Abstract
During chemotactic signaling by Escherichia coli, autophosphorylation of the histidine kinase CheA is coupled to chemoreceptor control by the CheW protein, which interacts with the C-terminal P5 domain of CheA. To identify P5 determinants important for CheW binding and receptor coupling control, we isolated and characterized a series of P5 missense mutants. The mutants fell into four phenotypic groups on the basis of in vivo behavioral and protein stability tests and in vitro assays with purified mutant proteins. Group 1 mutants exhibited autophosphorylation and receptor-coupling defects, and their CheA proteins were subject to relatively rapid degradation in vivo. Group 1 mutations were located at hydrophobic residues in P5 subdomain 2 and most likely caused folding defects. Group 2 mutants made stable CheA proteins with normal autophosphorylation ability but with defects in CheW binding and in receptor-mediated activation of CheA autophosphorylation. Their mutations affected residues in P5 subdomain 1 near the interface with the CheA dimerization (P3) and ATP-binding (P4) domains. Mutant proteins of group 3 were normal in all tests yet could not support chemotaxis, suggesting that P5 has one or more important but still unknown signaling functions. Group 4 mutant proteins were specifically defective in receptor-mediated deactivation control. The group 4 mutations were located in P5 subdomain 1 at the P3/P3′ interface. We conclude that P5 subdomain 1 is important for CheW binding and for receptor coupling control and that these processes may require substantial motions of the P5 domain relative to the neighboring P3 and P4 domains of CheA.
The histidine kinase CheA plays a central signaling role in bacterial chemotaxis pathways (see references 48 and 49 for recent reviews). In Escherichia coli, CheA forms signaling complexes with five membrane-associated chemoreceptors, known as methyl-accepting chemotaxis proteins (MCPs), that communicate with the flagellar motors by controlling CheA activity. CheA autophosphorylates at a histidine residue, using ATP as the phosphodonor, and subsequently donates those phosphoryl groups to two response regulators, CheY and CheB. Phosphorylated CheY (phospho-CheY) interacts with the switching machinery at the base of the flagellar motors to promote clockwise (CW) rotation, which causes random turning episodes or tumbles during swimming. Phospho-CheB functions in a feedback adaptation circuit that enables the cells to detect temporal changes in attractant and repellent concentrations as they move through spatial chemical gradients.
Chemoeffectors, sensed by the chemoreceptors, modulate CheA activity over a wide range. Unliganded receptors activate CheA several hundredfold over its basal, receptor-uncoupled autophosphorylation rate, whereas attractant-bound receptors deactivate CheA to about its basal activity level (8, 9, 26, 33, 45). CheA autophosphorylation control occurs in ternary complexes formed between the cytoplasmic signaling domains of the chemoreceptors, CheA, and the coupling protein CheW, which is essential for chemotactic behavior in vivo and for receptor-mediated activation of CheA in vitro. The role of CheW in receptor coupling control of CheA is poorly understood, owing to a paucity of structural information about the receptor signaling complex. However, in vitro CheW binds both to receptors (10, 17, 25, 29, 42) and to CheA (10, 16, 31, 38), so it may serve to couple ligand-induced conformational changes in the receptor to corresponding structural changes in CheA that allosterically regulate its activity. A better understanding of the CheA-CheW binding interaction could provide useful insights into the mechanism of receptor-mediated control of CheA.
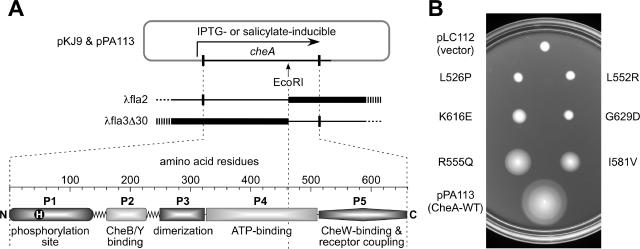
The CheA protein of E. coli functions as a homodimer; the 654-residue subunits have a modular architecture consisting of five functional domains (Fig. 1A). Autophosphorylation involves interaction of the N-terminal phosphorylation site domain (P1) in one subunit with the ATP-binding domain (P4) of the other subunit (46, 47, 51). The principal dimerization determinants lie in the central P3 domain (5, 23). The C-terminal P5 domain is not required for autophosphorylation but is essential for coupling CheA activity to chemoreceptor control (12) and contains determinants for a CheA-CheW binding interaction (11, 38). CheA molecules lacking part or all of the P5 domain cannot undergo activation by chemoreceptors (12) and fail to bind to CheW molecules bearing an N-terminal fluorescein (11) or glutathione S-transferase (GST) affinity handle (38). Thus, the P5 domain is most likely the binding target for CheW, but the CheW-binding determinants and the location of the binding surface on the P5 domain have not been established.
FIG. 1.
(A) CheA domain structure and genetic tools used in this study. Transducing phages λfla2 and λfla3Δ30 carry adjacent segments of the cheA gene, delineated by an EcoRI site. Thin lines denote cheA coding material carried in the phages; thick lines indicate deleted material. (B) Chemotaxis defects of CheA P5 missense mutants isolated in this study. Mutant derivatives of plasmid pPA113 were tested in strain RP9535 (ΔcheA) for ability to support chemotaxis on tryptone soft agar plates. The plates contained 0.4 μM sodium salicylate and 12.5 μg/ml chloramphenicol and were photographed after 8 h of incubation at 32.5°C. The L526P and L552R mutants are group 1 mutants, the K616E and G629D mutants are group 2 mutants, and the R555Q and I581V mutants are group 3 mutants. WT, wild type.
To better understand the CheA-CheW binding interaction that is correlated with ternary-complex formation and chemoreceptor control of CheA, we isolated and characterized P5 missense mutations that affect CheA function in vivo. In vitro studies of the mutant CheA proteins enabled us to identify P5 structural determinants important for CheW binding and for receptor-coupling control. The CheW-binding determinants defined by CheA P5 missense mutations are quite distinct from the binding surfaces previously predicted by Bilwes et al. (5) and Shimizu et al. (41) on the basis of less-direct evidence. The CheW-binding surface defined by our P5 mutant studies appears to be largely inaccessible in the X-ray structure of Thermotoga maritima CheA. We propose that, in solution at least, the P5 domain of E. coli CheA adopts a different orientation relative to the neighboring P3 and P4 domains. Moreover, relative movements of these domains probably play an important role in the assembly and subsequent operation of receptor signaling complexes.
MATERIALS AND METHODS
Bacterial strains and plasmids.
The following strains are closely related derivatives of E. coli K-12 strain RP437 (37): RP9535 (ΔcheA1643) (39), RP3098 [Δ(flhD-flhB)4] (44), RP9540 [ΔcheA1643 Δtsr-7028 Δ(tar-tap)5021 Δtrg-1] (28), RP9542 (ΔcheA1643 ΔcheZ6725) (28), and RP9543 [ΔcheA1643 Δtsr-7028 Δ(tar-tap)5021 Δtrg-1 ΔcheZ6725] (32). Strain RP526 carries the mutD5 mutator allele (14).
Plasmid pKJ9 confers ampicillin resistance and expresses functional CheA inducible by isopropyl-β-d-thiogalactopyranoside (IPTG) (15, 21). This high-copy plasmid was used for purification of CheA proteins for in vitro studies. Plasmid pPA113, obtained from Peter Ames (University of Utah), confers chloramphenicol resistance and expresses functional CheA inducible by sodium salicylate. It was constructed by inserting the PCR-amplified cheA gene between the NdeI and KpnI restriction sites of expression vector pLC112 (2). This plasmid was used for in vivo assays of CheA function.
Plasmid pCJ30 is an IPTG-inducible expression vector derived from pBR322 (4, 6); pJC3 is a derivative of pCJ30 that carries wild-type tsr (13); pAR1.CheY, kindly provided by Rick Stewart (University of Maryland), expresses cheY from an IPTG-inducible promoter; pPA770, obtained from Peter Ames (University of Utah), expresses cheW from an IPTG-inducible promoter; and pPA117 was generated by inserting cheW into pGEX-3X, a GST gene fusion vector obtained from Amersham Biosciences.
Growth media.
Chemotactic ability was assessed on semisolid tryptone medium (tryptone soft agar), consisting of tryptone broth (10 g tryptone, 5 g NaCl per liter) and 0.27% agar. L broth (tryptone broth plus 0.5% yeast extract) was generally used for growth of bacterial strains. H1 minimal salts medium (35) contained 1% Casamino Acids, 0.4% glycerol, and required amino acids (1 mM each) and was used as the growth medium for protein purification and protein stability assays. IPTG and sodium salicylate were purchased from Promega Corp. Ampicillin and chloramphenicol were purchased from Sigma Chemical Co. and used at 100 μg/ml and 25 μg/ml, respectively, in solid and liquid media, except in soft-agar chemotaxis assay plates, where their concentrations were halved.
CheA P5 mutant hunt.
Plasmids pKJ9 and pPA113 were subjected to random mutagenesis by propagation in RP526, a proofreading-deficient polymerase mutant (14). To identify RP526 clones that retained high mutator activity, single colonies were grown overnight in L broth at 37°C and approximately 108 cells were spread on L plates containing 30 μg/ml nalidixic acid. Cultures that exhibited high frequencies (at least 10−5) of nalidixic acid-resistant mutants were used as plasmid hosts.
Independent plasmid pools from RP526 were transferred to strain RP9535 by CaCl2 transformation. Samples of the transformation reaction mixture were added to an empty petri dish, mixed with 25 ml of tryptone broth containing 0.2% agar and selective antibiotic. These maxiswarm plates were allowed to stand at room temperature for 1 to 2 h to gel and then incubated at 35°C. After overnight growth, the plates were screened for small, nonchemotactic colonies embedded in a diffuse background of chemotactic cells that had spread throughout the plates. (The inoculum size was adjusted to yield about 5,000 to 10,000 transformant colonies per plate.) Candidate mutants were single colony purified and retested for chemotaxis defects on tryptone soft agar at 32.5°C for 8 h.
Crude mapping tests with λfla transducing phages (36) were used to identify mutant plasmids with mutations in the P5 coding segment of cheA. RP9535 cells carrying candidate plasmids were picked to tryptone soft agar plates containing ∼109 particles of λfla2 or λfla3Δ30 (Fig. 1) and scored for formation of chemotactic recombinants after incubation at 32.5°C for 8 h. Mutant plasmids that recombined with λfla3Δ30 but not λfla2 were kept for DNA sequencing, performed at the Protein-DNA Core Facility at the University of Utah.
Pseudotaxis assays for CheA function.
Mutant pPA113 plasmids were introduced by transformation into RP9540, RP9543, and RP9542, with selection on L plates containing 25 μg/ml chloramphenicol. After overnight incubation at 37°C, single colonies were picked to tryptone soft agar plates containing various concentrations of sodium salicylate and 12.5 μg/ml chloramphenicol. Colony sizes were compared after incubation for 12 to 17 h at 32.5°C.
CheA in vivo stability test.
Strain RP9535 carrying the mutant pKJ9 or pPA113 plasmid was grown to mid-log phase (optical density at 600 nm of 0.5) in H1 medium at 35°C. CheA expression was induced by addition of 1 mM IPTG (pKJ9) or 20 μM sodium salicylate (pPA113), and the cells were grown for an additional 2 hours at 35°C. Cells were then collected by centrifugation, washed once with 5 volumes of H1 medium containing no amino acid supplements or other carbon and energy sources, and resuspended in an equal volume of the same medium. For cells carrying pKJ9 plasmids, 100 μg/ml chloramphenicol was also added to further prevent protein synthesis. The cells were incubated at 35°C with shaking, and 1-ml samples were collected at various time points, flash frozen, and stored at −70°C until all samples had been collected. Cells were then broken by three cycles of freezing and boiling in sodium dodecyl sulfate (SDS) sample buffer (24). The cell extracts were subjected to 10% SDS-polyacrylamide gel electrophoresis (SDS-PAGE), and the amount of full-length CheA protein at each time point was determined by quantitative Western blotting using a polyclonal anti-CheA rabbit antiserum, provided by Phil Matsumura (University of Illinois—Chicago).
Purification of mutant CheA proteins.
Wild-type and all mutant CheA proteins were expressed from plasmid pKJ9 in strain RP3098 and purified by previously described methods (20). Briefly, cells were grown to mid-log phase (optical density at 600 nm of ∼0.5) in H1 medium at 35°C. CheA expression was induced with 1 mM IPTG, and the cells were grown for an additional 4 hours at 35°C. Cells were harvested by centrifugation, washed three times in 25 mM Tris (pH 7.5) plus 5 mM EDTA, and resuspended in TEDG-10 buffer (50 mM Tris-HCl [pH 7.5], 2 mM dithiothreitol, 0.5 mM EDTA, and 10% glycerol). After two cycles of disruption in a French press, cellular debris was removed by centrifugation for 25 min at 10,000 × g and the membranes were removed by centrifugation for 1 hour at 100,000 × g. The CheA protein in the clarified lysate was concentrated by a 25 to 40% ammonium sulfate precipitation and then purified on a 25-ml DEAE (DEAE cellulose) ion-exchange column with elution at ∼200 mM NaCl. The column fractions were collected and concentrated by ammonium sulfate precipitation and applied to a 400-ml S300 gel filtration column. CheA-containing fractions were concentrated again, dialyzed twice against 2 liters of TEDG-10 buffer for 12 h at 4°C, and stored at −70°C. Phenylmethylsulfonyl fluoride (1 mM) was used as a protease inhibitor throughout the purification process.
In vitro assays of mutant CheA proteins.
CheA autophosphorylation was assessed at a 1-μM concentration in phosphorylation buffer (50 mM Tris-HCl [pH 7.5], 50 mM KCl, 5 mM MgCl2) at room temperature as previously described (1). CheA activation in ternary signaling complexes was assessed with reaction mixtures containing purified CheW and CheY proteins, prepared as described previously (1, 30), and membranes containing the serine receptor Tsr, prepared as described previously (7). Coupling reaction mixtures contained 0.5 μM CheA, 2 μM CheW, 25 μM CheY, and 4 μM Tsr in membrane, and reactions were carried out as described previously (32). CheA deactivation was measured in the same manner as activation except that the reaction mixtures also contained 1 mM serine.
CheA-CheW binding assays.
GST-CheW and GST proteins were, respectively, expressed from pPA117 and pGEX-3X in strain RP3098 and purified according to the standard protocol provided by Amersham Biosciences. For CheA-CheW pull-down assays, glutathione-Sepharose beads were washed three times by being mixed with 10 volumes of phosphorylation buffer and pelleted in a table-top centrifuge. Washed beads (50 μl) were mixed with 50 μl of phosphorylation buffer containing either 430 μg GST-CheW or 250 μg GST and incubated at 4°C for at least 2 hours. Just before use, beads were washed three times with 10 volumes of wash buffer to remove unbound GST proteins. Washed beads were mixed with an equal volume of wash buffer, and the bead suspension was distributed to small centrifuge tubes in 10-μl aliquots. CheA samples (30 μl for each binding test) were adjusted to 50 μM with phosphorylation buffer, incubated at room temperature for 30 min, and then mixed with the beads and incubated at room temperature for 1 hour. Bead-plus-CheA mixtures were pelleted and then washed two to three times with 60 volumes of wash buffer. After removal of the final supernatant, beads were mixed with 30 μl of SDS sample buffer and boiled for 5 minutes. Samples were analyzed by SDS-PAGE, and proteins were detected by Coomassie staining or immunoblotting for quantification.
RESULTS
Isolation of CheA P5 missense mutations.
To identify determinants in the P5 domain essential for CheW binding, we first isolated an extensive collection of cheA missense mutations that impaired chemotactic ability, based on our assumption that the P5-dependent in vitro binding interaction with CheW would be important for CheA function in vivo. Mutations were isolated from two cheA plasmids, pKJ9 and pPA113. Most of them were obtained from pPA113, whose tightly controlled, salicylate-inducible CheA expression level made it more suitable for in vivo function tests. Both plasmids were mutagenized by passage through a mutD host (RP526), transformed into a ΔcheA tester strain (RP9535), and screened for chemotaxis defects by a variation of the miniswarm plate method (35). Altogether, 200 independent plasmid pools (50 for pKJ9 and 150 for pPA113) were screened for nonchemotactic mutants, which occurred at a frequency of about 10−3 after mutD mutagenesis. To identify possible P5 mutations, about 8 to 10 mutant isolates from each pool were subjected to mapping tests with two λfla transducing phages that carry different portions of the cheA coding region (Fig. 1A). We looked for plasmids that produced che+ recombinants with λfla3Δ30, which carries the entire P5 coding region and the C-terminal portion of the P4 coding region (36, 43). At most, one such plasmid was kept for DNA sequence analysis from each pool to ensure that all mutations had independent origins.
The P5 mutant hunt yielded a total of 95 independent isolates. Of these, eight carried P5 nonsense mutations, three carried double mutations, and nine had mutations in the P4 coding region. The remaining 75 isolates carried P5 missense (P5*) mutations representing 16 amino acid replacements at 14 different residues (Table 1, mutant groups 1 to 3). Because each mutant amino acid change was found independently at least three times and on average nearly six times, we conclude that the P5 mutant hunt is saturated for the chemotaxis-defective phenotypes we sought.
TABLE 1.
Summary of CheA P5* mutant properties
| Mutant group | CheA P5* mutant(s) | No. of isolatesa | Result of in vivo testb
|
Result of in vitro testc
|
||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Chemotaxis | Stability | Auto-P | Activation | Auto-P | Activation | Deactivation | CheW binding | |||
| 1 | L526P | 5 | − | − | ± | ± | NAd | NA | NA | NA |
| L542P | 7 | − | − | ± | ± | NA | NA | NA | NA | |
| L545P | 4 | − | − | ± | ± | NA | NA | NA | NA | |
| L552R | 7 | − | − | ± | ± | NA | NA | NA | NA | |
| L559P | 7 | − | − | − | − | NA | NA | NA | NA | |
| I584N | 3 | − | − | ± | ± | NA | NA | NA | NA | |
| Q641H | 6 | − | − | ± | ± | NA | NA | NA | NA | |
| 2 | K616E | 6 | − | + | ± | ± | + | − | NA | − |
| A622V | 3 | − | + | + | ± | + | − | NA | − | |
| G629D | 6 | − | + | + | ± | + | − | NA | − | |
| V631M | 3 | − | + | + | ± | + | − | NA | − | |
| 3 | R555Q, R555W | 5, 3 | ± | + | + | + | + | + | + | + |
| I581T, I581V | 9, 13 | ± | + | + | + | + | + | + | + | |
| G588S | 8 | − | + | + | + | + | + | + | + | |
| 4 | V606M | NA | + | + | + | + | + | + | − | + |
| G627D, G627S | NA | + | + | + | + | + | + | − | + | |
Number of independent isolates of a particular mutation.
The results of the in vivo function tests were scored as follows. For chemotaxis, −, less than 30% of the level of the wild-type control; ±, 30 to 50%; and +, greater than 50%. For protein stability, −, degradation rate at least twice that of the wild type and +, degradation rate not significantly different than that of the wild type. For autophosphorylation (auto-P), −, no detectable activity compared to that of the wild-type control; ±, partial activity; and +, activity. For receptor activation, −, no detectable activity compared to that of the wild-type control; ±, partial activity; and +, activity.
The results of the in vitro assays are given in Table 2 and are classified as follows. For autophosphorylation (auto-P), +, greater than 50% of the wild-type autophosphorylation rate. For activation, −, less than 50% of the wild-type level of activation and +, greater than 50%. For deactivation, −, deactivated level greater than 15% of the activated level and +, deactivated level less than 5% of the activated level. For CheW binding, −, less than 15% of the wild-type pull-down amount and +, more than 40%.
NA, not applicable.
We chose a representative for each mutant type and determined the DNA sequence of its entire cheA gene as well as the flanking upstream and downstream regions to confirm that each had only the identified base pair change in the P5 portion of the coding region. Because these mutants were obtained on the basis of a substantial chemotaxis defect, we also included for subsequent characterization three P5 mutants previously isolated as phenotypic suppressors of receptor defects (28). Those mutant proteins (V606M, G627D, and G627S) support about 70% of wild-type chemotactic ability and, consequently, were not found in our P5 mutant hunt. The signaling properties of all P5 missense mutants are summarized in Table 1 and are discussed in detail below.
Chemotactic ability of CheA P5 mutants.
Mutant pPA113 plasmids were used to express CheA P5 missense proteins in strain RP9535 (ΔcheA). Chemotactic ability of the strains was evaluated by colony morphology in tryptone soft agar containing 0.4 μM sodium salicylate, the inducer concentration that produced optimal complementation by wild-type pPA113. Representative mutant phenotypes are shown in Fig. 1B. All P5* mutants except the V606M, G627D, and G627S mutants were, as expected, significantly defective in supporting chemotaxis (Table 1). The nature of those chemotaxis defects was investigated with additional in vivo and in vitro functional tests.
In vivo autophosphorylation activity of P5* mutants.
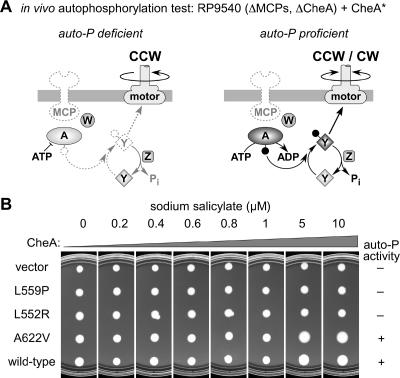
Two tests were used to evaluate the in vivo autophosphorylation ability of the mutant CheA proteins. Both tests exploited the phenomenon of pseudotaxis, which measures the ability of nonchemotactic cells to maneuver through soft agar by virtue of their unmodulated pattern of flagellar rotation. Cells that incessantly tumble (CW rotation) or that move only in forward runs (counterclockwise [CCW] rotation) spread more slowly than do cells that alternate running and tumbling episodes (3, 50). In the first test, mutant pPA113 plasmids were transferred into strain RP9540, which is deleted for the chromosomal cheA and four MCP chemoreceptor genes (Fig. 2). Although the aerotaxis transducer Aer is still present in RP9540, Aer alone does not detectably activate CheA, owing to its low abundance (27). Nevertheless, receptor-uncoupled CheA molecules that are able to autophosphorylate can contribute to phospho-CheY production in RP9540. At a sufficiently high expression level, basal CheA autophosphorylation activity generates enough phospho-CheY to cause a significant CW shift in flagellar rotation pattern, resulting in enhanced pseudotactic spreading (e.g., wild-type and A622V proteins [Fig. 2]). In contrast, autophosphorylation-deficient CheA proteins cause slow colony expansion at all expression levels (e.g., vector control and L559P and L552R proteins [Fig. 2]). The RP9540 test allowed us to make a provisional classification of the P5* mutants as either competent or incompetent for autophosphorylation.
FIG. 2.
In vivo test of CheA autophosphorylation. Mutant CheA plasmids (CheA*) were tested over a range of inducer concentrations for ability to produce pseudotactic changes in the colony size of strain RP9540 on tryptone soft agar. (A) Components of the chemotaxis signaling pathway present in RP9540 that can be used by autophosphorylation (auto-P)-proficient CheA molecules to induce CW flagellar rotation. (B) Examples of different test results and their relative auto-P scores used in compiling Table 1 (see the text). Plates were incubated at 32.5°C for 17 h.
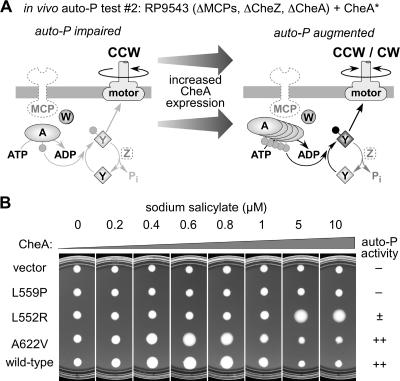
To identify CheA proteins with partial autophosphorylation activity, we also tested the P5* proteins in RP9543, a host that permits a more sensitive pseudotaxis test of autophosphorylation activity (Fig. 3). Like strain RP9540, RP9543 is deleted for the chromosomal cheA and MCP genes but in addition lacks cheZ function. The absence of CheZ raises the steady-state level of phospho-CheY by slowing its rate of dephosphorylation, thereby amplifying the effect of any CheA autophosphorylation activity that may be present. Mutant CheA proteins that were judged competent for autophosphorylation in the less sensitive RP9540 test typically caused an increase in pseudotactic spreading followed by a decrease at high expression levels in RP9543 (e.g., wild-type and A622V proteins [Fig. 3]). One autophosphorylation-incompetent protein remained so in the RP9543 test (L559P protein [Fig. 3]). However, the remainder of the seemingly autophosphorylation-incompetent P5* proteins produced detectable increases in pseudotactic spreading at elevated expression levels, demonstrating some autophosphorylation activity in the more sensitive RP9543 test (e.g., L552R protein [Fig. 3; Table 1]).
FIG. 3.
An in vivo CheA autophosphorylation (auto-P) test with sensitivity enhanced compared to the one shown in Fig. 2. In this test, mutant plasmids (CheA*) were examined for pseudotactic effects in strain RP9543, which also lacks CheZ. (A) Phosphorylation reactions and motor rotation effects expected of a partially active CheA. (B) Examples of different test results and their relative auto-P scores used to compile Table 1 (see the text). Plates were incubated at 32.5°C for 17 h.
In vivo receptor-mediated activation of P5* mutants.
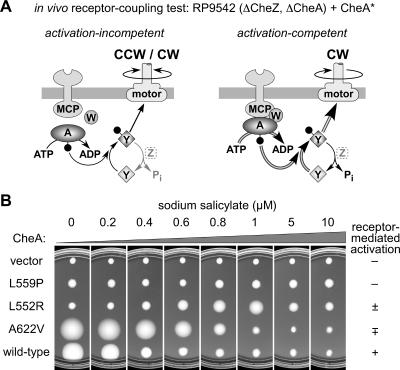
A third pseudotaxis test was devised to evaluate the ability of autophosphorylation-competent P5* proteins to undergo activation by receptors and CheW. The host for this test was RP9542, which carries chromosomal deletions in the cheA and cheZ genes but retains a full complement of chemoreceptors and CheW. As with RP9543, the absence of CheZ function makes the cellular phospho-CheY levels very sensitive to the CheA autophosphorylation rate. Accordingly, autophosphorylation-competent CheA proteins at very high expression levels should cause excessive CW rotation and reduced pseudotactic spreading as they do in the transducerless host RP9543, even if they are activation defective. However, mutant proteins that also undergo receptor-mediated activation should show maximal pseudotactic spreading at much lower expression levels, with a rapid decline in colony size as the cells become excessively CW biased at higher expression levels. Both the wild-type control and the A622V mutant illustrate such behavior (Fig. 4). Moreover, a comparison of the colony sizes produced by uninduced levels of these proteins in RP9543 (Fig. 3) and RP9542 (Fig. 4) clearly shows that the presence of transducers enhanced phospho-CheY production (as evidenced by pseudotactic spreading). However, the A622V mutant colony size declines much less rapidly than that of the wild type, indicating a substantial but evidently incomplete activation defect. The L552R mutant, which has less autophosphorylation activity than the A622V mutant (Fig. 3), also showed some activation ability in this test but at higher expression levels (Fig. 4).
FIG. 4.
In vivo test for CheA receptor-coupled activation. Mutant CheA plasmids (CheA*) were tested in strain RP9542. (A) Mutant CheA proteins able to form active ternary complexes with the receptor and CheW proteins of RP9542 should cause much higher levels of phospho-CheY and CW flagellar rotation than activation-incompetent CheA proteins. (B) Examples of different test results and their relative activation scores used to compile Table 1 (see the text). Plates were incubated at 32.5°C for 12 h.
Three other autophosphorylation-competent P5* mutants (Table 1, group 2) resembled the A622V mutant in the activation test (Fig. 4), suggesting that these four mutant proteins might be defective specifically in coupling CheA to chemoreceptor control. Five other autophosphorylation-competent mutants (Table 1, group 3) with moderate to severe chemotaxis defects nevertheless behaved normally in the activation test, indicating that their coupling control by receptors was not greatly impaired. Thus, the in vivo function tests provided no clues as to the nature of the group 3 P5* signaling defect.
In vivo stability of mutant CheA proteins.
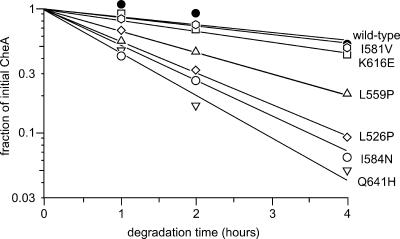
Because our mutant hunt emphasized loss-of-function phenotypes, some of the P5* proteins might be grossly misfolded or otherwise unstable in vivo. To identify unstable proteins, we tested their intracellular degradation rates after blocking protein synthesis in log-phase cultures, as detailed in Materials and Methods. We found that under these conditions wild-type CheA is degraded slowly, with over 70% of the initial molecules still present at 4 hours (Fig. 5). The mutant proteins in groups 2 to 4 (Table 1) had wild-type stabilities, but all group 1 proteins were rapidly degraded, declining to 30% or less of their initial levels (Fig. 5). The mutant residues in the group 1 proteins appear to be located predominantly at buried positions in the P5 structure (see Discussion), consistent with the possibility that these mutant proteins are misfolded.
FIG. 5.
Degradation rates of mutant CheA proteins. Cells containing plasmid-borne cheA mutations were grown with optimal induction to mid-log phase and then blocked for protein synthesis, as detailed in Materials and Methods. The cellular content of full-length CheA molecules was followed over time by quantitative immunoblotting. The lines represent least-squares best fits of the data points to a single-exponential-decay process. The examples presented here cover the range of decay rates observed with CheA P5* mutants.
In vitro activities of P5* proteins.
Mutant proteins of group 1 (Table 1), which were unstable and presumably misfolded in vivo, could not be adequately expressed or successfully purified. In contrast, mutant proteins from groups 2 to 4 (Table 1) were readily purified to at least 90% purity, as judged by SDS-PAGE and Coomassie staining. Nine representative P5* proteins were examined for various in vitro activities, with the results summarized in Table 2 and detailed below.
TABLE 2.
In vitro activities of CheA P5* mutants
| Wild type or mutant group | Wild type or CheA P5* mutant | Auto-Pa | Receptor-coupling control
|
CheW bindingd | |
|---|---|---|---|---|---|
| Activationb | Deactivationc | ||||
| Wild type | Wild type | 1.0 ± 0.1 | 191 ± 40 | 3 ± 0.5 | 100 |
| 2 | K616E | 1.1 ± 0.2 | 2.3 ± 0.7 | NA | 0 ± 4 |
| A622V | 1.9 ± 0.5 | 2.7 ± 0.3 | NA | 12 ± 4 | |
| G629D | 1.1 ± 0.2 | 1.5 ± 0.2 | NA | 7 ± 3 | |
| V631M | 1.0 ± 0.1 | 6.3 ± 4.0 | NA | 14 ± 12 | |
| 3 | R555Q | 1.2 ± 0.1 | 132 ± 15 | 4 ± 2.0 | 57 ± 6 |
| I581V | 1.2 ± 0.2 | 146 ± 18 | 3 ± 0.5 | 100 ± 5 | |
| G588S | 0.8 ± 0.3 | 123 ± 31 | 5 ± 1.3 | 42 ± 11 | |
| 4 | V606M | 1.0 ± 0.1 | 180 ± 21 | 37 ± 6 | 65 ± 7 |
| G627S | 1.1 ± 0.1 | 140 ± 13 | 26 ± 4 | 62 ± 25 | |
Pseudo-first-order rate of the receptor-uncoupled autophosphorylation (auto-P) reaction relative to that of wild-type CheA. The values are averages of two to four measurements ± standard errors of the means. The rate constant of the wild-type protein was 0.039 ± 0.004 s−1. At steady-state reaction conditions, all mutant proteins exhibited phosphorylation levels of 70% or more of the level of the wild-type protein (data not shown).
CheA activity (n-fold) relative to the receptor-uncoupled activity of the same protein. See Materials and Methods for experimental details. The values are averages of two to four independent measurements ± standard errors of the means.
CheA activity (n-fold) in the presence of 1 mM serine relative to the receptor-uncoupled activity of the same protein. The values are averages of two to four independent measurements ± standard errors of the means. NA, not applicable.
CheW binding was determined by GST-CheW pull-down assays, as detailed in Materials and Methods. Values give the percentages of the wild-type CheA pull-down amount and are averages of two to four independent measurements ± standard errors of the means.
Autophosphorylation.
All nine of the P5* proteins exhibited essentially normal autophosphorylation activity (Table 2). The autophosphorylation rate constant of the A622V protein was about twice that of the wild type; rates for the other eight P5* proteins were not significantly different from that of the wild type (Table 2). These findings are consistent with the results of the in vivo autophosphorylation tests (Table 1).
Receptor-mediated activation and deactivation.
P5* proteins were mixed with CheW and Tsr-containing membranes to evaluate their coupling control in ternary signaling complexes. Group 2 mutant proteins (K616E, A622V, G629D, and V631M), which exhibited activation defects in vivo, were also activation defective in vitro (Table 2). Whereas wild-type CheA was activated nearly 200-fold, the mutant proteins were activated at most a fewfold. In contrast, the group 3 and group 4 mutant proteins were activated 120-fold or more, consistent with their in vivo behavior. Receptor-mediated activation is reversed in the presence of attractant ligands. When tested for deactivation by serine-occupied Tsr, the group 3 mutant proteins (R555Q, I581V, and G588S) responded in the same manner as wild-type CheA, with autophosphorylation activities just a fewfold above the basal rates (Table 2). However, the group 4 mutant proteins (V606M and G627S) showed significantly less deactivation, with autophosphorylation rates more than 25-fold higher than uncoupled CheA activity (Table 2). The structural lesions in these proteins may specifically interfere with conformational changes that accompany ligand-induced down-regulation of CheA by receptor signaling complexes.
CheW binding.
P5* proteins were tested for CheW binding with a GST-CheW pull-down assay. Previous binding studies using surface plasmon resonance showed that this GST-CheW derivative bound to CheA with the same affinity as wild-type CheW and that binding was dependent on the P5 domain of CheA (38). In the pull-down assay, group 2 mutant proteins exhibited binding signals below 20% that of the wild type (Table 2). Thus, their inability to undergo receptor-mediated activation could be due to a CheW-binding defect that prevents ternary-complex formation. The group 3 and group 4 mutant proteins showed slightly reduced but nevertheless substantial binding in this assay (at least 40% of the wild-type signal). These minor deficits in CheW binding may be functionally insignificant because both groups of P5* proteins showed robust levels of receptor activation, implying normal ternary-complex formation. In this regard, the I581V protein is especially noteworthy because all of its in vitro activities, including CheW binding, are very close to those of the wild-type protein. In vivo, the I581V protein is stable and hence apparently properly folded, yet it cannot support normal chemotactic behavior. We conclude that the I581V protein is defective in a signaling function for which there is no in vitro assay at present.
DISCUSSION
Predicted structure of the E. coli CheA P5 domain.
The three-dimensional structure of the E. coli CheA P5 domain has not been experimentally determined. However, its primary structure is 32% identical to CheA P5 of Thermotoga maritima, for which an X-ray structure is available (5). Accordingly, we derived atomic coordinates for the P3-P4-P5 domains of E. coli CheA based on the corresponding T. maritima CheA coordinates, using the SWISS-MODEL protein modeling server (40). The modeled E. coli protein is not significantly different in structure from the T. maritima protein (∼0.2-Å root mean square deviation for backbone atoms), and we used the E. coli coordinates to compare the structural and functional features of the CheA P5 mutants.
Previously proposed CheW-binding regions in P5.
Two folding subdomains related by pseudo-twofold symmetry comprise the CheA P5 domain (5, 18, 19) (Fig. 6B). Both resemble the SH3 domain of human c-Src kinase, a structural motif that promotes protein-protein interactions in various contexts (22, 52). Interestingly, P5's binding partner, CheW, is also an SH3-like protein, closely related both evolutionarily and structurally to P5 (5, 18, 19). Two hydrophobic surface patches in subdomain 2 have been suggested, on the basis of rather indirect evidence, to be the binding surface for CheW (5, 41) (Fig. 6A). Neither of these two proposed CheW-binding pockets is consistent with our missense mutation study, as explained below.
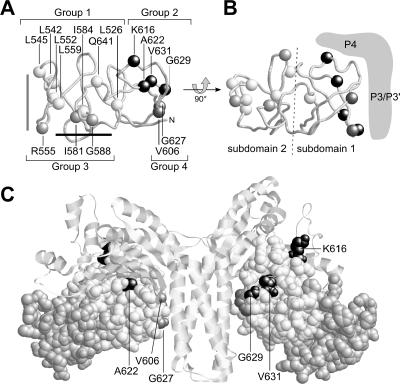
FIG. 6.
Structure-function relationships of CheA P5 mutants. (A) Alpha-carbon backbone of the P5 domain of E. coli CheA, modeled from the X-ray coordinates of the T. maritima P3-P4-P5 dimer (see text). The four groups of P5* mutations described in this study mainly define different regions in the P5 structure. The alpha-carbons of the mutant residues are shown in space-filled representation: white, group 1; light gray, group 3; dark gray, group 4; and black, group 2. Two previously proposed CheA-binding sites are indicated by light-gray (5) and black (41) lines near the left end of P5, distal to the N terminus (N). (B) P5 structure from panel A, rotated 90° to show the two folding subdomains and the approximate location of the adjoining P3 and P4 domains. (C) Modeled structure of the P3-P4-P5 dimer of E. coli CheA. The P3 and P4 domains are shown in ribbon representations; both P5 domains are shown in space-filled representation with different shading for the two subdomains. Residues defined by group 2 mutations (CheW binding defective) are shown in black, and group 4 mutation sites (deactivation defective) are shown in dark gray. The group 1 and group 3 sites are not shown in this panel.
Structure-function alterations in P5 mutants.
The four groups of P5 mutations characterized in this study define distinct structural as well as functional classes (Fig. 6).
(i) Group 1.
Group 1 lesions (Fig. 6A and B) destabilized P5, presumably through defects in folding or native structure. Although subject to rapid in vivo degradation, the mutant proteins had detectable autophosphorylation activity, implying that the misfolded P5 domains did not interfere with the functions of neighboring domains. These mutations most likely impair chemotaxis by drastically lowering steady-state CheA levels and, possibly, by disrupting normal receptor-coupling controls as well. Four of the seven sites identified by group 1 mutations are buried hydrophobic residues, three in subdomain 2 (L552, L559, and I584) and one (L526) in subdomain 1 at its interface with subdomain 2 (Fig. 6A). In all cases, the mutated amino acids have polar character and would be expected to disrupt core hydrophobic packing interactions. Two other hydrophobic group 1 residues (L542 and L545) are only partially buried and reside in the hydrophobic surface patch at the P3-distal end of P5 subdomain 2 (Fig. 6A). The final group 1 residue (Q641) also lies near the subdomain 1-subdomain 2 interface but at the C terminus of P5. This is the last residue of the T. maritima protein, but the E. coli protein has 14 more residues beyond this position, which might contribute additional stabilizing interactions for subdomain 2.
(ii) Group 3.
Group 3 mutations impaired the in vivo chemotactic function(s) of CheA, but the mutant proteins exhibited no significant in vitro defects. The group 3 mutations occurred at three residues in subdomain 2 (Fig. 6A and B). Although these mutant proteins cannot fully support chemotaxis, the nature of their functional defect is unclear and may not be related to their CheW-binding ability, which was ostensibly normal in the in vitro GST-CheW pull-down assay.
(iii) Group 4.
Group 4 CheA lesions did not impair chemotactic ability and were instead isolated as cheA mutations that could phenotypically suppress receptor mutants with CCW-biased signal states (28). The mutant proteins assembled activated ternary complexes in vitro that were not fully deactivated by attractant stimuli. Attenuated deactivation could account for their in vivo suppression effects, as this should enhance steady-state CheA activity and thereby offset the reduced activation ability of CCW-biased receptors. The group 4 residues (V606 and G627) are located in P5 subdomain 1, at the interface with P3/P3′ (Fig. 6).
(iv) Group 2.
Group 2 mutations caused substantial defects in CheW binding and receptor-mediated CheA activation. Although we did not directly assess ternary-complex formation in the present study, it seems likely that the activation defect of these mutant proteins reflects a failure to assemble ternary complexes, presumably due to their CheW-binding defect. The four group 2 residues are located in subdomain 1, near the interface with P3/P3′ (Fig. 6). Three (K616, G629, and V631) are surface exposed on the same face of P5 (Fig. 6C); the other (A622) is partly buried on an opposing face of P5 (Fig. 6C).
Location of the P5-dependent CheW-binding site.
The group 2 mutations, the only P5 lesions that affected CheW binding in vitro, must define important binding determinants. The most parsimonious interpretation is that group 2 mutations affect residues directly involved in the binding interaction, which implies that CheW binds primarily to P5 subdomain 1. Although much of the subdomain 1 surface seems to be inaccessible to CheW in the P3-P4-P5 crystal structure, CheA could conceivably undergo domain rearrangements in solution that expose the site proposed for CheW binding. In fact, the structure of a Thermotoga P4-P5/CheW complex, recently determined by Park et al. (34), is fully consistent with the proposed CheW interaction surface delineated by the group 2 P5 mutations in E. coli CheA.
Domain movements involved in CheA control.
The group 4 mutations, which are defective specifically in CheA deactivation, suggest that P5 domain movements also play an important role in receptor-mediated control. The group 4 residues (V606 and G627) reside directly at the P5-P3 interface (Fig. 6), where even subtle conformational changes might alter the spatial relationship between the two domains. The group 4 mutations introduce amino acids with larger side chains, which might also retard dynamic relative motions of the two domains.
In summary, we propose that CheW binds to subdomain 1 of the CheA P5 domain. Dynamic motion of the P5 domain relative to other parts of CheA may control CheW access as well as subsequent conformational changes in the ternary signaling complex that modulate CheA activity. The group 4 P5 lesions, which allow CheW binding and ternary-complex formation, specifically interfere with CheA deactivation, most likely by blocking movements of the P5 domain relative to the P4 and P3 domains. Although this model accounts for the in vitro defects of group 2 and group 4 P5 mutants, it cannot explain the in vivo defects of group 3 mutants, which have no apparent functional defects in vitro. These mutants indicate that additional or alternative P5 interactions, not currently measurable in vitro, are important for CheA signaling in vivo.
Acknowledgments
We thank Rebecca Reiser for assistance with the P5 mutant hunt and the protein degradation studies; Peter Ames, Rick Stewart, and Phil Matsumura for generously providing plasmids and antisera crucial to this study; Martin Horvath for valuable insights into protein structure; and Brian Crane for helpful discussions and communication of unpublished results.
This work was supported by research grant GM19559 from the National Institute of General Medical Sciences. The Protein-DNA Core Facility at the University of Utah receives support from National Cancer Institute grant CA42014 to the Huntsman Cancer Institute.
REFERENCES
- 1.Ames, P., and J. S. Parkinson. 1994. Constitutively signaling fragments of Tsr, the Escherichia coli serine chemoreceptor. J. Bacteriol. 176:6340-6348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ames, P., C. A. Studdert, R. H. Reiser, and J. S. Parkinson. 2002. Collaborative signaling by mixed chemoreceptor teams in Escherichia coli. Proc. Natl. Acad. Sci. USA 99:7060-7065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ames, P., Y. A. Yu, and J. S. Parkinson. 1996. Methylation segments are not required for chemotactic signalling by cytoplasmic fragments of Tsr, the methyl-accepting serine chemoreceptor of Escherichia coli. Mol. Microbiol. 19:737-746. [DOI] [PubMed] [Google Scholar]
- 4.Bibikov, S. I., R. Biran, K. E. Rudd, and J. S. Parkinson. 1997. A signal transducer for aerotaxis in Escherichia coli. J. Bacteriol. 179:4075-4079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bilwes, A. M., L. A. Alex, B. R. Crane, and M. I. Simon. 1999. Structure of CheA, a signal-transducing histidine kinase. Cell 96:131-141. [DOI] [PubMed] [Google Scholar]
- 6.Bolivar, F., R. Rodriguez, P. J. Greene, M. C. Betlach, H. L. Heyneker, and H. W. Boyer. 1977. Construction and characterization of new cloning vehicles. Gene 2:95-113. [PubMed] [Google Scholar]
- 7.Borkovich, K. A., and M. I. Simon. 1991. Coupling of receptor function to phosphate-transfer reactions in bacterial chemotaxis. Methods Enzymol. 200:205-214. [DOI] [PubMed] [Google Scholar]
- 8.Borkovich, K. A., and M. I. Simon. 1990. The dynamics of protein phosphorylation in bacterial chemotaxis. Cell 63:1339-1348. [DOI] [PubMed] [Google Scholar]
- 9.Bornhorst, J. A., and J. J. Falke. 2001. Evidence that both ligand binding and covalent adaptation drive a two-state equilibrium in the aspartate receptor signaling complex. J. Gen. Physiol. 118:693-710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Boukhvalova, M., R. VanBruggen, and R. C. Stewart. 2002. CheA kinase and chemoreceptor interaction surfaces on CheW. J. Biol. Chem. 277:23596-23603. [DOI] [PubMed] [Google Scholar]
- 11.Boukhvalova, M. S., F. W. Dahlquist, and R. C. Stewart. 2002. CheW binding interactions with CheA and Tar: importance for chemotaxis signaling in Escherichia coli. J. Biol. Chem. 277:22251-22259. [DOI] [PubMed] [Google Scholar]
- 12.Bourret, R. B., J. Davagnino, and M. I. Simon. 1993. The carboxy-terminal portion of the CheA kinase mediates regulation of autophosphorylation by transducer and CheW. J. Bacteriol. 175:2097-2101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chen, J. 1992. Genetic studies of transmembrane and intracellular signaling by a bacterial chemoreceptor. Ph.D. thesis. University of Utah, Salt Lake City.
- 14.Cox, E. C., and D. L. Horner. 1986. DNA sequence and coding properties of mutD(dnaQ) a dominant Escherichia coli mutator gene. J. Mol. Biol. 190:113-117. [DOI] [PubMed] [Google Scholar]
- 15.Garzón, A., and J. S. Parkinson. 1996. Chemotactic signaling by the P1 phosphorylation domain liberated from the CheA histidine kinase of Escherichia coli. J. Bacteriol. 178:6752-6758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gegner, J. A., and F. W. Dahlquist. 1991. Signal transduction in bacteria: CheW forms a reversible complex with the protein kinase CheA. Proc. Natl. Acad. Sci. USA 88:750-754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Gegner, J. A., D. R. Graham, A. F. Roth, and F. W. Dahlquist. 1992. Assembly of an MCP receptor, CheW, and kinase CheA complex in the bacterial chemotaxis signal transduction pathway. Cell 70:975-982. [DOI] [PubMed] [Google Scholar]
- 18.Griswold, I. J., and F. W. Dahlquist. 2002. The dynamic behavior of CheW from Thermotoga maritima in solution, as determined by nuclear magnetic resonance: implications for potential protein-protein interaction sites. Biophys. Chem. 101-102:359-373. [DOI] [PubMed] [Google Scholar]
- 19.Griswold, I. J., H. Zhou, M. Matison, R. V. Swanson, L. P. McIntosh, M. I. Simon, and F. W. Dahlquist. 2002. The solution structure and interactions of CheW from Thermotoga maritima. Nat. Struct. Biol. 9:121-125. [DOI] [PubMed] [Google Scholar]
- 20.Hess, J. F., R. B. Bourret, and M. I. Simon. 1991. Phosphorylation assays for proteins of the two-component regulatory system controlling chemotaxis in Escherichia coli. Methods Enzymol. 200:188-204. [DOI] [PubMed] [Google Scholar]
- 21.Jahreis, K., T. B. Morrison, A. Garzon, and J. S. Parkinson. 2004. Chemotactic signaling by an Escherichia coli CheA mutant that lacks the binding domain for phosphoacceptor partners. J. Bacteriol. 186:2664-2672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kishan, K. V., G. Scita, W. T. Wong, P. P. Di Fiore, and M. E. Newcomer. 1997. The SH3 domain of Eps8 exists as a novel intertwined dimer. Nat. Struct. Biol. 4:739-743. [DOI] [PubMed] [Google Scholar]
- 23.Kott, L., E. H. Braswell, A. L. Shrout, and R. M. Weis. 2004. Distributed subunit interactions in CheA contribute to dimer stability: a sedimentation equilibrium study. Biochim. Biophys. Acta 1696:131-140. [DOI] [PubMed] [Google Scholar]
- 24.Laemmli, U. K. 1970. Cleavage of structural proteins during assembly of the head of bacteriophage T4. Nature 227:680-685. [DOI] [PubMed] [Google Scholar]
- 25.Levit, M. N., and J. B. Stock. 2002. Receptor methylation controls the magnitude of stimulus-response coupling in bacterial chemotaxis. J. Biol. Chem. 277:36760-36765. [DOI] [PubMed] [Google Scholar]
- 26.Li, G., and R. M. Weis. 2000. Covalent modification regulates ligand binding to receptor complexes in the chemosensory system of Escherichia coli. Cell 100:357-365. [DOI] [PubMed] [Google Scholar]
- 27.Li, M., and G. L. Hazelbauer. 2004. Cellular stoichiometry of the components of the chemotaxis signaling complex. J. Bacteriol. 186:3687-3694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Liu, J. 1990. Molecular genetics of the chemotactic signaling pathway in Escherichia coli. Ph.D. thesis. University of Utah, Salt Lake City.
- 29.Liu, Y., M. Levit, R. Lurz, M. G. Surette, and J. B. Stock. 1997. Receptor-mediated protein kinase activation and the mechanism of transmembrane signaling in bacterial chemotaxis. EMBO J. 16:7231-7240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Matsumura, P., J. J. Rydel, R. Linzmeier, and D. Vacante. 1984. Overexpression and sequence of the Escherichia coli cheY gene and biochemical activities of the CheY protein. J. Bacteriol. 160:36-41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.McNally, D. F., and P. Matsumura. 1991. Bacterial chemotaxis signaling complexes: formation of a CheA/CheW complex enhances autophosphorylation and affinity for CheY. Proc. Natl. Acad. Sci. USA 88:6269-6273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Morrison, T. B., and J. S. Parkinson. 1997. A fragment liberated from the Escherichia coli CheA kinase that blocks stimulatory, but not inhibitory, chemoreceptor signaling. J. Bacteriol. 179:5543-5550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ninfa, E. G., A. Stock, S. Mowbray, and J. Stock. 1991. Reconstitution of the bacterial chemotaxis signal transduction system from purified components. J. Biol. Chem. 266:9764-9770. [PubMed] [Google Scholar]
- 34.Park, S., P. P. Borbat, G. Gonzalez-Bonet, J. Bhatagar, A. M. Pollard, J. H. Freed, A. M. Bilwes, and B. R. Crane. Nat. Struct. Mol. Biol., in press. [DOI] [PubMed]
- 35.Parkinson, J. S. 1976. cheA, cheB, and cheC genes of Escherichia coli and their role in chemotaxis. J. Bacteriol. 126:758-770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Parkinson, J. S. 1978. Complementation analysis and deletion mapping of Escherichia coli mutants defective in chemotaxis. J. Bacteriol. 135:45-53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Parkinson, J. S., and S. E. Houts. 1982. Isolation and behavior of Escherichia coli deletion mutants lacking chemotaxis functions. J. Bacteriol. 151:106-113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Rasmussen, R. P. 1998. Biochemical phenotypes of mutations in the chemotactic signaling complex of E. coli. Ph.D. thesis. University of Utah, Salt Lake City.
- 39.Sanatinia, H., E. C. Kofoid, T. B. Morrison, and J. S. Parkinson. 1995. The smaller of two overlapping cheA gene products is not essential for chemotaxis in Escherichia coli. J. Bacteriol. 177:2713-2720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Schwede, T., J. Kopp, N. Guex, and M. C. Peitsch. 2003. SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res. 31:3381-3385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Shimizu, T. S., N. Le Novere, M. D. Levin, A. J. Beavil, B. J. Sutton, and D. Bray. 2000. Molecular model of a lattice of signalling proteins involved in bacterial chemotaxis. Nat. Cell Biol. 2:792-796. [DOI] [PubMed] [Google Scholar]
- 42.Shrout, A. L., D. J. Montefusco, and R. M. Weis. 2003. Template-directed assembly of receptor signaling complexes. Biochemistry 42:13379-13385. [DOI] [PubMed] [Google Scholar]
- 43.Silverman, M., P. Matsumura, M. Hilmen, and M. Simon. 1977. Characterization of lambda Escherichia coli hybrids carrying chemotaxis genes. J. Bacteriol. 130:877-887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Smith, R. A., and J. S. Parkinson. 1980. Overlapping genes at the cheA locus of E. coli. Proc. Natl. Acad. Sci. USA 77:5370-5374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Sourjik, V., and H. C. Berg. 2004. Functional interactions between receptors in bacterial chemotaxis. Nature 428:437-441. [DOI] [PubMed] [Google Scholar]
- 46.Surette, M. G., M. Levit, Y. Liu, G. Lukat, E. G. Ninfa, A. Ninfa, and J. B. Stock. 1996. Dimerization is required for the activity of the protein histidine kinase CheA that mediates signal transduction in bacterial chemotaxis. J. Biol. Chem. 271:939-945. [DOI] [PubMed] [Google Scholar]
- 47.Swanson, R. V., R. B. Bourret, and M. I. Simon. 1993. Intermolecular complementation of the kinase activity of CheA. Mol. Microbiol. 8:435-441. [DOI] [PubMed] [Google Scholar]
- 48.Szurmant, H., and G. W. Ordal. 2004. Diversity in chemotaxis mechanisms among the bacteria and archaea. Microbiol. Mol. Biol. Rev. 68:301-319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Wadhams, G. H., and J. P. Armitage. 2004. Making sense of it all: bacterial chemotaxis. Nat. Rev. Mol. Cell Biol. 5:1024-1037. [DOI] [PubMed] [Google Scholar]
- 50.Wolfe, A. J., and H. C. Berg. 1989. Migration of bacteria in semisolid agar. Proc. Natl. Acad. Sci. USA 86:6973-6977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Wolfe, A. J., B. P. McNamara, and R. C. Stewart. 1994. The short form of CheA couples chemoreception to CheA phosphorylation. J. Bacteriol. 176:4483-4491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Xu, W., S. C. Harrison, and M. J. Eck. 1997. Three-dimensional structure of the tyrosine kinase c-Src. Nature 385:595-602. [DOI] [PubMed] [Google Scholar]