Abstract
In gram-negative organisms, high-affinity transport of iron substrates requires energy transduction to specific outer membrane receptors by the TonB-ExbB-ExbD complex. Vibrio cholerae encodes two TonB proteins, one of which, TonB1, recognizes only a subset of V. cholerae TonB-dependent receptors and does not facilitate transport through Escherichia coli receptors. To investigate the receptor specificity exhibited by V. cholerae TonB1, chimeras were created between V. cholerae TonB1 and E. coli TonB. The activities of the chimeric TonB proteins in iron utilization assays demonstrated that the C-terminal one-third of either TonB confers the receptor specificities associated with the full-length TonB. Single-amino-acid substitutions near the C terminus of V. cholerae TonB1 were identified that allowed TonB1 to recognize E. coli receptors and at least one V. cholerae TonB2-dependent receptor. This indicates that the very C-terminal end of V. cholerae TonB1 determines receptor specificity. The regions of the TonB-dependent receptors involved in specificity for a particular TonB protein were investigated in experiments involving domain switching between V. cholerae and E. coli receptors exhibiting different TonB specificities. Switching the conserved TonB box heptapeptides at the N termini of these receptors did not alter their TonB specificities. However, replacing the amino acid immediately preceding the TonB box in E. coli receptors with an aromatic residue allowed these receptors to use V. cholerae TonB1. Further, site-directed mutagenesis of the TonB box −1 residue in a V. cholerae TonB2-dependent receptor demonstrated that a large hydrophobic amino acid in this position promotes recognition of V. cholerae TonB1. These data suggest that the TonB box −1 position controls productive interactions with V. cholerae TonB1.
Iron is one of the most abundant elements in nature; however, acquiring enough iron for growth poses a significant problem for bacterial pathogens. Most iron sources used by gram-negative pathogens are too large (>600 Da) to pass through porins in the outer membrane or too scarce to bioaccumulate in any significant amount by diffusion. Rather, iron complexes are actively transported across the outer membrane via high-affinity outer membrane receptors that are specific for their substrates (8, 19, 56). The energy for this process is provided by the TonB, ExbB, and ExbD proteins, which form a complex in the inner membrane that facilitates energy transduction from the cytoplasmic membrane to the outer membrane receptors (3, 5, 45). By interacting directly with ligand-bound receptors, energized TonB is thought to induce a conformational change in these receptors that promotes internalization of the bound substrate (5, 45, 53).
TonB is composed of three functional domains. The N terminus of TonB is required for anchoring TonB in the cytoplasmic membrane (46) and is involved in the interactions with the accessory proteins ExbB and ExbD (25, 33, 35). From the cytoplasmic membrane, TonB extends into the periplasm (50) and may span the periplasmic space in order to directly contact outer membrane receptors (53). The central proline-rich domain of TonB appears to be critical for the ability of TonB to bridge the periplasmic space by allowing TonB to adopt a rigid extended conformation (34). TonB is predicted to associate with outer membrane receptors via its C-terminal domain. Loss of the C-terminal 48 residues rendered TonB inactive and prevented in vivo cross-linking to FepA, suggesting that residues within this region participate in the interaction with outer membrane receptors (32). Further, a soluble TonB fragment consisting of the C-terminal half of TonB was shown previously to inhibit all TonB-dependent transport when synthesized in vivo, most likely by outcompeting wild-type TonB for binding to outer membrane receptors or by inhibiting the formation of functional TonB dimers (29). The recently solved crystal structure of the C terminus of E. coli TonB showed that the region comprising the final 75 amino acids dimerizes to form a tightly intertwined cylinder-shaped structure (13). This structural feature of TonB may be responsible for productive interactions with outer membrane receptors.
TonB-dependent receptors from many different organisms have been characterized and exhibit significant overall sequence conservation. Based on structural information available for three TonB-dependent receptors from Escherichia coli, FepA (6), FhuA (15, 36), and FecA (14), it is predicted that all TonB-dependent receptors have a similar overall design: a transmembrane β-barrel porin structure with a unique N-terminal domain that fits tightly into the barrel like a plug from the periplasmic side. It is expected that all TonB-dependent receptors share a common binding motif for TonB, and several conserved receptor regions have been identified elsewhere (37), including a short stretch of amino acid homology near the N terminus termed the TonB box (30, 45, 47). The TonB box is located at the periplasmic face of the receptor (6, 14), where it may be accessible to TonB. For E. coli, both genetic and biochemical studies have implicated the TonB box in a direct interaction with TonB. Mutations in the TonB box that abolish transport via the TonB-dependent receptor for vitamin B12, BtuB, can be partially suppressed by mutations at residue Q160 in TonB (2, 21). Cross-linking studies have since indicated that the region of TonB containing the suppressor mutations may come into close contact with the BtuB TonB box (10). These data are consistent with a direct association between TonB and the TonB box; however, since a FhuA variant missing the N-terminal plug domain, and thus the TonB box, required TonB for optimal activity, it seems clear that TonB makes contacts with residues in the transmembrane β-barrel as well (4).
Vibrio cholerae, the causative agent of the diarrheal disease cholera, has a variety of high-affinity iron transport systems. Under iron-limiting conditions, V. cholerae secretes the siderophore vibriobactin (17). Vibriobactin uptake across the outer membrane occurs via the receptor ViuA (7, 54). V. cholerae can also use the E. coli siderophore enterobactin (52, 58). Two enterobactin receptors, IrgA and VctA, have been identified in V. cholerae (40). Other sources of iron used by V. cholerae include ferrichrome, transported via the receptor FhuA (49), and heme, transported via the receptors HutA, HutR, and HasR (22, 39).
Two sets of tonB-exbB-exbD genes have been identified in V. cholerae (24, 44). The two TonB systems have specific as well as overlapping functions (52), indicating that these systems are not completely redundant. While both TonB systems facilitate utilization of vibriobactin, ferrichrome, and heme (44), TonB2 is specifically required for transport of enterobactin through either VctA or IrgA (52) and for heme uptake through HasR (39). Differences between the two V. cholerae TonB systems are also observed when these are expressed in E. coli. The TonB2 system genes can complement an E. coli tonB mutant for transport of enterobactin and ferrichrome, implying a functional interaction between V. cholerae TonB2 and the E. coli receptors FepA and FhuA. However, V. cholerae TonB1 cannot substitute for the native E. coli TonB (22, 44), suggesting that TonB1 does not recognize the E. coli receptors. In this study we provide evidence that the specificity exhibited by TonB1 for a subset of V. cholerae receptors can be attributed to sequence features near the C terminus of TonB1. We also identify mutations near the TonB box of several receptors that alter their TonB specificities.
MATERIALS AND METHODS
Bacterial strains, plasmids, and media.
Bacterial strains (Table 1) were routinely grown at 37°C in Luria (L) broth or on L agar and maintained at −80°C in tryptic soy broth plus 20% glycerol. Iron-replete medium was prepared by the addition of 40 μM ferrous sulfate (FeSO4), and iron-depleted medium was prepared by the addition of ethylenediamine-di-(o-hydroxyphenylacetic acid) (EDDA), deferrated by the method of Rogers (48). Iron-free ferrichrome from Ustilago sphaerogena and bovine hemin were obtained from Sigma Chemical Company (St. Louis, Mo.). Overnight cultures of DH5α and CA401 were used as sources of enterobactin and vibriobactin, respectively. Antibiotics were used at the following concentrations for E. coli: 250 μg of carbenicillin per ml, 50 μg of kanamycin per ml, 30 μg of chloramphenicol per ml, 40 μg of nalidixic acid per ml, and 12.5 μg of tetracycline per ml. For V. cholerae, one-third of these concentrations of carbenicillin and kanamycin and one-fourth of these concentrations of nalidixic acid, chloramphenicol, and tetracycline were used.
TABLE 1.
Bacterial strains and plasmids used in this study
| Strain or plasmid | Description | Source or reference |
|---|---|---|
| Strains | ||
| V. cholerae | ||
| CA401 | V. cholerae classical strain | 16 |
| CA401S | CA401 Strr | 52 |
| CA40130 | Vibriobactin synthesis mutant of CA401 | 17 |
| CA40130N | CA40130 Nalr | 39 |
| DHH11 | CA40130 Nalr; defective in both TonB systems | 44 |
| ARM219 | DHH11 hutA::kan hutR::cam | 39 |
| ARM320 | CA40130N exbB2::cam | This study |
| ARM616 | CA40130N irgA::cam vctA::kan | 40 |
| E. coli | ||
| DH5α | Cloning strain | 20 |
| SM10 λpir | λpir recA Kanr; mobilizing strain for pGP704 derivatives | 41 |
| W3110 | F− IN(rrnD-rrnE)1 | 26 |
| KP1032 | W3110 tonB::kan | 33 |
| KP1032N | KP1032 Nalr | 44 |
| ARM100 | KP1032N entF::cam | This study |
| ARM107 | ARM100 fepAD11Y | This study |
| Plasmids | ||
| pACYC184 | Medium-copy-number cloning vector; Cmr Tcr | 12 |
| pHM5 | Suicide vector pGP704 carrying sacB; Cbr Sucs | 51 |
| pWSK29 | Low-copy-number cloning vector; Cbr | 57 |
| pWKS30 | Low-copy-number cloning vector; Cbr | 57 |
| pWKS30ΔX | pWKS30 with the XbaI site disrupted | Yuki Gleason |
| pAMC22 | pChuA encoding the HutA TonB box in place of the ChuA TonB box | This study |
| pAMC23 | pChuA encoding a T1F substitution in ChuA | This study |
| pAMF19 | pFepA encoding a D11Y substitution in FepA | This study |
| pAMH20 | pACYC184 carrying hutA; Tcr | This study |
| pAMH22 | pAMH20 encoding the ChuA TonB box in place of the HutA TonB box | This study |
| pAMI2 | pCAT121 encoding a D9Y substitution in IrgA | This study |
| pAMI3 | pCAT121 encoding a D9L substitution in IrgA | This study |
| pAMI4 | pCAT121 encoding a D9A substitution in IrgA | This study |
| pAMS7 | pHM5 carrying entF::cam | This study |
| pAMS9 | pHM5 carrying exbB2::cam | This study |
| pAMT12 | pvTonB1X carrying additionally vexbBD1 | This study |
| pAMT111 | pWKS30ΔX carrying etonBX vexbBD1 vhutBCD | This study |
| pAMT112 | pAMT12 carrying additionally vhutBCD | This study |
| pAMT113 | pAMT112 encoding eTonBX1-120/vTonB1X123-245 in place of vTonB1X | This study |
| pAMT114 | pAMT111 encoding vTonB1X1-122/eTonBX121-239 in place of eTonBX | This study |
| pAMT116 | pAMT112 encoding a P238T substitution in TonB1X | This study |
| pAMT117 | pAMT112 encoding a P238R substitution in TonB1X | This study |
| pAMT118 | pAMT112 encoding a P238A substitution in TonB1X | This study |
| pAMT121 | pAMT111 with a XhoI site introduced into etonBX at codon 172 (etonBXXh) | This study |
| pAMT122 | pAMT112 with a XhoI site introduced into vtonB1X at codon 178 (vtonB1XXh) | This study |
| pAMT123 | pAMT122 encoding eTonBXXh1-174/vTonB1XXh181-245 in place of vTonB1XXh | This study |
| pAMT124 | pAMT121 encoding vTonB1XXh1-180/eTonBXXh175-239 in place of eTonBXXh | This study |
| pAMT173 | pAMT112 encoding eTonBX1-158/vTonB1X158-245 in place of vTonB1X | This study |
| pAMT183 | pAMT112 encoding eTonBX1-158/vTonB1X165-245 in place of vTonB1X | This study |
| pCAT121 | pWKS30 carrying V. cholerae irgB irgA | 40 |
| pCHU101 | pACYC184 carrying chuA; Cmr | 55 |
| pChuA | pACYC184 carrying chuA; Tcr | This study |
| pEENTF::Cm | pWSc-1 (E. E. Wyckoff) carrying entF::cam | A. G. Torres |
| peTonBX | pWKS30ΔX carrying E. coli tonB with an XbaI site introduced at codon 119 | This study |
| pMS101 | pBR322 carrying entD fepA fes entF | C. F. Earhart |
| pMTLfepA | pMTL24 (11) carrying fepA | This study |
| pFepA | pACYC184 carrying fepA | This study |
| pOUT11 | pWSK29 carrying V. cholerae exbB2 exbD2 tonB2 | 44 |
| POUT13 | pWSc-1 (E. E. Wyckoff) carrying V. cholerae exbB2::cam | 44 |
| pvTonB1X | pWKS30ΔX carrying V. cholerae tonB1 with an XbaI site introduced at codon 121 | This study |
| pv1/eTonBX | pWKS30ΔX carrying the hybrid vTonB11-122/eTonB121-239 gene | This study |
Utilization of hemin and siderophores.
The ability of V. cholerae and E. coli strains to use various iron sources was tested in halo assays and in liquid culture growth assays as described previously (44, 52). All iron utilization assays were carried out with strains defective in siderophore production in order to reduce background growth. The concentrations of EDDA and of the iron sources used in the assays are indicated in each table or figure legend. All assays were repeated at least three times. In order to standardize assay conditions for all strains, antibiotics were not used in the assays.
PCR.
The oligonucleotide primers for PCR (Table 2) were purchased from IDT Inc. (Coralville, Iowa). PCR was performed with Taq polymerase (Qiagen) or Pfu polymerase (Stratagene) according to the manufacturer's instructions. Unless otherwise indicated, E. coli W3110 or V. cholerae CA401 cultures were used as template. All clones derived from PCR fragments were verified by sequencing.
TABLE 2.
Oligonucleotide primers used in this study
| Name | Primer sequence (5′ to 3′)a |
|---|---|
| chuA1 | AGAGATATCGAAGCTTGCAGAGC |
| chuA2 | CGCGATGATCATAGCCACGC |
| chuA3 | ACTGATGAAGTAGTAGTATCTACAACGGGGAATGCC |
| chuA4 | TGTAGATACTACTACTTCATCAGTAGCAAAAGCAAACG |
| chuA5 | GCTTTTGCTTTCGAAACCATGACCG |
| chuA6 | CATGGTTTCGAAAGCAAAAGCAAACG |
| e1 | AGATCTGCAACGGAAAGTCGACGTCTTTGTTAAGGCC |
| e2 | GTATTTTCAAACGGTGATGCCGGTCTAGACTCTACGG |
| e3 | GTAGAGTCTAGACCGGCATCACCGTTTGAAAATACGG |
| e4 | GTCGGATCCTTTTGACCTGCAGCTTACTGAATTTCGG |
| e5 | GCATTGCGCCTCGAGGGGCAGG |
| e6 | CCTGCCCCTCGAGGCGCAATGC |
| e9 | ACGGCTTAATGCGCGTGG |
| fepA1 | GCGTAATGAGCTCCGTGGAAGCG |
| fepA2 | TCATCGCCTAAACCTTGCG |
| fepA3 | TTCACATTACGATACTATTGTCG |
| fepA4 | ATAGTATCGTAATGTGAAACAGG |
| hutA1 | TCTCTCCCATGGATAGTTCACACCG |
| hutA2 | GCAGGATCCATGTTGAGTAAAACGC |
| hutA3 | ACTTGACTAGTAAATTATTAGC |
| hutA4 | ACTATTGTCGTTACCGCCACTCGCTTGAATACTCAAAT AACTG |
| hutA5 | GGCGGTAACGACAATAGTATCGAATGAGGCATAATCA TCCG |
| hutA6 | GAAACCATGACCGTTACGGCAACTCGCTTGAATACTC |
| hutA7 | TGCCGTAACGGTCATGGTTTCGAATGAGGC |
| irgA4 | TTCAATCGCTTGCAGTGGC |
| irgA5 | CGAAAACGTATGAAACCATGG |
| irgA6 | GTTTCATACGTTTTCGTCGC |
| irgA7 | CGAAAACGTTAGAAACCATGG |
| irgA8 | GTTTCTAACGTTTTCGTCGC |
| irgA9 | CGAAAACGGCTGAAACCATGG |
| irgA10 | GTTTCAGCCGTTTTCGTCGC |
| irgB3 | TTGTTCAATTCTTGCAACGC |
| v1 | TTCTCAAGGCAGTCGACATACAACAAGGGCTGCGC |
| v2 | GCCATTTCAGCGGTTCTAGACTCTACCTTCTCCGG |
| v3 | CCGGAGAAGGTAGAGTCTAGAACCGCTGAAATGGC |
| v4 | GCTGGTGTTGGAGTTGTTGTAACGGATCCATTAGCC |
| v5 | AACTGCAGGAGGGAATAATGGAATCGTTACAACAACT CC |
| v6 | CTTGGATCCATTAAGAGTGAGAAGG |
| v7 | GTCAATGGATCCTTCACCTTCTGCC |
| v8 | GCGGGCTCGAGGGCACTGTG |
| v9 | CACAGTGCCCTCGAGCCCGC |
| v12 | CCCGCTTTAGTGAGCGC |
| v13 | GTTCAACCACGTTATCCACG |
Restriction enzyme sites are underlined.
Construction of plasmids encoding hybrid TonB proteins.
Gene fusions between E. coli tonB and V. cholerae tonB1 were created by engineering unique restriction sites into the cloned E. coli and V. cholerae tonB1 genes by overlap extension PCR (27) and exchanging equivalent tonB fragments. To introduce the XbaI site into E. coli tonB, overlapping fragments containing an XbaI site were created from template pAML14 (28) with primer pairs e1-e2 and e3-e4 and primer pair e1-e4 to amplify the final etonBX product. (In this work, E. coli genes and gene products are preceded by the letter “e” and V. cholerae genes and gene products are preceded by the letter “v.”) An XbaI site was similarly introduced into V. cholerae tonB1 with primers v1-v2 and v3-v4 to create the overlapping fragments from template pHUT10 (23) and primers v1-v4 to amplify the final vtonB1X product. The XbaI mutation inserts a Ser residue in V. cholerae TonB1, but this insertion did not affect function compared with wild-type TonB1 in iron utilization assays (data not shown). The numbering scheme used throughout for V. cholerae TonB1 sequences reflects the Ser insertion. The etonB1X and the vtonB1X PCR products were digested with SalI and BamHI and cloned into pWKS30ΔX digested with SalI and BamHI to create peTonBX and pvTonB1X, respectively. The XbaI-BamHI fragments of peTonBX and pvTonB1X were exchanged to create pe/v1TonBX and pv1/eTonBX.
Plasmids carrying native or hybrid tonBX genes in the context of the entire V. cholerae tonB1 operon were constructed as follows. A fragment containing the 3′ half of the vtonB1X gene and the vexbBD1-hutBCD genes was amplified by PCR with primers v3 and v7. The resulting PCR product was digested with XbaI and BamHI and cloned into pvTonB1X or peTonBX digested with XbaI and BamHI to create pAMT112 and pAMT113, respectively. To create pAMT111 and pAMT114, a fragment containing only the vexbBD1 and hutBCD genes was amplified with primers v5 and v7. The v5 primer contains additionally a Shine-Dalgarno consensus sequence to ensure translation of V. cholerae genes cloned behind E. coli tonB. The PCR product was digested with PstI and BamHI and cloned into peTonBX or pv1/eTonBX digested with PstI and BamHI, respectively. Expression of the V. cholerae genes cloned downstream of E. coli tonB was verified by complementation of V. cholerae exbB1 or exbD1 mutations (data not shown).
To construct plasmids pAMT101 and pAMT102, the SalI-BamHI fragments from pAMT111 and pAMT112, respectively, were cloned into pACYC184 digested with SalI and BamHI. To introduce an XhoI site into etonBX, overlapping PCR products were created with primer sets e1-e6 and e5-v6 with pAMT101 as the template. The final product was amplified with primer set e1-v6. An XhoI site was similarly introduced into vtonB1X with use of primer sets v1-v9 and v8-v6 to create overlapping fragments from the pAMT102 template and primer set v1-v6 to amplify the final product. The final PCR products were digested with SalI and KpnI and cloned into pAMT101 or pAMT102 digested with SalI and KpnI to yield plasmids pAMT201 (etonBXXh) and pAMT202 (vtonB1XXh), respectively. The XhoI mutation resulted in an Ile-to-Leu change in both E. coli TonB (residue 172 in eTonBX) and V. cholerae TonB1 (residue 178 in vTonB1X); however, this change did not affect the function of either TonB in iron utilization assays (data not shown).
To create plasmid series pAMT121 to pAMT124, the SalI-BamHI fragments from pAMT201 and pAMT202 were first cloned into pWKS30ΔX digested with SalI and BamHI to create pAMT121 and pAMT122, respectively. Hybrid tonB constructs were created by exchanging XhoI fragments between pAMT121 and pAMT122 to give rise to pAMT123 and pAMT124.
The hybrid tonB genes contained in pAMT173 and pAMT183 were created by PCR amplification of a portion of the etonBX gene in pAMT111 with primers e1-e9 and amplification of a portion of the vtonB1 gene in pAMT112 with primer pair v12-v6 (pAMT173) or primer pair v13-v6 (pAMT183). The PCR products were 5′ phosphorylated, and the etonBX fragment was ligated to the appropriate vtonB1 fragment. The desired hybrid tonB ligation product was amplified directly from the ligation reaction mixture with primers e1-v6 and then digested with SalI and EcoRI and cloned into pAMT112 digested with SalI and EcoRI to create pAMT173 or pAMT183.
Construction of plasmids encoding outer membrane receptors.
All receptor mutations were created by overlap extension PCR (27). To replace the TonB box of HutA with that of ChuA, overlapping PCR fragments were created with primer sets P1-hutA7 and hutA6-hutA3 (ChuA TonB box) with pAMH20 as template. The overlap extension products were amplified with primer pair P1-hutA3 and were then digested with Bsu36I and SpeI and cloned into pAMH20 digested with Bsu36I and SpeI to create pAMH22.
To construct pChuA, the ∼2.7-kb EcoRV fragment from pCHU101 was cloned into pACYC184 digested with ScaI. To introduce the HutA TonB box or the T1F mutation into ChuA, primer sets chuA1-chuA4 and chuA3-chuA2 (HutA TonB box) or chuA1-chuA6 and chuA5-chuA2 (T1F) were used to amplify overlapping fragments from the pCHU101 template. The overlap extension products were amplified with primers chuA1-chuA2, digested with HpaI and KpnI, and ligated into pChuA digested with HpaI and KpnI to create pAMC22 and pAMC23, respectively.
To construct pFepA, the SspI-StuI fragment containing fepA from pMS101 was cloned into pACYC184 cut with ScaI and PvuII. The FepAD11Y mutation was introduced into FepA with the use of primer sets fepA1-fepA4 and fepA3-fepA2 to generate overlapping fragments from the pFepA template and primers fepA1-fepA2 to amplify the overlap extension product. The final PCR product was digested with Bsu36I and EcoRI and cloned into pFepA digested with Bsu36I and EcoRI to create pAMF19.
Mutations in IrgA were introduced by using primer sets irgB3-irgA6 and irgA5-irgA4 (D9Y), irgB3-irgA8 and irgA7-irgA4 (D9L), or irgB3-irgA10 and irgA9-irgA4 (D9A) to create overlapping fragments. The overlap extension products were amplified with primers irgB3 and irgA4, digested with MscI and BstEII, and cloned into pCAT121 digested with MscI and BstEII to create plasmids pAMI2, pAMI3, and pAMI4, respectively.
Construction of chromosomal mutations in E. coli and V. cholerae genes.
To construct allelic exchange plasmids pAMS7 and pAMS9, the SalI-EcoRV fragments containing the disrupted genes were excised from pEENTF::Cm and pOUT13, respectively, and cloned into the λpir-dependent suicide vector pHM5 digested with SalI and EcoRV. Allelic exchange was carried out as previously described (39).
Sequence analysis.
DNA sequencing was performed by the University of Texas Institute for Cellular and Molecular Biology DNA Core Facility using an ABI Prism 377 DNA sequencer. Analysis of DNA sequences was carried out with MacVector 7.0. Amino acid sequence alignments were carried out with the CLUSTAL-W program from within MacVector 7.0.
SDS-PAGE and immunoblotting.
Cells were grown to an optical density at 650 nm of 0.5 in L broth containing the appropriate antibiotics, and 10 μg of EDDA per ml was added to induce expression of iron-regulated genes. After further incubation for 1 h, samples containing an equivalent number of cells were treated with trichloroacetic acid (10% final concentration), washed with acetone, and resuspended in Laemmli solubilization buffer (31). The proteins were resolved by sodium dodecyl sulfate (SDS)-7.5% polyacrylamide gel electrophoresis (PAGE) and visualized by Coomassie brilliant blue staining or electroblotted for 1 h at 300 mA onto Hybond ECL nitrocellulose (Amersham Pharmacia Biotech, Little Chalfont, Buckinghamshire, England). Immunodetection of IrgA was carried out with rabbit polyclonal anti-IrgA antiserum (diluted 1:1,000; generous gift of R. K. Holmes) and horseradish peroxidase-conjugated goat anti-rabbit immunoglobulin G (Bio-Rad Laboratories, Hercules, Calif.) secondary antibody (diluted 1:10,000).
RESULTS
Construction and testing of hybrid tonB genes.
To investigate the basis for the narrow receptor specificity exhibited by V. cholerae TonB1 and to define the region(s) of the TonB proteins involved in the TonB-receptor interaction, a series of TonB chimeras between E. coli TonB and V. cholerae TonB1 was constructed.
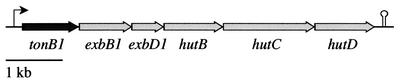
The hybrid tonB genes were created by exchanging equivalent E. coli tonB and V. cholerae tonB1 fragments by using unique restriction sites engineered into the genes by PCR. These restriction sites did not appear to alter the activity or the specificity of the TonB proteins compared with the corresponding wild-type TonB proteins in iron utilization assays (data not shown), and thus the nomenclature used for the TonB protein products has not been changed to reflect the small changes in the amino acid sequence due to the engineered restriction sites (see Materials and Methods). Further, no significant differences in TonB-dependent growth were observed between strains expressing TonB from a low-copy-number plasmid and strains expressing it from the chromosome (data not shown). The V. cholerae TonB1 N terminus does not recognize the native E. coli ExbB and ExbD proteins (reference 44 and data not shown), and it was necessary to include V. cholerae exbB1 and exbD1 in all constructs encoding the N-terminal domain of V. cholerae TonB1. In addition, heme uptake via HutA in E. coli was more efficient when the V. cholerae heme transport genes hutB, hutC, and hutD were expressed also (data not shown). Therefore, to maintain consistency, all native and chimeric tonB genes were inserted in place of V. cholerae tonB1 in the context of the entire V. cholerae tonB1-exbB1-exbD1-hutB-hutC-hutD operon (Fig. 1).
FIG. 1.
Organization of the tonB1 locus in V. cholerae.
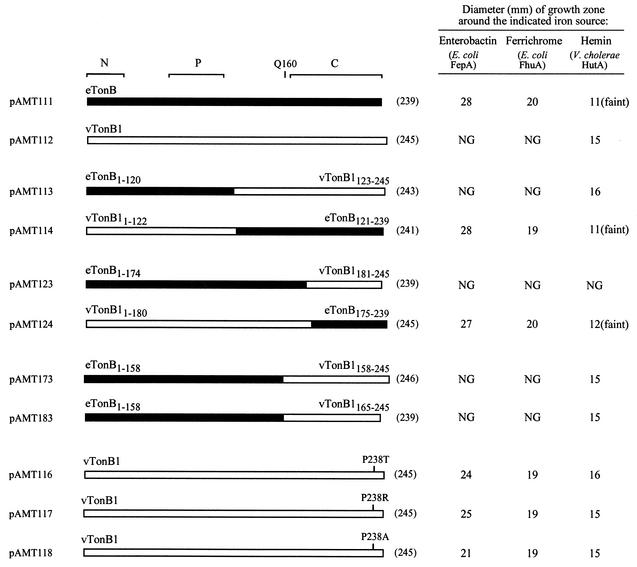
To determine the specificity of the native and hybrid TonB proteins in E. coli, the tonB constructs were expressed in the E. coli tonB entF mutant ARM100 carrying a plasmid, pAMH20, encoding the V. cholerae heme receptor hutA. Heme utilization in ARM100 was not observed in the absence of pAMH20 (data not shown). The tonB constructs were tested for the ability to facilitate heme utilization through V. cholerae HutA and enterobactin or ferrichrome utilization via the E. coli receptor FepA or FhuA, respectively. E. coli TonB, encoded by pAMT111, permitted uptake of enterobactin and ferrichrome but did not mediate efficient heme uptake, suggesting that E. coli TonB interacts poorly with HutA (Fig. 2). In contrast, V. cholerae TonB1, encoded by pAMT112, promoted heme uptake via HutA but was unable to facilitate transport through any of the E. coli receptors tested, demonstrating that V. cholerae TonB1 has an absolute specificity for V. cholerae receptors (Fig. 2). Similar results were obtained for E. coli TonB and V. cholerae TonB1 encoded by pAMT121 and pAMT122, respectively (data not shown). Thus, both E. coli TonB and V. cholerae TonB1 exhibit a degree of specificity for receptors of their own species, consistent with previous results (22, 44).
FIG. 2.
The specificity of TonB for outer membrane receptors is determined by the C terminus of TonB. E. coli TonB (eTonB) is represented by a solid bar; V. cholerae TonB1 (vTonB1) is represented by an open bar. The brackets above the eTonB bar delineate the three functional domains of E. coli TonB: the N-terminal domain (N), the central proline-rich region (P), and the C-terminal dimerization domain (C). Specific residues are shown above the relevant bar in the one-letter amino acid code. The length (in amino acid residues) of each protein is indicated in parentheses on the right. The receptor involved in the utilization of each iron source tested is listed in parentheses below the iron source. Heme utilization was tested in strains supplied with V. cholerae hutA on plasmid pAMH20. The plasmid encoding each TonB protein tested in the assay is listed on the left. E. coli ARM100 or ARM100/pAMH20 carrying the indicated tonB plasmid was seeded at 106 cells per ml into molten L agar containing 100 μg of EDDA per ml. Iron sources were spotted onto the solidified agar as follows: 20 μl of 10 mM FeSO4, 5 μl of 50 μM ferrichrome, 5 μl of DH5α overnight culture (enterobactin), and 5 μl of 50 μM hemin. Plates were incubated for 24 h at 37°C, and the diameter of each zone of growth was measured. Comparable growth around FeSO4 was observed for all strains (data not shown). NG, no growth. “Faint” indicates that this growth zone was less dense than the zones produced by other strains around the specified iron source. The assays were repeated at least three times. Although the relative differences between strains were consistent between experiments, the absolute values for each strain varied due to subtle changes in the medium and assay conditions, and thus only the results of one representative experiment are shown.
Replacing the carboxy-terminal 119 amino acids (121 to 239) of E. coli TonB with the equivalent region from V. cholerae TonB1 produced a functional TonB (encoded by pAMT113) that allowed transport through V. cholerae HutA but not through the E. coli receptors (Fig. 2). Conversely, replacing the carboxy-terminal 123 residues (123 to 245) of V. cholerae TonB1 with the equivalent region from E. coli TonB produced a functional TonB (encoded by pAMT114) that permitted transport through E. coli FepA and FhuA but did not mediate efficient heme uptake through V. cholerae HutA (Fig. 2). These data suggest that a region within the carboxy-terminal half of TonB is involved in determining the specificity of the interaction with outer membrane receptors.
Another set of TonB chimeras was created in which only the C-terminal 65 amino acids from each TonB protein were switched. This domain swap did not include the conserved Gln160 residue (Fig. 2) previously implicated in the interaction between E. coli TonB and the TonB box of the BtuB receptor in E. coli (2, 10). pAMT124, encoding a chimera composed of V. cholerae TonB1 residues 1 to 180 joined to E. coli TonB residues 175 to 239, conferred the ability to use enterobactin and ferrichrome but did not allow efficient heme utilization (Fig. 2). Thus, this chimera exhibited the specificities associated with E. coli TonB, indicating that the C-terminal 65 amino acids of E. coli TonB are sufficient for recognizing E. coli receptors. Chimeras containing fewer than 65 C-terminal E. coli TonB residues (23, 31, or 48 C-terminal residues) were nonfunctional (data not shown), suggesting that this region constitutes an important functional domain of TonB that may rely on specific intramolecular interactions to form properly.
The chimera encoded by pAMT123, consisting of the N-terminal 174 residues of E. coli TonB joined to the C-terminal 65 residues (181 to 245) from V. cholerae TonB1, was inactive in iron utilization assays (Fig. 2), suggesting that a domain(s) necessary for function was disrupted in this hybrid. Therefore, to explore further the minimal region of V. cholerae TonB1 necessary to confer the specificities associated with the TonB1 protein, hybrids were created between E. coli TonB and V. cholerae TonB1 that included more than 65 and progressively fewer than 123 TonB1 C-terminal residues. The chimera encoded by pAMT173, containing the N-terminal 158 residues of E. coli TonB joined to the C-terminal 88 residues (158 to 245) of V. cholerae TonB1, allowed hemin utilization via V. cholerae HutA but did not permit enterobactin utilization via E. coli FepA or ferrichrome utilization via E. coli FhuA, indicating that this chimera is specific for V. cholerae receptors (Fig. 2). Thus, this chimera contains the region of V. cholerae TonB1 involved in determining specificity.
The portion of TonB1 encoded by pAMT173 includes a short stretch of amino acids (residues 157 to 163) that does not align well with E. coli TonB or with V. cholerae TonB2 (Fig. 3A). To determine if specificity determinants are contained within this region of TonB1, a chimera lacking these residues was constructed by joining the C-terminal 81 amino acids (residues 165 to 245) of TonB1 to the N-terminal 158 amino acids of E. coli TonB. This chimera, encoded by pAMT183, exhibited all the expected TonB1 specificities and had wild type levels of TonB1 activity (Fig. 2), indicating that the missing residues were not required for TonB1 activity or specificity. These data demonstrate that the C-terminal 81 amino acids of TonB1 are sufficient to confer the specificities exhibited by V. cholerae TonB1.
FIG. 3.
(A) Amino acid sequence alignment of the C-terminal region from V. cholerae TonB1 (Vc TonB1) with the V. cholerae TonB2 (Vc TonB2) and E. coli TonB (Ec TonB) C termini. Identical residues are indicated by an asterisk, and conserved residues are indicated by a dot below the alignment. The numbers flanking the sequences refer to the first and last amino acids in the sequence. The numbering used for the Vc TonB1 sequence is based on the tonB1X sequence, which encodes an extra amino acid compared with the native V. cholerae TonB1 protein (see Materials and Methods). This insertion does not affect the alignment of the sequences shown. The V. cholerae TonB1 region included in the chimera encoded by pAMT173 but not in the chimera encoded by pAMT183 is underlined. The arrowhead marks the site of the C-terminal domain switch in the chimeras encoded by pAMT123 and pAMT124. The V. cholerae TonB1 Pro238 residue is shown in boldface. (B) Comparison of the predicted TonB box regions from E. coli and V. cholerae receptors. The TonB box heptapeptide is underlined. The TonB box −1 residue is shown in boldface.
Isolation of V. cholerae TonB1 point mutants capable of interacting with E. coli receptors.
To identify specific residues important for determining the narrow receptor specificity of V. cholerae TonB1, a selection scheme was devised to isolate mutations in the tonB1 gene that would permit productive interactions between V. cholerae TonB1 and E. coli receptors. E. coli ARM100 (EntF− TonB−) carrying a plasmid encoding the V. cholerae TonB1 complex was seeded into iron-limited medium containing enterobactin or ferrichrome as iron source, as described in the Fig. 2 legend. As expected, no halo of growth was observed around either of these iron substrates after 24 h; however, upon further incubation, a small number of colonies became detectable in the agar surrounding the iron sources. These colonies were further screened to identify those that could use both enterobactin and ferrichrome, suggesting that these strains expressed a V. cholerae TonB1 variant capable of interacting with multiple E. coli receptors. To verify that the mutation(s) was in V. cholerae tonB1, the plasmid encoding TonB1 was isolated and transformed into a fresh background, and the ability of the newly transformed ARM100 strain to use enterobactin and ferrichrome was confirmed (Fig. 2). Sequencing of the tonB1 gene on the plasmid (designated pAMT116) showed that it carried a mutation resulting in a Pro-to-Thr substitution at residue 238 in the TonB1 protein. Another plasmid, pAMT117, was isolated that encoded a TonB1 protein with a Pro-to-Arg substitution at position 238. The P238T and P238R mutations were both obtained several times in independent experiments (data not shown). No other tonB1 mutations were isolated through this selection scheme. P238 is very close to the C terminus of TonB1 (Fig. 3A). This suggests that either the extreme C terminus of TonB1 interacts with the receptor or mutations in this region influence the conformation of a domain needed for that interaction.
To establish whether the narrow receptor specificity of V. cholerae TonB1 is due to the absence of an appropriate residue (Thr or Arg) or the presence of an inappropriate residue (Pro) at the site defined above, a P238A mutation was created in TonB1. The Ala substitution resulted in the ability of TonB1 to facilitate both enterobactin and ferrichrome uptake in ARM100 (Fig. 2), suggesting that the presence of a Pro residue in this position in V. cholerae TonB1 restricts its interactions with E. coli receptors.
Functional analysis of V. cholerae and E. coli heme receptor chimeras involving the N-terminal TonB box domain.
A comparison of V. cholerae and E. coli TonB boxes reveals that the predicted V. cholerae HutA TonB box does not closely match those found in E. coli receptors (Fig. 3B). Thus, the specificity exhibited by V. cholerae TonB1 and E. coli TonB for certain receptors may be due in part to a requirement for a particular TonB box sequence (22). To investigate this hypothesis, chimeras were constructed between E. coli and V. cholerae receptors in which the TonB box heptapeptides were switched. The receptor chimeras were then tested for the ability to function with either V. cholerae TonB1 or E. coli TonB in iron utilization assays in E. coli ARM100.
The TonB box of V. cholerae HutA was replaced with the TonB box from the E. coli O157:H7 heme receptor ChuA. E. coli TonB-mediated hemin utilization was more efficient through this chimera than through wild-type HutA, as evidenced by a denser growth zone around hemin than previously observed (Fig. 4). However, V. cholerae TonB1 appeared to promote hemin utilization as efficiently through HutA as through HutA carrying the ChuA TonB box (Fig. 4). These data suggest that residues elsewhere in the HutA receptor contribute to the specificity of the interaction with V. cholerae TonB1.
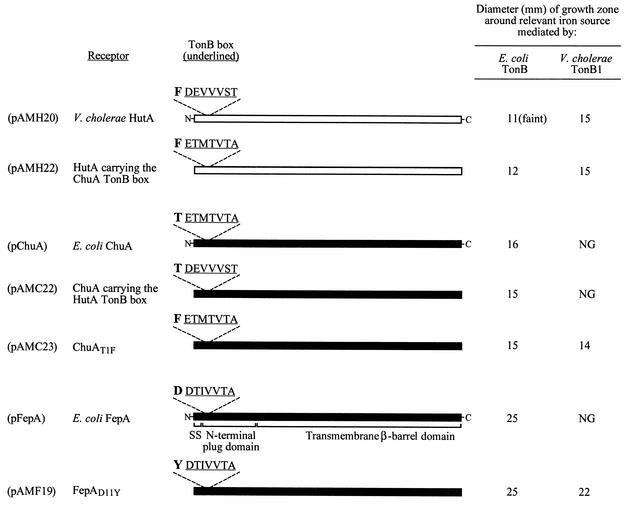
FIG. 4.
Analysis of receptor residues necessary for productive interactions with E. coli TonB or V. cholerae TonB1. Solid bars represent E. coli receptors; open bars represent V. cholerae receptors. The domain organization shown for FepA (signal sequence [SS], N-terminal plug domain, and β-barrel domain) is expected to be relevant for all the receptors shown. The TonB box sequence (underlined), as well as the residue immediately preceding the TonB box, is shown above each bar. The plasmid encoding each receptor protein is listed on the left. The numbering scheme used to designate the TonB box −1 position in FepA and ChuA reflects the amino acid position in the mature (cleaved) protein. E. coli ARM100 carrying the indicated receptor plasmid, and a plasmid encoding either E. coli TonB or V. cholerae TonB1, was tested as described in the legend to Fig. 2. The function of the HutA or ChuA receptors was tested with hemin as the iron source. The function of the FepA receptors was tested with enterobactin as the iron source. Comparable growth around FeSO4 was observed for all strains (data not shown). NG, no growth. “Faint” indicates that this growth zone was less dense than the zones produced by other strains around the specified iron source. The assays were repeated at least three times. Although the relative differences between strains were consistent between experiments, the absolute values for each strain varied due to subtle changes in the medium and assay conditions, and thus only results from one representative experiment are shown.
A reciprocal chimera was constructed in which the TonB box of E. coli ChuA was replaced with the TonB box from V. cholerae HutA. Figure 4 shows that the phenotype of this chimera is identical to that of the wild-type ChuA protein, which recognizes only E. coli TonB. This suggests that residues apart from, or in addition to, the HutA TonB box heptapeptide are required for the productive interactions with V. cholerae TonB1. Because the substitution did not eliminate productive interactions with E. coli TonB, the data further imply that the ChuA TonB box does not contain residues required for the specific recognition of E. coli TonB.
Isolation of an E. coli FepA mutant capable of interacting with V. cholerae TonB1.
To identify receptor residues involved in specific recognition of V. cholerae TonB1, mutants in E. coli FepA were selected that could function with V. cholerae TonB1. E. coli ARM100 carrying a plasmid encoding the V. cholerae TonB1 complex was seeded into iron-limited medium containing enterobactin as the sole source of iron, as described in the Fig. 2 legend. A small number of colonies were isolated and screened for those that could use enterobactin but not ferrichrome. A candidate fepA mutant strain, ARM107, was screened for loss of the plasmid encoding TonB1 and then retransformed with a plasmid encoding either V. cholerae TonB1 or E. coli TonB. Freshly transformed ARM107 used enterobactin in the presence of either V. cholerae TonB1 or E. coli TonB, indicating that the FepA mutation allowed for productive interactions with V. cholerae TonB1 without affecting the ability of FepA to interact with E. coli TonB (Table 3 and Fig. 4). In addition, the TonB chimera carrying just 81 C-terminal residues from V. cholerae TonB1 (encoded by pAMT183) promoted enterobactin utilization in ARM107 (data not shown) but not in ARM100 (Fig. 2), showing that, like TonB1, this chimera discriminated between wild-type FepA and the FepA variant expressed by ARM107. Sequencing of the fepA allele from ARM107 showed that it contained a mutation resulting in a D11Y substitution in the mature FepA protein. To verify that the D11Y mutation was responsible for the productive interaction of FepA with V. cholerae TonB1, the mutation was reconstructed within a wild-type fepA gene on a plasmid. The fepAD11Y plasmid pAMF19 conferred enterobactin utilization in ARM100 carrying the tonB1 plasmid pAMT112, whereas the wild-type fepA plasmid, pFepA, did not (Table 3 and Fig. 4). These data confirm the gain-of function phenotype of the FepAD11Y point mutant. Interestingly, the substituted residue in FepAD11Y immediately precedes the TonB box in the FepA amino acid sequence, strongly implicating this N-terminal receptor region in the interaction with TonB (Fig. 4).
TABLE 3.
TonB dependence of FepA and FepAD11Y in E. coli
| Strain | Relevant phenotypeb | Zone of growth (mm) after 24 ha
|
||
|---|---|---|---|---|
| FeSO4 | Enterobactin | Ferrichrome | ||
| ARM100/pWKS30ΔX | TonB− FepA+ | 20 | NG | NG |
| ARM100/pAMT111 | eTonB+ FepA+ | 22 | 25 | 15 |
| ARM100/pAMT112 | vTonB1+ FepA+ | 20 | NG | NG |
| ARM107/pAMT111 | eTonB+ FepAD11Y+ | 20 | 25 | 15 |
| ARM107/pAMT112 | vTonB1+ FepAD11Y+ | 21 | 22 | NG |
| ARM100/pAMT112/pFepA | vTonB1+ FepA+ | 23 | NG | NG |
| ARM100/pAMT112/pAMF19 | vTonB1+ FepA+ FepAD11Y+ | 22 | 23 | NG |
The indicated E. coli strains were seeded at 106 cells per ml into molten L agar containing 100 μg of EDDA per ml. Iron sources were spotted onto the solidified agar as follows: 20 μl of 10 mM FeSO4; 5 μl of 40 μM ferrichrome; and 5 μl of DH5α overnight culture (enterobactin). NG, no growth. Plates were incubated for 24 h at 37°C, and the diameter of each zone of growth was measured. The assays were repeated at least three times. Although the relative differences between strains were consistent between experiments, the absolute values for each strain varied due to subtle changes in the medium and assay conditions, and thus results of only one representative experiment are shown.
E. coli TonB is designated by eTonB; V. cholerae TonB1 is designated by vTonB1.
Analysis of an E. coli ChuA mutant capable of interacting with V. cholerae TonB1.
To investigate the relevance of the TonB box −1 position for TonB specificity, this region of the E. coli and V. cholerae receptors was compared. V. cholerae HutA, HutR, and FhuA, all of which use V. cholerae TonB1, contain a large hydrophobic residue immediately prior to the TonB box (Fig. 3B). In contrast, the V. cholerae receptors that fail to use TonB1, including HasR, IrgA, and VctA, all have an acidic residue in this position. Further, none of the E. coli receptors contains a large hydrophobic residue in the TonB box −1 position, and these receptors are not recognized by V. cholerae TonB1. To test the hypothesis that a large hydrophobic amino acid at the position immediately preceding the TonB box facilitates interactions with V. cholerae TonB1, a Phe residue was introduced into the E. coli O157:H7 heme receptor, ChuA, at this position. The ChuAT1F point mutant was tested for transport activity with E. coli TonB or V. cholerae TonB1. Figure 4 shows that, whereas V. cholerae TonB1 failed to promote heme transport through wild-type ChuA, TonB1 facilitated heme utilization through the ChuAT1F variant. This suggests that receptor recognition by V. cholerae TonB1 is strongly influenced by the residue immediately preceding the TonB box. As was observed for the D11Y mutation in FepA, the T1F mutation in ChuA did not abolish interactions with E. coli TonB; heme was used efficiently via both wild-type ChuA and ChuAT1F in the presence of E. coli TonB (Fig. 4 and data not shown). Thus, E. coli TonB appears to tolerate a wider range of residues in the TonB box −1 position.
Analysis of TonB1 specificity in V. cholerae.
It was important to establish whether the specificity exhibited by V. cholerae TonB1 in E. coli is relevant to the receptor bias displayed by TonB1 in V. cholerae. To answer this question, the V. cholerae TonB1P238T, TonB1P238R, and TonB1P238A variants, which recognize E. coli receptors, were tested for the ability to facilitate enterobactin transport in V. cholerae, a process that normally requires the TonB2 complex (52). In addition to carrying out all the wild-type TonB1 functions, such as facilitating use of ferrichrome, hemin, and vibriobactin, all three TonB1 Pro238 mutants conferred enterobactin utilization in the tonB mutant strain DHH11, presumably by interacting with one or both of the V. cholerae enterobactin receptors IrgA and VctA (Table 4). These data demonstrate that the C-terminal P238 substitutions effectively widen the range of receptors that TonB1 can interact with in V. cholerae. The data further imply that the molecular basis for receptor discrimination by TonB1 is similar in V. cholerae and in E. coli. Interestingly, none of the TonB1 P238 mutants mediated heme utilization in the hutA hutR mutant ARM219, suggesting that the heme receptor HasR specifically requires features of TonB2 that are not supplied by TonB1 or any of the TonB1 P238 mutants (Table 4).
TABLE 4.
Function of V. cholerae TonB1 deletion and point mutants in V. cholerae
| Strain and plasmid | TonB phenotype | Zone of growth (mm) after 24 ha
|
||||
|---|---|---|---|---|---|---|
| FeSO4 | Ferrichrome | Hemin | Vibriobactin | Enterobactin | ||
| DHH11 (Vib− TonB−)b | ||||||
| pWKS30 | No TonB | 21 | NG | NG | NG | NG |
| pAMT112 | TonB1 | 20 | 13 | 15 | 20 | NG |
| pOUT11 | TonB2 | 21 | 12 | 11 | 21 | 25 |
| pAMT112Δ | TonB1Δ | 21 | 13 | 15 | 20 | NG |
| pAMT116 | TonB1P238T | 20 | 13 | 14 | 21 | 20 |
| pAMT117 | TonB1P238R | 22 | 14 | 15 | 21 | 20 |
| pAMT118 | TonB1P238A | 22 | 13 | 15 | 20 | 18 |
| ARM219 (DHH11 HutA− HutR− HasR+) | ||||||
| pAMT112 | TonB1 | 22 | 13 | NG | 21 | NG |
| pOUT11 | TonB2 | 20 | 12 | 7 (faint)c | 21 | 25 |
| pAMT112Δ | TonB1Δ | 22 | 12 | NG | 22 | NG |
| pAMT116 | TonB1P238T | 22 | 12 | NG | 21 | 20 |
| pAMT117 | TonB1P238R | 22 | 13 | NG | 21 | 20 |
| pAMT118 | TonB1P238A | 21 | 13 | NG | 20 | 18 |
The V. cholerae strains were tested as described for Table 3. Five microliters of CA401 overnight culture was spotted as the source of vibriobactin. NG, no growth.
Vib− indicates that vibriobactin is not produced by this strain.
“Faint” indicates that this growth zone was less dense than the zones produced by other strains around this iron source.
V. cholerae TonB1 is longer than TonB2 by 38 amino acids, raising the possibility that the failure of TonB1 to recognize certain receptors in V. cholerae is related to its length, rather than to a lack of specific residues required for the productive interaction with these receptors. To test this hypothesis, a TonB1 mutant containing a 35-amino-acid deletion spanning the proline-rich region (52) was tested for the ability to mediate transport of various iron sources in V. cholerae. Strains expressing TonB1Δ used vibriobactin, ferrichrome, and hemin as sources of iron, indicating that the deletion did not adversely affect TonB1 function under the conditions tested. However, the TonB1Δ mutant, although comparable in length to TonB2, did not facilitate enterobactin utilization via IrgA or VctA or hemin utilization via HasR (Table 4). Thus, the inability of TonB1 to recognize these receptors in V. cholerae is probably not due solely to its being longer than TonB2.
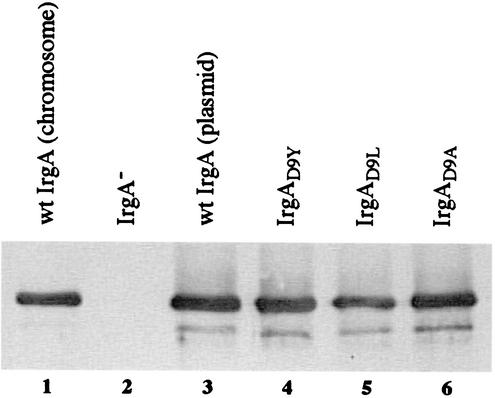
To verify that the TonB box −1 position is involved in determining TonB specificity in V. cholerae as well as in E. coli, changes at this position were created in the V. cholerae enterobactin receptor IrgA, which normally does not recognize TonB1. To analyze TonB1-dependent enterobactin utilization, plasmids encoding the IrgA point mutants were tested in ARM320 (Vib− TonB2−). Replacing D9 in IrgA with either Tyr or Leu resulted in IrgA variants that facilitated the use of enterobactin as an iron source in the presence of TonB1, albeit somewhat less efficiently than in the presence of TonB2 (Table 5). Replacing D9 with an Ala residue did not allow for TonB1-mediated enterobactin utilization (Table 5), indicating that TonB1 specificity is not determined exclusively by a restriction against the acidic residue found in this position in the TonB2-dependent V. cholerae receptors and in many E. coli receptors (Fig. 3B). These data support the results obtained with E. coli that the TonB box −1 receptor residue is important for determining productive interactions with V. cholerae TonB1. To ensure that the observed specificities were not due to aberrant expression of the plasmid-encoded IrgA mutants, Western immunoblotting was performed on V. cholerae ARM616 (IrgA− VctA−) carrying each of the irgA plasmids described in Table 5. As shown in Fig. 5, the three IrgA point mutants were expressed at levels comparable to that of plasmid-encoded wild-type IrgA. Importantly, the inability of the D9A mutant to facilitate TonB1-mediated enterobactin transport is not due to poor expression of the mutant protein. In addition, the D9A mutant restored enterobactin utilization in the enterobactin receptor mutant ARM616 (Table 5), confirming that IrgAD9A is a functional receptor and showing that the D9A mutation does not eliminate interactions with TonB2 (Table 5). This is likely the case for IrgAD9Y and IrgAD9L as well, since the level of enterobactin utilization via these receptors in ARM616 approached levels seen only for TonB2-expressing strains (Table 5). Taken together, these data demonstrate that, in contrast to TonB2, TonB1 may require a large hydrophobic residue immediately preceding the TonB box.
TABLE 5.
TonB dependence of IrgA TonB box −1 mutants in V. cholerae
| Strain and/or plasmid | IrgA phenotype | Zone of growth (mm) after 48 ha
|
||
|---|---|---|---|---|
| FeSO4 | Enterobactin | Vibriobactin | ||
| CA40130N (Vib−b) | IrgA+ | 29 | 27 | 24 |
| ARM320 (Vib− TonB2−) | ||||
| pWKS30ΔX | IrgA+ | 30 | NG | 23 |
| pCAT121 | IrgA+ | 30 | NG | 24 |
| pAMI2 | IrgA+; IrgAD9Y+ | 29 | 20 | 23 |
| pAMI3 | IrgA+; IrgAD9L+ | 29 | 25 | 23 |
| pAMI4 | IrgA+; IrgAD9A+ | 30 | NG | 24 |
| ARM616 (Vib− IrgA− VctA−) | ||||
| pWKS30ΔX | IrgA− | 30 | NG | 21 |
| pCAT121 | IrgA+ | 30 | 28 | 23 |
| pAMI2 | IrgAD9Y+ | 31 | 27 | 23 |
| pAMI3 | IrgAD9L+ | 30 | 28 | 24 |
| pAMI4 | IrgAD9A+ | 29 | 28 | 23 |
FIG. 5.
Expression of wild-type (wt) IrgA and IrgA point mutants in V. cholerae. Samples representing 5 × 106 cells were prepared as described in Materials and Methods and resolved by SDS-7.5% PAGE. Immunoblotting was performed with polyclonal anti-IrgA antiserum. Lane 1, CA40130N; lane 2, ARM616/pWKS30; lane 3, ARM616/pCAT121; lane 4, ARM616/pAMI2; lane 5, ARM616/pAMI3; lane 6, ARM616/pAMI4.
DISCUSSION
Despite the differences in receptor specificity between V. cholerae TonB1 and E. coli TonB, their comparable lengths and overall sequence and domain organization similarities made it feasible to construct functional chimeras between these two TonB proteins. We show here that, while the N terminus of the TonB protein determines specificity for a particular ExbB-ExbD complex, the C terminus controls the specificity of the interaction with TonB-dependent receptors. A chimera composed of the C-terminal 65 residues (175 to 239) of E. coli TonB fused to the N terminus and central region of V. cholerae TonB1 exhibited all the receptor specificities associated with wild-type E. coli TonB. These 65 residues may represent the minimal region required to reconstitute a functional E. coli TonB C-terminal dimerization domain. Although at least 75 C-terminal residues (164 to 239) of E. coli TonB were determined by crystallography to participate in dimerization (13), the first 11 of these (164 to 174) may not be required to form the dimer. Alternatively, the equivalent 11 residues contributed by V. cholerae TonB1 may be similar enough to the substituted E. coli TonB residues to permit the formation of a functional dimer. Between 66 and 81 C-terminal V. cholerae TonB1 residues were required to form a functional chimera with TonB1-associated receptor specificities. Thus, the putative V. cholerae TonB1 C-terminal dimer may involve a greater number of residues or have more stringent amino acid requirements than the E. coli TonB dimer.
Residues surrounding Q160 in E. coli TonB have been implicated previously in the direct interaction with the TonB box of the BtuB receptor (10). However, since the Q160 residue, as well as some of the neighboring amino acids, is conserved in V. cholerae TonB1, it is unlikely that this region is responsible for the differences in receptor specificity observed between E. coli TonB and V. cholerae TonB1. This is supported by data presented here showing that a chimera containing the E. coli TonB C terminus, but not the Q160 region, was able to interact productively with all the E. coli receptors tested. This does not, however, exclude a role for these residues in the TonB-receptor interaction(s), since they may participate in receptor interactions that are conserved in both E. coli TonB and V. cholerae TonB1. Although the Q160 region is not required for dimerization, its proximity to the dimerization domain suggests that it could form part of a C-terminal receptor-binding site. Alternatively, the Q160 region may interact with the receptor independently of the C-terminal dimerization domain.
Given the tightly intertwined nature of the E. coli TonB C-terminal dimer (13), it is perhaps not surprising that none of the chimeras involving fusions within the core of this domain were functional (data not shown). Each TonB molecule contributes three β-strands to the large antiparallel β-sheet covering most of the dimer surface, and these strands are connected by an extensive network of hydrogen bonds (13). It is therefore unlikely that a large number of amino acid substitutions would be tolerated within this region. Interestingly, in a study by Zhao and Poole (59), functional chimeras between Pseudomonas aeruginosa TonB1 and E. coli TonB involving fewer than 65 E. coli TonB C-terminal residues were created; however, E. coli TonB is more similar to P. aeruginosa TonB1 (44% identity; 62% similarity) than to V. cholerae TonB1 (17% identity; 35% similarity) in the C-terminal dimerization region.
The observed lack of activity of TonB chimeras involving the putative C-terminal dimerization domain could be due to poor expression and/or to degradation of the hybrid protein, rather than to nonproductive receptor interactions. It was determined by Western blot analyses that all the chimeric proteins were expressed but were significantly less stable than wild-type E. coli TonB in whole-cell extracts (data not shown). However, there was no correlation between TonB activity in iron utilization assays and apparent stability of the hybrid protein, and many of the hybrids that displayed wild-type TonB activity had similar degradation profiles as those that were functionally inactive. Nevertheless, degradation of chimeric proteins poses a concern for the interpretation of negative results. In this study, conclusions about specificity were founded on the ability of each hybrid TonB to display wild-type activities towards a minimum of two different outer membrane receptors, at least one of which is chromosomally encoded. It seems unlikely, therefore, that the failure of an active TonB hybrid to recognize a particular receptor is due solely to insufficient levels of the mature TonB protein. For the same reason, it seems unlikely that negative results are due to dominant-negative effects on the TonB-receptor interaction from TonB degradation products, since this would be expected to inhibit all TonB-dependent transport.
The specificity of V. cholerae TonB1 towards receptors in both V. cholerae and E. coli appears to be determined by the very C-terminal end of the TonB molecule. A P238T, P238R, or P238A substitution in V. cholerae TonB1 conferred upon TonB1 the ability to recognize at least one TonB2-dependent V. cholerae receptor, as well as multiple E. coli receptors. Based on a CLUSTAL-W alignment (Fig. 3), the V. cholerae TonB1 P238 residue corresponds to R198 in V. cholerae TonB2. This suggests that an Arg residue in this position may be optimal for interactions with certain TonB2-dependent receptors. However, a more important consideration could be that the Pro residue is inhibitive to certain receptor interactions. The TonB1 P238 residue aligns with N227 in E. coli TonB, which is positioned on the β-sheet face of the cylindrical E. coli TonB dimer in such a way that the two N227 residues, one from each TonB molecule, are in close proximity to each other and to the cleft formed by the two α-helices on the opposite face of the dimer. The α-helix cleft was proposed previously as a binding site for outer membrane receptors, since this region exhibits significant flexibility and forms a structural motif that could feasibly accommodate a receptor domain (13). In TonB1, the Pro residues may distort the binding pocket in such a way that only certain receptors are accommodated. Although receptor binding through the TonB C-terminal dimerization domain is an attractive hypothesis, it should be noted that the involvement of the TonB dimer in receptor interactions has not yet been demonstrated. Thus, it is not clear whether the C-terminal dimer does, in fact, represent the receptor binding conformation of TonB.
The P238 substitutions in V. cholerae TonB1 did not appear to interfere with any of the known TonB1 functions. Rather, the effect of each mutation was to widen the range of receptors that TonB1 can productively interact with in V. cholerae. It is perhaps surprising that none of these mutations have been selected for during the evolution of V. cholerae as a pathogen. One possibility is that such mutations permit TonB1 to functionally interact with the enterobactin receptor IrgA, which normally only recognizes TonB2. Since IrgA is one of the most abundantly expressed TonB-dependent receptors in V. cholerae, interactions with this receptor might sequester TonB molecules to such an extent that transport of other substrates, including heme and vibriobactin, would be diminished. It is also conceivable that there is no selective advantage of these mutations due to the presence of TonB2 or that the P238 mutations in TonB1 have an adverse effect on TonB1 function that is not detectable under the assay conditions used. Nutritional assays, although extremely sensitive at low levels of transport activity, are not very quantitative at higher levels of activity, and decreases in TonB1 activity due to the P238 mutations may not have been apparent.
The TonB box is an important functional domain in TonB-dependent receptors. Several mutations have been identified within this domain of E. coli FepA and BtuB that eliminate substrate transport without affecting substrate binding (1, 2, 9, 32). A comparison of E. coli and V. cholerae receptor TonB boxes suggested that this region might contain residues involved in determining the specificity of the TonB-receptor interactions. However, domain-switching experiments showed that the TonB box heptapeptide does not represent a specificity module. E. coli ChuA carrying the V. cholerae HutA TonB box exhibited the same specificities as did wild-type ChuA. Replacing the TonB box of HutA with that of ChuA appeared to strengthen the weak interactions with E. coli TonB, but interactions with V. cholerae TonB1 were unaffected, suggesting that the specificity of the receptor had not changed. These data are consistent with earlier observations that the TonB box imparts a particular conformation, rather than specific residues, needed for the interaction with TonB (9, 18, 32).
While the TonB box itself does not determine specificity for a particular TonB protein, the amino acid residue immediately preceding the TonB box domain had a profound impact on the ability of E. coli receptors to recognize V. cholerae TonB1. Insertion at this position of a Tyr residue in E. coli FepA, or a Phe residue in E. coli ChuA, allowed these receptors to interact productively with V. cholerae TonB1, suggesting that TonB1 may specifically require an aromatic residue directly prior to the TonB box. Indeed, the V. cholerae heme receptors HutA and HutR, both of which recognize TonB1, carry aromatic residues at the TonB box −1 position. However, since insertion of a Leu residue at this position in the V. cholerae enterobactin receptor IrgA also allowed for productive TonB1 interactions, the requirement may be for a large hydrophobic residue. This hypothesis is supported by the finding that the V. cholerae ferrichrome receptor FhuA, which uses both TonB1 and TonB2, carries a Leu in the predicted TonB box −1 position. Taken together, these findings may necessitate a redefinition of the TonB box functional domain to include the TonB box −1 position.
The data presented here strongly imply that the extreme N terminus of the receptor is involved in determining the specificity of the TonB interactions. It is likely, therefore, that this receptor region contacts TonB directly. The relevance of this interaction for transport is not currently known. TonB interacts preferentially with ligand-bound receptors (42), and it is conceivable that TonB discriminates between the bound and unbound states by sensing changes at the periplasmic face of the receptor. Such changes at the periplasmic face of a TonB-dependent receptor in response to ligand binding were first reported for FhuA (43), and structural comparisons of liganded and unliganded FhuA later showed that, when FhuA binds its substrate, a periplasmically exposed region of the N-terminal plug domain undergoes dramatic conformational shifts (36). Studies of the vitamin B12 receptor BtuB have further demonstrated that ligand binding causes the TonB box heptapeptide to unwind and adopt an extended conformation that may reach into the periplasm (38). These data suggest that the receptor region containing the TonB box domain could be involved in signaling substrate binding to TonB (36, 38, 43).
TonB1 exhibited the same specificities in both V. cholerae and E. coli. Thus, the receptor discrimination displayed by TonB1 likely represents a true requirement for particular interaction sites. Such a specificity is consistent with a more specialized role for TonB1 in the life cycle of V. cholerae. While TonB2 is encoded on the large chromosome along with the V. cholerae housekeeping genes, the TonB1 system genes are located on the smaller replicon and were probably acquired by horizontal transfer. The tonB1 locus may have been maintained during the evolution of this pathogen in order to cope with specific environments such as the marine environment or the human intestinal tract.
Acknowledgments
We gratefully acknowledge Deborah A. Wrona, Elizabeth E. Wyckoff, and Douglas P. Henderson for helpful discussions and Elizabeth E. Wyckoff and Laura Runyen-Janecky for critical reading of the manuscript. We thank Kathleen Postle for many helpful suggestions and for generously providing strains and E. coli TonB antisera. We are greatly indebted to Randall K. Holmes for supplying the V. cholerae IrgA antiserum. We also thank Charles F. Earhart, Yuki Gleason, and Alfredo G. Torres for providing plasmids.
This work was supported by a grant from the Foundation for Research and by grant AI50669 from the National Institutes of Health.
REFERENCES
- 1.Barnard, T. J., M. E. Watson, and M. A. McIntosh. 2001. Mutations in the Escherichia coli receptor FepA reveal residues involved in ligand binding and transport. Mol. Microbiol. 41:527-536. [DOI] [PubMed] [Google Scholar]
- 2.Bell, P. E., C. D. Nau, J. T. Brown, J. Konisky, and R. J. Kadner. 1990. Genetic suppression demonstrates interaction of TonB protein with outer membrane transport proteins in Escherichia coli. J. Bacteriol. 172:3826-3829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bradbeer, C. 1993. The proton motive force drives the outer membrane transport of cobalamin in Escherichia coli. J. Bacteriol. 175:3146-3150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Braun, M., H. Killmann, and V. Braun. 1999. The β-barrel domain of FhuΔ5-160 is sufficient for TonB-dependent FhuA activities of Escherichia coli. Mol. Microbiol. 33:1037-1049. [DOI] [PubMed] [Google Scholar]
- 5.Braun, V. 1995. Energy-coupled transport and signal transduction through the gram-negative outer membrane via TonB-ExbB-ExbD-dependent receptor proteins. FEMS Microbiol. Rev. 16:295-307. [DOI] [PubMed] [Google Scholar]
- 6.Buchanan, S. K., B. S. Smith, L. Venkatramani, D. Xia, L. Esser, M. Palnitkar, R. Chakraborty, D. van der Helm, and J. Deisenhofer. 1999. Crystal structure of the outer membrane active transporter FepA from Escherichia coli. Nat. Struct. Biol. 6:56-63. [DOI] [PubMed] [Google Scholar]
- 7.Butterton, J. R., J. A. Stoebner, S. M. Payne, and S. B. Calderwood. 1992. Cloning, sequencing, and transcriptional regulation of viuA, the gene encoding the ferric vibriobactin receptor of Vibrio cholerae. J. Bacteriol. 174:3729-3738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Byers, B. R., and J. E. Arceneaux. 1998. Microbial iron transport: iron acquisition by pathogenic microorganisms. Met. Ions Biol. Syst. 35:37-66. [PubMed] [Google Scholar]
- 9.Cadieux, N., C. Bradbeer, and R. J. Kadner. 2000. Sequence changes in the Ton box region of BtuB affect its transport activities and interaction with TonB protein. J. Bacteriol. 182:5954-5961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cadieux, N., and R. J. Kadner. 1999. Site-directed disulfide bonding reveals an interaction site between energy-coupling protein TonB and BtuB, the outer membrane cobalamin transporter. Proc. Natl. Acad. Sci. USA 96:10673-10678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chambers, S. P., S. E. Prior, D. A. Barstow, and N. P. Minton. 1988. The pMTL nic− cloning vectors. I. Improved pUC polylinker regions to facilitate the use of sonicated DNA for nucleotide sequencing. Gene 68:139-149. [DOI] [PubMed] [Google Scholar]
- 12.Chang, A. C. Y., and S. N. Cohen. 1978. Construction and characterization of amplifiable multicopy DNA cloning vehicles derived from the P15A cryptic miniplasmid. J. Bacteriol. 134:1141-1156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chang, C., A. Mooser, A. Pluckthun, and A. Wlodawer. 2001. Crystal structure of the dimeric C-terminal domain of TonB reveals a novel fold. J. Biol. Chem. 276:27535-27540. [DOI] [PubMed] [Google Scholar]
- 14.Ferguson, A. D., R. Chakraborty, B. S. Smith, L. Esser, D. van der Helm, and J. Deisenhofer. 2002. Structural basis of gating by the outer membrane transporter FecA. Science 295:1715-1719. [DOI] [PubMed] [Google Scholar]
- 15.Ferguson, A. D., E. Hofmann, J. W. Coulton, K. Diederichs, and W. Welte. 1998. Siderophore-mediated iron transport: crystal structure of FhuA with bound lipopolysaccharide. Science 282:2215-2220. [DOI] [PubMed] [Google Scholar]
- 16.Gardner, E. W., S. T. Lyles, and C. E. Lankford. 1964. A comparison of virulence in Vibrio cholerae strains for embryonated egg. J. Infect. Dis. 114:412-416. [DOI] [PubMed] [Google Scholar]
- 17.Griffiths, G. L., S. P. Sigel, S. M. Payne, and J. B. Neilands. 1984. Vibriobactin, a siderophore from Vibrio cholerae. J. Biol. Chem. 259:383-385. [PubMed] [Google Scholar]
- 18.Gudmundsdottir, A., P. E. Bell, M. D. Lundrigan, C. Bradbeer, and R. J. Kadner. 1989. Point mutations in a conserved region (TonB box) of Escherichia coli outer membrane protein BtuB affect vitamin B12 transport. J. Bacteriol. 171:6526-6533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Guerinot, M. L. 1994. Microbial iron transport. Annu. Rev. Microbiol. 48:743-772. [DOI] [PubMed] [Google Scholar]
- 20.Hanahan, D. 1983. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 166:557-580. [DOI] [PubMed] [Google Scholar]
- 21.Heller, K., R. J. Kadner, and K. Günther. 1988. Suppression of the btuB451 mutation by mutations in the tonB gene suggests a direct interaction between TonB and TonB-dependent receptor proteins in the outer membrane of Escherichia coli. Gene 64:147-153. [DOI] [PubMed] [Google Scholar]
- 22.Henderson, D. P., and S. M. Payne. 1994. Characterization of the Vibrio cholerae outer membrane heme transport protein HutA: sequence of the gene, regulation of expression, and homology to the family of TonB-dependent proteins. J. Bacteriol. 176:3269-3277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Henderson, D. P., and S. M. Payne. 1993. Cloning and characterization of the Vibrio cholerae genes encoding the utilization of iron from haemin and haemoglobin. Mol. Microbiol. 7:461-469. [DOI] [PubMed] [Google Scholar]
- 24.Henderson, D. P., and S. M. Payne. 1994. Vibrio cholerae iron transport systems: role of heme and siderophore iron transport in virulence and identification of a gene associated with multiple iron transport systems. Infect. Immun. 62:5120-5125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Higgs, P. I., P. S. Myers, and K. Postle. 1998. Interactions in the TonB-dependent energy transduction complex: ExbB and ExbD form homomultimers. J. Bacteriol. 180:6031-6038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hill, C., and B. Harnish. 1981. Inversions between ribosomal RNA genes of Escherichia coli. Proc. Natl. Acad. Sci. USA 78:7069-7072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ho, S. N., H. D. Hunt, R. M. Horton, J. K. Pullen, and L. R. Pease. 1989. Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene 77:51-59. [DOI] [PubMed] [Google Scholar]
- 28.Hong, M., Y. Gleason, E. E. Wyckoff, and S. M. Payne. 1998. Identification of two Shigella flexneri chromosomal loci involved in intercellular spreading. Infect. Immun. 66:4700-4710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Howard, S. P., C. Herrmann, C. W. Stratilo, and V. Braun. 2001. In vivo synthesis of the periplasmic domain of TonB inhibits transport through the FecA and FhuA iron siderophore transporters of Escherichia coli. J. Bacteriol. 183:5885-5895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kadner, R. J. 1990. Vitamin B12 transport in Escherichia coli: energy coupling between membranes. Mol. Microbiol. 4:2027-2033. [DOI] [PubMed] [Google Scholar]
- 31.Laemmli, U. K. 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680-685. [DOI] [PubMed] [Google Scholar]
- 32.Larsen, R. A., D. Foster-Hartnett, M. A. McIntosh, and K. Postle. 1997. Regions of Escherichia coli TonB and FepA proteins essential for in vivo physical interactions. J. Bacteriol. 179:3213-3221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Larsen, R. A., M. G. Thomas, G. E. Wood, and K. Postle. 1994. Partial suppression of an Escherichia coli TonB transmembrane domain mutation (delta V17) by a missense mutation in ExbB. Mol. Microbiol. 13:627-640. [DOI] [PubMed] [Google Scholar]
- 34.Larsen, R. A., G. E. Wood, and K. Postle. 1993. The conserved proline-rich motif is not essential for energy transduction by Escherichia coli TonB protein. Mol. Microbiol. 10:943-953. [DOI] [PubMed] [Google Scholar]
- 35.Letain, T. E., and K. Postle. 1997. TonB protein appears to transduce energy by shuttling between the cytoplasmic membrane and the outer membrane in Escherichia coli. Mol. Microbiol. 24:271-283. (Erratum, 25:617.) [DOI] [PubMed]
- 36.Locher, K. P., B. Rees, R. Koebnik, A. Mitschler, L. Moulinier, J. P. Rosenbusch, and D. Moras. 1998. Transmembrane signaling across the ligand-gated FhuA receptor: crystal structures of free and ferrichrome-bound states reveal allosteric changes. Cell 95:771-778. [DOI] [PubMed] [Google Scholar]
- 37.Lundrigan, M. D., and R. J. Kadner. 1986. Nucleotide sequence of the gene for the ferrienterochelin receptor FepA in Escherichia coli. Homology among outer membrane receptors that interact with TonB. J. Biol. Chem. 261:10797-10801. [PubMed] [Google Scholar]
- 38.Merianos, H. J., N. Cadieux, C. H. Lin, R. J. Kadner, and D. S. Cafiso. 2000. Substrate-induced exposure of an energy-coupling motif of a membrane transporter. Nat. Struct. Biol. 7:205-209. [DOI] [PubMed] [Google Scholar]
- 39.Mey, A. R., and S. M. Payne. 2001. Haem utilization in Vibrio cholerae involves multiple TonB-dependent haem receptors. Mol. Microbiol. 42:835-849. [DOI] [PubMed] [Google Scholar]
- 40.Mey, A. R., E. E. Wyckoff, A. G. Oglesby, E. Rab, R. K. Taylor, and S. M. Payne. 2002. Identification of the Vibrio cholerae enterobactin receptors VctA and IrgA: IrgA is not required for virulence. Infect. Immun. 70:3419-3426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Miller, V. L., and J. J. Mekalanos. 1988. A novel suicide vector and its use in construction of insertion mutations: osmoregulation of outer membrane proteins and virulence determinants in Vibrio cholerae requires toxR. J. Bacteriol. 170:2575-2583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Moeck, G. S., J. W. Coulton, and K. Postle. 1997. Cell envelope signaling in Escherichia coli. Ligand binding to the ferrichrome-iron receptor FhuA promotes interaction with the energy-transducing protein TonB. J. Biol. Chem. 272:28391-28397. [DOI] [PubMed] [Google Scholar]
- 43.Moeck, G. S., P. Tawa, H. Xiang, A. A. Ismail, J. L. Turnbull, and J. W. Coulton. 1996. Ligand-induced conformational change in the ferrichrome-iron receptor of Escherichia coli K-12. Mol. Microbiol. 22:459-471. [DOI] [PubMed] [Google Scholar]
- 44.Occhino, D. A., E. E. Wyckoff, D. P. Henderson, T. J. Wrona, and S. M. Payne. 1998. Vibrio cholerae iron transport: haem transport genes are linked to one of two sets of tonB, exbB, exbD genes. Mol. Microbiol. 29:1493-1507. [DOI] [PubMed] [Google Scholar]
- 45.Postle, K. 1993. TonB protein and energy transduction between membranes. J. Bioenerg. Biomembr. 25:591-601. [DOI] [PubMed] [Google Scholar]
- 46.Postle, K., and J. T. Skare. 1988. Escherichia coli TonB protein is exported from the cytoplasm without proteolytic cleavage of its amino terminus. J. Biol. Chem. 263:11000-11007. [PubMed] [Google Scholar]
- 47.Pressler, U., H. Staudenmaier, L. Zimmermann, and V. Braun. 1988. Genetics of the iron dicitrate transport system of Escherichia coli. J. Bacteriol. 170:2716-2724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Rogers, H. J. 1973. Iron-binding catechols and virulence in Escherichia coli. Infect. Immun. 7:445-456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Rogers, M. B., J. A. Sexton, G. J. DeCastro, and S. B. Calderwood. 2000. Identification of an operon required for ferrichrome iron utilization in Vibrio cholerae. J. Bacteriol. 182:2350-2353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Roof, S. K., J. D. Allard, K. P. Bertrand, and K. Postle. 1991. Analysis of Escherichia coli TonB membrane topology by use of PhoA fusions. J. Bacteriol. 173:5554-5557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Runyen-Janecky, L. J., M. Hong, and S. M. Payne. 1999. Virulence plasmid-encoded impCAB operon enhances survival and induced mutagenesis in Shigella flexneri after exposure to UV radiation. Infect. Immun. 67:1415-1423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Seliger, S., A. Mey, A. Valle, and S. Payne. 2001. The two TonB systems of Vibrio cholerae: redundant and specific functions. Mol. Microbiol. 39:801-812. [DOI] [PubMed] [Google Scholar]
- 53.Skare, J. T., B. M. Ahmer, C. L. Seachord, R. P. Darveau, and K. Postle. 1993. Energy transduction between membranes. TonB, a cytoplasmic membrane protein, can be chemically cross-linked in vivo to the outer membrane receptor FepA. J. Biol. Chem. 268:16302-16308. [PubMed] [Google Scholar]
- 54.Stoebner, J. A., J. R. Butterton, S. B. Calderwood, and S. M. Payne. 1992. Identification of the vibriobactin receptor of Vibrio cholerae. J. Bacteriol. 174:3270-3274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Torres, A. G., and S. M. Payne. 1997. Haem iron-transport system in enterohaemorrhagic Escherichia coli O157:H7. Mol. Microbiol. 23:825-833. [DOI] [PubMed] [Google Scholar]
- 56.van der Helm, D. 1998. The physical chemistry of bacterial outer-membrane siderophore receptor proteins. Met. Ions Biol. Syst. 35:355-401. [PubMed] [Google Scholar]
- 57.Wang, R. F., and S. R. Kushner. 1991. Construction of versatile low-copy-number vectors for cloning, sequencing and gene expression in Escherichia coli. Gene 100:195-199. [PubMed] [Google Scholar]
- 58.Wyckoff, E. E., A.-M. Valle, S. L. Smith, and S. M. Payne. 1999. A multifunctional ABC transporter system from Vibrio cholerae transports vibriobactin and enterobactin. J. Bacteriol. 181:7588-7596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Zhao, Q., and K. Poole. 2002. Mutational analysis of the TonB1 energy coupler of Pseudomonas aeruginosa. J. Bacteriol. 184:1503-1513. [DOI] [PMC free article] [PubMed] [Google Scholar]