Abstract
Ribosome biogenesis is a complex process that requires >150 transacting factors, many of which form macromolecular assemblies as big and complex as the ribosome itself. One of those complexes, the SSU processome, is required for pre-18S rRNA maturation. Although many of its components have been identified, the endonucleases that cleave the pre-18S rRNA have remained mysterious. Here we examine the role of four previously uncharacterized PINc domain proteins, which are predicted to function as nucleases, in yeast ribosome biogenesis. We also included Utp23, a protein homologous to the PINc domain protein Utp24, in our analysis. Our results demonstrate that Utp23 and Utp24 are essential nucleolar proteins and previously undescribed components of the SSU processome. In that sense, both Utp23 and Utp24 are required for the first three cleavage steps in 18S rRNA maturation. In addition, single-point mutations in the conserved putative active site of Utp24 but not Utp23 abrogate its function in ribosome biogenesis. Our results suggest that Utp24 might be the elusive endonuclease that cleaves the pre-rRNA at sites A1 and/or A2.
Keywords: ribosome biogenesis, small subunit processome, endonuclease, yeast, RNA processing
Ribosomes translate mRNA into proteins and, thereby, govern cell growth and survival. In Saccharomyces cerevisiae, ribosome biogenesis starts with the transcription of the rDNA into a polycistronic rRNA precursor by RNA polymerase I in the nucleolus. The primary transcript, the 35S pre-rRNA, encodes the small subunit (SSU) rRNA, 18S, the large subunit rRNAs, 25S and 5.8S, and also external and internal transcribed spacer regions (5′ETS, ITS1, ITS2, and 3′ETS; Fig. 1). This precursor is extensively endo- and exonucleolytically cleaved to mature the rRNAs. Maturation begins with endonucleolytic cleavages at sites A0, A1, and A2, the latter separating the 20S pre-rRNA, the direct precursor to the mature 18S rRNA, and the 27SA2 pre-rRNA, the precursor to the large subunit rRNAs. The 20S pre-rRNA is exported into the cytoplasm as part of the 43S preribosome, where the 18S rRNA is matured by cleavage at site D.
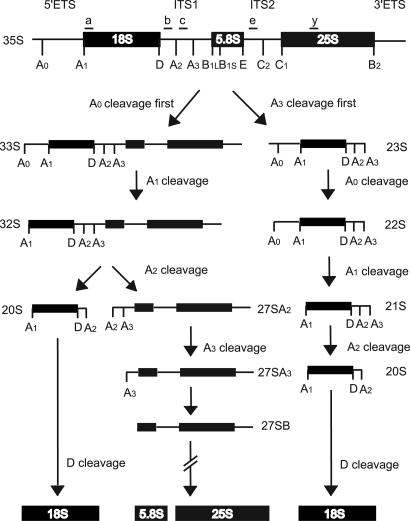
Fig. 1.
Pre-rRNA processing in S. cerevisiae. The 35S pre-rRNA encoding the 18S, 5.8S, and 25S rRNAs is processed at sites A0, A1, A2, and A3 via several intermediates to yield the 20S pre-rRNA. The 20S pre-rRNA cleavage at site D matures the 18S rRNA, whereas numerous endo- and exonucleolytic cleavages in the nucleolus mature the 5.8S and 25S rRNAs from the 27SA2 or 27SA3 precursor. Small letters above the primary transcripts indicate the oligonucleotides used. The maturation of the large subunit rRNAs downstream of the 27SA3 pre-rRNA is the same for each pathway.
The A0, A1, and A2 pre-rRNA cleavages require a large ribonucleoprotein complex, the SSU processome/90S preribosome, which contains at least 40 different proteins and numerous small nucleolar RNAs (snoRNAs) (1–4). This complex likely corresponds to the terminal knobs seen in Miller chromatin spreads (1, 5). Depletion of essential SSU processome components prevents cleavages at sites A0–A2 but not A3. Although this results in accumulation of the 23S pre-rRNA and inhibition of 18S synthesis, maturation of the 25S and 5.8S rRNAs can occur normally (Fig. 1).
Despite the identification of numerous proteins involved in early pre-18S rRNA processing by large-scale purifications, the endonuclease(s) and possible candidates that carry out these cleavages have remained elusive. Recently, an interesting group of proteins possessing a PIN domain (PilT N terminus, PINc domain in SMART database) has been implicated as potential nucleases in eukaryotic RNA metabolism (6, 7).
In S. cerevisiae, seven PINc domain proteins are classified in the SMART database. Only three have been previously characterized, including Nmd4, a protein involved in nonsense-mediated mRNA decay, Rrp44, an exosome component, and Nob1. Nob1 is a component of the 43S preribosome and has been suggested to be the endonuclease that cleaves the 20S pre-rRNA at site D in the cytoplasm (8, 9). Although the nuclease function of eukaryotic PIN domain proteins has not yet been demonstrated in vitro, the function of a subset of them in RNA metabolism and the conservation of the putative active site residues make them attractive candidates.
Here, we investigated the role of the remaining four yeast PINc domain proteins, YDR339c, YIL151c, YKR096w, and YOR166c, as well as the potential PINc domain protein YOR004w in ribosome biogenesis. Our results demonstrate that two of these proteins, YOR004w and YDR339c, encode essential nucleolar proteins that are required for processing of the pre-rRNA at sites A0–A2 as previously undescribed components of the SSU processome. Consequently, we named these proteins Utp23 and Utp24 (for U three-associated protein), respectively. Furthermore, point mutations in the putative active site of Utp24 are required for its function in A1 and A2 cleavage but dispensable for cleavage at site A0, thereby uncoupling these cleavage events. In contrast, mutations in conserved acidic residues of Utp23 did not interfere with rRNA maturation and cell viability, which is consistent with the idea that an intact PINc-like domain in Utp23 is not required for function, and, hence, Utp23 is unlikely to be a bona fide PINc domain protein.
Results
Utp24 Contains a Conserved PINc Domain and Shows Significant Sequence Similarity to Utp23.
In a search for potential nucleases involved in pre-rRNA processing in S. cerevisiae, we identified four previously uncharacterized proteins that are predicted to contain a PINc domain in the SMART database: YDR339c (Utp24), YIL151c, YKR096w, and YOR166c. Most of the PINc domain proteins share four conserved acidic amino acid residues that are predicted to be involved in metal ion binding during catalysis as well as a conserved threonine or serine residue upstream of the last conserved acidic amino acid residue (refs. 6 and 7; Fig. 2; see also Fig. 7, which is published as supporting information on the PNAS web site). In addition to the classified PINc domain proteins, homology searches with Utp24 revealed that the protein sequence of Utp23 (YOR004w) is 24% identical and 47% similar to that of Utp24, and two of the four acidic amino acid residues as well as the serine/threonine residue defining the PINc domain are conserved in Utp23 (Figs. 2 and 7). Intriguingly, protein sequence alignment of Utp23 homologs in different species demonstrates that a total of four acidic amino acid residues, including the above, are conserved from low to high eukaryotes (Fig. 8, which is published as supporting information on the PNAS web site). Therefore, we hypothesized that Utp23 also might contain a PINc domain and included it in our analysis.
Fig. 2.
Schematic of the classical PINc domain with conserved putative active-site residues. Residue numbers below the scheme represent corresponding amino acids in Utp23 and Utp24 based on the detailed alignment shown in Fig. 7.
Utp23 and Utp24 Are Essential for SSU Biogenesis.
To analyze the function of the uncharacterized PINc domain proteins, yeast strains were generated that express the respective protein under the control of a conditional tetracycline repressible promoter (tetO7). Depletion of these proteins with doxycyline revealed that Utp23 and Utp24 are essential for growth, with a major shift in growth rate after 10 h of depletion (Fig. 3A), whereas depletion of YOR166c, YIL151c, and YKR096w did not impair growth at all tested temperatures (17°C, 23°C, 30°C, and 35°C; data not shown). This effect is consistent with results from a large-scale deletion project that found YKR096w and YOR166c dispensable for cell viability (10).
Fig. 3.
Utp23 and Utp24 are essential for SSU pre-rRNA processing. (A) Depletion of Utp23 and Utp24 impairs cell growth. Growth of wild-type cells (squares), tetO7::Utp23 cells (circles), and tetO7::Utp24 cells (triangles) was compared in the presence (open symbols) or absence (filled symbols) of doxycycline. The OD600 was measured up to 40 h after the addition of doxycycline. (B) Depletion of Utp23 and Utp24 affects SSU biogenesis. Twenty micrograms of total RNA from cells depleted and undepleted of either Utp23 or Utp24, respectively, was separated on 1.25% formaldehyde/agarose gels. RNA from depleted and undepleted cells was run on the same gel and subjected to the same exposure. Pre-rRNAs and mature rRNAs were detected by Northern blotting with oligonucleotides C (detects 35S, 33/32S, 27SA2, and 23S pre-rRNAs), B and E (also label the 27SA3, 27SB, and 20S pre-rRNAs), or A and Y (label mature 18S and 25S rRNAs). (C) Depletion of Utp23 or Utp24 impairs pre-rRNA processing but does not affect rRNA stability. Yeast cells depleted of Utp23 or Utp24 for 10 h with doxycycline were labeled metabolically with 3H-uracil for 2 min and incubated with excess of unlabeled uracil for 0, 3, 12, 30, and 60 min. Twenty thousand counts per million of RNA were separated as in B, transferred to Hybond N+ membrane, and detected by autoradiography. The parental strain YPH499 also was grown in 2 μg/ml doxycycline for 10 h.
To test whether the severe growth phenotype of Utp24 depletion is due to impaired ribosome biogenesis and to characterize the previously described 18S maturation defect for Utp23 in greater detail, we analyzed steady-state levels of pre-rRNAs and mature rRNAs in cells depleted of Utp23 or Utp24 (11). Depletion of either protein resulted in accumulation of the 35S pre-rRNA, which was accompanied by a decrease in the levels of the 27SA2 and 20S pre-rRNAs and the mature 18S rRNA (Fig. 3B, compare lanes 2 and 5, 3 and 6, and 4 and 7 for Utp23 and lanes 9 and 12, 10 and 13, and 11 and 14 for Utp24). Depletion of Utp24 resulted in a stronger processing phenotype because it also led to accumulation of the 23S pre-rRNA. In contrast, the large subunit rRNAs, 25S, 5.8S, and 5S, were not significantly reduced, resulting in a decline in the 18S/25S rRNA ratio upon depletion of either Utp23 or Utp24 (Fig. 3B; see also Fig. 9 A and B, which are published as supporting information on the PNAS web site). Importantly, the levels of the U3 and U14 snoRNAs, two box C/D type snoRNAs directly involved in pre-rRNA processing, did not decrease, excluding indirect effects caused by affecting box C/D snoRNA biogenesis or stability (Fig. 9B). To substantiate our results, we metabolically labeled yeast cells that were depleted of Utp23 or Utp24 with [3H]uracil (Fig. 3C). Processing of SSU pre-rRNAs into the 20S pre-rRNA and into the mature 18S rRNA was inefficient in yeast depleted of Utp23 or Utp24 as compared with the parental strain and resulted in accumulation of the 35S and 23S pre-rRNAs, whereas processing of 27SA3/B pre-rRNAs into the 25S rRNA was unaffected. In contrast to Utp23 and Utp24, depletion of the nonessential proteins YOR166c, YIL151c, and YKR096w did not affect pre-rRNA processing (data not shown). Together, these results strongly argue for a direct role for Utp23 and Utp24 in SSU biogenesis, specifically in processing of the pre-rRNA at sites A0, A1, and A2.
Utp23 and Utp24 Localize to the Nucleolus.
Ribosome biogenesis starts in the nucleolus, and many ribosome maturation factors localize to this subcellular compartment. Therefore, we asked whether Utp23 and Utp24 also localize to the nucleolus. Indirect immunofluorescence experiments were performed to analyze the cellular distribution of hemagglutinin (HA)-tagged Utp23 and Utp24, respectively. Both proteins and Utp8, a positive control protein, localized to a crescent-shaped compartment next to the nucleus and colocalized with Mpp10, a nucleolar marker protein (Fig. 4). Therefore, we conclude that Utp23 and Utp24 are mainly nucleolar, being consistent with our biochemical data indicating a role of both proteins in 18S maturation.
Fig. 4.
Utp23 and Utp24 are nucleolar proteins. HA-tagged Utp23, Utp24, and the positive control protein Utp8 were localized with α-HA antibodies and rhodamine-conjugated α-mouse IgG. Mpp10, a nucleolar marker protein, was detected with polyclonal antibodies against Mpp10 and fluorescein-conjugated α-rabbit IgG. Nuclei were stained with DAPI. The parental strain YPH499 was used as a negative control.
Utp23 and Utp24 Are Components of the SSU Processome.
The endonucleolytic cleavage events at sites A0–A2 require the SSU processome, an 80S complex that assembles on the 35S pre-rRNA. Because Utp23 and Utp24 also are required for these cleavages, we tested whether these proteins represent previously undescribed components of the SSU processome. Immunoprecipitation experiments using Utp23 or Utp24 epitope-tagged proteins as bait resulted in weak but consistent coimmunoprecipitation of Mpp10, an SSU processome-specific protein, and the U3 snoRNA above background levels (mock; Figs. 5A and B). HA-tagged Utp8, a protein previously shown to be part of the SSU processome, was used as a positive control. The low immunoprecipitation efficiency of Utp23 and Utp24 as compared with Utp8 might result from an only transient association of these proteins with the SSU processome or of a reduced accessibility of the epitope tag. Furthermore, both Utp23 and Utp24 migrate at 80S, the size of the SSU processome, in sucrose density gradients (data not shown).
Fig. 5.
Utp23 and Utp24 are associated with the SSU processome. (A) Utp23 and Utp24 coimmunoprecipitate Mpp10, an SSU processome protein. α-HA antibodies were used to immunoprecipitate HA-tagged Utp23, Utp24, and Utp8. Immunoprecipitates (IP) and 5% of total input (T) were separated by SDS/PAGE, and Mpp10 was detected by Western blotting. (B) Utp23 and Utp24 coimmunoprecipitate the U3 snoRNA. RNA was extracted from IPs (performed as in A) and 5% T, separated in 8% urea/acrylamide gels, and detected by Northern blotting. (C) Depletion of either Utp23 or Utp24 results in failure to form SSU processomes on nascent rRNA transcripts. Shown are representative rRNA genes from chromatin spreads of cells undepleted or 18 h depleted of Utp23 or Utp24. Small arrows indicate typical SSU processome formation on nascent transcripts, brackets indicate shorter transcripts after cotranscriptional cleavage in ITS1, and open arrows indicate the lack of SSU processome formation.
The SSU processome has been suggested to correspond to the terminal knobs capping the nascent rRNA transcript in chromatin spreads. To examine the role of Utp23 and Utp24 in SSU processome assembly and terminal knob formation, we compared chromatin spreads from yeast undepleted or depleted of Utp23 or Utp24 (Fig. 5C). Genes from undepleted cells show typical SSU processome formation on nascent transcripts and shorter transcripts at the 3′ end of the gene resulting from cotranscriptional cleavage in ITS1 (5). After depletion of either Utp23 or Utp24, the SSU processome does not form and transcripts are full-length when they reach the end of the genes. Thus, our results substantiate our hypothesis that Utp23 and Utp24 are specifically required for pre-rRNA cleavages as part of the SSU processome.
The Putative Active Site of Utp24, but Not Utp23, Is Essential for Protein Function.
The putative active site of PINc domain proteins is characterized by four conserved acidic amino acids that are involved in divalent metal ion binding and enzymatic activity. In Utp23, they include D31, E101, D123, and E149. In Utp24, these residues are D68, E105, D138, and D157 (Figs. 2, 7, and 8). We tested whether these residues are essential for Utp23 and Utp24 protein function, as would be expected if these proteins were indeed PINc domain proteins. For that, TAP-tagged Utp23 and Utp24 were cloned into the constitutive low-copy yeast expression vector p415GPD. We mutated the conserved acidic amino acid residues, substituting aspartic acid with asparagine (D31N and D123N in Utp23 and D68N and D138N in Utp24) and glutamic acid with glutamine (E101Q and E149Q in Utp23). Such mutations have been shown to interfere with protein function, most likely by disrupting metal ion coordination without strongly affecting other biophysical properties (12). Western blot analysis demonstrated that both wild-type and mutant Utp23 and Utp24 proteins were expressed at least at the levels of the endogenously TAP-tagged proteins from control strains (data not shown).
Next, we tested whether expression of TAP-tagged wild-type and mutant Utp23 or Utp24 could rescue the growth and rRNA processing defects in cells depleted of the respective endogenous proteins. Surprisingly, wild-type and all mutant Utp23 proteins were able to support growth of Utp23-depleted cells at all tested temperatures (Fig. 6A). Thus, our results are consistent with the current SMART database classification that does not include Utp23 into the PINc domain protein family.
Fig. 6.
Mutations in the putative active site of Utp24 disrupt protein function. (A) Acidic residues in Utp24 but not Utp23 are essential for in vivo protein function. Ten-fold serial dilutions of yeast cells containing empty vector (−), wild-type (WT), or indicated mutant Utp23 or Utp24 proteins were analyzed for growth in the absence (−Dox) and presence (+Dox) of doxycycline at 17°C, 23°C, 30°C, and 35°C. The pictures were modified so that the dots align. (B) Expression of Utp24 mutants results in accumulation of the 22S pre-rRNA. Cells transformed with empty vector or expressing WT Utp24 or the Utp24 mutants D68N or D138N from p415GPD were harvested 0, 10, and 20 h after growth in media with (depleted) or without (undepleted) doxycycline. Total RNA was extracted and equal amounts were separated on 1.25% formaldehyde/agarose gels. Pre-rRNAs were detected by Northern blotting with oligos B and E.
In contrast, the wild-type TAP-tagged Utp24, but neither of the mutant Utp24 proteins, D68N and D138N, was able to rescue the growth defect observed in Utp24-depleted cells (+Dox; Fig. 6A). Interestingly, expression of the D68N and especially the D138N mutant Utp24 also conferred a dominant negative phenotype in the presence of the endogenous wild-type Utp24 protein (−Dox). This effect was most prominent at 17°C and 35°C.
To test the effect of the Utp24 mutants on pre-rRNA processing, we analyzed the rRNA precursors by Northern blotting. As expected, cells expressing empty p415GPD vector showed a typical SSU rRNA processing defect after depletion of endogenous Utp24 with accumulation of the 35S and 23S and reduction of the 27SA2 and 20S pre-rRNAs (Fig. 6B). This processing defect could be rescued successfully by expression of wild-type Utp24 from the p415GPD plasmid. Interestingly, expression of either the D68N or D138N Utp24 mutants resulted in accumulation of the 22S pre-rRNA, an intermediate extending from sites A0 to A3, which hybridized to oligonucleotides complementary to sequences between A0 and A1 as well as between A2 and A3 (Fig. 6B and data not shown). Intriguingly, the 22S pre-rRNA also was present in Utp24 undepleted yeast that express the mutant Utp24 proteins. This delay in processing at sites A1 and A2 correlates with the dominant negative growth phenotype observed in these cells.
Therefore, we conclude that conserved acidic residues in the putative active site of Utp24 are essential for pre-rRNA processing at sites A1 and A2, whereas the presence of the Utp24 protein itself also is required for cleavage at A0. Despite its sequence similarity to Utp24, Utp23 likely is not a PIN domain protein because the conserved acidic amino residues are not essential for its function.
Discussion
PINc domain proteins have been implicated to function as ribonucleases (6). A subset of these proteins, Nob1, Nmd4, and Rrp44, are involved in RNA metabolism, including ribosome biogenesis. With this rationale, we investigated whether additional members of the PINc domain family in yeast, Utp24 (YDR339c), YIL151c, YKR096w, YOR166c, and the potential PIN domain protein Utp23 (YOR004w) function in ribosome biogenesis and, if so, whether the putative active site is required for their function in vivo. Our results demonstrate that only Utp23 and Utp24 are essential for SSU biogenesis. Consistent with the predicted enzymatic activity of PINc domain proteins, mutations in the putative active site of Utp24 are incompatible with viability and SSU maturation. Interestingly, these mutations do not impair cleavage at A0 of the pre-rRNA. This result suggests that although Utp24 is essential for cleavages at A0–A2 of the pre-rRNA, the predicted enzymatic function of the PINc domain itself is only critical to cleavage at sites A1 and A2.
More than 40 proteins have been found to be required for the early steps in SSU pre-rRNA processing, mostly by purification of preribosomal particles (1, 2, 13). However, the list of proteins involved in early steps of SSU biogenesis is not complete, as new components of early pre-rRNA processing complexes continue to be identified (4, 14–17). Here we add two previously undescribed proteins to that list, Utp23 and Utp24, based on the following lines of evidence: First, conditional depletion of either Utp23 or Utp24 specifically inhibits the A0–A2 cleavages essential for SSU biogenesis (Fig. 3 B and C). Second, Utp23 and Utp24 are nucleolar proteins (Fig. 4). Third, both Utp23 and Utp24 associate at least transiently with Mpp10 and the U3 snoRNA, two components of the SSU processome (Figs. 5 A and B). Fourth, depletion of Utp23 or Utp24 results in the loss of SSU processomes on nascent transcripts (Fig. 5C). Consistent with our results, a very recent large-scale proteome survey found Utp23 (YOR004w) copurifying with Mrd1, Rrp5, and Utp22 (18). All of the aforementioned proteins have been reported to be involved in early steps of 18S rRNA biogenesis, and Rrp5 and Utp22 have been purified as components of the SSU processome/90S preribosome (1, 2, 13, 18–20). Thus, Utp23 and Utp24 represent two previously undescribed bona fide SSU processome components.
In addition to Utp24, we also investigated the role of three other classified PINc domain proteins, YIL151c, YKR096w, and YOR166c, in ribosome biogenesis. Conditional depletion of these proteins does not affect yeast viability or rRNA processing (data not shown). However, we cannot exclude that these proteins perform nonessential functions in ribosome biogenesis, especially YKR096w, which has been reported to localize to the nucleus and cytoplasm and has been found to affinity purify with Utp18, an SSU processome component (18, 21–23).
Bioinformatic analysis of numerous PIN domain proteins recently suggested homology of the PIN domain proteins with the family of 5′ nuclease domain proteins that include T4 RNase H, the 5′ to 3′ exonuclease domain of Taq polymerase I, T5 5′ exonuclease, and the Flap endonuclease Fen-1 (6). Strikingly, both families share four conserved acidic residues that are essential for either divalent metal ion binding or catalysis in 5′ nucleases, and the conserved acidic residues in the archeal PIN domain protein PAE2754 structurally align with the catalytic site residues of T4 RNase H (7). This observation predicts that these conserved residues also would be essential for the enzymatic function of PIN domain proteins. Indeed, we demonstrate that single-point mutations in either of the two tested conserved acidic residues predicted to be in the active site of Utp24 are essential for its in vivo function in SSU biogenesis (Fig. 6). Intriguingly, yeast cells expressing mutant Utp24 protein slightly differed in their rRNA processing defects as compared with depletion of the endogenous Utp24 protein: Although depletion of endogenous Utp24 impairs cleavages at sites A0–A2 of the pre-rRNA, the mutant Utp24 proteins are able to rescue the cleavage defect at site A0, but not A1 and A2. We hypothesize that Utp24 plays two distinct roles in early pre-rRNA processing: (i) it is either a structural component of the A0 cleavage complex or essential for its formation, which is independent of an intact PINc domain, and (ii) it is required for cleavages at sites A1 and A2, which depends on the PINc domain and the proposed catalytic activity of Utp24. In addition to the lack of complementation of the wild-type protein function, we also observed a dominant negative phenotype when expressing the mutant Utp24 proteins. Dominant negative phenotypes usually result from competition of wild-type and mutant proteins for the same substrate, thereby trapping it in nonfunctional complexes. Together, these results suggest the PINc domain protein Utp24 functions as an enzyme in ribosome biogenesis.
Previous work by Fatica et al. (8) demonstrated that Nob1, one member of the yeast PINc protein family, is essential for SSU biogenesis. In contrast to Utp24, it is not required for the first three nucleolar cleavage steps at site A0–A2 but for the final cleavage step at site D in the cytoplasm that matures the 18S rRNA from the 20S pre-rRNA. Consistent with this function, Nob1 is not associated with the SSU processome/90S preribosome but instead is a component of the 43S preribosome (20). Interestingly, of the two mutations in the predicted active site of Nob1, one abolished Nob1 protein function in vivo (9). Therefore, Nob1 was suggested to be the endonuclease that cleaves the 20S pre-rRNA at site D. However, the proposed nuclease activity of Nob1 has so far not been demonstrated in vitro.
Is Utp24 the endonuclease that processes the 35S pre-rRNA into the 20S pre-rRNA in the nucleolus? Maturation of the 18S rRNA requires four cleavage events at sites A0–A2 and D. These processing events are likely to be endonucleolytic, because the cleaved transcribed spacers accumulate in yeast cells defective of the 5′ to 3′ exoribonucleases Rat1 and Xrn1 (24). The enzymes that perform these cleavages, however, have remained elusive. Our result concerning the in vivo function of the PINc domain protein Utp24, especially the effect of its point-mutant proteins, are consistent with the idea that Utp24 might be the endonuclease cleaving the pre-rRNA at sites A1 and/or A2. However, we cannot rule out that the putative active site of Utp24 is essential for formation of the A1/A2 cleavage competent complex rather than performing the catalysis itself. Yet another possibility is that Utp24 functions as an exonuclease in SSU biogenesis, although there is no direct evidence that either the 5′ or 3′ end of the SSU pre-rRNAs are exonucleolytically modified. That PIN domain proteins can function as exonucleases recently has been demonstrated by Arcus et al. (7) in vitro for the archeal protein PAE2754. Intriguingly, 5′ nucleases like Fen-1 are able to function as both endo- and exonucleases, suggesting that PIN domain proteins might be able to perform multiple different nucleolytic events (25).
It is interesting to note that in addition to Nob1 and Utp24, a third PINc domain protein, Rrp44, is involved in ribosome biogenesis. As a nucleolar exosome component, Rrp44 has 3′ to 5′ exonuclease activity that has been demonstrated in vitro (26). Because Rrp44 also contains a ribonuclease II domain, the contribution of the PINc domain to the 3′ to 5′ exonucleolytic activity remains elusive. Experimental proof that Utp24, whose PINc domain is required for processing at sites A1 and A2, does function as a nuclease in SSU biogenesis and, if so, whether it functions as an endonuclease or exonuclease remains a possible subject for future in vitro studies.
Materials and Methods
Strains and Media.
S. cerevisiae strains are derived from YPH499 (MATa ura3-52 lys2-80 ade2-101 trp1-Δ63 his3-Δ200 leu2-Δ1). Cells were grown in 1% yeast extract/2% peptone/2% glucose media. Cells containing the p415GPD plasmid were grown in SD-LEU media. For depletion of proteins under the control of the tetO7 promoter, 2 μg/ml doxycycline was added to the media. HA-tagged, TAP-tagged, or tetO7-depletion strains were created as described in refs. 27–29.
Growth Analysis.
Growth was monitored at 30°C in liquid culture in the absence and presence of 2 μg/ml doxycycline by measuring the OD600 up to 40 h. Cells were maintained in early log phase. For growth analysis on plates, 400 μl of cells with an OD600 of 0.5 were harvested and 10-fold serially diluted, spotted onto media plates with or without doxycycline, and grown at the indicated temperatures.
RNA and Protein Manipulations.
Steady-state RNA analysis of cells grown to mid-log phase was performed as described in ref. 30 with the following modifications: 20 μg of total RNA was separated on 1.25% formaldehyde/agarose gels or 8% urea/polyacrylamide gels and transferred to Hybond N+ membrane. Oligonucleotides hybridizing to pre-rRNAs and mature rRNAs are detailed in refs. 31 and 32 except for oligonucleotide C (CCT CTG GGC CCC GAT TGC TCG AA; Fig. 1).
For metabolic labeling, 40 ml of cells grown in SD-URA to an OD600 of 0.5 were harvested and pulsed with 100 μCi [3H]uracil (1 Ci = 37 GBq) for 2 min. Cells were diluted 10-fold into SD media and chased for the indicated time periods. RNA was extracted and separated in 1.25% agarose/formaldehyde gels as described in ref. 30.
For Western blot analysis, proteins were separated by SDS/PAGE on 10% gels, transferred to Millipore membrane, and incubated with anti-Mpp10 and anti-rabbit IgG (Amersham Pharmacia) at 1:10,000 or peroxidase anti-peroxidase antibodies (α-TAP; Sigma) at 1:5,000 dilutions (30).
Immunofluorescence.
Subcellular localization of HA-tagged proteins was performed by indirect immunofluorescence as described in (33) with the following modifications: monoclonal anti-HA antibodies (12CA5) and rhodamine conjugated polyclonal anti-mouse IgG were diluted 1:1,000 and 1:10,000, respectively. The polyclonal rabbit anti-Mpp10 antibodies and fluorescein-conjugated polyclonal anti-rabbit IgG were used at dilutions of 1:2,000 and 1:3,000 (30).
Immunoprecipitations.
Twenty milliliters of cell culture at an OD600 of 0.5 were harvested, lysed with acid treated glass beads, and subjected to immunoprecipitation experiments (30, 34). Two hundred microliters of α-HA (culture supernatant; 12CA5) were used. Five percent of the input and immunoprecipitated RNA and proteins were analyzed by Northern and Western blotting as described above.
Miller Chromatin Spreads.
Miller chromatin spreads were done as described in ref. 5 by using 11 mM KCl for chromatin dispersal.
Cloning and Mutagenesis.
C-terminally TAP-tagged Utp23 and Utp24 genes were amplified by PCR from genomic DNA adding BamHI and PstI restriction sites to the 5′ end of the gene and the 3′ of the TAP-tag, respectively. PCR products were cloned into pCR4-TOPO and subsequently subcloned into p415GPD by using BamHI and PstI. Point mutations were introduced into Utp23 and Utp24 in the p415GPD vector with the QuikChange Site-Directed Mutagenesis Kit (Stratagene) and verified by sequencing (W. M. Keck Facility, Yale University).
Supplementary Material
Acknowledgments
We thank Madhu Nandineni and Anna Szekely for help with the project and immunofluorescence imaging. This work is supported by a Boehringer Ingelheim Foundation Ph.D. Scholarship (to F.B.), The Anna Fuller Research Fellowship (to S.G.), National Science Foundation Grant MCB448171 (to Y.N.O. and A.L.B.), and National Institutes of Health Grant GM52581 (to S.J.B.).
Abbreviations
- HA
hemagglutinin
- SSU
small subunit.
Footnotes
Conflict of interest statement: No conflicts declared.
References
- 1.Dragon F., Gallagher J. E., Compagnone-Post P. A., Mitchell B. M., Porwancher K. A., Wehner K. A., Wormsley S., Settlage R. E., Shabanowitz J., Osheim Y., et al. Nature. 2002;417:967–970. doi: 10.1038/nature00769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Grandi P., Rybin V., Bassler J., Petfalski E., Strauss D., Marzioch M., Schafer T., Kuster B., Tschochner H., Tollervey D., et al. Mol. Cell. 2002;10:105–115. doi: 10.1016/s1097-2765(02)00579-8. [DOI] [PubMed] [Google Scholar]
- 3.Hoang T., Peng W. T., Vanrobays E., Krogan N., Hiley S., Beyer A. L., Osheim Y. N., Greenblatt J., Hughes T. R., Lafontaine D. L. Mol. Cell. Biol. 2005;25:5523–5534. doi: 10.1128/MCB.25.13.5523-5534.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Lebaron S., Froment C., Fromont-Racine M., Rain J. C., Monsarrat B., Caizergues-Ferrer M., Henry Y. Mol. Cell. Biol. 2005;25:9269–9282. doi: 10.1128/MCB.25.21.9269-9282.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Osheim Y. N., French S. L., Keck K. M., Champion E. A., Spasov K., Dragon F., Baserga S. J., Beyer A. L. Mol. Cell. 2004;16:943–954. doi: 10.1016/j.molcel.2004.11.031. [DOI] [PubMed] [Google Scholar]
- 6.Clissold P. M., Ponting C. P. Curr. Biol. 2000;10:R888–R890. doi: 10.1016/s0960-9822(00)00858-7. [DOI] [PubMed] [Google Scholar]
- 7.Arcus V. L., Backbro K., Roos A., Daniel E. L., Baker E. N. J. Biol. Chem. 2004;279:16471–16478. doi: 10.1074/jbc.M313833200. [DOI] [PubMed] [Google Scholar]
- 8.Fatica A., Oeffinger M., Dlakic M., Tollervey D. Mol. Cell. Biol. 2003;23:1798–1807. doi: 10.1128/MCB.23.5.1798-1807.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Fatica A., Tollervey D., Dlakic M. RNA. 2004;10:1698–1701. doi: 10.1261/rna.7123504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Giaever G., Chu A. M., Ni L., Connelly C., Riles L., Veronneau S., Dow S., Lucau-Danila A., Anderson K., Andre B., et al. Nature. 2002;418:387–391. doi: 10.1038/nature00935. [DOI] [PubMed] [Google Scholar]
- 11.Wu L. F., Hughes T. R., Davierwala A. P., Robinson M. D., Stoughton R., Altschuler S. J. Nat. Genet. 2002;31:255–265. doi: 10.1038/ng906. [DOI] [PubMed] [Google Scholar]
- 12.Baker T. A., Luo L. Proc. Natl. Acad. Sci. USA. 1994;91:6654–6658. doi: 10.1073/pnas.91.14.6654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Krogan N. J., Peng W. T., Cagney G., Robinson M. D., Haw R., Zhong G., Guo X., Zhang X., Canadien V., Richards D. P., et al. Mol. Cell. 2004;13:225–239. doi: 10.1016/s1097-2765(04)00003-6. [DOI] [PubMed] [Google Scholar]
- 14.Bernstein K. A., Gallagher J. E., Mitchell B. M., Granneman S., Baserga S. J. Eukaryotic Cell. 2004;3:1619–1626. doi: 10.1128/EC.3.6.1619-1626.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Peng W. T., Krogan N. J., Richards D. P., Greenblatt J. F., Hughes T. R. Nucleic Acids Res. 2004;32:1993–1999. doi: 10.1093/nar/gkh518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Combs D. J., Nagel R. J., Ares M., Jr., Stevens S. W. Mol. Cell. Biol. 2006;26:523–534. doi: 10.1128/MCB.26.2.523-534.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Leeds N. B., Small E. C., Hiley S. L., Hughes T. R., Staley J. P. Mol. Cell. Biol. 2006;26:513–522. doi: 10.1128/MCB.26.2.513-522.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Gavin A. C., Aloy P., Grandi P., Krause R., Boesche M., Marzioch M., Rau C., Jensen L. J., Bastuck S., Dumpelfeld B., et al. Nature. 2006;440:631–636. doi: 10.1038/nature04532. [DOI] [PubMed] [Google Scholar]
- 19.Jin S. B., Zhao J., Bjork P., Schmekel K., Ljungdahl P. O., Wieslander L. J. Biol. Chem. 2002;277:18431–18439. doi: 10.1074/jbc.M112395200. [DOI] [PubMed] [Google Scholar]
- 20.Schafer T., Strauss D., Petfalski E., Tollervey D., Hurt E. EMBO J. 2003;22:1370–1380. doi: 10.1093/emboj/cdg121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Huh W. K., Falvo J. V., Gerke L. C., Carroll A. S., Howson R. W., Weissman J. S., O’Shea E. K. Nature. 2003;425:686–691. doi: 10.1038/nature02026. [DOI] [PubMed] [Google Scholar]
- 22.Ho Y., Gruhler A., Heilbut A., Bader G. D., Moore L., Adams S. L., Millar A., Taylor P., Bennett K., Boutilier K., et al. Nature. 2002;415:180–183. doi: 10.1038/415180a. [DOI] [PubMed] [Google Scholar]
- 23.Samanta M. P., Liang S. Proc. Natl. Acad. Sci. USA. 2003;100:12579–12583. doi: 10.1073/pnas.2132527100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Venema J., Tollervey D. Annu. Rev. Genet. 1999;33:261–311. doi: 10.1146/annurev.genet.33.1.261. [DOI] [PubMed] [Google Scholar]
- 25.Ceska T. A., Sayers J. R. Trends Biochem. Sci. 1998;23:331–336. doi: 10.1016/s0968-0004(98)01259-6. [DOI] [PubMed] [Google Scholar]
- 26.Mitchell P., Petfalski E., Shevchenko A., Mann M., Tollervey D. Cell. 1997;91:457–466. doi: 10.1016/s0092-8674(00)80432-8. [DOI] [PubMed] [Google Scholar]
- 27.Knop M., Siegers K., Pereira G., Zachariae W., Winsor B., Nasmyth K., Schiebel E. Yeast. 1999;15:963–972. doi: 10.1002/(SICI)1097-0061(199907)15:10B<963::AID-YEA399>3.0.CO;2-W. [DOI] [PubMed] [Google Scholar]
- 28.Gavin A. C., Bosche M., Krause R., Grandi P., Marzioch M., Bauer A., Schultz J., Rick J. M., Michon A. M., Cruciat C. M., et al. Nature. 2002;415:141–147. doi: 10.1038/415141a. [DOI] [PubMed] [Google Scholar]
- 29.Gari E., Piedrafita L., Aldea M., Herrero E. Yeast. 1997;13:837–848. doi: 10.1002/(SICI)1097-0061(199707)13:9<837::AID-YEA145>3.0.CO;2-T. [DOI] [PubMed] [Google Scholar]
- 30.Dunbar D. A., Wormsley S., Agentis T. M., Baserga S. J. Mol. Cell. Biol. 1997;17:5803–5812. doi: 10.1128/mcb.17.10.5803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wehner K. A., Gallagher J. E., Baserga S. J. Mol. Cell. Biol. 2002;22:7258–7267. doi: 10.1128/MCB.22.20.7258-7267.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Wehner K. A., Baserga S. J. Mol. Cell. 2002;9:329–339. doi: 10.1016/s1097-2765(02)00438-0. [DOI] [PubMed] [Google Scholar]
- 33.Dunbar D. A., Dragon F., Lee S. J., Baserga S. J. Proc. Natl. Acad. Sci. USA. 2000;97:13027–13032. doi: 10.1073/pnas.97.24.13027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lee S. J., Baserga S. J. Mol. Cell. Biol. 1999;19:5441–5452. doi: 10.1128/mcb.19.8.5441. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.