Abstract
Vibrio fischeri colonizes the squid Euprymna scolopes in a mutualistic symbiosis. Hatchling squid lack these bacterial symbionts, and V. fischeri strains must compete to occupy this privileged niche. We cloned a V. fischeri gene, designated pilA, that contributes to colonization competitiveness and encodes a protein similar to type IV-A pilins. Unlike its closest known relatives, Vibrio cholerae mshA and vcfA, pilA is monocistronic and not clustered with genes associated with pilin export or assembly. Using wild-type strain ES114 as the parent, we generated an in-frame pilA deletion mutant, as well as pilA mutants marked with a kanamycin resistance gene. In mixed inocula, marked mutants were repeatedly outcompeted by ES114 (P < 0.05) but not by an unmarked pilA mutant, for squid colonization. In contrast, the ratio of mutant to ES114 CFUs did not change during 70 generations of coculturing. The competitive defect of pilA mutants ranged from 1.7- to 10-fold and was more pronounced when inocula were within the range estimated for V. fischeri populations in Hawaiian seawater (200 to 2,000 cells/ml) than when higher densities were used. ES114 also outcompeted a pilA mutant by an average of twofold at lower inoculum densities, when only a fraction of the squid became infected, most by only one strain. V. fischeri strain ET101, which was isolated from Euprymna tasmanica and is outcompeted by ES114, lacks pilA; however, 11 other diverse V. fischeri isolates apparently possess pilA. The competitive defect of pilA mutants suggests that cell surface molecules may play important roles in the initiation of beneficial symbioses in which animals must acquire symbionts from a mixed community of environmental bacteria.
All animals and plants are hosts to a native microbiota. These microbial symbionts often contribute to the normal health and development of their respective hosts in exchange for a relatively privileged niche. Many plant and animal symbionts are transmitted horizontally, through the environment after embryogenesis, and are not directly inherited through germplasm from the previous generation. Therefore, with each host generation, environmental bacteria must compete to colonize the empty niches within new host individuals. In perhaps the best-studied example of such competition, mixed communities of rhizobia compete for access to the roots and nodules of leguminous plants. Many complex traits, including antibiotic production, motility, and specific cell surface attributes contribute to the nodulation competitiveness of rhizobial strains (3, 9, 41, 46). Less is known about the competition between environmental bacteria for colonization of animal hosts, especially the natural modes of infection by the native, nonpathogenic microbiota.
The light organ symbiosis between the luminescent bacterium Vibrio fischeri and the nocturnal Hawaiian squid Euprymna scolopes serves as a model chronic, mutualistic association between extracellular bacteria and animal epithelia (32, 43). At the time of hatching, the light organs of juvenile squid are not colonized by V. fischeri but they rapidly acquire these symbionts from the surrounding seawater (28). The nascent light organ possesses elaborate ciliated appendages that entrain and concentrate planktonic V. fischeri in a mucus matrix, facilitating initial colonization (31). During this process, motile V. fischeri cells move from pores on the surface of the light organ, through ducts, into epithelium-lined crypt spaces (12, 31). Within hours after infection by V. fischeri, the squid also display a diurnal behavior in which they expel approximately 90% of their light organ symbionts each morning and supply the remaining bacteria with sufficient nutrients to repopulate the light organ by nightfall (6, 13). Therefore, V. fischeri strains may compete during initial entry into the light organ, as well as during subsequent daily expulsion and regrowth. Increased populations of V. fischeri in squid habitats appear to be a direct result of occupation of this symbiotic niche (24), illustrating an ecological advantage for strains that compete effectively for light organ colonization.
Although most, if not all, wild-type V. fischeri isolates can colonize E. scolopes, isolates native to this host outcompete isolates from other sources for light organ colonization in mixed inoculations. For example, when presented to E. scolopes juveniles in equal ratios, strain ES114, which was isolated from E. scolopes, was more than 10-fold more competitive than strains EM17 and ET101, which were isolated from light organs of Euprymna morsei and Euprymna tasmanica, respectively (30). V. fischeri isolates not associated with squid may be even less competitive for colonization of E. scolopes (22). It is likely that several factors contribute to colonization competitiveness, and mutant analyses provide a means by which to reveal them. For example, Visick and Ruby demonstrated that a katA mutant, deficient in the production of a periplasmic catalase, was outcompeted by the parent strain (44).
McFall-Ngai and colleagues found that addition of a mannose analog to seawater, or treatment of V. fischeri with extracellular protease, could partially block infection of E. scolopes and suggested that a V. fischeri mannose-binding cell surface protein contributes to the initiation of the V. fischeri-E. scolopes light organ symbiosis (16, 27). We therefore became interested in testing the role of cell surface molecules in colonization and colonization competitiveness. In this paper, we report the cloning and characterization of V. fischeri pilA, a gene that encodes a protein with similarity to type IV-A pilins and that contributes to the competitiveness of V. fischeri during colonization of the E. scolopes light organ.
MATERIALS AND METHODS
Bacteria, media, and reagents.
Wild-type V. fischeri strain ES114 (4) isolated from E. scolopes was the parent strain for mutant construction and was the source of DNA for the cloning of pilA. The other V. fischeri strains used were ES12, ES213 (5), ET401, ET101, EM17 (30), SR5, SA1 (10), CG101, MJ11, WH1 (21), H905 (23), and ATCC 7744. Plasmids were maintained in Escherichia coli strain DH5α (14), with the exception of pEVS104, which was maintained in strain CC118λpir (17). E. coli was grown in LB medium (29), and V. fischeri was grown in either a seawater-based complex medium (SWT) (4) or a Tris-buffered, high-salt, rich, complex medium (LBS) (35). For analysis of competition in culture, Instant Ocean (Aquarium Systems, Mentor, Ohio) was substituted for the seawater in SWT. Agar (15 mg ml−1) was added to solidify the media for plating experiments. Chemicals were obtained from Sigma Chemical Co. (St. Louis, Mo.). Restriction enzymes and DNA ligase were obtained from New England Biolabs (Beverly, Mass.). AmpliTaq DNA polymerase was obtained from Perkin-Elmer (Branchburg, N.J.). Oligonucleotides were synthesized by Operon Technologies Inc. (Alameda, Calif.). When added to LB medium for selection of E. coli, ampicillin, chloramphenicol, and kanamycin were used at concentrations of 100, 20, and 40 μg ml−1, respectively. When added to LBS medium for selection or screening in V. fischeri, chloramphenicol and kanamycin were used at concentrations of 5 and 100 μg ml−1, respectively.
Molecular genetic techniques and sequence analysis.
Southern blotting and dot blotting were performed with the DIG DNA Labeling and Detection kit and the DIG Oligonucleotide Tailing kit (Roche Biochemicals, Indianapolis, Ind.) and a labeled 60-mer oligonucleotide complementary to pilA (5′-CCT GCT CGT TTC AAT TCA AGA AGT GGA TGG TCA ACA GCA ACT GTA TAT CGG ATT TGC ACA-3′) as the probe. For Southern analyses, 5 μg of HindIII-digested genomic DNA was electrophoresed and transferred to nylon membrane, blots were hybridized with the probe overnight at 55°C, and membranes were washed under medium-stringency conditions (two 10-min washes at 55°C in 3 mM sodium citrate-30 mM NaCl-0.1% sodium dodecyl sulfate, pH 7.0) and developed by using the chemiluminescent substrate CDP-Star (Boehringer Mannheim).
Mobilizable vectors were transferred to ES114 by triparental mating with conjugal helper plasmid pEVS104 (36). Plasmids were purified by using the PerfectPrep Plasmid kit (5 prime-3 prime, Inc., Boulder, Colo.). Between restriction and ligation reactions, DNA was recovered with the Wizard DNA Cleanup kit (Promega Corp., Madison, Wis.). PCRs were prepared as previously described (10), with some modifications (see below). DNA sequencing was conducted on an ABI automated DNA sequencer at the University of Hawaii Biotechnology/Molecular Biology Instrumentation and Training Facility. Deposited sequence data were generated by sequencing both DNA strands. Sequence analysis, e.g., identification of open reading frames (ORFs), was performed by using DNA Strider 1.2, and comparisons of ORF and protein sequences were conducted with either the CLUSTAL W (40) or the BLASTP (1) algorithm and the BLOSUM62 scoring matrix (15).
Cloning, sequence analysis, and disruption of pilA.
pilA was identified during the cloning of hvnB (35); however, the plasmid clone used in that study lacked sufficient upstream DNA for allelic replacement. We therefore used a PCR with primers EVS47 (5′-CGA GAG CTC ATA TTA AAG CTC CTT CGT G-3′) and EVS48 (5′-CGT CGT CGT ATT GAG GCG ATG CGT GAG-3′), which amplify pilA, to screen an existing library (44) of ES114 genomic DNA cloned in pBluescript KS (Stratagene Inc., La Jolla, Calif.). pilA was identified in increasingly less complex pools of clones, and we ultimately isolated plasmid pEVS56, which carries pilA on a 7-kb EcoRI fragment.
Partial digestion of pEVS56 with EcoRV, followed by self-ligation, resulted in pEVS56EV, which contained a 4.5-kb pilA-containing fragment. The insert in pEVS56EV was moved into the polylinker of pEVS79 (36), a mobilizable derivative of pBCSK+ (Stratagene), generating pEVS85. pEVS85 was subjected to in vitro transposon mutagenesis (Epicentre, Madison, Wis.). One insertion mutant was digested with XhoI and self-ligated, deleting the mini-Tn5 transposon and about 500 bp of V. fischeri DNA, generating plasmid pEVS85-19ΔXho, which contained pilA centered within a 4-kb insert. Oligonucleotides LNK1 (5′-TAA GGT CGA CA-3′) and LNK2 (5′-AGC TTG TCG ACC-3′) were annealed to each other, and this synthetic insert was cloned into pEVS85-19ΔXho, replacing the BlpI-HindIII pilA fragment. In the resulting construct, pEVS90, residues 21 to 127 in the PilA ORF were replaced with Val-Asp and a SalI site was introduced. The SalI kanR cassette from pUC4K (42) was cloned into the SalI site of pEVS90, resulting in pEVS90K1 and pEVS90K2, which carried the kanR gene in the same or the opposite orientation relative to pilA, respectively.
The ΔpilA::kanR alleles of pEVS90K1 and pEVS90K2 were crossed into the chromosome of ES114 by marker exchange (confirmed by PCR and Southern blot analyses), generating strains EVS401 and EVS402, respectively. The ΔpilA allele of pEVS90 was exchanged into the chromosome of ES114 by first selecting for chloramphenicol-resistant single recombinants and then screening subsequent chloramphenicol-sensitive double recombinants by PCR with primers EVS47 and EVS48. One putative mutant was designated strain EVS400, and the absence of pilA in this strain was confirmed by Southern blotting.
Symbiosis assays.
The ability of V. fischeri strains to initiate colonization of the E. scolopes light organ was measured as described previously (33). Juvenile E. scolopes squid were exposed to an inoculum for 12 to 14 h before being rinsed in inoculum-free seawater. Prior to dilution in seawater, V. fischeri inocula were grown unshaken in 5 ml of SWT medium in 50-ml conical tubes at 28°C such that the optical density at 600 nm (OD600) was between 0.28 and 0.95. The relationship between the number of CFU per milliliter and the OD600 could be expressed as follows: number of CFU per milliliter = [(8.7 × 108) × OD600] + (4.6 × 107) (r2 = 0.81). This relationship was used to estimate the desired inoculum dilution, but inocula were also plated after serial dilution to determine CFU concentrations.
To determine the ratio of kanR-marked to unmarked strains in competition experiments, cells in a culture, inoculum, or infected squid were first plated onto nonselective LBS medium and then patched onto solidified LBS with and without kanamycin. At least 66, 200, or 50 colonies were patched to determine the strain ratios in a culture, an inoculum, or a squid, respectively. To assess whether marked strains were significantly outcompeted by unmarked strains in mixed infections, a Student t test was applied to log-transformed relative competitiveness indexes (RCIs) to determine P values. To assess whether EVS401 was outcompeted by ES114 in clonal (or clonally skewed) infections that arose from low doses of mixed inocula (Table 1), the number of animals infected with >90% of one strain or the other was compared by a binomial sign test (34) to determine probability values.
TABLE 1.
Competition between a pilA mutant and the wild type at low inoculum doses
| Expt | Inoculum
|
No. of squid infected/no. inoculated (%) | No. of squid with:
|
Avg ratio of EVS401 to ES114 in squid | RCI | ||||
|---|---|---|---|---|---|---|---|---|---|
| No. of CFU/ml | No. of CFU/squid | Ratio of EVS401 to ES114 | >90% ES114 | 11-90% of each | >90% EVS401 | ||||
| 1 | 33 | 84 | 1.08:1 | 25/39 (64) | 18 | 2 | 5 | 0.316:1 | 0.29 |
| 2 | 30 | 176 | 1.44:1 | 11/17 (65) | 6 | 3 | 2 | 0.408:1 | 0.28 |
| 3 | 14 | 56 | 1.17:1 | 19/46 (41) | 12 | 0 | 7 | 0.613:1 | 0.52 |
| 4 | 34 | 136 | 1.17:1 | 4/12 (33) | 3 | 1 | 0 | 0.266:1 | 0.23 |
| 5 | 38 | 200 | 0.923:1 | 18/36 (50) | 11 | 1 | 6 | 0.538:1 | 0.58 |
| 6 | 55 | 275 | 1.33:1 | 16/36 (47) | 4 | 4 | 8 | 1.38:1 | 1.0 |
| Total | 14-55 | 56-275 | 1.16:1a | 93/186 (50) | 54 | 11 | 28 | 0.611:1a | 0.53a |
Values represent averages for the 93 animals infected, not the averages of the six experiments weighted equally.
We generated mixed inocula at roughly 1:1 ratios, stored them at −80°C in 20% glycerol, and then revived portions of these stocks to set up inocula in later experiments. The ratio of marked to unmarked strains did not change during cryopreservation. Similarly, when infected squid were frozen at −80°C, the number of V. fischeri bacteria recovered upon thawing, homogenization, and plating was similar to that recovered when squid were homogenized and plated immediately.
To collect bacteria vented from the light organ, colonized animals were rinsed three times between 12 and 24 h after inoculation with cultured bacteria and the seawater was plated to confirm the absence of the original inoculum, and expelled bacteria were collected 36 to 40 h after initial inoculation. Expelled bacteria were then diluted at least 10-fold in fresh seawater, and newly hatched squid were exposed to this inoculum approximately 12 h after the expelled bacteria were collected.
Nucleotide sequence accession numbers.
The nucleotide sequences reported here have been submitted to the GenBank database and assigned accession no. AF487524 and AF206718.
RESULTS
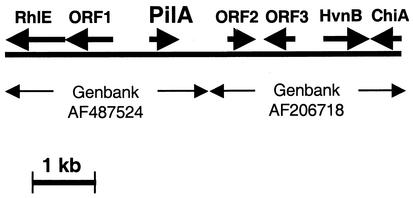
Bacterial cell surface adhesins and pili may play important roles in determining the colonization proficiency of V. fischeri strains as they compete to occupy the light organs of hatchling Hawaiian bobtail squid. We identified a putative pilin-encoding V. fischeri gene, which we designated pilA (Fig. 1). On the basis of the locations and orientations of surrounding ORFs, pilA appears to be monocistronic. In contrast to mshA (19, 26) and many similar pilin genes, pilA is not flanked by genes normally associated with pilin export or assembly. Rather, the region extending more than 2 kb upstream or downstream from pilA appears to encode an RNA helicase (RhlE), an NADase (HvnB), a ferredoxin-like protein (ORF2), and proteins of unknown function common to other eubacteria (ORF1 and ORF3) (Fig. 1).
FIG. 1.
Sequence analysis of the pilA region. The solid line and single-headed arrows represent V. fischeri genomic DNA and ORFs, respectively. The double-headed arrows flanking the accession numbers mark the sequences submitted to the GenBank database.
The PilA ORF encodes a 14.1-kDa, 128-amino-acid protein that, by analogy to pilin systems (37), may be processed to a 13.4-kDa, 122-amino-acid pilin. The PilA ORF is similar to pilins that comprise type IV-A pili, with the highest degree of similarity at the N termini of these proteins (Fig. 2) and low overall similarity, as is typical in comparisons of type IV-A pili (37). The closest known relatives to PilA are VcfA (20) and MshA (19, 26) from V. cholerae. PilA and MshA were 33% identical and 52% similar over 56 N-terminal residues. The conserved phenylalanine (F) residue found at the N terminus of a mature pilin is conserved in the V. fischeri PilA sequence; however, the preceding residue is not a glycine (G) as is found in other type IV-A pili (37).
FIG. 2.
N-terminal comparison of V. fischeri PilA and type IV-A pilins. (A) N-terminal sequence of V. fischeri (V.f.) PilA. (B) Alignment of type-IV-A pilins relative to V. fischeri PilA. Bold white letters (on a black background) and bold black letters indicate residues that are identical or similar to those of V.f. PilA, respectively. Similarity was assigned by using the BLOSUM62 scoring matrix. The asterisk marks the N-terminal phenylalanine (F) of mature pilins. Abbreviations (GenBank database accession numbers): V.c., Vibrio cholerae (CAA54430); V.v., Vibrio vulnificus (AAL32049); P.a., Pseudomonas aeruginosa (AAK68038); N.g., Neisseria gonorrhoeae (CAA73472); A.sp., Azoarcus species (AAC27898); X.c., Xanthomonas citri (https://ncbi.nlm.nih.gov/nucleotide/AAB97527); M.x., Myxococcus xanthus (S70530).
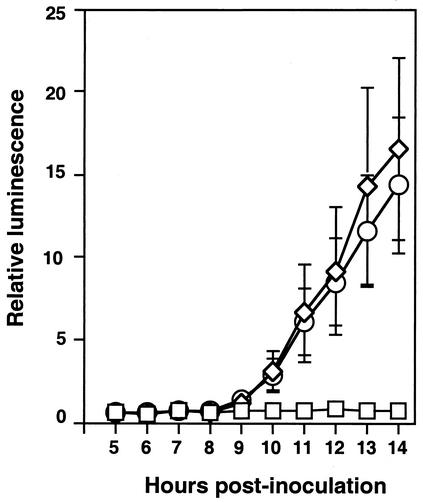
To test whether pilA contributes to the process of infecting E. scolopes, we generated pilA mutant strains and tested them in squid colonization experiments. The pilA mutants, generated in the wild-type ES114 parent, included an unmarked in-frame pilA deletion mutant, strain EVS400, as well as mutants bearing kanR, strains EVS401 and EVS402 (both are ΔpilA::kanR mutants, but they have opposite kanR orientations). In single-inoculation experiments, EVS400, EVS401, and EVS402 were able to infect E. scolopes, and animals colonized by these strains produced amounts of bioluminescence (which correlate well with the symbiont population sizes) similar to those produced by animals colonized with ES114 for at least 3 days after infection (Fig. 3 and data not shown).
FIG. 3.
Colonization of squid by a V. fischeri pilA mutant. Twenty juvenile squid were inoculated with approximately 1,500 CFU of wild-type strain ES114 or strain EVS400 (ΔpilA) per ml. Twenty control animals received no inoculation. Luminescence was measured with an automated luminometer at hourly intervals, and the average values for animals inoculated with ES114 (circles), EVS400 (diamonds), or no V. fischeri (squares) are plotted. Error bars indicate standard errors.
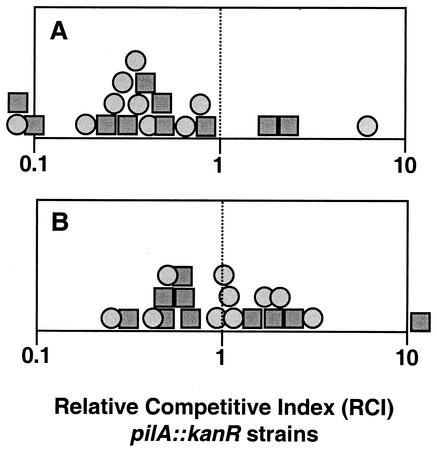
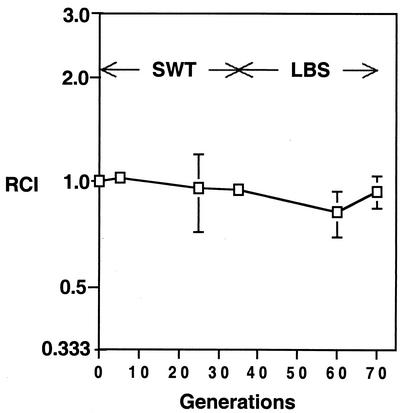
We measured the colonization competitiveness of pilA mutants as an RCI, which is defined as the ratio of marked to unmarked strain in squid divided by the ratio of these strains in the inoculum. An RCI of <1 indicates a competitiveness defect of the marked strain. When kanR strain EVS401 or EVS402 was coinoculated with parent strain ES114 at a nearly 1:1 ratio, the representation of the strains in infected squid was significantly (P < 0.05) dominated by the wild type (Fig. 4A). EVS401 and EVS402 were also significantly outcompeted by the wild type in four subsequent experiments (P < 0.05). Assuming modest V. fischeri death rates within the colonized light organ, we estimated that the colonizing bacteria would go through fewer than 30 generations during these experiments. When EVS401 was cocultured with ES114 for >30 generations, in either of two different media, there was no significant change in the ratio of strains (Fig. 5). Thus, the competitive defect of EVS401 that is apparent in the squid environment is absent in culture.
FIG. 4.
Relative competitiveness of pilA mutants. Juvenile squid were exposed to a mixed inoculum of marked (kanR) and unmarked strains at a total concentration of approximately 3,000 CFU/ml, and the RCIs of the marked strains were determined for each animal (see Materials and Methods) and plotted as either circles (EVS401) or squares (EVS402). Marked pilA::kanR mutant strains EVS401 and EVS402 competed against wild-type ES114 (A) or unmarked ΔpilA mutant strain EVS400 (B). Symbols to the left or right of the frames indicate datum points where the RCI was less than 0.1 or greater than 10, respectively.
FIG. 5.
Relative competitiveness of pilA mutant EVS401 in culture. Marked pilA::kanR mutant EVS401 was cocultured with wild-type ES114 during growth in SWT, followed by growth in LBS. Error bars (some too small to visualize) indicate standard errors (n = 3).
In contrast to competition against the wild type, when either EVS401 or EVS402 was coinoculated with the unmarked ΔpilA mutant EVS400, the representation of the two strains in infected squid was roughly equivalent to the ratio in the inoculum (Fig. 4B), indicating that the presence or orientation of the kanR cassette does not account for the loss of competitiveness of the pilA::kanR mutants. In one of the four subsequent experiments, EVS401 was also outcompeted by EVS400 (P < 0.05), and the kanR marker itself may contribute somewhat to a decrease in competitive fitness. However, EVS401 and EVS402 were not consistently outcompeted by EVS400 and it is not unlikely that one experiment in five would show a significant effect at this confidence level merely by chance. As a whole, these experiments indicate that EVS401 and EVS402 show a greater disadvantage when forced to compete with the wild type than when forced to compete with the unmarked pilA deletion strain.
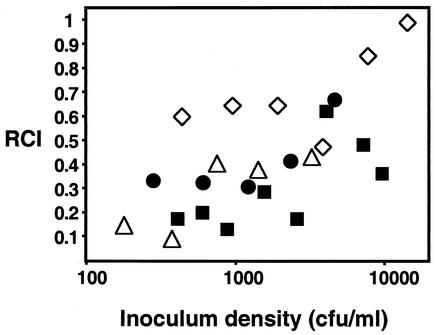
Over the course of several experiments, we found that strain EVS401 was outcompeted by the wild type between 1.7- and 10-fold, as illustrated by RCIs between 0.6 and 0.1 (Fig. 6 and data not shown). We considered the possibility that pilA might be expressed preferentially in the host and that differences between the wild type and the pilA mutant might be more pronounced if host-derived cells, rather than cultured cells, were used as inocula. However, similar results were obtained when either cultured cells or cells vented (expelled) from colonized animals were used to inoculate hatchling squid (Fig. 6). Interestingly, the competitive defect of EVS401 was most pronounced when the total inoculum density was less than 2,000 cells/ml and was attenuated by higher inoculum densities (Fig. 6).
FIG. 6.
Dose dependence and variability of the EVS401 competition defect. Squid were exposed to serially diluted mixtures of EVS401 and ES114, and the RCI of EVS401 was determined for each animal. For two experiments (filled circles and squares), animals were inoculated with cultured V. fischeri, and for two other experiments (open triangles and diamonds), V. fischeri cells vented from E. scolopes were used to inoculate fresh animals. Each datum point represents the mean value obtained by analysis of 15 to 20 animals.
We also measured colonization competitiveness after diluting the strain mixtures such that only a fraction of the inoculated squid became infected. The rationale behind these experiments was that infections arising from low doses might be clonal, and any difference in the ratio of strains in virtually clonal infections relative to the ratio of strains in the inoculum would represent competition to establish clonal infections rather than interstrain competition within the light organ crypts. We found that when EVS401 and ES114 competed at low inoculum doses (14 to 55 CFU/ml and 56 to 275 CFU/squid), approximately half of the juvenile squid became infected and most of these were dominated (usually to the limit of detection) by only one of the two inoculated strains (Table 1). This distribution toward clonal (or heavily weighted) infections stands in contrast to the distribution of infections at higher inoculum doses (Fig. 4). At low inoculum doses, EVS401 displayed an RCI that was below 1.0 in five out of six experiments (Table 1). The apparent defect of EVS401 in competing to establish clonal infections was only significant (P < 0.05) in the experiment with the most animals, experiment 1 (P = 0.003); however, if the remaining five smaller experiments are combined and considered a second repetition, they also show a significant (P = 0.02) defect of the mutant. Furthermore, combination of the data from all six experiments showed that EVS401 was defective relative to the wild type (P = 0.0003) for establishing clonally skewed infections and was outcompeted by approximately twofold, as indicated by an average RCI of 0.53 (Table 1).
Finally, to test whether the lack or presence of pilA may be a contributing factor in the relatively poor competitiveness of certain natural V. fischeri isolates, we used both PCR and Southern hybridization to detect pilA in their genomic DNA and obtained similar results with both methods (data not shown). Strain ET101, isolated from E. tasmanica, lacked pilA; however, pilA was present in strains ES12, ES213, ET101, ET401, EM17, SR5, SA1, CG101, MJ11, H905, WH1, and ATCC 7744. Therefore, the lack of pilA may contribute to the competitive deficiency of ET101 (30) but our data do not support the hypothesis that pilA is a unique attribute of E. scolopes isolates or that its presence is the sole reason for their relative competitiveness, although variations in the sequence or the expression of pilA could, in part, explain competitive differences between E. scolopes isolates and other strains.
DISCUSSION
In near-shore Hawaiian waters, V. fischeri strains compete to occupy the privileged niche within the light organs of E. scolopes hatchlings, and bacterial cell surface components such as pili may contribute to symbiont-host recognition and colonization competitiveness. We have cloned a gene, designated pilA, from V. fischeri that encodes an ORF resembling type IV-A pili, the nearest known relatives of which are the mannose-sensitive pilins VcfA (20) and MshA (19, 26) of V. cholerae. Because previous studies (16, 27) suggested a role of mannose-recognizing adhesins in the establishment of this symbiosis, we assessed the symbiotic competence of pilA mutants.
As is the case for wild-type V. fischeri strains from diverse origins (22, 30) and a katA mutant (44), pilA mutants were able to colonize E. scolopes (e.g., Fig. 3) but showed colonization defects when forced to compete with parent ES114 cells (Fig. 4 and 6). This competitive defect could be directly due to the loss of pilA or to secondary effects associated with the mutant pilA alleles (e.g., general deleterious effects of producing a truncated PilA peptide); however, the trait was symbiosis specific in that no defect of the pilA mutant was observed after 70 generations of coculture with the wild type in liquid media (Fig. 5).
Assays of competitiveness are inherently more sensitive than single-inoculation assays and have been used to detect small changes in fitness (7, 25). Competition assays can detect subtle phenotypes in part because the variability between host individuals is compensated for by exposing both bacterial strains to the same set of hosts, and in part because mutations subtly affecting the rate at which strains grow and persist in a specific environment can be detected, even though the strains may reach the same final population levels if inoculated singly. In separate experiments, we found that strain EVS401 (pilA::kanR) was outcompeted by ES114 between 1.7- and 10-fold (Fig. 6). Notably, the competitive defect of EVS401 tended to be greater when the total inoculum was within the range (200 to 2,000 cells/ml) estimated for natural V. fischeri populations in E. scolopes habitats (23, 24) and was attenuated by successively higher inoculum densities (Fig. 6), underscoring the importance of using natural-infection models. The sixfold variability between experiments could be due to differences in the source of host animals, as both the squid genotype and the embryonic age of the animals were subject to variation in this study.
This observed defect in competitiveness would have a dramatic impact on the abundance of pilA strains in the environment on an evolutionary time scale. Assuming (i) a conservative 1.7-fold relative decrease in the population level per squid generation, (ii) an average squid generation time of 6 months, and (iii) that, as extrapolated by Lee and Ruby (24), the ability to colonize E. scolopes is a primary factor determining environmental population levels of V. fischeri in Hawaii, a pilA mutant would be selected against 106-fold in just 15 years. Therefore, although the symbiotic defect of pilA mutants may appear subtle in these laboratory studies, this locus may be a critical fitness determinant for V. fischeri in the environment.
Strains may be outcompeted for entry into the light organ (31), during growth in the crypt spaces, and/or during subsequent daily expulsion and regrowth (6). Our data suggest that competition during the initial stages of infection may contribute to the pilA competition defect. Specifically, at low inoculum densities, at which only half of the juvenile squid became infected, most of the squid were colonized by only one of the two strains in the mixed inoculum yet ES114 still outcompeted EVS401 an average of twofold (Table 1). The observation of a competition defect in clonally skewed infections suggests competition for light organ entry rather than competition between strains in an ongoing mixed infection of the crypt spaces. This supposition is supported by the observation that within a group of inoculated squid, the RCI did not change appreciably when assayed at 24, 48, or 72 h postinfection (data not shown), which suggests that no further competitive selection occurred after the first day. The initial stages of infection are marked by aggregation and adherence of V. fischeri outside the light organ, followed by motility through an animal-derived mucus matrix into the light organ (31). Because pili are known to mediate both adherence and twitching motility, pilA might contribute to either or both of these early infection processes. The behavior of bacteria and their interactions with the host during early infection events were recently elucidated by Nyholm et al. (31) through the microscopic observation of strains labeled with green fluorescent protein (GFP). Similarly, dual labeling of ES114 and pilA mutants with distinct fluorescent markers (e.g., GFP and red fluorescent protein) would allow a visual analysis of interstrain competition during the initiation of symbiotic infection.
mshA mutants of V. cholerae are unaffected both in virulence assays and in colonization of the human intestine (2, 38, 39) but are compromised in the ability to colonize invertebrate zooplankton hosts (8). While both mshA and V. fischeri pilA apparently contribute to the colonization of invertebrates by cells of Vibrio spp., these genes are distinct in five ways. First, V. fischeri pilA does not appear to be clustered with genes involved in pilin biogenesis (Fig. 1). Second, V. fischeri PilA lacks the glycine residue present at the site of prepilin processing in other pili, suggesting a unique mechanism of prepilin processing (Fig. 2). Third, in contrast to mshA mutants (18, 45), pilA mutants did not appear compromised in the ability to attach to either borosilicate or mannose-coated agarose beads (data not shown). These discrepancies could indicate a different function for pilA or could be due to the presence of other genes functionally redundant with pilA. Many bacteria, including V. cholerae (11), have multiple type IV pili, and preliminary data indicate that V. fischeri ES114 possesses at least one gene other than pilA that encodes a type IV pilin (B. Feliciano and E. Ruby, Abstr. 99th Gen. Meet. Am. Soc. Microbiol. 1999, abstr. N-75, p. 462, 1999). Fourth, while V. cholerae pilins may be the known proteins most similar to PilA, there is considerable sequence divergence between them. Finally, while the mshA gene has clearly been linked to a pilus structure in V. cholerae, further experiments are necessary to determine whether the V. fischeri pilA gene produces a surface-exposed pilin protein, as its sequence suggests.
Several avenues of research will help define the role(s) of pilA and pili in the V. fischeri-E. scolopes symbiosis. Comparisons of light organ infections by GFP-labeled wild-type and pilA mutant strains by fluorescence microscopy may reveal the stage at which PilA contributes to symbiotic competence. Elucidation of PilA structure and expression will help determine its function and the conditions under which it is advantageous to V. fischeri. Elucidation of the role (if any) of pilA or other genes (e.g., a homologue of pilT) in twitching motility could establish a relationship between this mode of locomotion and host colonization. Perhaps most importantly, we expect that genomic analysis of V. fischeri ES114 will lead to the discovery of other potential pilins and adhesins that can be targeted for mutation and through the generation of multiple mutants the presence of functional overlap in these systems can be assessed. Together, these experimental approaches will help elucidate the mechanisms by which bacteria compete to colonize their hosts in mutualistic bacterium-animal interactions.
Acknowledgments
We thank Katanuya Kapeli for technical assistance, Michelle Nishiguchi for providing strains, and Janice Flory, Deborah Millikan, and Lawrence Shimkets for insightful comments on the manuscript.
This work was supported by National Science Foundation grant IBN-9904601 and by National Research Service award GM20041 from the National Institutes of Health.
REFERENCES
- 1.Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. [DOI] [PubMed] [Google Scholar]
- 2.Attridge, S. R., P. A. Manning, J. Holmgren, and G. Jonson. 1996. Relative significance of mannose-sensitive hemagglutinin and toxin-coregulated pili in colonization of infant mice by Vibrio cholerae El Tor. Infect. Immun. 64:3369-3373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bittinger, M. A., J. L. Milner, B. J. Saville, and J. Handelsman. 1997. rosR, a determinant of nodulation competitiveness in Rhizobium etli. Mol. Plant-Microbe Interact. 10:180-186. [DOI] [PubMed] [Google Scholar]
- 4.Boettcher, K. J., and E. G. Ruby. 1990. Depressed light emission by symbiotic Vibrio fischeri of the sepiolid squid Euprymna scolopes. J. Bacteriol. 172:3701-3706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Boettcher, K. J., and E. G. Ruby. 1994. Occurrence of plasmid DNA in the sepiolid squid symbiont Vibrio fischeri. Curr. Microbiol. 29:279-286. [Google Scholar]
- 6.Boettcher, K. J., E. G. Ruby, and M. J. McFall-Ngai. 1996. Bioluminescence in the symbiotic squid Euprymna scolopes is controlled by a daily biological rhythm. J. Comp. Physiol. 179:65-73. [Google Scholar]
- 7.Brandl, M. T., and S. E. Lindow. 1998. Contribution of indole-3-acetic acid production to the epiphytic fitness of Erwinia herbicola. Appl. Environ. Microbiol. 64:3256-3263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chiavelli, D. A., J. W. Marsh, and R. K. Taylor. 2001. The mannose-sensitive hemagglutinin of Vibrio cholerae promotes adherence to zooplankton. Appl. Environ. Microbiol. 67:3220-3225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Dowling, D. N., and W. J. Broughton. 1986. Competition for nodulation of legumes. Annu. Rev. Microbiol. 40:131-157. [DOI] [PubMed] [Google Scholar]
- 10.Fidopiastis, P. M., S. Von Boletzky, and E. G. Ruby. 1998. A new niche for Vibrio logei, the predominant light organ symbiont of squids in the genus Sepiola. J. Bacteriol. 180:59-64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Fullner, K. J., and J. J. Mekalanos. 1999. Genetic characterization of a new type IV-A pilus gene cluster found in both classical and El Tor biotypes of Vibrio cholerae. Infect. Immun. 67:1393-1404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Graf, J., P. V. Dunlap, and E. G. Ruby. 1994. Effect of transposon-induced motility mutations on colonization of the host light organ by Vibrio fischeri. J. Bacteriol. 176:6986-6991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Graf, J., and E. G. Ruby. 1998. Host-derived amino acids support the proliferation of symbiotic bacteria. Proc. Natl. Acad. Sci. USA 95:1818-1822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hanahan, D. 1983. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 166:557-580. [DOI] [PubMed] [Google Scholar]
- 15.Henikoff, S., and J. G. Henikoff. 1992. Amino acid substitution matrices from protein blocks. Proc. Natl. Acad. Sci. USA 89:10915-10919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hensey, S., and M. McFall-Ngai. 1992. A surface peptide of the bacterium Vibrio fischeri plays a key role in specificity and recognition in the symbiosis with the squid Euprymna scolopes. Am. Zool. 32:37. [Google Scholar]
- 17.Herrero, M., V. De Lorenzo, and K. N. Timmis. 1990. Transposon vectors containing non-antibiotic resistance selection markers for cloning and stable chromosomal insertion of foreign genes in gram-negative bacteria. J. Bacteriol. 172:6557-6567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jonson, G., J. Holmgren, and A. M. Svennerholm. 1991. Identification of a mannose-binding pilus on Vibrio cholerae El Tor. Microb. Pathog. 11:433-441. [DOI] [PubMed] [Google Scholar]
- 19.Jonson, G., M. Lebens, and J. Holmgren. 1994. Cloning and sequencing of Vibrio cholerae mannose-sensitive haemagglutinin pilin gene: localization of mshA within a cluster of type 4 pilin genes. Mol. Microbiol. 13:109-118. [DOI] [PubMed] [Google Scholar]
- 20.Kuroki, H., C. Toma, N. Nakasone, T. Yamashiro, and M. Iwanaga. 2001. Gene analysis of Vibrio cholerae NAGV14 pilus and its distribution. Microbiol. Immunol. 45:417-424. [DOI] [PubMed] [Google Scholar]
- 21.Lee, K.-H. 1994. Ecology of Vibrio fischeri, the light organ symbiont of the Hawaiian sepiolid squid Euprymna scolopes. Ph.D. thesis. University of Southern California, Los Angeles.
- 22.Lee, K.-H., and E. G. Ruby. 1994. Competition between Vibrio fischeri strains during initiation and maintenance of a light organ symbiosis. J. Bacteriol. 176:1985-1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lee, K.-H., and E. G. Ruby. 1992. Detection of the light organ symbiont, Vibrio fischeri, in Hawaiian seawater using lux gene probes. Appl. Environ. Microbiol. 58:942-947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lee, K.-H., and E. G. Ruby. 1994. Effect of the squid host on the abundance and distribution of symbiotic Vibrio fischeri in nature. Appl. Environ. Microbiol. 60:1565-1571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lenski, R. E. 1992. Relative Fitness: its estimation and its significance for environmental applications of microorganisms, p. 183-198. In M. A. Levin, R. J. Seidler, and M. Rogul (ed.), Microbial ecology: principles, methods, and applications. McGraw-Hill Book Co., New York, N.Y.
- 26.Marsh, J. W., D. Sun, and R. K. Taylor. 1996. Physical linkage of the Vibrio cholerae mannose-sensitive hemagglutinin secretory and structural subunit gene loci: identification of the mshG coding sequence. Infect. Immun. 64:460-465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.McFall-Ngai, M., C. Brennan, V. Weis, and L. Lamarcq. 1998. Mannose adhesin-glycan interactions in the Euprymna scolopes-Vibrio fischeri symbiosis, p. 273-276. In L. Le Gal and H. O. Halvorson (ed.), New developments in marine biotechnology. Plenum Press, New York, N.Y.
- 28.McFall-Ngai, M. J., and E. G. Ruby. 1991. Symbiont recognition and subsequent morphogenesis as early events in an animal-bacterial symbiosis. Science 254:1491-1494. [DOI] [PubMed] [Google Scholar]
- 29.Miller, J. H. 1992. A short course in bacterial genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 30.Nishiguchi, M. K., E. G. Ruby, and M. J. McFall-Ngai. 1998. Competitive dominance among strains of luminous bacteria provides an unusual form of evidence for parallel evolution in sepiolid squid-vibrio symbioses. Appl. Environ. Microbiol. 64:3209-3213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Nyholm, S. V., E. V. Stabb, E. G. Ruby, and M. J. McFall-Ngai. 2000. Establishment of an animal-bacterial association: recruiting symbiotic vibrios from the environment. Proc. Natl. Acad. Sci. USA 97:10231-10235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ruby, E. G. 1996. Lessons from a cooperative bacterial-animal association: the Vibrio fischeri-Euprymna scolopes light organ symbiosis. Annu. Rev. Microbiol. 50:591-624. [DOI] [PubMed] [Google Scholar]
- 33.Ruby, E. G., and L. M. Asato. 1993. Growth and flagellation of Vibrio fischeri during initiation of the sepiolid squid light organ symbiosis. Arch. Microbiol. 159:160-167. [DOI] [PubMed] [Google Scholar]
- 34.Siegal, S. 1956. Nonparametric statistics for the behavior sciences. McGraw-Hill Book Co., New York, N.Y.
- 35.Stabb, E. V., K. A. Reich, and E. G. Ruby. 2001. Vibrio fischeri genes hvnA and hvnB encode secreted NAD+-glycohydrolases. J. Bacteriol. 183:309-317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Stabb, E. V., and E. G. Ruby. 2002. New RP4-based plasmids for conjugation between Escherichia coli and members of the Vibrionaceae. Methods Enzymol. 358:413-426. [DOI] [PubMed]
- 37.Strom, M. S., and S. Lory. 1993. Structure-function and biogenesis of the type IV pili. Annu. Rev. Microbiol. 47:565-596. [DOI] [PubMed] [Google Scholar]
- 38.Tacket, C. O., R. K. Taylor, G. Losonsky, Y. Lim, J. P. Nataro, J. B. Kaper, and M. M. Levine. 1998. Investigation of the roles of toxin-coregulated pili and mannose-sensitive hemagglutinin in the pathogenesis of Vibrio cholerae O139 infection. Infect. Immun. 66:692-695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Thelin, K. H., and R. K. Taylor. 1996. Toxin-coregulated pilus, but not mannose-sensitive hemagglutinin, is required for colonization by Vibrio cholerae O1 El Tor biotype and O139 strains. Infect. Immun. 64:2853-2856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Thompson, J. D., D. G. Higgins, and T. J. Gibson. 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22:4673-4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Triplett, E. W., and M. J. Sadowsky. 1992. Genetics of competition for nodulation of legumes. Annu. Rev. Microbiol. 46:399-428. [DOI] [PubMed] [Google Scholar]
- 42.Vieira, J., and J. Messing. 1982. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene 19:259-268. [DOI] [PubMed] [Google Scholar]
- 43.Visick, K. L., and M. J. McFall-Ngai. 2000. An exclusive contract: specificity in the Vibrio fischeri-Euprymna scolopes partnership. J. Bacteriol. 182:1779-1787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Visick, K. L., and E. G. Ruby. 1998. The periplasmic, group III catalase of Vibrio fischeri is required for normal symbiotic competence and is induced both by oxidative stress and approach to stationary phase. J. Bacteriol. 180:2087-2092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Watnick, P. I., K. J. Fullner, and R. Kolter. 1999. A role for the mannose-sensitive hemagglutinin in biofilm formation by Vibrio cholerae El Tor. J. Bacteriol. 181:3606-3609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.You, Z., X. Gao, M. M. Ho, and D. Borthakur. 1998. A stomatin-like protein encoded by the slp gene of Rhizobium etli is required for nodulation competitiveness on the common bean. Microbiology 144:2619-2627. [DOI] [PubMed] [Google Scholar]