19 | >99 | 3 |
| pBM93 | >99 | 78 | >99 | 15 |
After 30 generations in the absence of selection. Approximately 30 generations are needed to form a colony from a single cell (14).
Structural characterization of MBPRepA proteins.
In order to determine how the mutations might affect the protein structure, stability, or functionality, we carried out the purification of RepA as a chimera with the MBP (MBPRepA). Details of the construction and purification of MBPRepA fusions are given in Materials and Methods. Attempts to release the MBP part to yield the RepA moiety by factor Xa protease digestion failed due to low digestion rates and the low solubility of the RepA protein (data not shown). In any case, MBPRepA fusions are soluble and have been previously shown to conserve their regulatory activity in vivo (16) and their DNA binding functionality in vitro (19).
In view of the RepA mutations reported in Table 1, which include some nonconservative changes, the possibility existed that they could alter the structure of RepA, inducing a conformational change that might affect its functionality. However, circular dichroism spectra show no significant changes between mutant and wild-type MBPRepA proteins (data not shown), strongly suggesting that the mutations do not affect the secondary structure to a significant extent. Another possibility could be that the structure was unaffected but the stability had changed, perhaps accounting for an increased or decreased in vivo stability of the complexes with the DNA or with the host range factors involved in the initiation of replication. For this reason, we performed thermal stability experiments by monitoring the change of circular dichroism signal with temperature. Typical traces were biphasic, with two transitions corresponding to the independent unfolding of MBP and RepA domains, and no significant differences were found in the denaturation temperatures of the wild-type and the mutant RepA moieties (data not shown), demonstrating that the thermal stabilities of all RepA variants are comparable.
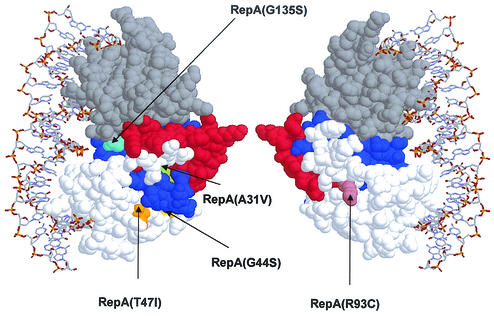
Further structural information can be deduced from the high sequence similarity between RepA and the replication protein RepE from the F plasmid. The three-dimensional structure of the monomeric form of RepE bound to iterons has been recently solved (27). This allows the modeling of RepA with the structure of RepE as a template, by threading procedures with the Swiss-Model utilities included in the Swiss-PDB viewer (23). Figure 3 shows a model of wild-type RepA, and the proposed structural domains and the positions mutated in this work are highlighted. Based in this model, it is noteworthy that all the mutations accumulate in regions far away from the DNA recognition patches and that they are deeply buried in the protein except for Arg93 and, to a much lesser degree, Thr47. Calculated accessibilities for Arg93, Thr47, Gly135, Ala31, and Gly44 are, respectively, 50, 17, 11, 6, and <5%.
FIG. 3.
Two views (differing in 180° rotation) of the modeled three-dimensional structure of pPS10 RepA protein based on the amino acid similarity with the RepE initiation replication protein of F plasmid, showing the mutated positions. The picture shows the N- and C-terminal winged-helix domains in white and gray, respectively, as well as the connecting turn (red) and the LZ motif (blue). DNA is shown only for presentation purposes. Drawing was performed with the RASMOL program (http://www.umass.edu/microbio/rasmol/index2.htm).
Binding of RepA to the iterons of the replication origin.
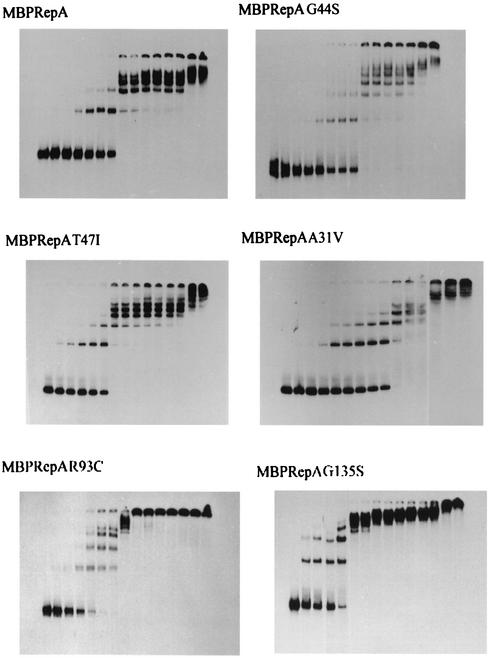
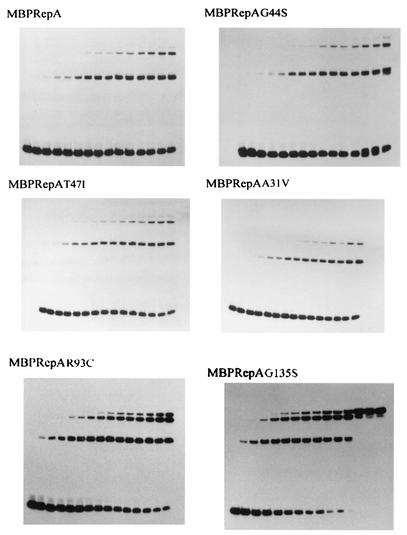
One of the first steps in plasmid replication is the binding of the initiator protein to the origin of replication, and some plasmid Rep mutants with increased initiation activity have been described (26, 49, 50). We have previously shown that wild-type MBPRepA binds to the four iterons present in the origin (17, 19). The efficiency of mutant MBPRepA proteins in binding to pPS10 iterons was analyzed by EMSA experiments with a DNA probe containing the four iterons and an increasing amount of MBPRepA. The results show that the binding of MBPRepA(A31V), MBPRepA(G44S), and MBPRepA(T47I) is very similar to that of MBPRepA, as roughly the same amount of protein is necessary to deplete the free probe (Fig. 4). On the other hand, both MBPRepA(R93C) and MBPRepA(G135S) show an increased efficiency of binding to the same probe and also display an increased cooperativity in binding, as high-molecular-weight complexes (arising from binding to multiple iterons) are formed with much less protein than the wild type or the three other MBPRepA mutants (Fig. 4).
FIG. 4.
Binding of MBPRepA proteins to the oriV region. Increasing quantities of protein (0, 0.43, 0.72, 1.44, 2.88, 4.32, 5.76, 7.20, 8.65, 10.09, 11.53, 12.97, 14.41, 21.61, and 28.82 pmol from left to right) were incubated with 24.5 fmol of the labeled origin probe.
Binding of RepA to the inverted repeats of the repA gene operator.
The RepA-mediated initiation of replication is also dependent on the intracellular levels of the monomeric form. To check whether the highest interaction of MBPRepA(R93C) and MBPRepA(G135S) with the iterons of the origin might reside in an alteration of the dimerization constant Kd of RepA mutants, we performed EMSA experiments with a DNA probe containing the inverted repeats of the repA operator. As the dimeric form is required for RepA binding to this probe, the assay should reveal the possible effect of the mutations on the dimer/monomer ratio.
Similarly to what was observed with the iterons, the strongest DNA binding corresponds to MBPRepA(R93C) and MBPRepA(G135S), whereas the differences between the rest of the mutants and the wild type are hardly noticeable (Fig. 5). This makes unlikely the hypothesis of a better interaction due merely to an alteration in Kd of the RepA mutants. In addition, the result indicates that these mutations affect positively a common element involved in the binding of RepA to both the iterons of the origin and the repA operator region.
FIG. 5.
Binding of MBPRepA proteins to the promoter operator region. Increasing quantities of protein (0, 0.07, 0.14, 0.29, 0.43, 0.57, 0.72, 0.86, 1.01, 1.29, 1.44, 1.80, 2.16, 2.52, and 2.88 pmol from left to right) were incubated with 8.6 fmol of the labeled operator probe.
Binding of RepA to E. coli DnaA protein.
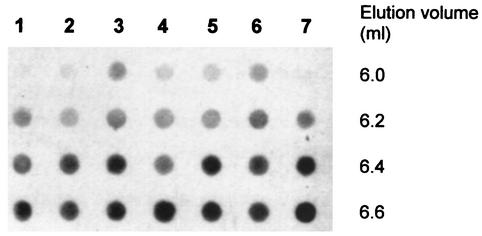
A previous work (30) characterized an E. coli chromosomal mutant (LE403) that allowed the efficient replication of wild-type pPS10. The LE403 strain codes for a mutant DnaA protein (DnaA403) which seems to be the main host protein responsible for the establishment of the plasmid. This DnaA mutant forms a complex with wild-type MBPRepA that can be detected by gel filtration chromatography (30). To check whether the mutant MBPRepA proteins reported in this work were capable of interacting with wild-type DnaA, we performed a similar chromatography experiment, the results of which are shown in Fig. 6. Visible complexes are detected in the exclusion volume of a Sephadex G-75 column for all MBPRepA mutants, but not for the wild type, which displays a similar pattern as that of the control lane (lane 7) without MBPRepA. An accurate affinity quantitation must wait for a complete biophysical study. Even so, the signal corresponding to MBPRepA(G135S) is very faint, and given the peculiar characteristics of the protein and the marginal stability of the pSBM135 mutant (see above), this could reflect the formation of a relatively unstable RepA-DnaA complex compared to the rest of the mutants. In any case, this result suggests that mutations have all strengthened a RepA-DnaA interaction that, if it exists, is very weak in the case of wild-type RepA.
FIG. 6.
Dot blot analysis of the interaction of DnaA with wild-type and mutant MBPRepA fusions by gel-filtration chromatography. The exclusion volume of the column is 6.0 ml. Extracts were obtained from E. coli LE392(pUC392) and CC118(pMALRepA). Lane 1, extracts containing DnaA plus MBPRepA wild type; lane 2, DnaA plus MBPRepA(G135S); lane 3, DnaA plus MBPRepA(G44S); lane 4, DnaA plus MBPRepA(R93C); lane 5, DnaA plus MBPRepA(T47I); lane 6, DnaA plus MBPRepA(A31V); lane 7, DnaA without MBPRepA.
DISCUSSION
In this work we have attempted to gain deeper insights into plasmid host range by searching for mutants of the Pseudomonas plasmid pPS10 that can be established in E. coli at 30 or 37°C. All the mutants tested presented a single mutation in the repA gene, coding for the replication initiator RepA, clearly pointing to this protein as a major host range determinant factor, as previously proposed (14). We have investigated several properties of the mutants, namely, stability, copy number in P. aeruginosa and E. coli, establishment in other species, and structural integrity of RepA proteins, which bind DNA regularly upon mutation compared to wild-type RepA, with the exception of an improved interaction with DnaA that is displayed by all mutants and not by the wild-type protein (Fig. 6). This result, together with the fact that a similar enhanced RepA-DnaA interaction has been recently described for an E. coli mutant that allows the establishment of wild-type pPS10 (30), prompts us to hypothesize that an effective communication between the replication protein RepA and other host factors such as DnaA is a crucial determinant of the host range of the plasmid.
This finding opens new ways to further analyze the DnaA-RepA interactions in the mutants in order to understand the host range phenotype. In addition to improving these interactions, the pBM93 and pBM31 mutants increase the copy number of the replicon, and furthermore, the pMB93 mutant improves interactions of RepA with specific sequences of the basic replicon (both the repA operator and the iterons). The first effect implies that the mutations prevent the possible negative effect of ori-RepA-RepA-ori interactions in initiation of replication, the so-called “handcuffing” mechanism (31). The second effect implies an additional influence on the second winged-helix domain interacting with the specific sequences of the basic replicon (20). It is possible that these effects could also contribute to the host range phenotype. Alternatively, they might constitute only side effects arising from a common mechanism, i.e., the improvement of the interaction between RepA and host factors.
Inspection of the RepA protein at the structural level would be highly valuable to explain how the mutations have altered the interactions with other molecules, although the three-dimensional structure of the protein has unfortunately not yet been solved. According to the mechanism proposed by Giraldo et al. (20), RepA monomers display in solution an elongated conformation with the leucine zipper and the two winged-helix domains loosely packed between them. This conformation allows the two domains of the monomer to interact with the DNA. Although the host range mutations shown here do not seem to induce large structural changes in the secondary structure of the unbound protein (data not shown), they might be of more importance once the complex with DNA or with host factors is formed. In this work we have made a comparative analysis with the three-dimensional structure of the monomeric form of the replication protein RepE from the F plasmid, bound to iterons (27). The high sequence similarity between RepE and RepA allows modeling of the latter with the structure of the former as a template. Figure 3 shows a model of wild-type RepA. The positions mutated are localized far from the DNA binding region and appear, in general, to be buried in the core of the protein, but the effects of mutations may be transmitted to this or other functional parts of the protein. It is possible that substitution of glycines 44 and 135 for serine in the RepA(G44S) and RepA(G135S) proteins modifies the flexibility of these regions since glycine is an amino acid that allows great conformational freedom. The position 135 is especially important since it is located at the end of the flexible linker between the two winged-helix domains, and loss of conformational fluctuations may help to fix the complex of the protein with DNA, causing an enhancement in binding of MBPRepA4(G135S) as shown in Fig. 4 and 5. On the other hand, replacement of the polar amino acids Arg93 and Thr47 by hydrophobic residues (Cys and Ile, respectively) is a more drastic event. Due to their solvent-exposed situation far from the DNA binding site, they may be involved in protein-protein interactions with host factors or with another RepA molecule. It is noteworthy that Arg93 is located in the model very close to Val98, the counterpart of which is Arg118 in RepE, a residue involved in RepE-RepE interactions (27).
An examination of the RepE-based RepA model also suggests interesting functionalities for the leucine zipper region. This sequence is unequivocally involved in RepA dimerization (17). The fact that the A31V is a host range mutation prompted us to speculate about direct interactions between the zipper and the host factors (17, 20). However, as shown in Fig. 3, if RepA behaves like RepE, the leucine zipper might internalize in the protein when bound to iterons, establishing many hydrophobic contacts within the protein core. Since the A31V mutation does not affect Kd significantly (17), it could be possible that this mutation exerts its effect only when the protein forms the initiation complex through inner conformational changes that are transmitted to other parts of the protein, i.e., the ones involved in true RepA-host factor interactions.
Acknowledgments
We thank Walter Messer and Harald Seitz for their kind supplying of anti-DnaA antibodies; Cruz Morenilla for help in Western blotting experiments; and Rafael Giraldo, Marc Lemmonier, and Michel Faelen for critical reading of the manuscript and helpful discussions. We are also grateful to A. Serrano and C. Pardo for excellent technical assistance.
This work was supported by Contrato-Programa de la C.A.M. Grupos Estratégicos 2000-2003, grants BIO98-0106 and QL K2-CT-2000-00634 from the EC, and grants BIO99-0859-CO1 and PM 99-0096 from the Spanish MCYT. B.M. was supported by an FPI predoctoral fellowship from the Spanish Ministerio de Educación y Ciencia and a short-term EMBO fellowship.
REFERENCES
- 1.Bagdasarian, M. M., and K. N. Timmis. 1982. Host-vector systems for gene cloning in Pseudomonas. Curr. Top. Microbiol. Immunol. 96:47-67. [DOI] [PubMed] [Google Scholar]
- 2.Benveniste, R., and J. Davies. 1973. Mechanisms of antibiotic resistance in bacteria. Annu. Rev. Biochem. 42:471-506. [DOI] [PubMed] [Google Scholar]
- 3.Cereghino, J. L., and D. R. Helinski. 1993. Essentiality of the three carboxyl-terminal amino acids of the plasmid RK2 replication initiation protein TrfA for DNA binding and replication activity in gram-negative bacteria. J. Biol. Chem. 268:24926-24932. [PubMed] [Google Scholar]
- 4.del Solar, G., J. C. Alonso, M. Espinosa, and R. Díaz-Orejas. 1996. Broad-host-range plasmid replication: an open question. Mol. Microbiol. 21:661-666. [DOI] [PubMed] [Google Scholar]
- 5.del Solar, G., R. Giraldo, M. J. Ruiz-Echevarria, M. Espinosa, and R. Díaz-Orejas. 1998. Replication and control of circular bacterial plasmids. Microbiol. Mol. Biol. Rev. 62:434-464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Díaz, E., M. Munthali, V. de Lorenzo, and K. N. Timmis. 1994. Universal barrier to lateral spread of specific genes among microorganisms. Mol. Microbiol. 13:855-861. [DOI] [PubMed] [Google Scholar]
- 7.Diaz, R., and W. L. Staudenbauer. 1982. Replication of the broad host range plasmid RSF1010 in cell-free extracts of Escherichia coli and Pseudomonas aeruginosa. Nucleic Acids Res. 10:4678-4702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Doran, K. S., D. R. Helinski, and I. Konieczny. 1999. Host-dependent requirement for specific DnaA boxes for plasmid RK2 replication. Mol. Microbiol. 33:490-498. [DOI] [PubMed] [Google Scholar]
- 9.Doran, K. S., I. Konieczny, and D. R. Helinski. 1998. Replication origin of the broad host range plasmid RK2. Positioning of various motifs is critical for initiation of replication. J. Biol. Chem. 273:8447-8453. [DOI] [PubMed] [Google Scholar]
- 10.Dower, W. I., J. F. Miller, and C. W. Ragsdale. 1988. High efficiency transformation of E. coli by high voltage electroporation. Nucleic Acids Res. 16:6127-6145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Durland, R. H., and D. R. Helinski. 1987. The sequence encoding the 43-kilodalton TrfA protein is required for efficient replication or maintenance of minimal RK2 replicons in Pseudomonas aeruginosa. Plasmid 18:164-169. [DOI] [PubMed] [Google Scholar]
- 12.Fang, F. C., and D. R. Helinski. 1991. Broad-host-range properties of plasmid RK2: importance of overlapping genes encoding the plasmid replication initiation protein TrfA. J. Bacteriol. 173:5861-5868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Fasman, G. D. (ed.). 1976. Handbook of biochemistry and molecular biology. Section A. Proteins, 3rd ed., vol. III. CRC Press, Inc., Cleveland, Ohio.
- 14.Fernández-Tresguerres, E., M. Martín, D. García de Viedma, R. Giraldo, and R. Díaz-Orejas. 1995. Host growth temperature and a conservative amino acid substitution in the replication protein of pPS10 influence plasmid host range. J. Bacteriol. 177:4377-4384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fried, M., and D. M. Crothers. 1981. Equilibria and kinetics of lac repressor-operator interactions by polyacrylamide gel electrophoresis. Nucleic Acids Res. 9:6505-6525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.García de Viedma, D. 1994. Análisis de la proteína RepA del plásmido pPS10: interacciones y motivos implicados en autorregulacion e inicio de replicación. Doctoral thesis. Universidad de Alcalá de Henares, Madrid, Spain.
- 17.García de Viedma, D., R. Giraldo, G. Rivas, E. Fernández-Tresguerres, and R. Díaz-Orejas. 1996. A leucine-zipper motif determines different functions in a DNA replication protein. EMBO J. 15:925-934. [PMC free article] [PubMed] [Google Scholar]
- 18.García de Viedma, D., R. Giraldo, M. J. Ruíz-Echevarría, R. Lurz, and R. Díaz-Orejas. 1995. Transcription of repA, the gene of the initiation protein of the Pseudomonas plasmid pPS10, is autoregulated by interactions of the RepA protein at a symmetrical operator. J. Mol. Biol. 247:211-223. [DOI] [PubMed] [Google Scholar]
- 19.García de Viedma, D., A. Serrano-López, and R. Díaz-Orejas. 1995. Specific binding of the replication protein of plasmid pPS10 to direct and inverted repeats is mediated by an HTH motif. Nucleic Acids Res. 23:5048-5054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Giraldo, R., J. M. Andreu, and R. Díaz-Orejas. 1998. Protein domains and conformational changes in the activation of RepA, a DNA replication initiator. EMBO J. 17:4511-4526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Giraldo, R., M. Martín, E. Fernández-Tresguerres, C. Nieto, and R. Díaz. 1992. Mutations within the minimal replicon of plasmid pPS10 increase its host range, p. 225-237. In P. Hughes, E. Fanning, and M. Kohiyama (ed.), DNA replication: the regulatory mechanisms. Springer-Verlag, Berlin, Germany.
- 22.Giraldo, R., C. Nieto, E. Fernández-Tresguerres, and R. Díaz. 1989. Bacterial zipper. Nature 342:866.. [DOI] [PubMed] [Google Scholar]
- 23.Guex, N., and M. C. Peitsch. 1997. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18:2714-2723. [DOI] [PubMed] [Google Scholar]
- 24.Herrero, H., V. de Lorenzo, and K. N. Timmis. 1990. Transposon vectors containing non-antibiotic resistance selection markers for cloning and stable chromosomal insertion of foreign genes in gram-negative bacteria. J. Bacteriol. 172:6557-6567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Humphreys, G. O., G. A. Willshaw, H. R. Smith, and E. S. Anderson. 1976. Mutagenesis of plasmid DNA with hydroxylamine: isolation of mutants of multi-copy plasmids. Mol. Gen. Genet. 145:101-108. [DOI] [PubMed] [Google Scholar]
- 26.Kawasaki, Y., C. Wada, and T. Yura. 1991. Mini-F plasmid mutants able to replicate in the absence of σ32: mutations in the repE coding region producing hyperactive initiator protein. J. Bacteriol. 173:1064-1072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Komori, H., F. Matsunaga, Y. Higuchi, M. Ishiai, Y. Wada, and K. Miki. 1999. Crystal structure of a prokaryotic replication initiator protein bound to DNA at 2.6 Å resolution. EMBO J. 18:4597-4607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Laemmli, U. K. 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680-685. [DOI] [PubMed] [Google Scholar]
- 29.Lederberg, E. M., and S. N. Cohen. 1974. Transformation of Salmonella typhimurium by plasmid deoxyribonucleic acid. J. Bacteriol. 119:1072-1074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Maestro, B., J. M. Sanz, M. Faelen, M. Couturier, R. Diaz-Orejas, and E. Fernandez-Tresguerres. 2002. Modulation of pPS10 host range by DnaA. Mol. Microbiol. 46:223-234. [DOI] [PubMed] [Google Scholar]
- 31.McEachern, M. J., M. A. Bott, P. A. Tooker, and D. R. Helinski. 1989. Negative control of plasmid R6K replication: possible role of intermolecular coupling of replication origins. Proc. Natl. Acad. Sci. USA 86:7942-7946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Nieto, C. 1988. Desarrollo de vectores de clonaje a partir del plásmido pPS10 de Pseudomonas savastanoi y caracterización de su replicón básico. Doctoral thesis. Universidad Autónoma de Madrid, Madrid, Spain.
- 33.Nieto, C., E. Fernández-Tresguerres, N. Sánchez, M. Vicente, and R. Díaz. 1990. Cloning vectors, derived from a naturally occurring plasmid of Pseudomonas savastanoi, specifically tailored for genetic manipulations in Pseudomonas. Gene 87:145-149. [DOI] [PubMed] [Google Scholar]
- 34.Nieto, C., R. Giraldo, E. Fernández-Tresguerres, and R. Díaz. 1992. Genetic and functional analysis of the basic replicon of pPS10, a plasmid specific for Pseudomonas isolated from Pseudomonas syringae pathovar savastanoi. J. Mol. Biol. 223:415-426. [DOI] [PubMed] [Google Scholar]
- 35.Nordstrom, K., S. Molin, and H. Aagard-Hansen. 1980. Partitioning of plasmid R1 in Escherichia coli. Plasmid 4:215-227. [DOI] [PubMed] [Google Scholar]
- 36.Phillips, J. H., and D. M. Brown. 1967. The mutagenic action of hydroxylamine. Prog. Nucleic Acids Res. Mol. Biol. 7:349-368. [Google Scholar]
- 37.Projan, S. J., S. Carleton, and R. P. Novick. 1983. Determination of plasmid copy number by fluorescence densitometry. Plasmid 9:182-190. [DOI] [PubMed] [Google Scholar]
- 38.Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 39.Scherzinger, E., V. Haring, R. Lurz, and S. Otto. 1991. Plasmid RSF1010 DNA replication in vitro promoted by purified RSF1010 RepA, RepB and RepC proteins. Nucleic Acids Res. 19:1203-1211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Shah, D. S., M. A. Cross, D. Porter, and C. M. Thomas. 1995. Dissection of the core and auxiliary sequences in the vegetative replication origin of promiscuous plasmid RK2. J. Mol. Biol. 254:608-622. [DOI] [PubMed] [Google Scholar]
- 41.Shingler, V., and C. M. Thomas. 1984. Analysis of the trfA region of broad-host-range plasmid RK2 by transposon mutagenesis and identification of polypeptide products. J. Mol. Biol. 175:229-249. [DOI] [PubMed] [Google Scholar]
- 42.Siddique, A., and D. H. Figurski. 2002. The active partition gene incC of IncP plasmids is required for stable maintenance in a broad range of hosts. J. Bacteriol. 184:1788-1793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Smith, A. W., and B. H. Iglewski. 1989. Transformation of Pseudomonas aeruginosa by electroporation. Nucleic Acids Res. 17:10509.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Southern, E. M. 1975. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J. Mol. Biol. 98:503-517. [DOI] [PubMed] [Google Scholar]
- 45.Thomas, C. M. (ed.). 1989. Promiscuous plasmids of gram-negative bacteria. Academic Press, London, United Kingdom.
- 46.Uhlin, B. E., and K. Nordstrom. 1977. R plasmid gene dosage effects in Escherichia coli K-12: copy mutants of the R plasmid R1drd-19. Plasmid 1:1-7. [DOI] [PubMed] [Google Scholar]
- 47.Wechsler, J. A., V. Nusslein, B. Otto, A. Klein, F. Bonhoeffer, R. Hezzman, L. S. Gloger, and H. Schaller. 1973. Isolation and characterization of thermosensitive Escherichia coli mutants defective in deoxyribonucleic acid replication. J. Bacteriol. 113:1381-1388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Woese, C. R. 1987. Bacterial evolution. Microbiol. Rev. 51:221-271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Xia, G., D. Manem, Y. Yu, and L. Caro. 1993. In vivo and in vitro studies of a copy number mutation of the RepA replication protein of plasmid pSC101. J. Bacteriol. 175:4165-4175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.York, D., and M. Filutowicz. 1993. Autoregulation-deficient mutants of the plasmid R6K-encoded Π protein distinguish between palindromic and nonpalindromic binding sites. J. Biol. Chem. 268:21854-21861. [PubMed] [Google Scholar]