Abstract
The transfer RNA gene downstream from the HMR locus in S. cerevisiae functions as part of a boundary (barrier) element that restricts the spread of heterochromatic gene silencing into the downstream region of chromosome III. A genetic screen for identifying additional genes that, when mutated, allow inappropriate spreading of silencing from HMR through the tRNA gene was performed. YTA7, a gene containing bromodomain and ATPase homologies, was identified multiple times. Previously, others had shown that the bromodomain protein Bdf1p functions to restrict silencing at yeast euchromatin-heterochromatin boundaries; therefore we deleted nonessential bromodomain-containing genes to test their effects on heterochromatin spreading. Deletion of RSC2, coding for a component of the RSC chromatin-remodeling complex, resulted in a significant spread of silencing at HMR. Since the bromodomain of YTA7 lacks a key tyrosine residue shown to be important for acetyllysine binding in other bromodomains, we confirmed that a GST-Yta7p bromodomain fusion was capable of binding to histones in vitro. Epistasis analysis suggests that YTA7 and the HMR-tRNA function independently to restrict the spread of silencing, while RSC2 may function through the tRNA element. Our results suggest that multiple bromodomain proteins are involved in restricting the propagation of heterochromatin at HMR.
PROPER regulation of gene expression is of utmost importance for development and survival of all forms of life, as regulated transcription of complex genomes is necessary for cellular energy economy and the precise patterns of gene expression required for cellular differentiation. Transcription of genomes is regulated at multiple levels, which can involve large regions of chromosomes, individual nucleosomes, or direct effects on the transcription machinery at individual promoters (Kimura and Horikoshi 2004). At the level of chromatin in eukaryotes, active euchromatic and predominately inactive heterochromatic chromosomal domains alternate along chromosomes, and mechanisms must exist to prevent regulatory elements from each domain from influencing gene expression patterns of adjacent domains. Chromatin boundary elements (Donze and Kamakaka 2002; West et al. 2002; Donze 2004; Kimura and Horikoshi 2004) serve this purpose, as boundaries can prevent either a distal enhancer from activating gene expression inappropriately (insulators) or the unrestricted spreading of heterochromatin (barriers).
The budding yeast Saccharomyces cerevisiae contains limited heterochromatic regions at its telomeric and silent mating loci, and these chromosomal regions are important model systems for studying the establishment and propagation of silenced chromatin (Rusche et al. 2003) and also for studying chromatin boundaries and global mechanisms that restrict heterochromatic propagation (Bi and Broach 1999; Donze et al. 1999; Fourel et al. 1999; Kimura et al. 2002; Suka et al. 2002; Ladurner et al. 2003; Meneghini et al. 2003; Tackett et al. 2005). Silenced chromatin in yeast is maintained by the targeted regional action of the silent information regulator (Sir) proteins, where the Sir2p histone deacetylase establishes the histone acetylation state required for heterochromatin formation as part of a proposed histone code (Jenuwein and Allis 2001), although the specificity of such a code is currently being debated (Kurdistani et al. 2004; Dion et al. 2005). Deacetylation of histones allows higher affinity binding of Sir3p (Carmen et al. 2002), which then recruits the Sir4p/Sir2p complex, allowing heterochromatin to propagate along nucleosomes (Hoppe et al. 2002; Rusche et al. 2002).
At the HMR silent mating locus, a transfer RNA gene and its associated RNA polymerase III complex functions as part of a boundary element that prevents the spreading of Sir protein-mediated heterochromatin (Donze et al. 1999; Donze and Kamakaka 2001). However, we found that while the HMR-tRNA alone is sufficient to act as a barrier to silencing in a plasmid-based assay (Donze and Kamakaka 2001), when integrated back into the chromosome, the tRNA alone shows only partial barrier function (our unpublished results), suggesting that additional proteins are required to completely block silencing. Other genes coding for chromatin-associated proteins that can lead to a partial spreading of silencing through the tRNA have been identified and include HTZ1, BDF1, and YTA7 (Ladurner et al. 2003; Meneghini et al. 2003; Tackett et al. 2005), and the global level of histone acetylation regulated by SIR2 and SAS2 influences silencing spread (Kimura et al. 2002; Suka et al. 2002). Taken together, these results suggest that both local boundary-associated factors and global chromatin modifications affect the formation of a true euchromatin-heterochromatin boundary (Kimura and Horikoshi 2004).
To assess what other chromatin proteins participate in boundary formation, we used transposon-mediated mutagenesis and direct gene knockouts in a yeast boundary-element-dependent reporter strain to identify mutations that led to a spread of silencing through the HMR-tRNA boundary element, which included ∼400 bp at either side of the tRNA. In addition to identifying several factors known to be involved in regulating the spread of silencing, we identified a novel bromodomain-containing gene, YTA7, which additionally contains two separate AAA family ATPase domains. We also find that mutation of several different bromodomain genes leads to a general loss of boundary function. This suggests that bromodomains may be key effectors that recognize and maintain the euchromatic histone modification patterns necessary for boundary integrity. Epistasis analysis implicates independent roles for YTA7 and HMR-tRNA in restricting silencing, while RCS2 may be functioning through the RNA polymerase III complex.
MATERIALS AND METHODS
All yeast strains used in this study (Table 1) were isogenic to W303-1a. The parent boundary reporter strain used in the mating assays (DDY277) was derived from a cross of a similar strain, ROY962, previously described (Donze et al. 1999). The yeast transposon mutagenized library was obtained from Mike Snyder (Yale University), and mutagenesis was performed as described (Burns et al. 1994; Ross-Macdonald et al. 1997, 1999). All transposon insertion strains were created in DDY277, and identification of mutagenized genes was performed using the vectorette PCR method as described (Ross-Macdonald et al. 1999). Mating assay strains containing specific mutations were constructed by direct homologous recombination-mediated replacement using standard methods or by crossing previously constructed mutant alleles into the DDY277 background. Mating assays were performed as described (Donze et al. 1999).
TABLE 1.
Strains of S. cerevisiae generated for this study
| Strain | Genotype |
|---|---|
| DDY277 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 |
| DDY282 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMRΔI |
| DDY799 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:yta7 |
| DDY687 | MATα ADE2 his3 leu2 LYS2 trp1 ura3 HMR-E-Boundary-a1 sas2Δ∷TRP1 |
| DDY695 | MATα ADE2 his3 leu2 LYS2 trp1 ura3 HMR-E-Boundary-a1 gcn5∷TRP1 |
| DDY800 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:sas5 |
| DDY802 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:sas5 |
| DDY803 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:yta7 |
| DDY811 | MATaade2 his3 leu2 LYS2 trp1 ura3 HMR-trnaΔ-ADE2 |
| DDY814 | MATaade2 his3 leu2 LYS2 trp1 ura3 HMR-ADE2 |
| DDY1174 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:yta7 |
| DDY1309 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:yta7 |
| DDY1313 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:sas4 |
| DDY1316 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:yta7 |
| DDY1318 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:yta7 |
| DDY1340 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:yta7 |
| DDY1344 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:rpd3 |
| DDY1345 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:rpd3 |
| DDY1347 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:yta7 |
| DDY1665 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 rsc2Δ∷TRP1 |
| DDY1893 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 rsc1Δ∷URA3 |
| DDY1997 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 yta7Δ∷TRP1 |
| DDY2078 | MATα ADE2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 Tn∷LEU2:yta7 sir2Δ∷TRP1 |
| DDY2093 | MATα ade2 his3 leu2 lys2Δ trp1 ura3 HMR-ADE2 Tn:LEU2:rpd3 |
| DDY2106 | MATα ade2 his3 leu2 lys2Δ trp1 ura3 HMR-ADE2 Tn:LEU2:sas5 |
| DDY2114 | MATaade2 his3 leu2 LYS2 trp1 ura3 hmrΔ∷ADE2 |
| DDY2128 | MATaade2 his3 leu2 lys2Δ trp1 ura3 Tn:LEU2:rpd3 hmrΔ∷ADE2 |
| DDY2142 | MATaade2 his3 leu2 LYS2 trp1 ura3 Tn:LEU2:sas5 hmrΔ∷ADE2 |
| DDY2143 | MATaade2 his3 leu2 LYS2 trp1 ura3 Tn:LEU2:rpd3 HMR trnaΔ-ADE2 |
| DDY2156 | MATaade2 his3 leu2 LYS2 trp1 ura3 Tn:LEU2:sas5 HMR trna Δ-ADE2 |
| DDY2198 | MATaade2 his3 leu2, LYS2 trp1 ura3 hmrΔ∷ADE2 yta7Δ∷TRP1 |
| DDY2200 | MATaade2 his3 leu2, LYS2 trp1 ura3 HMR trnaΔ-ADE2 yta7Δ∷TRP |
| DDY2205 | MATα ade2 his3 leu2, lys2Δ trp1 ura3 HMR-ADE2 yta7Δ∷TRP1 |
| DDY2450 | MATα ade2 his3 leu2 LYS2 trp1 ura3 hmrΔ∷ADE2 rsc2Δ∷TRP1 |
| DDY2489 | MATaade2 his3 leu2 LYS2 trp1 ura3 HMR trna Δ-ADE2 rsc2∷TRP1 |
| DDY2496 | MATα ade2 his3 leu2 lys2Δ trp1 ura3 HMR-ADE2 rsc2Δ∷TRP1 |
| DDY2514 | MATα ade2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 spt7Δ∷URA3 |
| DDY2596 | MATα ade2 his3 leu2 LYS2 trp1 ura3 HMR-E-Boundary-a1 bdf1Δ∷HIS3 |
| DDY2601 | MATα ade2 his3 leu2 lys2Δ trp1 ura3 HMR-E-Boundary-a1 bdf2Δ∷LEU2 |
| DDY2659 | MATα ade2 his3 leu2 LYS2 trp1 ura3 HMR-E-Boundary-a1 snf2∷LEU2 |
| DDY2716 | MATα ADE2 his3 leu2 LYS2 trp1 ura3 HMR-E-Boundary-a1 htz1Δ∷KanMX |
The schematic of YTA7 and amino acid positions of AAA ATPase and bromodomains was based on query of the Conserved Domain Database (Marchler-Bauer et al. 2005). The alignment of S. cerevisiae bromodomains was created using ClustalW (Chenna et al. 2003) and was manually adjusted to fit a structurally based alignment of bromodomains (Zeng and Zhou 2002). The GST-Yta7p bromodomain fusion was made by PCR cloning the coding sequence of YTA7 from amino acids 1000 to 1101 into the glutathione-S-transferase (GST) vector pGEX-2TK (Amersham Biosciences), and the fusion protein was expressed and purified according to the manufacturer's instructions. Purified chicken core histones were purchased from Upstate (Lake Placid, NY). Histone pull-down assays were performed as described (Ladurner et al. 2003), and the recovered samples were run on an 18% SDS-PAGE gel and stained with Bio-Safe Coomassie stain (Bio-Rad Laboratories, Hercules, CA). Briefly, core histones were dissolved in water at 1 mg/ml, and 500-μl binding reactions contained 1× HEMG buffer (25 mm HEPES-KOH, pH 7.6, 0.2 mm EDTA, 12.5 mm MgCl2, 10% glycerol, diluted from a 5× stock), 50-μg core histones, 150 mm KCl, 15 μl glutathione-Sepharose (Amersham Biosciences) equilibrated in 1× HEMG plus 150 mm KCl, and 15 μg GST-Yta7p fusion protein or an equimolar amount of GST in the controls. Binding reactions were incubated for 4 hr at 4°, and beads were collected by centrifugation and then washed four times in 500 μl 1× HEMG, 150 mm KCl, 0.01% NP-40. A total of 125 μl 1× Laemmli loading buffer was added directly to the beads for gel analysis, and unbound fractions were precipitated with trichloroacetic acid, redissolved in 25 μl 0.1 n NaOH, and brought to 125 μl in 1× Laemmli loading buffer.
To construct the HMR-ADE2 reporter strains, a plasmid containing the 8.2-kb SacI-SalI HMR fragment was mutagenized to create a BamHI site 180 bp downstream of the HMR-tRNA (pDD657). The 2.5-kb BglII fragment of ADE2 (Stotz and Linder 1990) was cloned into this site such that ADE2 was transcribed away from HMR to create plasmid pDD659. HMR-trnaΔ-ADE2 was constructed by site-directed mutagenesis of pDD659 to delete the tRNA coding sequence (pDD661). Plasmid pDD833, hmrΔ-ADE2 was created by subcloning the SacI-SalI fragment of pDD659 into Bluescript cut with SacI and XhoI to destroy the polylinker XhoI site creating pDD833. The resulting plasmid was digested with SnaBI and XhoI to remove HMR, ends blunted with Klenow polymerase, and then religated to create pDD837. Each of these ADE2 plasmids was linearized and transformed into yeast strain DDY142, which contains an hmrΔ∷URA3 allele. Transformants were selected on minimal media lacking adenine, and individual colonies were patched and replica plated onto minimal media lacking uracil and media containing 5-FOA to identify isolates that had become Ade+ and uracil auxotrophic. Proper integration was then confirmed by Southern blot analysis.
Mutant alleles of genes affecting the spread of silencing through the boundary were then crossed into each of these ADE2 parent strains (DDY811, DDY814, and DDY2114). Colony color assays were performed by streaking representative strains onto minimal media containing 3 μg/ml adenine, which is 10% of the normal level in this medium. Colonies were grown for 3 days and held at 4° for 1 week prior to photography on a dissecting microscope equipped with a digital imaging system. Colony color was assessed for three independent isolates for each mutant; one representative strain is shown for each.
RESULTS
We used a MATα boundary reporter strain (DDY277) that harbors a modified HMR locus deleted for the HMR-I silencer and that also contains the downstream boundary sequence cloned into the HMRa2 gene (Figure 1, top). This strain is nonmating, as the boundary blocks the spread of silencing from HMR-E, allowing the a1 gene to be expressed in the MATα background (Donze et al. 1999). To identify genes involved in restricting the spread of silencing, we subjected this strain to a transposon mutagenesis protocol (Ross-Macdonald et al. 1999) and selected transposon insertion mutants that acquired an α-mating phenotype, indicating that silencing was now able to spread through the boundary into a1. A total of 20,000 Leu+ transformants were replica plated onto MATa tester lawns to identify mutants that lost boundary function at the modified HMR locus as judged by their ability to mate.
Figure 1.
Mating assay to identify extragenic mutations that lead to spreading of silencing through the HMR right boundary. (Top) The 1.0-kb region downstream of HMR containing the boundary tRNA was cloned into the a2 gene, and the resulting construct was integrated back into chromosome III in a MATα strain. The boundary element blocks the spread of silencing into a1, resulting in a nonmating phenotype (DDY277). The mating assay was performed essentially as described previously (Donze et al. 1999). DDY277 was mutagenized by transformation with a yeast genomic DNA library containing random transposon-LEU2 insertions (Ross-Macdonald et al. 1999), and Leu+ recombinants were tested for mating. (Bottom) Thirteen independent isolates that showed increased silencing were confirmed, representing mutations in four different genes, YTA7, SAS5, SAS4, and RPD3. All strain genotypes are listed in Table 1.
α-Mating isolates were confirmed by backcrossing to a MATa strain containing the same reporter construct, and only tetrad isolates in which the transposon-encoded LEU2 marker completely cosegregated with the mating phenotype (20 tetrads analyzed) were studied further. After this analysis, mutations in only four separate genes were confirmed as leading to a spread of silencing: SAS4 and SAS5, encoding subunits of the something about silencing (SAS) acetyltransferase complex; RPD3, a histone deacetylase; and YTA7, a little studied gene containing a weak bromodomain homology and two domains of AAA family ATPase homology. The mating phenotypes of each independent mutant isolate are shown in Figure 1. A single mutant of the SAS4 gene resulted in increased mating, along with two independent insertions into SAS5 and two independent insertions into RPD3. Mutations in the SAS complex were previously shown to increase the silencing efficiency of a defective hmra-e** locus (Ehrenhofer-Murray et al. 1997; Xu et al. 1999), and sas2 mutations were first identified as leading to increased silencing in sir1 mutants (Reifsnyder et al. 1996). Since hmra-e** lacks the boundary element, it is likely that mutations in the SAS complex globally affect the robustness of silencing by affecting cellular levels of histone H4 lysine 16 acetylation (Meijsing and Ehrenhofer-Murray 2001; Kimura et al. 2002; Suka et al. 2002; Shia et al. 2005), as opposed to specifically affecting the integrity of the boundary element. Mutations in RPD3 are also well documented as leading to a global increase of silencing at all three yeast heterochromatin loci (mating loci, rDNA, and telomeres) by an unknown mechanism (Rundlett et al. 1996; Vannier et al. 1996; Kim et al. 1999; Smith et al. 1999; Sun and Hampsey 1999).
Interestingly, eight independent isolates containing transposon insertions in YTA7, representing five different insertion sites, were isolated in the screen. It is not clear why yta7 mutants would occur so frequently in the library compared to other genes. It should be noted that the screen, while calculated to be saturating, was not, as mutations in genes previously known to induce a mating phenotype in this reporter system, such as the acetyltransferase subunit of SAS, SAS2, and the bromodomain-acetyltransferase GCN5, were not recovered (Donze and Kamakaka 2001). Additionally, insertions into other genes known to be required for restriction of silencing at HMR, BDF1 (Ladurner et al. 2003), and HTZ1 (Meneghini et al. 2003) were also not recovered. However, as discussed below, direct deletion of BDF1 or HTZ1 does lead to a spread of silencing in this reporter assay (Figure 3). The Sir protein dependence of the observed spread of silencing is shown for a yta7 mutant (DDY2078, Figure 1) and was also confirmed for rpd3 and SAS mutants (our unpublished data).
Figure 3.
Mutation of multiple bromodomain-containing genes results in spreading of silencing through the HMR boundary. Specific deletions or insertions were created or crossed into DDY277 to test their effects on boundary function in the mating assay described in Figure 1. Null mutations in YTA7, BDF1, RSC2, and SAS2 led to a significant spread of silencing, while mutation of GCN5, RSC1, BDF1, and HTZ1 showed a weaker phenotype. Mutation of SPT7 or SNF2 showed no phenotype.
YTA7 contains AAA family ATPase domains and a single bromodomain module, schematically depicted in Figure 2A. Since bromodomains have been demonstrated to be acetyllysine-binding modules (Dhalluin et al. 1999; Hudson et al. 2000; Jacobson et al. 2000; Owen et al. 2000) and have been suggested to mediate their functions by binding to acetylated histones and other chromatin-associated proteins (Zeng and Zhou 2002 ; Yang 2004; de la Cruz et al. 2005), it seemed likely that Yta7p may be functioning through histone recognition. The bromodomain protein Bdf1p is also known to have an antisilencing function at HMR, as bdf1 mutants show decreased gene expression and increased Sir3p association in regions adjacent to yeast silenced loci (Ladurner et al. 2003). Alignment of the YTA7 bromodomain with other yeast bromodomains (Figure 2B), however, reveals that YTA7 lacks a critical conserved tyrosine residue required for acetyllysine recognition by P/CAF, Gcn5p, and Bdf1p via a key hydrogen bond formation within the binding pocket (Dhalluin et al. 1999; Owen et al. 2000; Ladurner et al. 2003). Yta7p does contain a serine and threonine at the same location.
Figure 2.
YTA7 encodes a histone-binding bromodomain protein. (A) Schematic of the predicted YTA7 open reading frame, showing the relative locations of the two AAA family ATPase domains (amino acids 449–580 and 771–957) and the single bromodomain (amino acids 1003–1091) as determined by query of the Conserved Domain Database (Marchler-Bauer et al. 2005). (B) Alignment of yeast bromodomain sequences reveals that YTA7 lacks a key conserved tyrosine residue (arrow, red) that is critical for histone binding in other bromodomains, but contains a tandem serine-threonine in the same position (green). (C) The GST-Yta7p pull-down assay reveals that the bromodomain of Yta7p specifically retains histones H3 and H4. Reactions and washes were carried out in 150 mm KCl (see materials and methods). Increasing KCl to 500 mm in binding and wash steps reduced the level of histone binding by ∼50% (data not shown).
Due to the absence of this key tyrosine residue, we wanted to test whether the Yta7p bromodomain module was able to bind to histones. We cloned the coding sequence of the YTA7 bromodomain into a GST fusion vector and purified the GST-Yta7p fusion protein from Escherichia coli cells. This fusion protein was used in a pull-down assay with purified chicken erythrocyte core histones. Figure 2C shows that GST-Yta7p efficiently binds to histones H3 and H4, compared to no binding of GST alone to the histones. This result shows that, despite lacking the conserved tyrosine, the Yta7p bromodomain is a functional histone-binding domain in vitro.
The bromodomain protein Bdf1p has also been shown to restrict the spread of silencing in yeast (Ladurner et al. 2003), and the histone-binding domain of the transcription factor CTF-1 is essential for its heterochromatin barrier activity (Ferrari et al. 2004), suggesting a role for histone-binding proteins in blocking silencing. The S. cerevisiae genome contains 10 genes with bromodomain homologies (aligned in Figure 2B). Two of these genes, the RSC complex genes STH1 and RSC4, are essential for viability. Since loss of Bdf1p and Yta7p functions lead to an apparent spreading of silencing, we asked if bromodomain proteins in general contain boundary and/or antisilencing functions. We created a set of strains containing deletions or disruptions of each nonessential bromodomain-containing gene in strain DDY277 to assess the spreading of silencing across the boundary element using the mating assay described in Figure 1. The results shown in Figure 3 demonstrate that loss of function of several bromodomain proteins leads to a spreading of silencing. Deletion of RSC2, YTA7, or BDF1 leads to the greatest loss of boundary function, while deletion of RSC1, GCN5, or BDF2 shows a weaker phenotype. Deletion of SPT7 or SNF2 do not show spreading of silencing as determined by this assay. As controls for spreading of silencing at HMR, we assayed mutations in SAS2 and HTZ1 and observed the expected spreading of silencing as indicated by increased mating compared to the control strain DDY277 (Figure 3). The htz1 mutants showed a consistently variegated phenotype in this assay, with patches derived from some colonies mating and others not.
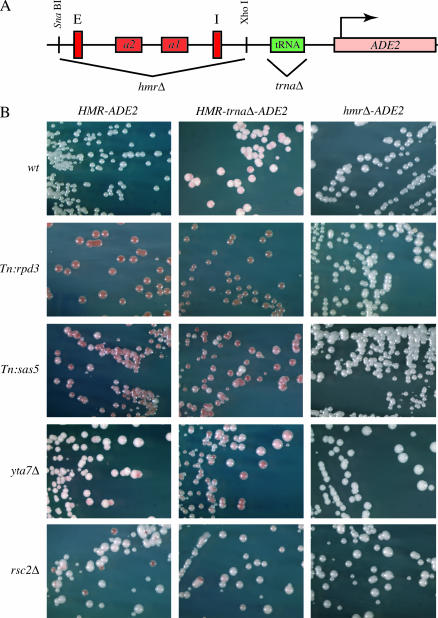
We next wanted to test epistatic relationships among bromodomain factors and the HMR-tRNA boundary element. Since deletion of the HMR-tRNA leads to a near-complete mating phenotype in the mating assay, we developed a second system for assessing the effects of tRNA deletion on heterochromatin spread at HMR. The S. cerevisiae ADE2 gene was integrated downstream of HMR (referred to here as HMR-ADE2), and isogenic strains that were HMR-trnaΔ-ADE2 or hmrΔ-ADE2 were created. Repression of ADE2 expression leads to the accumulation of a red pigment derived from the Ade2p substrate (Fisher 1969). Phenotypes of these parent strains are shown in the top row of Figure 4B: strains containing a normal HMR locus or a deletion of HMR (including both silencers) give rise to all-white colonies, demonstrating expression of ADE2 at this integration site; deletion of only the HMR-tRNA gives rise to a variegated phenotype with some sectored pink colonies arising due to epigenetic silencing of ADE2.
Figure 4.
YTA7 and the HMR-tRNA function through different pathways to restrict the spread of silencing at HMR. (A) The ADE2 gene was inserted into chromosome III downstream of HMR in S. cerevisiae (HMR-ADE2). Isogenic strains lacking the HMR-tRNA (HMR-trnaΔ-ADE2) or lacking HMR (hmrΔ-ADE2) were also constructed. (B) Each strain was crossed into rpd3, sas5, yta7, or rsc2 mutant backgrounds, and silencing was assessed by the degree of pigmentation exhibited by the presence or absence of ADE2 expression. The combination of yta7Δ trnaΔ leads to a more silenced phenotype (more and darker red colonies), suggesting that each functions independently. No difference in phenotype is seen between strains containing rsc2Δ and rsc2Δ trnaΔ. Strains depicted are: first row [wild type (wt)], DDY814, DDY811, and DDY2114; second row (rpd3), DDY2093, DDY2143, and DDY2128; third row (sas5), DDY2106, DDY2156, and DDY2142; fourth row (yta7), DDY2205, DDY2200, and DDY2198; and fifth row (rsc2), DDY2496, DDY2489, and DDY2450.
These ADE2 reporter strains were crossed to strains containing mutations that resulted in spreading of silencing in the mating assay. Mutation of RPD3 resulted in completely red colonies with or without the tRNA present. This complete penetrance of the rpd3 phenotype prevented any epistatic analysis using this assay. Mutation of SAS5 resulted in a variegated but mostly pink-red colony phenotype, which was unchanged when the HMR-tRNA gene was deleted. Again, on the basis of the qualitative nature of this assay, it was impossible to determine epistatic effects of sas5Δ trnaΔ mutations.
Deletion of YTA7 in HMR-ADE2 strains yielded a variegated pink-white phenotype, similar in magnitude to the HMR-trnaΔ-ADE2 strain; however, in yta7Δ trnaΔ strains, a darker red color was observed in a greater percentage of colonies, including some completely red colonies, demonstrating a more robust spread of silencing when both mutations are present (Figure 4B, fourth row, center). This additive effect suggests that YTA7 functions independently of the HMR-tRNA in restricting the spread of silencing. RSC2 deletion shows a weakly variegated phenotype in the HMR-ADE2 background, with rare silenced colonies that were somewhat darker red. Deletion of the HMR-tRNA in the rsc2Δ background did not increase the severity of the phenotype, suggesting that RSC2 and the HMR-tRNA may function through the same pathway in restricting the spread of silencing. We attempted to determine epistatic relationships among bromodomain-containing genes, but we were unable to obtain yta7-rsc2 or yta7-bdf1 double mutants by crosses of single-mutant strains or by homologous recombination, suggesting a synthetic lethality of these mutant combinations in our W303 background (our unpublished results).
Importantly, when HMR was deleted in each of these mutant backgrounds (Figure 4B, right column), all-white colonies were observed. This confirms that phenotypes observed for each of these mutations are due to the spread of silencing from HMR and not due to global histone acetylation effects or spreading of silencing from the telomere of chromosome III. Deletion of SIR2 or SIR4 in these backgrounds also results in completely white colonies, confirming the Sir dependence of the phenotype (our unpublished data). Also, it should be noted that the spreading of silencing observed in all experiments described here is due to normal levels of Sir proteins and that no overexpression of Sir3p was employed to obtain the observed phenotypes.
DISCUSSION
YTA7 encodes a histone-binding chromatin boundary protein:
The mutational analysis of boundary function described here revealed that the AAA family ATPase/bromodomain protein Yta7p plays a role in the integrity of the euchromatin/heterochromatin boundary at the S. cerevisiae HMR locus. YTA7 contains a region of bromodomain homology from amino acids 1003 to 1091 and two separate AAA family ATPase domains from amino acids 449 to 580 and 771 to 957 (Figure 2A). Comparison of the homology of the YTA7 bromodomain to other yeast bromodomains (Figure 2B) revealed that YTA7 lacks a key tyrosine residue (conserved in most bromodomains), which makes a critical water-mediated hydrogen bond to acetyllysine in Gcn5p and is required for the interaction of acetylated histones with Bdf1p (Dhalluin et al. 1999; Owen et al. 2000; Ladurner et al. 2003). However, the results shown in Figure 2C demonstrate that the YTA7 bromodomain is a functional histone-binding module. The YTA7 sequence contains tandem serine and threonine residues at the position of the conserved tyrosine, which may contribute the necessary hydrogen-bonding potential within the bromodomain-binding pocket. Bioinfomatic analysis of YTA7 orthologs in the related yeast species S. paradoxus, S. mikatae, and S. bayanus and the less-related Debaryomyces hansenii and Yarrowia lipolytica revealed that these YTA7 orthologs have the same serine-threonine substitutions in their respective bromodomains (Christie et al. 2004), and the same substitution is seen in both human and mouse Tif1β (Zeng and Zhou 2002). Yta7p is also a confirmed nuclear protein, as determined by a genome-wide analysis of yeast protein localization (Huh et al. 2003).
While this work was in progress, Tackett et al. (2005) published a proteomic study that also implicated Yta7p in boundary function. They found that Yta7p associated with complexes containing Dpb4p, a component of the DNA polymerase ε and ISW2/CHRAC chromatin complexes, and that both of these complexes are involved in regulating telomeric silencing (Iida and Araki 2004). Tackett et al. (2005) also showed that deletion of YTA7 led to spreading of silencing at HMR, and most significantly, they localized Dpb4p-Yta7p complexes to sites near HMR, HML, chromosome ends, and the conditionally expressed and epigenetically regulated FLO genes, suggesting a specific targeted role for Yta7p in restricting repression from multiple loci. The results presented in this article confirm the role of Yta7p in preventing the spread of silencing and extend these results to demonstrate independent functions of Yta7p and the HMR-tRNA with an epistasis analysis (discussed below).
Additionally, the study by Tackett et al. (2005) identified roles for DNA polymerase ε, Isw2, and Sas3-Spt16 chromatin-remodeling complexes in boundary function. We previously tested mutations in SAS3 (Donze and Kamakaka 2001) and SPT16 (our unpublished results) and saw no spreading of silencing in our mating assay. Tackett et al.'s (2005) analysis of the effect of SAS3 deletion showed a reduction in transcript levels of genes adjacent to HMR (GIT1 and YCR095C), but it is unknown if this reduction is Sir protein dependent, as expression of these genes was not reported in a sirΔ or hmrΔ background.
Our genetic analysis also uncovered a role for remodels the structure of chromatin (RSC) complex proteins in boundary function. RSC is a multisubunit, highly abundant, and essential chromatin-remodeling complex first purified from S. cerevisiae cells (Cairns et al. 1996). Genome-wide analysis of the location of the RSC complex has demonstrated that RSC is targeted to many tRNA genes in vivo (Ng et al. 2002), suggesting a potentially direct connection between the function of the HMR-tRNA and the RSC complex in heterochromatin boundary function. Our epistasis analysis demonstrating no difference in phenotype between rsc2Δ and rsc2Δ trnaΔ strains shown in Figure 4 is consistent with this proposed connection.
Yta7p appears to function independently of the HMR-tRNA, as deletion of both the cis-element and the trans-factor leads to a more silenced phenotype (Figure 4). We could not determine the epistatic relationships between the HMR-tRNA and BDF1, BDF2, RSC1, or GCN5 in this assay, as strains containing deletions of these genes grew as all-white colonies in the HMR-ADE2 background and had no apparent effect on colony color when in the HMR-trnaΔ-ADE2 background (our unpublished results). Mutations in BDF2, RSC1, and GCN5 all exhibited a weaker phenotype in the mating assay, an effect that may be too weak to affect the ADE2 promoter. However, we observed a strong increase in silencing upon deletion of BDF1 in the mating assay, but this mutation did not give rise to pigmented colonies in the HMR-ADE2 background. It should be noted that bdf1 strains are very slow growing, and it is uncertain how growth rate might affect the accumulation of pigment in ADE2 silenced strains.
The role of multiple chromatin-associated proteins in maintaining euchromatin-heterochromatin boundaries:
Since the discovery and characterization of a discrete heterochromatin boundary downstream of the HMR locus (Loo and Rine 1994; Donze et al. 1999; Donze and Kamakaka 2001), many mutations have been uncovered that alter the integrity of the convergence point of heterochromatin with downstream euchromatin at this locus. As described here and in other studies, in addition to mutation of the HMR-tRNA or its associated RNA polymerase III complex proteins, loss of function of chromatin-associated complexes containing Yta7p, Rsc2p, Rpd3p, Sas2, -4, or -5 proteins, Htz1p, or Bdf1p leads to spreading of silencing through the HMR boundary element. These effects either appear to be targeted to the boundary region in the case of the RNA polymerase III complex, Yta7p, and Bdf1p (Donze and Kamakaka 2001; Ladurner et al. 2003; Tackett et al. 2005) or may act more globally to affect histone acetylation levels, as has been suggested for the SAS complex (Kimura et al. 2002; Suka et al. 2002).
Several recent studies have demonstrated that direct targeting of transcription factors or components of chromatin-modifying complexes can act as a barrier, preventing the spread of SIR-mediated heterochromatin in yeast (Donze and Kamakaka 2001; Fourel et al. 2001; Chiu et al. 2003; Jacobson and Pillus 2004; Oki et al. 2004). The barrier effect created by acetyltransferase targeting was not necessarily due to the enzymatic activity of acetyltransferase fusions, but did require the presence of other components of the respective complexes (Chiu et al. 2003; Jacobson and Pillus 2004; Oki et al. 2004), suggesting that targeting of a specific factor results in the recruitment of multiple proteins. Global models of heterochromatin barrier function suggest that acetyltransferases and deacetylases establish “negotiable borders” or “fuzzy boundaries” by convergence of the effects of their activities, without the need for a specific barrier element or localized targeting (Fourel et al. 2004; Kimura and Horikoshi 2004). However, while both targeted and negotiable models are consistent with current data, the effects of global genome alteration (such as in a sas2 background) on the normal targeting of chromatin-associated proteins is not well defined. Any natural targeting of proteins occurs within the context of normal cellular chromatin acetylation. Therefore, while domains of global histone acetylation may be negotiable in mutant strains, such genome-wide effects may lead to specific alterations in acetylation-state-dependent localization of proteins, such as the targeting of bromodomains.
Taken together, the current evidence suggests that euchromatin-heterochromatin boundaries are determined by both site-specific interactions and global influences. Sir-protein-dependent silencing in yeast appears to propagate by successive Sir2p deacetylation, Sir3p binding to deacetylated nucleosomes, and recruitment of the Sir4p-Sir2p complex, allowing spreading of the heterochromatic chromatin along a nucleosomal array after initial targeting of the SIR complex at silencers or telomeres (Hoppe et al. 2002; Rusche et al. 2002). In certain cases, simply excluding nucleosomes by binding of LexA to multimerized sites can block the spread of Sir-protein-mediated heterochromatin propagation in S. cerevisiae (Bi et al. 2004), perhaps by simply distancing the next nucleosome from the previous Sir2-3-4-nucleosome complex. Part of the function of the HMR-tRNA may involve creating such a nucleosomal gap, as fully assembled RNA polymerase III complexes footprint close to 150 bp (Chedin et al. 1998; Kassavetis et al. 1998), approximately the length of the DNA occupied by a nucleosome. However, since the HMR-tRNA alone cannot provide full boundary function in a chromosomal context (our unpublished results), other chromatin-associated factors are clearly involved.
Additional studies will be required to identify the roles of bromodomains and other histone-binding proteins like CTF-1 (Ferrari et al. 2004) in establishing heterochromatin-euchromatin boundaries. Bdf1p has been shown to protect histone H4 from Sir2p deacetylation in vitro (Ladurner et al. 2003), suggesting one possible mechanism of bromodomain action in restricting silencing. However, bromodomains are known to bind to acetylated proteins other than histones (Yang 2004; de la Cruz et al. 2005); therefore the effects on boundary establishment reported in this study may involve more than bromodomain-histone interactions. The results presented here suggest that the RSC complex may be acting through the HMR-tRNA, which is supported by the fact that RSC preferentially associates with tRNA genes in vivo (Ng et al. 2002). However, it is currently unknown whether RSC is targeted to assembled RNA polymerase III genes or if prior RSC binding promotes RNA polymerase III complex assembly. Yta7p appears to be targeted to a limited set of chromosomal sites (Tackett et al. 2005), and while the mechanism of this targeting is unknown, it may involve the histone acetylation state near the boundary. The variability of bromodomain mutant phenotypes on boundary function may reflect a varied array of target proteins for bromodomain binding or could be a function of varied effects on the different multisubunit complexes in which most bromodomain factors have been identified. It will be of interest to determine the acetylation requirements for Yta7p binding at heterochromatin boundaries and how mutations that cause specific or global alterations in the acetylation states of histones, or possibly other chromatin-associated proteins, affect the targeting and function of boundary-associated bromodomain factors.
Acknowledgments
We thank David Stillman (University of Utah Health Sciences Center) for the gcn5 allele, Rohinton Kamakaka (National Institute of Child Health and Human Development, Bethesda, MD) for the snf2 and htz1 alleles, and Andreas Ladurner (European Molecular Biology Laboratory, Heidelberg, Germany) for the bdf1 and bdf2 alleles. Research in the Donze laboratory is generously supported by grants from the Human Frontier Science Program (RGY11/2002) and the National Science Foundation (MCB-0342113).
References
- Bi, X., and J. R. Broach, 1999. UASrpg can function as a heterochromatin boundary element in yeast. Genes Dev. 13: 1089–1101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bi, X., Q. Yu, J. J. Sandmeier and Y. Zou, 2004. Formation of boundaries of transcriptionally silent chromatin by nucleosome-excluding structures. Mol. Cell. Biol. 24: 2118–2131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burns, N., B. Grimwade, P. B. Ross-Macdonald, E. Y. Choi, K. Finberg et al., 1994. Large-scale analysis of gene expression, protein localization, and gene disruption in Saccharomyces cerevisiae. Genes Dev. 8: 1087–1105. [DOI] [PubMed] [Google Scholar]
- Cairns, B. R., Y. Lorch, Y. Li, M. Zhang, L. Lacomis et al., 1996. RSC, an essential, abundant chromatin-remodeling complex. Cell 87: 1249–1260. [DOI] [PubMed] [Google Scholar]
- Carmen, A. A., L. Milne and M. Grunstein, 2002. Acetylation of the yeast histone H4 N terminus regulates its binding to heterochromatin protein SIR3. J. Biol. Chem. 277: 4778–4781. [DOI] [PubMed] [Google Scholar]
- Chedin, S., M. L. Ferri, G. Peyroche, J. C. Andrau, S. Jourdain et al., 1998. The yeast RNA polymerase III transcription machinery: a paradigm for eukaryotic gene activation. Cold Spring Harbor Symp. Quant. Biol. 63: 381–389. [DOI] [PubMed] [Google Scholar]
- Chenna, R., H. Sugawara, T. Koike, R. Lopez, T. J. Gibson et al., 2003. Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res. 31: 3497–3500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiu, Y. H., Q. Yu, J. J. Sandmeier and X. Bi, 2003. A targeted histone acetyltransferase can create a sizable region of hyperacetylated chromatin and counteract the propagation of transcriptionally silent chromatin. Genetics 165: 115–125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Christie, K. R., S. Weng, R. Balakrishnan, M. C. Costanzo, K. Dolinski et al., 2004. Saccharomyces Genome Database (SGD) provides tools to identify and analyze sequences from Saccharomyces cerevisiae and related sequences from other organisms. Nucleic Acids Res. 32: D311–D314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de la Cruz, X., S. Lois, S. Sanchez-Molina and M. A. Martinez-Balbas, 2005. Do protein motifs read the histone code? BioEssays 27: 164–175. [DOI] [PubMed] [Google Scholar]
- Dhalluin, C., J. E. Carlson, L. Zeng, C. He, A. K. Aggarwal et al., 1999. Structure and ligand of a histone acetyltransferase bromodomain. Nature 399: 491–496. [DOI] [PubMed] [Google Scholar]
- Dion, M. F., S. J. Altschuler, L. F. Wu and O. J. Rando, 2005. Genomic characterization reveals a simple histone H4 acetylation code. Proc. Natl. Acad. Sci. USA 102: 5501–5506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donze, D., 2004. Breaking the histone code of silence: the propagation and blocking of heterochromatin. Curr. Org. Chem. 8: 211–221. [Google Scholar]
- Donze, D., and R. T. Kamakaka, 2001. RNA polymerase III and RNA polymerase II promoter complexes are heterochromatin barriers in Saccharomyces cerevisiae. EMBO J. 20: 520–531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donze, D., and R. T. Kamakaka, 2002. Braking the silence: how heterochromatic gene repression is stopped in its tracks. BioEssays 24: 344–349. [DOI] [PubMed] [Google Scholar]
- Donze, D., C. R. Adams, J. Rine and R. T. Kamakaka, 1999. The boundaries of the silenced HMR domain in Saccharomyces cerevisiae. Genes Dev. 13: 698–708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ehrenhofer-Murray, A. E., D. H. Rivier and J. Rine, 1997. The role of Sas2, an acetyltransferase homologue of Saccharomyces cerevisiae, in silencing and ORC function. Genetics 145: 923–934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferrari, S., K. C. Simmen, Y. Dusserre, K. Muller, G. Fourel et al., 2004. Chromatin domain boundaries delimited by a histone-binding protein in yeast. J. Biol. Chem. 279: 55520–55530. [DOI] [PubMed] [Google Scholar]
- Fisher, C. R., 1969. Enzymology of the pigmented adenine-requiring mutants of Saccharomyces and Schizosaccharomyces. Biochem. Biophys. Res. Commun. 34: 306–310. [DOI] [PubMed] [Google Scholar]
- Fourel, G., E. Revardel, C. E. Koering and E. Gilson, 1999. Cohabitation of insulators and silencing elements in yeast subtelomeric regions. EMBO J. 18: 2522–2537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fourel, G., C. Boscheron, E. Revardel, E. Lebrun, Y. F. Hu et al., 2001. An activation-independent role of transcription factors in insulator function. EMBO Rep. 2: 124–132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fourel, G., F. Magdinier and E. Gilson, 2004. Insulator dynamics and the setting of chromatin domains. BioEssays 26: 523–532. [DOI] [PubMed] [Google Scholar]
- Hoppe, G. J., J. C. Tanny, A. D. Rudner, S. A. Gerber, S. Danaie et al., 2002. Steps in assembly of silent chromatin in yeast: Sir3-independent binding of a Sir2/Sir4 complex to silencers and role for Sir2-dependent deacetylation. Mol. Cell. Biol. 22: 4167–4180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hudson, B. P., M. A. Martinez-Yamout, H. J. Dyson and P. E. Wright, 2000. Solution structure and acetyl-lysine binding activity of the GCN5 bromodomain. J. Mol. Biol. 304: 355–370. [DOI] [PubMed] [Google Scholar]
- Huh, W. K., J. V. Falvo, L. C. Gerke, A. S. Carroll, R. W. Howson et al., 2003. Global analysis of protein localization in budding yeast. Nature 425: 686–691. [DOI] [PubMed] [Google Scholar]
- Iida, T., and H. Araki, 2004. Noncompetitive counteractions of DNA polymerase epsilon and ISW2/yCHRAC for epigenetic inheritance of telomere position effect in Saccharomyces cerevisiae. Mol. Cell. Biol. 24: 217–227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacobson, S., and L. Pillus, 2004. Molecular requirements for gene expression mediated by targeted histone acetyltransferases. Mol. Cell. Biol. 24: 6029–6039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacobson, R. H., A. G. Ladurner, D. S. King and R. Tjian, 2000. Structure and function of a human TAFII250 double bromodomain module. Science 288: 1422–1425. [DOI] [PubMed] [Google Scholar]
- Jenuwein, T., and C. D. Allis, 2001. Translating the histone code. Science 293: 1074–1080. [DOI] [PubMed] [Google Scholar]
- Kassavetis, G. A., A. Kumar, E. Ramirez and E. P. Geiduschek, 1998. Functional and structural organization of Brf, the TFIIB-related component of the RNA polymerase III transcription initiation complex. Mol. Cell. Biol. 18: 5587–5599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim, S., A. Benguria, C. Y. Lai and S. M. Jazwinski, 1999. Modulation of life-span by histone deacetylase genes in Saccharomyces cerevisiae. Mol. Biol. Cell 10: 3125–3136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimura, A., and M. Horikoshi, 2004. Partition of distinct chromosomal regions: negotiable border and fixed border. Genes Cells 9: 499–508. [DOI] [PubMed] [Google Scholar]
- Kimura, A., T. Umehara and M. Horikoshi, 2002. Chromosomal gradient of histone acetylation established by Sas2p and Sir2p functions as a shield against gene silencing. Nat. Genet. 32: 370–377. [DOI] [PubMed] [Google Scholar]
- Kurdistani, S. K., S. Tavazoie and M. Grunstein, 2004. Mapping global histone acetylation patterns to gene expression. Cell 117: 721–733. [DOI] [PubMed] [Google Scholar]
- Ladurner, A. G., C. Inouye, R. Jain and R. Tjian, 2003. Bromodomains mediate an acetyl-histone encoded antisilencing function at heterochromatin boundaries. Mol. Cell 11: 365–376. [DOI] [PubMed] [Google Scholar]
- Loo, S., and J. Rine, 1994. Silencers and domains of generalized repression. Science 264: 1768–1771. [DOI] [PubMed] [Google Scholar]
- Marchler-Bauer, A., J. B. Anderson, P. F. Cherukuri, C. DeWeese-Scott, L. Y. Geer et al., 2005. CDD: a conserved domain database for protein classification. Nucleic Acids Res. 33: D192–D196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meijsing, S. H., and A. E. Ehrenhofer-Murray, 2001. The silencing complex SAS-I links histone acetylation to the assembly of repressed chromatin by CAF-I and Asf1 in Saccharomyces cerevisiae. Genes Dev. 15: 3169–3182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meneghini, M. D., M. Wu and H. D. Madhani, 2003. Conserved histone variant H2A.Z protects euchromatin from the ectopic spread of silent heterochromatin. Cell 112: 725–736. [DOI] [PubMed] [Google Scholar]
- Ng, H. H., F. Robert, R. A. Young and K. Struhl, 2002. Genome-wide location and regulated recruitment of the RSC nucleosome-remodeling complex. Genes Dev. 16: 806–819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oki, M., L. Valenzuela, T. Chiba, T. Ito and R. T. Kamakaka, 2004. Barrier proteins remodel and modify chromatin to restrict silenced domains. Mol. Cell. Biol. 24: 1956–1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Owen, D. J., P. Ornaghi, J. C. Yang, N. Lowe, P. R. Evans et al., 2000. The structural basis for the recognition of acetylated histone H4 by the bromodomain of histone acetyltransferase gcn5p. EMBO J. 19: 6141–6149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reifsnyder, C., J. Lowell, A. Clarke and L. Pillus, 1996. Yeast SAS silencing genes and human genes associated with AML and HIV-1 Tat interactions are homologous with acetyltransferases. Nat. Genet. 14: 42–49. [DOI] [PubMed] [Google Scholar]
- Ross-Macdonald, P., A. Sheehan, G. S. Roeder and M. Snyder, 1997. A multipurpose transposon system for analyzing protein production, localization, and function in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 94: 190–195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ross-Macdonald, P., A. Sheehan, C. Friddle, G. S. Roeder and M. Snyder, 1999. Transposon mutagenesis for the analysis of protein production, function, and localization. Methods Enzymol. 303: 512–532. [DOI] [PubMed] [Google Scholar]
- Rundlett, S. E., A. A. Carmen, R. Kobayashi, S. Bavykin, B. M. Turner et al., 1996. HDA1 and RPD3 are members of distinct yeast histone deacetylase complexes that regulate silencing and transcription. Proc. Natl. Acad. Sci. USA 93: 14503–14508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rusche, L. N., A. L. Kirchmaier and J. Rine, 2002. Ordered nucleation and spreading of silenced chromatin in Saccharomyces cerevisiae. Mol. Biol. Cell 13: 2207–2222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rusche, L. N., A. L. Kirchmaier and J. Rine, 2003. The establishment, inheritance, and function of silenced chromatin in Saccharomyces cerevisiae. Annu. Rev. Biochem. 72: 481–516. [DOI] [PubMed] [Google Scholar]
- Shia, W. J., S. Osada, L. Florens, S. K. Swanson, M. P. Washburn et al., 2005. Characterization of the yeast trimeric-SAS acetyltransferase complex. J. Biol. Chem. 280: 11987–11994. [DOI] [PubMed] [Google Scholar]
- Smith, J. S., E. Caputo and J. D. Boeke, 1999. A genetic screen for ribosomal DNA silencing defects identifies multiple DNA replication and chromatin-modulating factors. Mol. Cell. Biol. 19: 3184–3197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stotz, A., and P. Linder, 1990. The ADE2 gene from Saccharomyces cerevisiae: sequence and new vectors. Gene 95: 91–98. [DOI] [PubMed] [Google Scholar]
- Suka, N., K. Luo and M. Grunstein, 2002. Sir2p and Sas2p opposingly regulate acetylation of yeast histone H4 lysine16 and spreading of heterochromatin. Nat. Genet. 32: 378–383. [DOI] [PubMed] [Google Scholar]
- Sun, Z. W., and M. Hampsey, 1999. A general requirement for the Sin3-Rpd3 histone deacetylase complex in regulating silencing in Saccharomyces cerevisiae. Genetics 152: 921–932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tackett, A. J., D. J. Dilworth, M. J. Davey, M. O'Donnell, J. D. Aitchison et al., 2005. Proteomic and genomic characterization of chromatin complexes at a boundary. J. Cell Biol. 169: 35–47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vannier, D., D. Balderes and D. Shore, 1996. Evidence that the transcriptional regulators SIN3 and RPD3, and a novel gene (SDS3) with similar functions, are involved in transcriptional silencing in S. cerevisiae. Genetics 144: 1343–1353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- West, A. G., M. Gaszner and G. Felsenfeld, 2002. Insulators: many functions, many mechanisms. Genes Dev. 16: 271–288. [DOI] [PubMed] [Google Scholar]
- Xu, E. Y., S. Kim, K. Replogle, J. Rine and D. H. Rivier, 1999. Identification of SAS4 and SAS5, two genes that regulate silencing in Saccharomyces cerevisiae. Genetics 153: 13–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang, X. J., 2004. Lysine acetylation and the bromodomain: a new partnership for signaling. BioEssays 26: 1076–1087. [DOI] [PubMed] [Google Scholar]
- Zeng, L., and M. M. Zhou, 2002. Bromodomain: an acetyl-lysine binding domain. FEBS Lett. 513: 124–128. [DOI] [PubMed] [Google Scholar]