Abstract
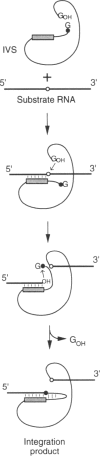
Group I introns in rRNA genes are clustered in highly conserved regions that include tRNA and mRNA binding sites. This pattern is consistent with insertion of group I introns by direct interaction with exposed regions of rRNA. Integration of the Tetrahymena group I intron (or intervening sequence, IVS) into large subunit rRNA via reverse splicing was investigated using E. coli 23S rRNA as a model substrate. The results show that sequences homologous to the splice junction in Tetrahymena are the preferred site of integration, but that many other sequences in the 23S rRNA provide secondary targets. Like the original splice junction, many new reaction sites are in regions of stable secondary structure. Reaction at the natural splice junction is observed in 50S subunits and to a lesser extent in 70S ribosomes. These results support the feasibility of intron transposition to new sites in rRNA genes via reverse splicing.
Full text
PDF
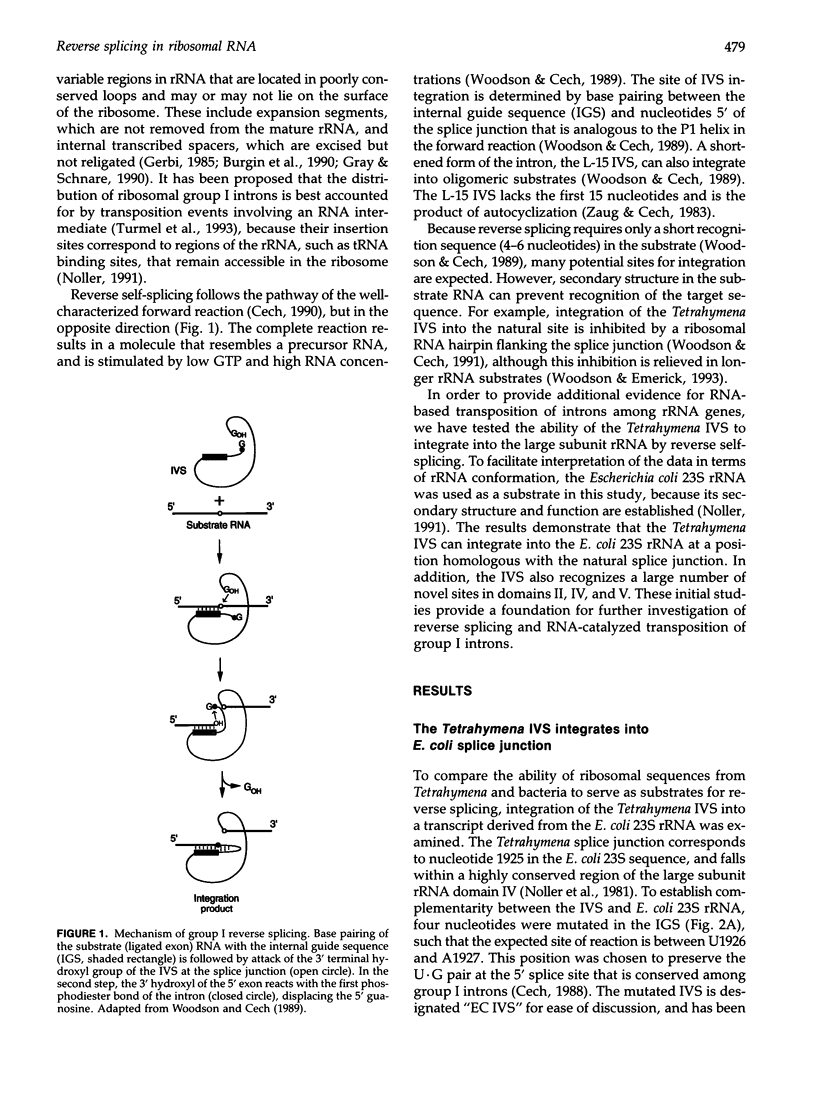


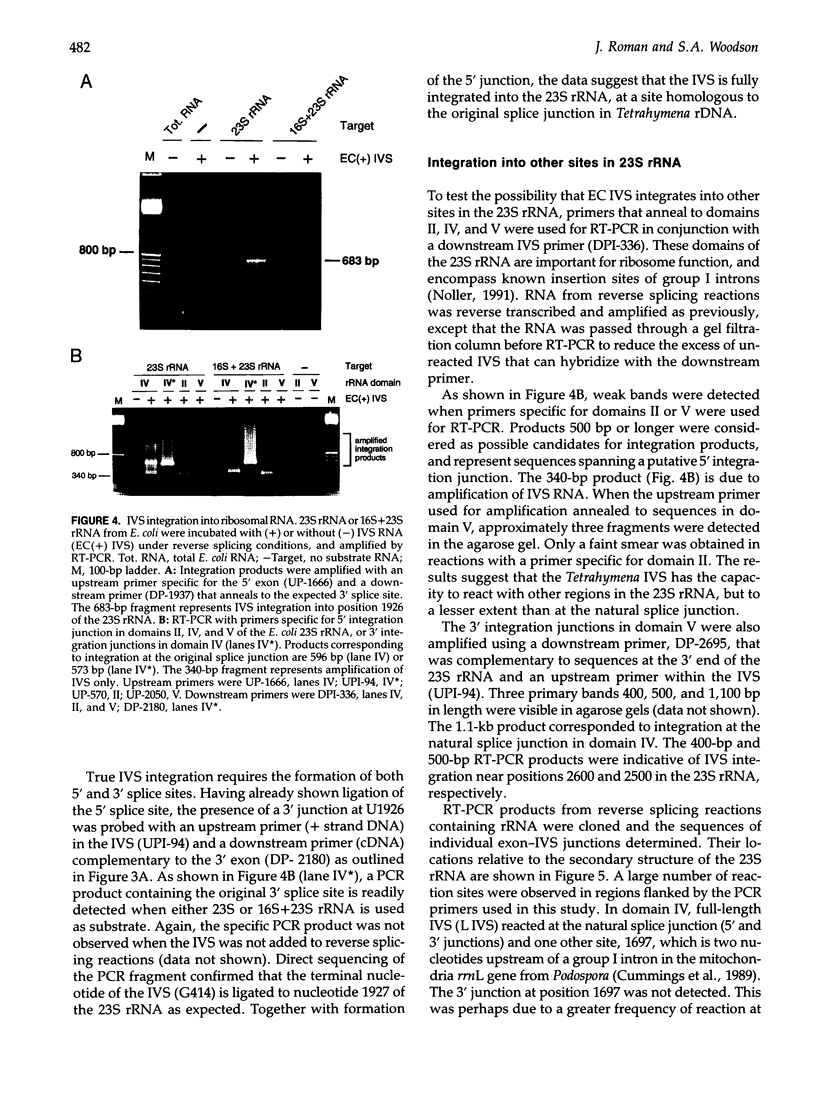
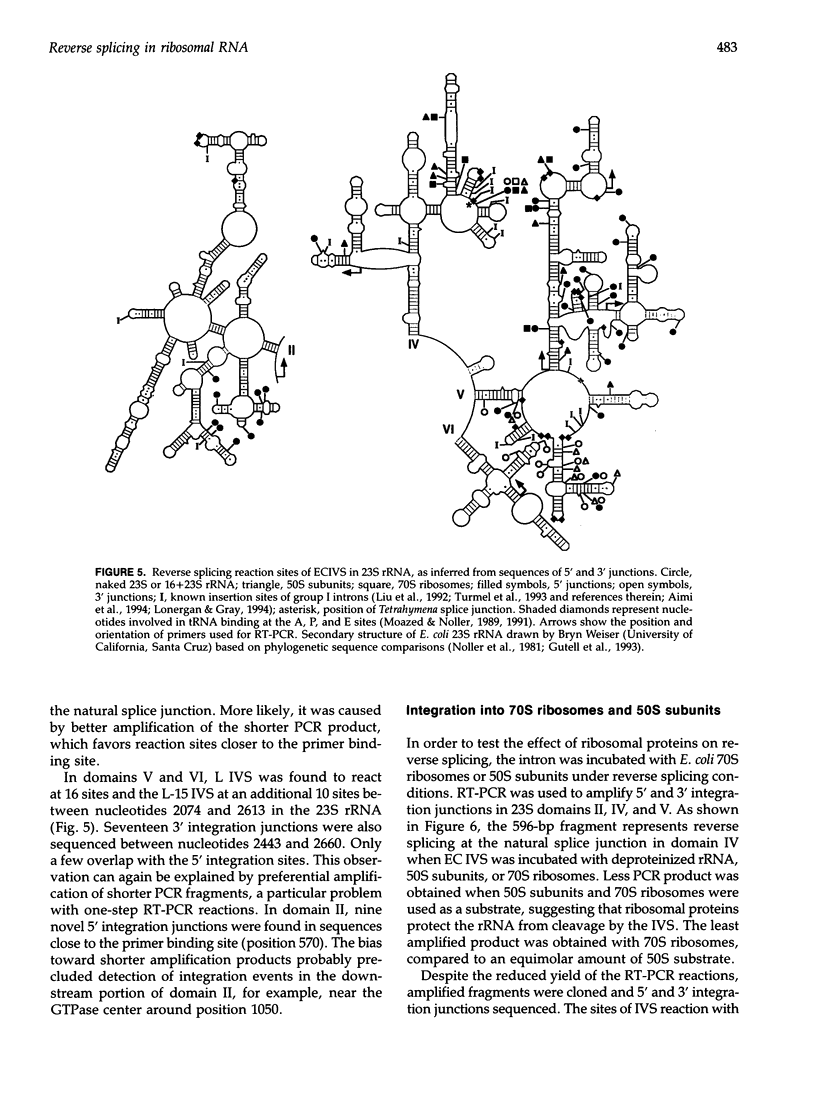

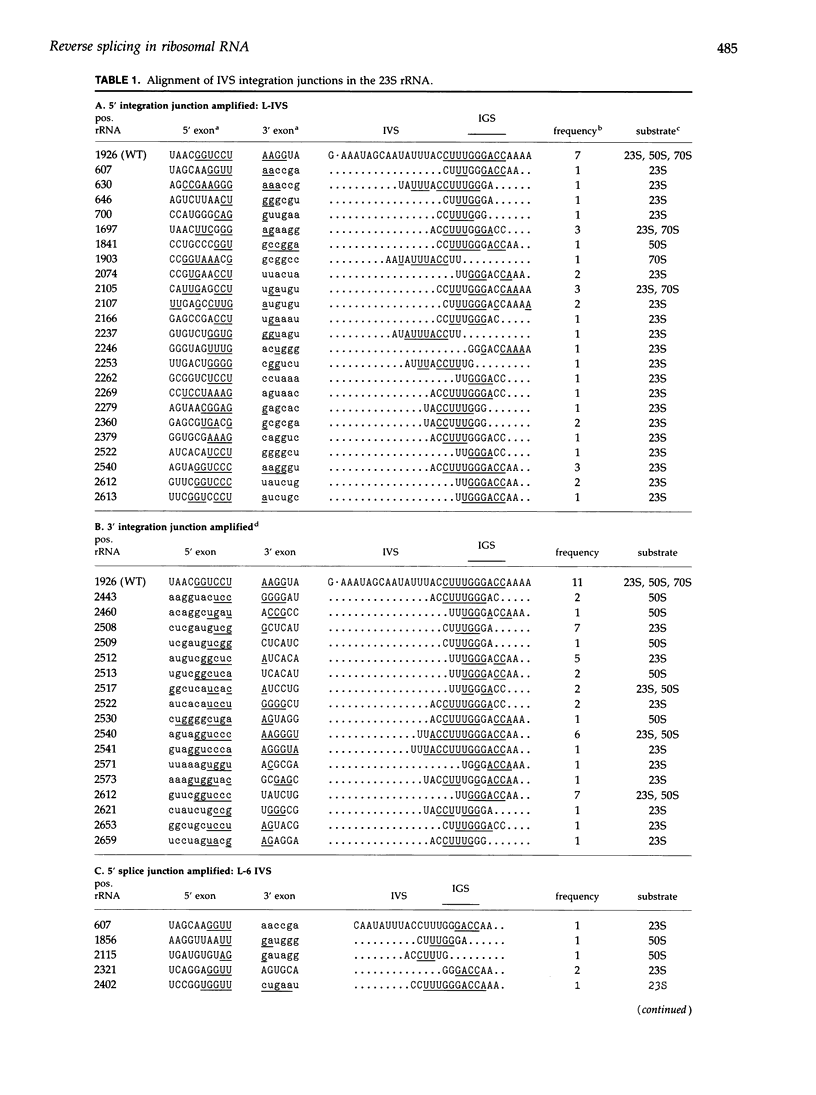

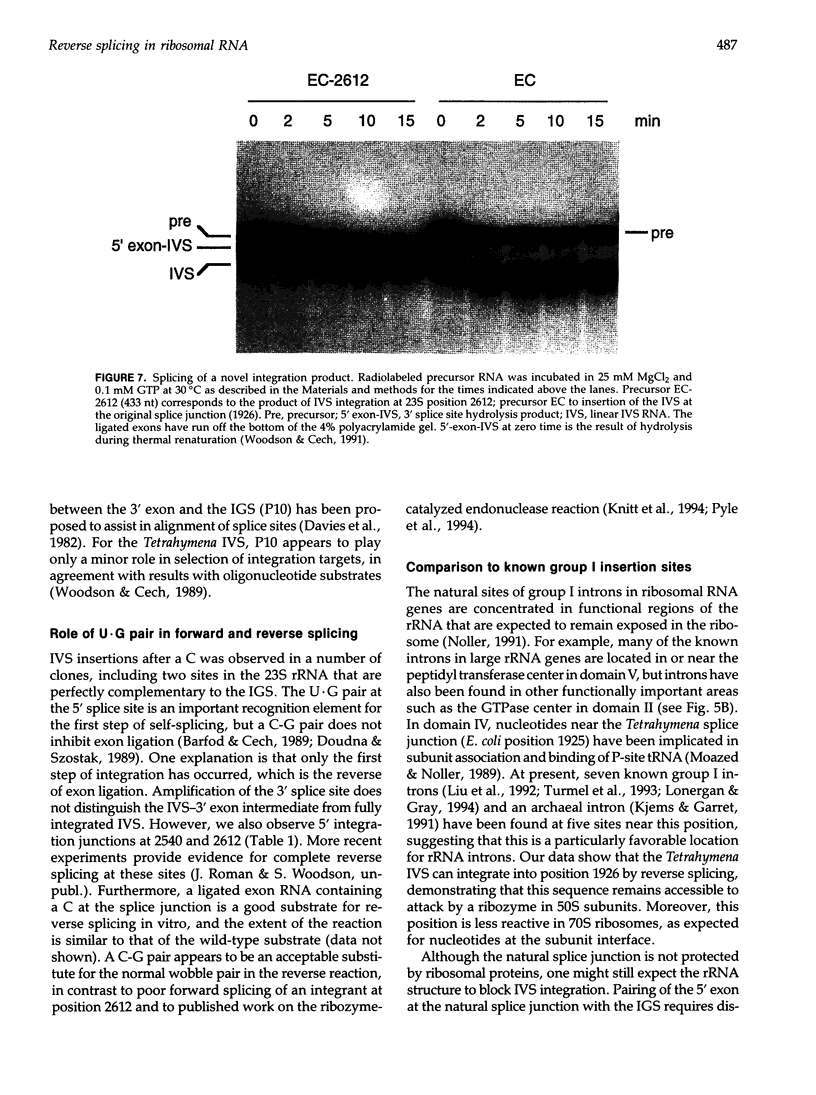



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aimi T., Yamada T., Yamashita M., Murooka Y. Characterization of the nuclear large-subunit rRNA-encoding gene and the group-I self-splicing intron from Chlorella ellipsoidea C-87. Gene. 1994 Jul 22;145(1):139–144. doi: 10.1016/0378-1119(94)90337-9. [DOI] [PubMed] [Google Scholar]
- Barfod E. T., Cech T. R. The conserved U.G pair in the 5' splice site duplex of a group I intron is required in the first but not the second step of self-splicing. Mol Cell Biol. 1989 Sep;9(9):3657–3666. doi: 10.1128/mcb.9.9.3657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Been M. D., Cech T. R. One binding site determines sequence specificity of Tetrahymena pre-rRNA self-splicing, trans-splicing, and RNA enzyme activity. Cell. 1986 Oct 24;47(2):207–216. doi: 10.1016/0092-8674(86)90443-5. [DOI] [PubMed] [Google Scholar]
- Been M. D., Cech T. R. Selection of circularization sites in a group I IVS RNA requires multiple alignments of an internal template-like sequence. Cell. 1987 Sep 11;50(6):951–961. doi: 10.1016/0092-8674(87)90522-8. [DOI] [PubMed] [Google Scholar]
- Belfort M., Ehrenman K., Chandry P. S. Genetic and molecular analysis of RNA splicing in Escherichia coli. Methods Enzymol. 1990;181:521–539. doi: 10.1016/0076-6879(90)81149-o. [DOI] [PubMed] [Google Scholar]
- Belfort M. Phage T4 introns: self-splicing and mobility. Annu Rev Genet. 1990;24:363–385. doi: 10.1146/annurev.ge.24.120190.002051. [DOI] [PubMed] [Google Scholar]
- Burgin A. B., Parodos K., Lane D. J., Pace N. R. The excision of intervening sequences from Salmonella 23S ribosomal RNA. Cell. 1990 Feb 9;60(3):405–414. doi: 10.1016/0092-8674(90)90592-3. [DOI] [PubMed] [Google Scholar]
- Cech T. R. Conserved sequences and structures of group I introns: building an active site for RNA catalysis--a review. Gene. 1988 Dec 20;73(2):259–271. doi: 10.1016/0378-1119(88)90492-1. [DOI] [PubMed] [Google Scholar]
- Cech T. R. Self-splicing RNA: implications for evolution. Int Rev Cytol. 1985;93:3–22. doi: 10.1016/s0074-7696(08)61370-4. [DOI] [PubMed] [Google Scholar]
- Cech T. R. Self-splicing of group I introns. Annu Rev Biochem. 1990;59:543–568. doi: 10.1146/annurev.bi.59.070190.002551. [DOI] [PubMed] [Google Scholar]
- Cummings D. J., Domenico J. M., Nelson J. DNA sequence and secondary structures of the large subunit rRNA coding regions and its two class I introns of mitochondrial DNA from Podospora anserina. J Mol Evol. 1989 Mar;28(3):242–255. doi: 10.1007/BF02102482. [DOI] [PubMed] [Google Scholar]
- Davies R. W., Waring R. B., Ray J. A., Brown T. A., Scazzocchio C. Making ends meet: a model for RNA splicing in fungal mitochondria. Nature. 1982 Dec 23;300(5894):719–724. doi: 10.1038/300719a0. [DOI] [PubMed] [Google Scholar]
- Doudna J. A., Cormack B. P., Szostak J. W. RNA structure, not sequence, determines the 5' splice-site specificity of a group I intron. Proc Natl Acad Sci U S A. 1989 Oct;86(19):7402–7406. doi: 10.1073/pnas.86.19.7402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doudna J. A., Szostak J. W. RNA-catalysed synthesis of complementary-strand RNA. Nature. 1989 Jun 15;339(6225):519–522. doi: 10.1038/339519a0. [DOI] [PubMed] [Google Scholar]
- Dujon B. Group I introns as mobile genetic elements: facts and mechanistic speculations--a review. Gene. 1989 Oct 15;82(1):91–114. doi: 10.1016/0378-1119(89)90034-6. [DOI] [PubMed] [Google Scholar]
- Garrett R. A., Dalgaard J., Larsen N., Kjems J., Mankin A. S. Archaeal rRNA operons. Trends Biochem Sci. 1991 Jan;16(1):22–26. doi: 10.1016/0968-0004(91)90011-j. [DOI] [PubMed] [Google Scholar]
- Gutell R. R., Gray M. W., Schnare M. N. A compilation of large subunit (23S and 23S-like) ribosomal RNA structures: 1993. Nucleic Acids Res. 1993 Jul 1;21(13):3055–3074. doi: 10.1093/nar/21.13.3055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacquier A., Dujon B. An intron-encoded protein is active in a gene conversion process that spreads an intron into a mitochondrial gene. Cell. 1985 Jun;41(2):383–394. doi: 10.1016/s0092-8674(85)80011-8. [DOI] [PubMed] [Google Scholar]
- Kennell J. C., Moran J. V., Perlman P. S., Butow R. A., Lambowitz A. M. Reverse transcriptase activity associated with maturase-encoding group II introns in yeast mitochondria. Cell. 1993 Apr 9;73(1):133–146. doi: 10.1016/0092-8674(93)90166-n. [DOI] [PubMed] [Google Scholar]
- Kjems J., Garrett R. A. Ribosomal RNA introns in archaea and evidence for RNA conformational changes associated with splicing. Proc Natl Acad Sci U S A. 1991 Jan 15;88(2):439–443. doi: 10.1073/pnas.88.2.439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knitt D. S., Narlikar G. J., Herschlag D. Dissection of the role of the conserved G.U pair in group I RNA self-splicing. Biochemistry. 1994 Nov 22;33(46):13864–13879. doi: 10.1021/bi00250a041. [DOI] [PubMed] [Google Scholar]
- Lambowitz A. M., Belfort M. Introns as mobile genetic elements. Annu Rev Biochem. 1993;62:587–622. doi: 10.1146/annurev.bi.62.070193.003103. [DOI] [PubMed] [Google Scholar]
- Liu Y., Rocourt M., Pan S., Liu C., Leibowitz M. J. Sequence and variability of the 5.8S and 26S rRNA genes of Pneumocystis carinii. Nucleic Acids Res. 1992 Jul 25;20(14):3763–3772. doi: 10.1093/nar/20.14.3763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lonergan K. M., Gray M. W. The ribosomal RNA gene region in Acanthamoeba castellanii mitochondrial DNA. A case of evolutionary transfer of introns between mitochondria and plastids? J Mol Biol. 1994 Jun 17;239(4):476–499. doi: 10.1006/jmbi.1994.1390. [DOI] [PubMed] [Google Scholar]
- Macreadie I. G., Scott R. M., Zinn A. R., Butow R. A. Transposition of an intron in yeast mitochondria requires a protein encoded by that intron. Cell. 1985 Jun;41(2):395–402. doi: 10.1016/s0092-8674(85)80012-x. [DOI] [PubMed] [Google Scholar]
- Moazed D., Noller H. F. Interaction of tRNA with 23S rRNA in the ribosomal A, P, and E sites. Cell. 1989 May 19;57(4):585–597. doi: 10.1016/0092-8674(89)90128-1. [DOI] [PubMed] [Google Scholar]
- Moazed D., Noller H. F. Sites of interaction of the CCA end of peptidyl-tRNA with 23S rRNA. Proc Natl Acad Sci U S A. 1991 May 1;88(9):3725–3728. doi: 10.1073/pnas.88.9.3725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mueller M. W., Allmaier M., Eskes R., Schweyen R. J. Transposition of group II intron aI1 in yeast and invasion of mitochondrial genes at new locations. Nature. 1993 Nov 11;366(6451):174–176. doi: 10.1038/366174a0. [DOI] [PubMed] [Google Scholar]
- Noller H. F., Kop J., Wheaton V., Brosius J., Gutell R. R., Kopylov A. M., Dohme F., Herr W., Stahl D. A., Gupta R. Secondary structure model for 23S ribosomal RNA. Nucleic Acids Res. 1981 Nov 25;9(22):6167–6189. doi: 10.1093/nar/9.22.6167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noller H. F. Ribosomal RNA and translation. Annu Rev Biochem. 1991;60:191–227. doi: 10.1146/annurev.bi.60.070191.001203. [DOI] [PubMed] [Google Scholar]
- Pyle A. M., Moran S., Strobel S. A., Chapman T., Turner D. H., Cech T. R. Replacement of the conserved G.U with a G-C pair at the cleavage site of the Tetrahymena ribozyme decreases binding, reactivity, and fidelity. Biochemistry. 1994 Nov 22;33(46):13856–13863. doi: 10.1021/bi00250a040. [DOI] [PubMed] [Google Scholar]
- Saiki R. K., Gelfand D. H., Stoffel S., Scharf S. J., Higuchi R., Horn G. T., Mullis K. B., Erlich H. A. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science. 1988 Jan 29;239(4839):487–491. doi: 10.1126/science.2448875. [DOI] [PubMed] [Google Scholar]
- Sellem C. H., Lecellier G., Belcour L. Transposition of a group II intron. Nature. 1993 Nov 11;366(6451):176–178. doi: 10.1038/366176a0. [DOI] [PubMed] [Google Scholar]
- Sharp P. A. On the origin of RNA splicing and introns. Cell. 1985 Sep;42(2):397–400. doi: 10.1016/0092-8674(85)90092-3. [DOI] [PubMed] [Google Scholar]
- Shub D. A., Gott J. M., Xu M. Q., Lang B. F., Michel F., Tomaschewski J., Pedersen-Lane J., Belfort M. Structural conservation among three homologous introns of bacteriophage T4 and the group I introns of eukaryotes. Proc Natl Acad Sci U S A. 1988 Feb;85(4):1151–1155. doi: 10.1073/pnas.85.4.1151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strobel S. A., Cech T. R. Minor groove recognition of the conserved G.U pair at the Tetrahymena ribozyme reaction site. Science. 1995 Feb 3;267(5198):675–679. doi: 10.1126/science.7839142. [DOI] [PubMed] [Google Scholar]
- Sullivan F. X., Cech T. R. Reversibility of cyclization of the Tetrahymena rRNA intervening sequence: implication for the mechanism of splice site choice. Cell. 1985 Sep;42(2):639–648. doi: 10.1016/0092-8674(85)90121-7. [DOI] [PubMed] [Google Scholar]
- Turmel M., Gutell R. R., Mercier J. P., Otis C., Lemieux C. Analysis of the chloroplast large subunit ribosomal RNA gene from 17 Chlamydomonas taxa. Three internal transcribed spacers and 12 group I intron insertion sites. J Mol Biol. 1993 Jul 20;232(2):446–467. doi: 10.1006/jmbi.1993.1402. [DOI] [PubMed] [Google Scholar]
- Walstrum S. A., Uhlenbeck O. C. The self-splicing RNA of Tetrahymena is trapped in a less active conformation by gel purification. Biochemistry. 1990 Nov 20;29(46):10573–10576. doi: 10.1021/bi00498a022. [DOI] [PubMed] [Google Scholar]
- Woodson S. A., Cech T. R. Alternative secondary structures in the 5' exon affect both forward and reverse self-splicing of the Tetrahymena intervening sequence RNA. Biochemistry. 1991 Feb 26;30(8):2042–2050. doi: 10.1021/bi00222a006. [DOI] [PubMed] [Google Scholar]
- Woodson S. A., Cech T. R. Reverse self-splicing of the tetrahymena group I intron: implication for the directionality of splicing and for intron transposition. Cell. 1989 Apr 21;57(2):335–345. doi: 10.1016/0092-8674(89)90971-9. [DOI] [PubMed] [Google Scholar]
- Woodson S. A., Emerick V. L. An alternative helix in the 26S rRNA promotes excision and integration of the Tetrahymena intervening sequence. Mol Cell Biol. 1993 Feb;13(2):1137–1145. doi: 10.1128/mcb.13.2.1137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaug A. J., Grabowski P. J., Cech T. R. Autocatalytic cyclization of an excised intervening sequence RNA is a cleavage-ligation reaction. Nature. 1983 Feb 17;301(5901):578–583. doi: 10.1038/301578a0. [DOI] [PubMed] [Google Scholar]
- Zhang F., Ramsay E. S., Woodson S. A. In vivo facilitation of Tetrahymena group I intron splicing in Escherichia coli pre-ribosomal RNA. RNA. 1995 May;1(3):284–292. [PMC free article] [PubMed] [Google Scholar]