Abstract
An efficient, nanoliter-scale microfabricated bioprocessor integrating all three Sanger sequencing steps, thermal cycling, sample purification, and capillary electrophoresis, has been developed and evaluated. Hybrid glass–polydimethylsiloxane (PDMS) wafer-scale construction is used to combine 250-nl reactors, affinity-capture purification chambers, high-performance capillary electrophoresis channels, and pneumatic valves and pumps onto a single microfabricated device. Lab-on-a-chip-level integration enables complete Sanger sequencing from only 1 fmol of DNA template. Up to 556 continuous bases were sequenced with 99% accuracy, demonstrating read lengths required for de novo sequencing of human and other complex genomes. The performance of this miniaturized DNA sequencer provides a benchmark for predicting the ultimate cost and efficiency limits of Sanger sequencing.
Keywords: capillary electrophoresis, genetic analysis, microfluidic
The Sanger method of DNA sequencing is remarkable in its ability to generate long and accurate sequence reads. Alternative low-cost methodologies aimed at displacing Sanger sequencing currently produce shorter and less accurate sequence reads, a drawback that could limit their usefulness in de novo sequencing of complex genomes and in the detection of structural variations. Comparative analysis of mammalian genomes is focusing attention on the importance of structural variations (insertions, inversions, deletions, and duplications) in speciation, evolution, disease, and cancer (1–3). Consequently, new near-term technologies for low-cost genome sequencing and eventual ultra-low-cost sequencing technologies (4) must be evaluated in the context of their ability to generate accurate short-range as well as long-range genome assemblies.
Emerging near-term technologies include two cyclic array methods that produce highly parallel, short-read data (5, 6). Shendure et al. (5) used DNA ligation to generate 26-base reads that were aligned to 70.5% of the 3.3-Mb Escherichia coli genome with high confidence. Using a pyrophosphate-based approach, Margulies et al. (6) generated read-lengths of ≈110 bases for sequencing and de novo assembly of 90.4% of the 2.1-Mb Streptococcus pneumoniae genome. Significantly, it was necessary to exclude repetitive DNA sequences from the assembly. Such repeat sequences are vital to genome structure and regulation, given that over half of the human genome is composed of repetitive DNA (7); repetitive sequences are ubiquitous and functionally important in prokaryotic and eukaryotic genomes (8), and gene duplications are evident in disease (9), speciation (2) and cancer (10). Longer read-lengths with enhanced accuracy would increase sequence coverage and improve de novo genome assemblies, but extending cyclic sequencing reads is challenging because the process is serial, requiring a linearly increasing number of enzymatic steps. Necessarily, genomes sequenced thus far by using cyclic methods are more than three orders of magnitude smaller than the human genome.
Because Sanger sequencing is the only demonstrated technology for complete sequencing of plant and animal genomes (7, 11–13), it is, thus, critical to ask: What are the ultimate limits of Sanger sequencing and how can these limits be achieved? The feasibility of performing completely integrated, nanoliter-scale Sanger sequencing from a femtomole of template has been discussed in ref. 14. A significant challenge in creating such a system is the integration of diverse biochemical processes and high-performance electrophoretic separation onto a single microdevice. Sequencing separations alone were demonstrated on a single-channel glass microdevice in 1995 and subsequently scaled to 96 and 384 channels (15–17). However, in each case, sample production and purification was performed by using conventional microliter-scale processing that generates a 1,000-fold more product than is needed for analysis (18). Because a significant cost in Sanger sequencing is associated with reagent consumption (19) as well as equipment expense and labor, conversion to an integrated lab-on-a-chip technology (20–23) could revolutionize Sanger sequencing by efficiently linking miniaturized analysis to ultra-low-volume sample generation and purification.
Inspiration for solving this process integration challenge is provided by the sample analysis systems developed recently for pathogen and extraterrestrial life detection (24, 25), which use a hybrid glass–polydimethylsiloxane (PDMS) microdevice construction (26, 27). Incorporation of a microvalve-forming PDMS membrane into a multilayer glass wafer structure enables a level of complexity, integration, performance, and functionality previously unattainable in microfabricated systems. Here, we explore the practical limits of Sanger sequencing by using this hybrid glass–PDMS microdevice structure to create an efficient, nanoliter-volume bioprocessor capable of read-lengths equivalent to those used in the initial sequencing of the human genome (7).
Results
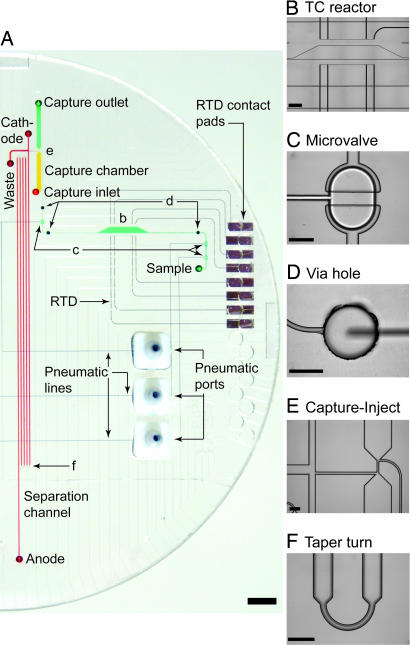
The bilaterally symmetric bioprocessor shown in Fig. 1A contains two independent DNA sequencing systems. Device elements are microfabricated on three 100-mm-diameter glass wafers and assembled with a featureless PDMS membrane. The top two glass wafers are thermally bonded to form enclosed all-glass reaction chambers and capillary electrophoresis (CE) channels. Contact bonding between the fused top wafers, PDMS membrane, and bottom manifold wafer forms three-dimensional interconnects and microvalves (Fig. 1B). The microvalve structures comprise a discontinuous fluidic channel on the bottom side of the resistive temperature detector (RTD)/valve wafer, the PDMS membrane, and a displacement chamber on the top side of the manifold wafer (26). Vacuum applied to the manifold lines deflects the PDMS membrane into the displacement chamber, creating a fluidic connection.
Fig. 1.
Integrated nanoliter-scale nucleic acid bioprocessor for Sanger DNA sequencing. (A) Top view of the assembled bioprocessor containing two sets of thermal cycling reactors, purification/concentration chambers, CE channels (black), RTDs (red), microvalves/pumps (green), pneumatic manifold channels (blue), and surface heaters (orange). (B) Expanded view, showing microdevice layers. Rim colors indicate the surface on which the respective features are fabricated. The top two glass wafers are thermally bonded and then assembled with a featureless PDMS membrane and manifold wafer.
Fig. 2A shows one half of the bioprocessor filled with distinct dyes to indicate reagent locations during operation. Each DNA sequencing system comprises three functional units corresponding to the three steps in Sanger sequencing. The thermal cycling unit consists of a 250-nl reactor with RTDs (Fig. 2B), microvalves (Fig. 2C), and a surface heater. Via holes (Fig. 2D) allow reagent to move between the enclosed all-glass layer and the glass–PDMS microvalve layer. Three microvalves for each reactor perform the dual role of sealing the chamber during thermal cycling and forming a self-priming diaphragm pump system for postprocessing. Thermal-cycled sequencing reagent is processed by the capture/purification unit (Fig. 2E Right) before injection (Fig. 2E Left) into the CE separation unit. Tapered turns (Fig. 2F) fold the 30-cm capillary onto the compact device while minimizing turn-induced dispersion (28).
Fig. 2.
Bioprocessor components. (A) Photograph of the microdevice, showing one of two complete nucleic acid processing systems. Colors indicate the location of sequencing reagent (green), capture gel (yellow), separation gel (red), and pneumatic channels (blue). (Scale bar, 5 mm.) B–F correspond to the following component microphotographs. (B) A 250-nl thermal cycling reactor with RTDs. (Scale bar, 1 mm.) (C) A 5-nl displacement volume microvalve. (D) A 500-μm-diameter via hole. (E) Capture chamber and cross injector. (F) A 65-μm-wide tapered turn. (Scale bars, 300 μm.) All features are etched to a depth of 30 μm.
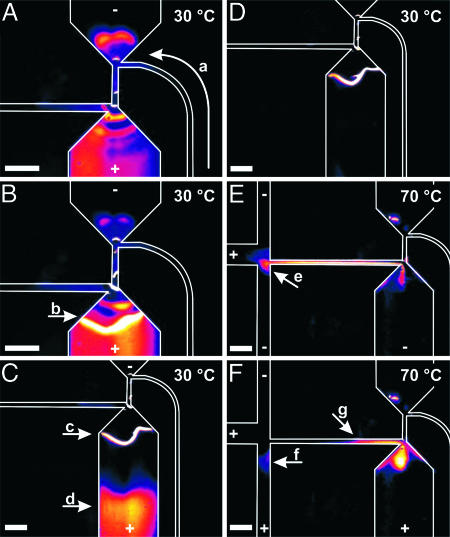
To demonstrate bioprocessor functionality, we performed integrated sequencing on 1 fmol of a 750-bp pUC18 amplicon. In the sequencing process, energy-transfer dye-terminator sequencing reaction reagent containing DNA template and −47 M13 sequencing primer is rapidly thermal-cycled in the 250-nl reaction chamber (35 cycles of 95°C for 12 s and 60°C for 55 s). The mixture is then pumped by the microvalves into the upper capture/purification chamber (Fig. 3A). A 33-V/cm electric field applied between the capture outlet and inlet ports drives the charged reaction mixture into the capture gel (Fig. 3B). The gel comprises a sparsely cross-linked polyacrylamide matrix containing 40 μM covalently bound oligonucleotide that is complementary to the DNA sequence immediately 3′ of the sequencing primer (29). Sanger extension fragments containing 12 or more complementary bases selectively hybridize at 30°C and concentrate at the gel/buffer interface, whereas residual salts, primer, template, and unincorporated nucleotides that would interfere with sample analysis pass through to the capture inlet port (Fig. 3C).
Fig. 3.
False-color fluorescence images of the capture, purification, and injection steps. Cooler colors indicate lower intensity; warmer colors indicate higher intensity. Fluidic channels are outlined in white. Relative electric potentials are indicated by + and −. (Scale bars, 300 μm.) (A) Thermally cycled dye-terminator sequencing reagent is pumped into the buffer-filled upper chamber (a). Simultaneously, an electric field drives the sequencing mixture into the lower, capture-gel-filled chamber. (B) Fluorescently labeled DNA extension fragments selectively hybridize and concentrate at the gel/buffer interface (b). (C) Desired extension fragments (c) are bound to the capture gel interface, and contaminants (d) continue to electrophorese out of the gel. (D) Purified and concentrated sample is ready for injection. (E) Extension fragments are released from the capture gel at 70°C and injected into the channel junction (e). Pinching potentials constrain the plug size. (F) A ≈1-nl sample plug is injected onto the separation capillary at (f), and reverse potentials pull away uninjected material (g).
After capture is complete, the microvalve pump system is turned off, and the electric potential is maintained to allow all residual reagents to electrophorese out of the capture chamber (Fig. 3D). Fluorescence quantitation indicates that 25 fmol of extension fragments are retained in a ≈1.2-nl band of capture gel at ≈20 μM, a concentration factor of 200×. Given a starting quantity of 1 fmol of template in a 35-cycle linear amplification reaction, total efficiency for extension fragment production and purification is 71%. This efficiency is consistent with observed PCR efficiencies and is dictated primarily by the sequencing reaction because capture is quantitative.
Extension fragment affinity for the capture gel is reversed by increasing the temperature beyond the melting point of the duplex (Tm = 54.7°C). Heating the microdevice to ≥67°C completely releases the hybridized extension fragments and raises the separation capillary to an optimal temperature for DNA sequencing. A ≈200-μm-long plug of concentrated and purified extension fragments is cross-injected onto the separation capillary (Fig. 3E). Sample separation is initiated by applying a 167-V/cm field between the cathode and anode (Fig. 3F), and small reverse potentials at the waste and capture inlet ports prevent uninjected material from leaking into the separation capillary (see Materials and Methods). Approximately 3.6 fmol of product is injected, as shown in Fig. 3F, giving an injection efficiency of 15% and a total sample-delivery efficiency of 10% for 1 fmol of template.
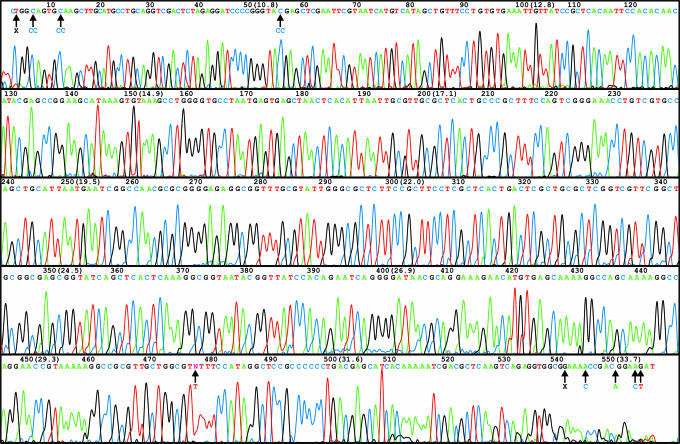
Four-color sequence data were collected on a radial scanner (18) in <35 min and processed with the cimarron 3.12 base caller and phred 0.020425.c to generate base calls with accuracy estimates (30). Although phred quality scores are calibrated for commercial sequencing instruments, they were in good agreement with observed accuracies in this study and are useful for relative evaluation of run conditions and other prototype instruments. Fig. 4 presents high-quality sequence data generated on the integrated bioprocessor. Primer and capture sequences are internal to the 750-bp amplicon, resulting in a maximum possible read length of 564 bases. Comigrating unincorporated fluorescently labeled terminators are absent in the sequencing trace, indicating their complete removal by the capture/purification process. Using a 99% accuracy cutoff with the known pUC18 sequence, cimarron and phred base calls produced a read length of 556 bases.
Fig. 4.
High-quality sequence data generated on the integrated bioprocessor. Sanger sequencing extension fragments from a 750-bp pUC18 PCR amplicon are resolved at 73°C, 167 V/cm with linear polyacrylamide separation gel (3.5% linear polyacrylamide/5% DMSO/3 M urea/1× TTE buffer) in 34 min. Automatic base calls by the program phred and base numbers are indicated above the electropherogram. Elapsed time in minutes from injection is indicated parenthetically every 50 bases. The base position where no call could be made is assigned “N.” Compared with the known pUC18 sequence, total read length is 556 bases with an accuracy of 99%. Black arrows indicate base call errors; correct calls are given below. X indicates an inserted base.
A range of injection potentials, electrophoretic separation fields, separation temperatures, and separation gels were investigated in 18 separate experiments to optimize run parameters (see Materials and Methods). The average 99% accuracy phred-generated read length for all runs was 427 bases (σ = 85). Consistent increases in read length expected with increased temperature or decreased separation field strength (31) were not observed, indicating an additional, overriding factor affecting read length. Inexact positioning of the capture band relative to the cross-injector with concomitant changes in injection timing is believed to be a significant source of this read-length variability. Our best sample-injection conditions generated five runs of >500 bases (μ = 540, σ = 17), with separation potentials ranging from 150 to 167 V/cm and temperatures ranging from 70°C to 73°C.
Fig. 5 graphs raw phred quality scores for each base (gray) and predicted cumulative read accuracy (black) for the data presented in Fig. 4. Base call accuracy rises rapidly after the first 25 bases to a predicted 1 in 1,000 error rate. Poorly resolved sequential Cs early in the run are correctly assigned low scores, causing dips in quality. Between 100 and 450 bases, error rates average 1 in 100,000. Accuracy drops at base 477, where phred is unable to make a base call in a typically problematic homopolymeric domain. Subsequent calls are accurate until base 541, after which point, signal strength drops substantially, making further base calling increasingly difficult. Aggregate phred error rates correctly estimate a read length of 556 bases with an accuracy of 99%.
Fig. 5.
Base-call accuracies and sequence read length as predicted by phred. Percent accuracy is related to the phred quality score by: 100 × (1 − Pe), where Pe, the probability that the base call is incorrect, is equal to 1/10Q/10. A one-in-a-hundred error rate is indicated by the dashed line. The gray line plots phred quality scores at each base position. The black line charts predicted read accuracy at each base position: 100 × (Basei − ΣPei)/Basei.
Discussion
The DNA sequencing bioprocessor presented here incorporates a range of advanced lab-on-a-chip technologies, including miniaturized temperature sensing, nanoliter-scale Sanger extension reactions, microvalves/pumps, DNA affinity-capture, and high-performance CE. Several innovations were necessary during development to integrate the disparate elements of the Sanger sequencing process into a single microdevice. The use of combined electrophoretic, pneumatic, and thermal sample control enabled efficient manipulation of minute reagent quantities. Earlier design iterations that relied solely on electrophoretic control were inefficient because of slow movement of sequencing sample in weak-field regions present in interlayer via holes. Transitioning from a monolithic substrate to a hybrid glass–PDMS assembly was critical. Multilayer construction enables greater design complexity and allows crossing of fluidic and pneumatic lines (25), a capability that is important for scaling feature density and parallel processing. At the same time, the durable hybrid construction preserves the all-glass structures necessary for single-base-resolution DNA separation. A single microdevice has been used for >40 complete runs without any indication of microvalve or CE channel degradation.
Development of optimized separation and capture gels and improved sample injection methods will be central in further improving read length and reproducibility. Previous work with the affinity-capture gel system used 2 μl of off-chip-prepared sequencing sample (29). Scaling the sequencing reaction down to 250 nl eliminated sample excess, making the injection timing more sensitive to thermal gel expansion and electrophoretic movement of the charged capture gel. Sparsely cross-linking the capture gel and reducing the capture temperature ameliorated these effects but resulted in increased purification/concentration times. As expected, the resolution of commonly used dye-terminator samples proved more challenging than idealized dye-primer samples typically tested in prototype systems. Extension of the separation channel from 16 to 30 cm increased the resolving power of the system, resulting in error rates of ≈1 in a million between 100 and 300 bases. Appropriately tuned separation gels should extend the >99% accuracy range by uniformly resolving peaks over greater amplicon lengths (32).
The core DNA sequencing engine demonstrated here establishes a performance benchmark from which to evaluate the feasibility of miniaturizing high-throughput Sanger sequencing. Practical limits to reaction volume reduction are determined by the sensitivity of the detection system and the efficiency of sample processing. In a capillary array scanning system, 1 million fluorophores per band significantly exceed the limit of detection (18) and could be reduced 10-fold with the improved scanning and detection technologies now available. By comparison, in this study, each band contained ≈4 million fluorophores. A reaction containing 1 fmol of template generates ≈26 times more product than is needed for detection (6 × 108 template molecules × 35 cycles × 71% efficiency/564 bands ≈ 26 × 106 molecules per band). More efficient injection techniques under investigation would eliminate the wasteful cross-injection methods used in this study, reducing template requirements 10-fold to 100 attomoles. Combined with scanner sensitivity improvements, the ultimate minimal template quantity extrapolates to ≈10 attomoles.
If we conservatively adopt a 10-fold reduction in template from 1 fmol to 100 attomoles, a sequencing reaction performed at standard concentrations in an easily fabricated 25-nl reactor represents a 400-fold reduction in current sequencing reagent consumption [Department of Energy Joint Genome Institute (2004) www.jgi.doe.gov/sequencing/protocols/prots_production.html] with 800-fold less DNA template. Such ultra-low-volume/template reactions should make direct Sanger sequencing of clonal PCR amplicons (33) on 5-μm-diameter controlled pore glass microbeads possible. To achieve this ultimate limit of Sanger sequencing, we are working toward a Microbead Integrated DNA Sequencer (MINDS) that parses PCR-colony beads into discrete thermal cycling chambers coupled to purification and electrophoretic separation to produce a fully integrated genome-center-on-a-chip.
In addition to DNA sequencing, the bioprocessor functional units are capable of performing three fundamental biomedical and biological research operations: DNA amplification, nucleic acid concentration/purification, and DNA fragment-size determination. Consequently, the bioprocessor is suitable for a range of applications including genotyping, microsatellite analysis, single nucleotide polymorphism screening, and loss of heterozygosity detection. Integrated, low-volume analysis could be vital in a wide range of applications, such as tumor biopsy screening, forensic analysis, or pathogen detection, where sample is limited and error-free operation is essential.
Sanger sequencing has undergone a remarkable evolution since its radioactive slab-gel beginnings ≈30 years ago. The prevalence and longevity of this core technique results from its unique ability to generate both long reads and highly accurate base calls. It is essential that we drive the cost, efficiency, and scalability limits of this robust sequencing technology to its ultimate limit. Based on our results, template/reagent requirements can be reduced an additional 100-fold, and a fully integrated microfluidic genome sequencing system should also lead to significant infrastructure and labor savings. The bioprocessor demonstrated here is a key step toward the development of a bench-top platform capable of de novo sequencing of plant and animal genomes. The ability to determine long-range genome structure and not simply local sequence uniqueness is critical in complex genome sequencing and is likely to be central in fully understanding the subtle effects of genome organization on development, speciation, and disease.
Materials and Methods
Fabrication and Device Control.
The bioprocessor is constructed from three patterned 100-mm-diameter glass wafers (D263; Schott, Duryea, PA) and a featureless 254-μm-thick PDMS membrane (Bisco, Schaumberg, IL). Patterns are photolithographically transferred to the glass wafers and etched to a depth of 30 μm in 49% HF by using a 2,000-Å-thick amorphous silicon hard mask (34). Holes are drilled for fluidic, electric, pneumatic, and interlayer access. A 1.1-mm-thick wafer containing channel and chamber features etched on the bottom side forms the top of the bioprocessor. Four-point RTDs are fabricated on the top side of a 550-μm-thick wafer coated with 200 Å/2,000 Å Ti/Pt. The bottom side of this wafer is patterned with fluidic lines and microvalve seats in register with the RTDs by using backside alignment lithography (KS Aligner; Süss MicroTec, Santa Clara, CA). The two wafers are thermally bonded in a vacuum furnace at 570°C for 6 h, forming all-glass enclosed features. The bottom side of the bonded stack is UV-ozone-cleaned for 3 min. Similar treatment is performed on the top surface of the PDMS membrane, and the two are assembled immediately. The final layer of the device is a 550-μm-thick wafer containing pneumatic channels and microvalve displacement chambers (26). The wafer and PDMS membrane are UV-ozone-cleaned and assembled as before. The fully assembled device is left overnight under a 2-kg weight, ensuring a permanent bond between the glass layers and the PDMS membrane. Access holes are punched in the PDMS membrane and 1/16-inch barbed pneumatic ports are epoxied to the top surface.
Thermal cycling and microvalve actuation are controlled by a labview graphical interface (National Instruments). A 10-mm-diameter surface heater (7.8 Ω; Minco, Minneapolis) is mounted underneath the thermal cycling reactor. RTDs are contacted by using electrical pins, and 1 mA is passed through the outer leads. The voltage drop across the inner leads as a function of temperature is measured in real-time by using a signal conditioner (5B31-01; Analog Devices) and a data acquisition board (National Instruments). The output drives a proportional integral derivative (PID) controller that regulates 0–8 V for heating and pressurized air for active cooling. Microvalves are closed or opened by applying 5 psi (1 psi = 6.89 kPa) pressure or vacuum, respectively, to the pneumatic ports.
Bioprocessor Preparation.
Before operation, the channel and chamber walls are derivatized after a modified Hjerten coating (35). The system is primed by first filling capture gel [5% linear polyacrylamide/1 × 10−4% bis-acrylamide/40 μM methacrylate-modified DNA/5′-Acrydite-ACTGGCCGTCGTTTTACAA-3′ (Operon Technologies)/1× TTE buffer] to the point of chamber constriction through the capture inlet port. The oligonucleotide sequence is complementary to the 19 bases immediately 3′ of the sequencing primer. Separation gel [in-house-prepared matrix (3.5% linear polyacrylamide/5% DMSO/3 M urea/1× TTE buffer), MegaBACE Long Read Matrix (Amersham Pharmacia), or CEQ Separation Gel LPA-1 (Beckman)], is pushed into the CE unit through the anode port at 600 psi by using a high-pressure loader. Excess gel flows out the waste and cathode ports as well as into the top portion of the capture chamber. Water flushed through the sample inlet port and out the capture outlet port at 73°C removes excess separation gel from the capture chamber. Anode, cathode, waste, capture inlet, and capture outlet ports are filled with 5 μl of 1× TTE buffer.
DNA Sequencing.
Energy-transfer dye-terminator sequencing reagent (1× DYEnamic ET mix (Amersham Pharmacia)/250-nM −47 M13 forward sequencing primer (5′-CGCCAGGGTTTTCCCAGTCACGAC-3′)/4 nM 750-bp pUC18 amplicon/250 ng/μl BSA) is loaded into the 250-nl thermal cycling chamber through the sample port. The chamber is sealed by applying 5 psi pressure to the three pneumatic ports and rapidly thermal cycled for 35 cycles of 95°C (12 s) and 60°C (55 s). Three serial valves used for input, displacement, and output form a diaphragm pump system that is cycled as follows: open input and close output valves (4 s), open displacement valve (200 ms), close input valve (200 ms), open output valve (200 ms), close displacement valve (200 ms). As the reaction mixture enters the upper portion of the capture chamber, a continuously applied 33-V/cm electric field between the capture inlet and outlet ports drives charged extension fragments, template DNA, excess primers, unreacted nucleotides, and salts into the lower, capture-gel-filled chamber held at 30°C. After capture is completed in 20 min, the pumping is turned off, and the electric field is maintained for an additional 5 min to wash the captured products free of residual reagents. Extension fragments are released from the capture gel at 67–75°C and electrokinetically injected for 3–4 s by applying 750 V to the waste port while maintaining the cathode, anode, and capture inlet ports at 650, 250, and 450 V, respectively. Fragment separation occurs at 125–167 V/cm, and 75- and 100-V pull-back potentials at the waste and capture inlet, respectively, prevent leakage of uninjected product. Bands are detected by laser-induced fluorescence on the Berkeley confocal rotary scanner (18).
Sequence Analysis.
Raw electropherogram data were processed with the cimarron 3.12 base-caller (NNIM, Sandy, UT). Additional data processing to obtain base-call accuracy estimates was performed with phred 0.020425.c (30). The “ET Terminators” chemistry profile was specified, causing phred to derive error probabilities from an internal lookup table calibrated for the Molecular Dynamics MegaBACE instrument. phred quality scores relate to the base-call error probability as −10 × log10(Pe), where Pe is the probability that the base call is incorrect. Percent accuracy plots were generated by extrapolating Pe from the phred quality scores and graphing: (Basei − ΣPei)/Basei vs. Basei. Base calls were aligned to the known pUC18 sequence by using the Needleman–Wunsch algorithm (+3 for a correct match reward, −1 for a mismatch penalty, −3 for a gap initiation penalty, and −2 for a gap extension penalty). Read lengths were determined by finding the maximum number of aligned bases with 99% identity.
Acknowledgments
We acknowledge discussions with Brian Paegel on sample purification; with William Grover and Alison Skelley on microvalve design; and with Charles Emrich, Teris Liu, and Nicholas Toriello on microdevice fabrication. Microfabrication was carried out at the University of California, Berkeley, Microfabrication Laboratory. This work was supported by National Institutes of Health (NIH) Grants HG01399 and HG003583 via Microchip Biotechnologies, Inc. R.G.B. was supported by NIH Trainee Fellowship HG00047 from the Berkeley Program in Genomics.
Abbreviations
- CE
capillary electrophoresis
- PDMS
polydimethylsiloxane
- RTD
resistive temperature detector.
Footnotes
Conflict of interest statement: R.A.M. is a consultant with Microchip Biotechnologies Inc. (MBI) and has a financial interest in MBI. MBI is working on the commercialization of microchip sequencing technologies and may benefit from the results of this research.
References
- 1.Zhao S. Y., Shetty J., Hou L. H., Delcher A., Zhu B. L., Osoegawa K., de Jong P., Nierman W. C., Strausberg R. L., Fraser C. M. Genome Res. 2004;14:1851–1860. doi: 10.1101/gr.2663304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Cheng Z., Ventura M., She X. W., Khaitovich P., Graves T., Osoegawa K., Church D., DeJong P., Wilson R. K., Paabo S., et al. Nature. 2005;437:88–93. doi: 10.1038/nature04000. [DOI] [PubMed] [Google Scholar]
- 3.Check E. Nature. 2005;437:1084–1086. doi: 10.1038/4371084a. [DOI] [PubMed] [Google Scholar]
- 4.National Institutes of Health. Near-Term Technology Development for Genome Sequencing. Bethesda: Natl. Inst. of Health; 2004. Feb 12, RFA-HG-04-002. [Google Scholar]
- 5.Shendure J., Porreca G. J., Reppas N. B., Lin X. X., McCutcheon J. P., Rosenbaum A. M., Wang M. D., Zhang K., Mitra R. D., Church G. M. Science. 2005;309:1728–1732. doi: 10.1126/science.1117389. [DOI] [PubMed] [Google Scholar]
- 6.Margulies M., Egholm M., Altman W. E., Attiya S., Bader J. S., Bemben L. A., Berka J., Braverman M. S., Chen Y. J., Chen Z. T., et al. Nature. 2005;437:376–380. doi: 10.1038/nature03959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lander E. S., Linton L. M., Birren B., Nusbaum C., Zody M. C., Baldwin J., Devon K., Dewar K., Doyle M., FitzHugh W., et al. Nature. 2001;409:860–921. doi: 10.1038/35057062. [DOI] [PubMed] [Google Scholar]
- 8.Shapiro J. A., von Sternberg R. Biol. Rev. 2005;80:227–250. doi: 10.1017/s1464793104006657. [DOI] [PubMed] [Google Scholar]
- 9.Gonzalez E., Kulkarni H., Bolivar H., Mangano A., Sanchez R., Catano G., Nibbs R. J., Freedman B. I., Quinones M. P., Bamshad M. J., et al. Science. 2005;307:1434–1440. doi: 10.1126/science.1101160. [DOI] [PubMed] [Google Scholar]
- 10.Lengauer C., Kinzler K. W., Vogelstein B. Nature. 1998;396:643–649. doi: 10.1038/25292. [DOI] [PubMed] [Google Scholar]
- 11.C. elegans Sequencing Consortium. Science. 1998;282:2012–2018. doi: 10.1126/science.282.5396.2012. [DOI] [PubMed] [Google Scholar]
- 12.Arabidopsis Genome Initiative. Nature. 2000;408:796–815. doi: 10.1038/35048692. [DOI] [PubMed] [Google Scholar]
- 13.Adams M. D., Celniker S. E., Holt R. A., Evans C. A., Gocayne J. D., Amanatides P. G., Scherer S. E., Li P. W., Hoskins R. A., Galle R. F., et al. Science. 2000;287:2185–2195. doi: 10.1126/science.287.5461.2185. [DOI] [PubMed] [Google Scholar]
- 14.Paegel B. M., Blazej R. G., Mathies R. A. Curr. Opin. Biotechnol. 2003;14:42–50. doi: 10.1016/s0958-1669(02)00004-6. [DOI] [PubMed] [Google Scholar]
- 15.Woolley A. T., Mathies R. A. Anal. Chem. 1995;67:3676–3680. doi: 10.1021/ac00116a010. [DOI] [PubMed] [Google Scholar]
- 16.Paegel B. M., Emrich C. A., Weyemayer G. J., Scherer J. R., Mathies R. A. Proc. Natl. Acad. Sci. USA. 2002;99:574–579. doi: 10.1073/pnas.012608699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Aborn J. H., El-Difrawy S. A., Novotny M., Gismondi E. A., Lam R., Matsudaira P., McKenna B. K., O'Neil T., Streechon P., Ehrlich D. J. Lab Chip. 2005;5:669–674. doi: 10.1039/b501104c. [DOI] [PubMed] [Google Scholar]
- 18.Shi Y. N., Simpson P. C., Scherer J. R., Wexler D., Skibola C., Smith M. T., Mathies R. A. Anal. Chem. 1999;71:5354–5361. doi: 10.1021/ac990518p. [DOI] [PubMed] [Google Scholar]
- 19.Nakane J., Broemeling D., Donaldson R., Marziali A., Willis T. D., O'Keefe M., Davis R. W. Genome Res. 2001;11:441–447. doi: 10.1101/gr.164401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Dittrich P. S., Manz A. Anal. Bioanal. Chem. 2005;382:1771–1782. doi: 10.1007/s00216-005-3335-9. [DOI] [PubMed] [Google Scholar]
- 21.Gardeniers J. G. E., van den Berg A. Anal. Bioanal. Chem. 2004;378:1700–1703. doi: 10.1007/s00216-003-2435-7. [DOI] [PubMed] [Google Scholar]
- 22.Hansen C., Quake S. R. Curr. Opin. Struct. Biol. 2003;13:538–544. doi: 10.1016/j.sbi.2003.09.010. [DOI] [PubMed] [Google Scholar]
- 23.Krishnan M., Namasivayam V., Lin R. S., Pal R., Burns M. A. Curr. Opin. Biotechnol. 2001;12:92–98. doi: 10.1016/s0958-1669(00)00166-x. [DOI] [PubMed] [Google Scholar]
- 24.Lagally E. T., Scherer J. R., Blazej R. G., Toriello N. M., Diep B. A., Ramchandani M., Sensabaugh G. F., Riley L. W., Mathies R. A. Anal. Chem. 2004;76:3162–3170. doi: 10.1021/ac035310p. [DOI] [PubMed] [Google Scholar]
- 25.Skelley A. M., Scherer J. R., Aubrey A. D., Grover W. H., Ivester R. H. C., Ehrenfreund P., Grunthaner F. J., Bada J. L., Mathies R. A. Proc. Natl. Acad. Sci. USA. 2005;102:1041–1046. doi: 10.1073/pnas.0406798102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Grover W. H., Skelley A. M., Liu C. N., Lagally E. T., Mathies R. A. Sens. Actuator B-Chem. 2003;89:315–323. [Google Scholar]
- 27.Karlinsey J. M., Monahan J., Marchiarullo D. J., Ferrance J. P., Landers J. P. Anal. Chem. 2005;77:3637–3643. doi: 10.1021/ac048115z. [DOI] [PubMed] [Google Scholar]
- 28.Paegel B. M., Hutt L. D., Simpson P. C., Mathies R. A. Anal. Chem. 2000;72:3030–3037. doi: 10.1021/ac000054r. [DOI] [PubMed] [Google Scholar]
- 29.Paegel B. M., Yeung S. H. I., Mathies R. A. Anal. Chem. 2002;74:5092–5098. doi: 10.1021/ac0203645. [DOI] [PubMed] [Google Scholar]
- 30.Ewing B., Hillier L., Wendl M. C., Green P. Genome Res. 1998;8:175–185. doi: 10.1101/gr.8.3.175. [DOI] [PubMed] [Google Scholar]
- 31.Kotler L., He H., Miller A. W., Karger B. L. Electrophoresis. 2002;23:3062–3070. doi: 10.1002/1522-2683(200209)23:17<3062::AID-ELPS3062>3.0.CO;2-G. [DOI] [PubMed] [Google Scholar]
- 32.Doherty E. A. S., Kan C. W., Paegel B. M., Yeung S. H. I., Cao S. T., Mathies R. A., Barron A. E. Anal. Chem. 2004;76:5249–5256. doi: 10.1021/ac049721x. [DOI] [PubMed] [Google Scholar]
- 33.Dressman D., Yan H., Traverso G., Kinzler K. W., Vogelstein B. Proc. Natl. Acad. Sci. USA. 2003;100:8817–8822. doi: 10.1073/pnas.1133470100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Simpson P. C., Roach D., Woolley A. T., Thorsen T., Johnston R., Sensabaugh G. F., Mathies R. A. Proc. Natl. Acad. Sci. USA. 1998;95:2256–2261. doi: 10.1073/pnas.95.5.2256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hjerten S. J. Chromatogr. 1985;347:191–198. [Google Scholar]