Abstract
The amount and distribution of variation in the genomic region containing the genes in the lytic-lysogenic genetic switch and the sequence that determines the integration site into the host chromosome were analyzed for 38 P2-like phages from Escherichia coli. The genetic switch consists of two convergent mutually exclusive promoters, Pe and Pc, and two repressors, C and Cox. The immunity repressor C blocks the early Pe promoter, leading to the establishment of lysogeny. The Cox repressor blocks expression of Pc, allowing lytic growth. Phylogenetic analyses showed that the C and Cox proteins were distributed into seven distinct classes. The phylogenetic relationship differed between the two proteins, and we showed that homologous recombination plays a major role in creating alterations in the genetic switch, leading to new immunity classes. Analyses of the host integration site for these phages resulted in the discovery of a previously unknown site, and there were at least four regular integration sites. Interestingly, we found no case where phages of the same immunity class had different host attachment sites. The evolution of immunity and integration sites is complex, since it involves interactions both between the phages themselves and between phages and hosts, and often, both regulatory proteins and target DNA must change.
P2-like phages are a group of related temperate phages that grow on γ-proteobacteria and share common traits such as morphology, control of lytic versus lysogenic growth, and noninducibility by UV light (for a recent review, see reference 26). Temperate phages have the ability to reproduce by two alternative life cycles: the lytic or the lysogenic cycle. In the latter life cycle, the phage genome integrates into a specific location on the host chromosome, and most phage genes are turned off by the phage-encoded immunity repressor C. P2-like phages are prevalent in Escherichia coli strains; about 30% of the strains in the ECOR collection (28) contain P2-like prophages (27). P2-like phages that are found in other γ-proteobacteria are more distantly related to P2 than those found in E. coli, and it seems as if the evolution of the P2-like phages tracks the evolution of their respective hosts (26; A. S. Nilsson, unpublished data). An analysis of the DNA sequence of the late structural genes of 18 P2-like isolates that grow on E. coli showed that these genes are at least 96% identical to the genes of P2 (25). Thus, these P2-like coliphages might be considered different isolates of P2, but they have been shown to have different immunities, based on their capacity to grow on bacteria lysogenized with different P2-like phages (6, 9, 14) and to integrate at at least two different sites in the host chromosome (22, 38). Phage 186 is a more distantly related E. coli phage, and its immunity repressor, cI, differs in size and sequence from the C repressors of the P2-like coliphages in this study. In fact, both cI and Apl (the equivalent of Cox) of phage 186 are more related to the cI and Cox repressors of Haemophilus influenzae phages HP1 and HP2 and Pseudomonas aeruginosa phage K139 (26) and are therefore excluded from this study.
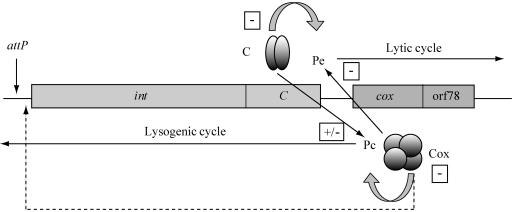
Since P2-like phages are frequently found in E. coli strains, they compete with each other when present in the same host cell, either after superinfection of a lysogen or after mixed infection. This most likely has driven evolution towards different immunities and integration sites. The transcriptional switch of P2-like phages that controls the lytic versus the lysogenic growth cycle contains two face-to-face-located promoters and two repressors, C and Cox (Fig. 1). The immunity repressor C, encoded by the first gene of the Pc promoter, blocks transcription from the early promoter Pe, leading to the formation of lysogeny. The integration of the phage genome into the host chromosome is promoted by the phage integrase, which is encoded by the int gene located downstream of the C gene. The first gene of the Pe operon is cox, whose product is a repressor of Pc. In this way, the two pathways are mutually exclusive (26). In the lysogenic stage, the immunity repressor C will also block the lytic growth of a superinfecting P2-like phage belonging to the same immunity group. Under these conditions, the superinfecting phage must integrate into the host chromosome to be stably maintained. However, if they have the same host integration site, it would lead to prophages integrated in tandem, which is an unstable state for phage P2 due to the expression of the int gene when prophages are integrated in this way (7, 8, 9). However, the superinfecting phage genome can become integrated into secondary integration sites under this condition (2, 10). In the P2-like phages studied so far, there is a coupling between the developmental switch and the site-specific recombination that leads to the integration or excision of the phage genome in or out of the chromosome. The Cox protein is, in addition to being a repressor of Pc, a directionality factor for site-specific recombination. It inhibits integrative recombination but is required for excision (39).
FIG. 1.
Schematic drawing of the control region of the P2-like phages. The C gene encodes the immunity repressor that is a transcriptional repressor of the Pe transcript. In addition, it controls its own Pc promoter. The cox gene encodes the Cox protein that functions as a repressor of Pc, and at high concentrations, it also reduces the activity of Pe. It is also a directionality factor that blocks integration and promotes excision by binding to attP (indicated by the dashed arrow). The int gene encodes the integrase that is needed for the site-specific recombination between the attP site of the phage and the attB site on the chromosome. orf78 is a well-conserved open reading frame of unknown function.
In this work, we have addressed the evolution of the immunities and integration sites in the P2-like coliphages by sequencing pertinent regions of the P2-like prophages found in the ECOR collection and by performing phylogenetic analyses of the C and cox genes. Our data support the hypothesis that recombination is an important mechanism for generating new immunity classes and that the divergence of immunities follows the emergence of new integration sites.
MATERIALS AND METHODS
Biological materials.
The bacterial strains, phages, and plasmids used in this study are listed in Table 1.
TABLE 1.
Bacterium, bacteriophage, and plasmid data
| Organism | Origin/pertinent feature(s) | Reference(s) or source | EMBL accession no. |
|---|---|---|---|
| Bacteria | |||
| C-1a | Prototrophic E. coli C strain | 35 | |
| C-1920 | C-1a lysogenized with WΦ | 18 | |
| TD204 | C-1a lysogenized with HK109 | 14 | |
| MG1655 | E. coli K-12 strain | 11 | NC_000913 |
| SCRI1043 | E. carotovora subsp. atroseptica strain | 3 | NC_0045447 |
| ECOR4 | Human, Iowa | 28 | AM159080 |
| ECOR5 | Human, Iowa | 28 | AM159064 |
| ECOR7 | Orangutan, Wash. | 28 | AM159054 |
| ECOR10 | Human, Sweden | 28 | AM159065 |
| ECOR12 | Human, Sweden | 28 | AM159076 |
| ECOR15 | Human, Sweden | 28 | AM159078 |
| ECOR30 | Bison, Canada | 28 | AM159055 |
| ECOR31 | Leopard, Wash. | 28 | AM159056 |
| ECOR44 | Cougar, Wash. | 28 | AM159067 |
| ECOR45 | Pig, Indonesia | 28 | AM159057 |
| ECOR46 | Ape, Wash. | 28 | AM159079 |
| ECOR48 | Human, Sweden | 28 | AM159058 |
| ECOR53 | Human, Iowa | 28 | AM159070 |
| ECOR58 | Lion, Wash. | 28 | AM159059 |
| ECOR59 | Human, Mass. | 28 | AM159071 |
| ECOR61 | Human, Sweden | 28 | AM159072 |
| ECOR62 | Human, Sweden | 28 | AM159073 |
| ECOR64 | Human, Sweden | 28 | AM159074 |
| ECOR67 | Goat, Indonesia | 28 | AM159066 |
| Bacteriophages | |||
| 18 | E. coli clinical isolate, Paris, France | 20 | AM159068 |
| 299 | E. coli clinical isolate, Paris, France | 17, 20 | AM159053 |
| L-413C | NAa | Elliot et al.b | NC_004745 |
| HK109 | E. coli clinical isolate, Hong Kong | 14 | AM159069 |
| HK111 | E. coli clinical isolate, Hong Kong | 14 | AM159062 |
| HK113 | E. coli clinical isolate, Hong Kong | 14 | AM159081 |
| HK114 | E. coli clinical isolate, Hong Kong | 14 | AM159063 |
| HK239 | Cow dung, Hong Kong | 14 | AM159082 |
| HK240 | Cow dung, Hong Kong | 14 | AM159051 |
| HK241 | Human feces, Hong Kong | 14 | AM159052 |
| P2 | Lisbonne and Carrère E. coli strain | 5 | AF063097 |
| P2 Hy dis | Recombinant between P2 and a defective P2-like prophage in an E. coli B strain | 4, 12, 13 | AJ306907 |
| PK | E. coli strain K235 | 21 | AM159049 |
| WΦ | E. coli strain W | 18, 29 | NC_005056 |
| ΦD124 | Clinical isolate, Los Angeles, Calif. | 9 | AM159060 |
| ΦD145 | Clinical isolate, Los Angeles, Calif. | 9 | AM159077, |
| AM158280 | |||
| ΦD160 | Clinical isolate, Los Angeles, Calif. | 9 | AM159075 |
| ΦD252 | Clinical isolate, Los Angeles, Calif. | 9 | AM159061 |
| ΦD266 | Clinical isolate, Los Angeles, Calif. | 9 | AM159050 |
| Plasmids | |||
| pET-16b | Expression vector | Novagen Inc. | |
| pEE900 | pET-16b derivative with the WΦ C gene | 22 | |
| pEE2050 | pET-16b derivative with the P2-EC4 C gene | This work |
NA, not applicable.
J. M. Elliott, A. A. Filippov, V. V. Kutyrev, A. G. Bobrov, O. A. Kirillina, V. L. Motin, P. S. Chain, and E. Garcia, unpublished data.
Plasmid constructions.
Constructions were made according to standard procedures (34). Escherichia coli C-1a (35) was used as a recipient strain for transformations.
The P2-EC4 C gene was cloned into the pET-16b (Novagen Inc.) expression vector under the control of the T7 promoter, resulting in plasmid pEE2050. The C gene was amplified from genomic DNA of the ECOR4 isolate containing the P2-like prophage P2-EC4 by using primers EC4 C-C and EC4 C-N (Table 2). The 0.3-kb PCR product generated was cleaved with NcoI and BamHI and inserted between the NcoI and BamHI sites of the vector. Subsequent DNA sequencing was used to confirm the in-frame ligation.
TABLE 2.
Oligonucleotide primers
| Oligonucleotide | Sequence (5′-3′) |
|---|---|
| 79.0 L | GGGTGATGGTGAAGTCAATC |
| 76.1 R | GCCACCAATAGTAGGCACTT |
| 76.3 R | GTAGGCACTTGTGATGAATA |
| 73.5 R | CGTCTGAAGCGAGGGTAATC |
| C rev-Wphi | GGTAACAAATGGTGAAAATG |
| HK109 C rev | GCGGATGCGTTTTCCGTCGCGACCGCG |
| HK109 cox rev | CCTGCCAGCGAGGGACGGCAAGATGC |
| P2 C start NcoI L | CATGCCATGGGGATGTCAAACACGATAAGCGAGAAGATAG |
| WPhi int armR | CATTCTTTCTGGCTGGCGTAGGCG |
| EC64 cox R | GCCCTTCAGCCTCACGCAAGGCC |
| EC44 cox R | GACTGTTCCTAGAGCTATCCCGC |
| HK240 C L | CCGGGCAAATTGCGCCCGCTCTCG |
| LocI R | AGGTTATCATCCGTTTCGCTGA |
| LocI L | CCAATACTACACCGAAGCCG |
| K12 core R | CGCCGCCCGACTGACCAGCC |
| K12 core L | GCGTGACGTGACGCAATGGC |
| Wf12-R | GCAGCGCCACGTTCATCACTTATG |
| Erw attL-L | GACGTCATTACCCGTACCTTCACACAI |
| Fd attL-R | TGTGGTCAATGTGTGGACGTGACCAG |
| Fd attR-L | TCCATAAACCAGAGTTAACTAGAG |
| K-12 attR-R | TCCAGCAGTTCATCAATCACCGGATC |
| EC4 C-C | CGGGATCCTTAGCCAATCCTTTTCGCGGAG |
| EC4 C-N | CGCCATGGCAAATATCGCACAGAAATTAAGAGC |
DNA extraction, amplification, and sequencing.
Bacterial strains lysogenic for different P2-like phages (Table 1) were propagated in LB medium at 37°C overnight, and the DNA was either extracted using the QIAGEN blood and cell culture kit or amplified with the GenomiPhi DNA amplification kit (GE Healthcare). Ready-to-Go PCR beads (GE Healthcare) were used together with oligonucleotide primers (DNA Technology, Aarhus, Denmark) (Table 2) for amplification of pertinent regions. PCR products were purified with the QIAquick PCR purification kit or purified from gels with the QIAEX II gel extraction kit, both from QIAGEN. The amplification products were separated in Tris-borate-EDTA-1% agarose gels stained with ethidium bromide. The amplified fragments, containing the complete C and cox genes together with the intervening Pe-Pc control region, were sequenced for 19 prophages in E. coli from the ECOR collection and for 15 other P2-like coliphage isolates listed in Table 1. In addition, the ΦD145 integrase gene and attP region were sequenced, and the sequences of four P2-like phages were downloaded from the GenBank database, i.e., P2, L-413C, P2 Hy dis, and WΦ. Nucleotide sequencing was performed by Macrogen Inc. (Seoul, Korea) or MWG-Biotech (Ebersberg, Germany).
Sequence alignment and analysis.
Similarity searches were performed at the website of the National Center for Biotechnology Information, where the sequences were compared to each other and to other sequences in the GenBank database with the programs ORF Finder (http://www.ncbi.nlm.nih.gov/gorf/), BLAST (http://www.ncbi.nlm.nih.gov/BLAST/), and bl2seq (http://www.ncbi.nlm.nih.gov/BLAST/bl2seq/wblast2.cgi).
The amino acid sequences for the C and the Cox proteins were aligned with the ClustalX multiple sequence alignment program (version 1.81; IGBMC, University of Strasbourg, France [ftp://ftp-igbmc.u-strasbg.fr/pub/ClustalX/]). Sequence editing was performed with the Jalview alignment editor (version 2.02; School of Life Sciences, University of Dundee, Scotland, United Kingdom [http://www.jalview.org/]), and final refinement was done manually. When nucleotide sequences were used in the phylogenetic analyses, they were aligned and edited as described above and then adjusted by hand to match amino acid alignments.
Phylogenetic analyses.
The PAUP* program (version 4.0b10; Sinauer Associates, Inc., Sunderland, Mass. [http://paup.csit.fsu.edu/about.html]) was used for the phylogenetic analyses under maximum-parsimony criteria and with the heuristic search option due to the large numbers of taxa and some homoplastic characters. The program was used with all the default settings except that the stepwise addition of taxa was randomized 10 times per run, keeping the two shortest trees of each run. To assess the degree of confidence for the resulting shortest trees, they were tested in bootstrap analyses, each with 1,000 replicates.
The history of homologous recombination between taxa was evaluated with three different methods: (i) visualization of the relationship between taxa with the SplitsTree program (version 4; Algorithms in Bioinformatics, University of Tübingen, Germany [http://www-ab.informatik.uni-tuebingen.de/software/jsplits/welcome.html]), (ii) inspection of the distribution of the parsimoniously informative (PI) characters (characters present in at least two but not all taxa) that are homoplastic in the shortest trees, and (iii) statistical testing of the distribution of character differences between taxa with the program Geneconv (version 1.81; Department of Mathematics, Washington University in St. Louis, St. Louis, Mo. [http://www.math.wustl.edu/∼sawyer/geneconv/]).
The SplitsTree program tree-building algorithm does not presume a bifurcating tree. Instead, it detects conflicting phylogenetic signals and tries to construct a figure that reveals these more complex relationships (1). If the program detects that recombination has occurred, affected taxa will be connected by more than one node, and the resulting tree will include one or more networks.
To visualize possible homologous recombination, the distribution of homoplasies in an aligned set of PI sites for all of the genes was assessed according to a method described previously by Nilsson and Haggård-Ljungquist (25). Since excess homoplasy is regarded as a hallmark of recombination (24), homoplasy can be used to detect regions that have undergone recombination. An uneven distribution of homoplastic PI sites signals recombination but could also be caused by rapid recent directional selection. The influence of selection was assessed with the program K-estimator (version 6.1; Genome Informatics Laboratory, Department of Biological Sciences, Indiana University, Ind. [http://www.biology.uiowa.edu/comeron/index_files/Page322.htm]) as described below.
The Geneconv program extends the methods of statistical tests for detecting gene conversions previously described by Sawyer (36). Basically, the program tests whether a pair of sequences in the alignment contains significantly longer identical or almost identical fragments than other pairs in the alignment, which results in a global probability for the actual pair. The probability is corrected for both total alignment length and the number of possible pairs, which grows exponentially in an alignment with the number of taxa. Thus, Bonferroni-corrected Karlin-Altschul P values of <0.05 were considered significant for global fragments. The program also tests whether a fragment within a pair is longer than expected in the absence of recombination. These pairwise P values are also corrected for total sequence length and were considered significant if they where below 0.01. The aligned nucleotide sequences of concatenated coding regions (start of int-C-cox-start of orf78) of all possible pairs (21 pairs in all) of the seven immunity classes were analyzed by applying the default settings, except that the mismatch penalty was set to 1 (gscale = 1). This setting allows for the possibility of mutations taking place within the recombined regions after a recombination has occurred.
The possibility that observed homoplasies were caused by selection was evaluated by comparing the number of synonymous nucleotide substitutions per synonymous site (Ks) with the number of nonsynonymous nucleotide substitutions per nonsynonymous site (Ka), which was carried out by the program K-estimator. A Ka/Ks ratio below 1 is a sign of purifying or stabilizing selection, whereas a Ka/Ks ratio of >1 indicates directional selection.
Nucleotide sequence accession numbers.
The nucleotide sequences between int and orf78 of the 34 prophages have been submitted to the EMBL databases under accession numbers AM159049 to AM159082, and the complete sequence of ΦD145 int can be found under accession number AM158280 (Table 1).
RESULTS
Phylogenetic analysis.
The phylogenetic analyses of amino acid matrices of the C and Cox proteins for the 38 taxa resulted in seven groups, but the phylogenetic relationships of these groups differed between the C and Cox proteins (Fig. 2). Within groups, these genes seemed to have a long common evolutionary history, but the evolution of the groups themselves differed in both topology and the relative rate of evolution between some groups. This is also obvious in light of the amino acid sequence identities, where the identity for the C protein within each class was at least 95%, and often 100%, while the identities between sequences of different classes varied between 38 and 78%. The identity within each class of the Cox protein was at least 97%, and the identities between the classes varied between 30 and 86% (Table 3). The major differences in tree topology were the position of the clusters of ΦD160 and P2-EC12, ΦD145, P2-EC46 and P2-EC15 and the position of P2-EC4. The two gene trees could be largely reconciled if all of these were removed. The support for both gene phylogenies was generally good; most nodes had a 100% bootstrap support, so it seems likely that the differences in tree topology reflect a real difference in the evolution of C and Cox for these phages.
FIG. 2.
Phylogenetic trees of the C (left) and cox (right) genes and a SplitsTree graph of the immunity classes (top, center). The phylogenetic trees were constructed by applying maximum-parsimony criteria and generated by the PAUP program. The trees are unrooted bootstrap consensus trees based on the amino acid sequences of the two genes C and cox. The branch lengths are shown above branches, where applicable, and the bootstrap percentages are given within parentheses. All bootstrap values were based on 1,000 replicates, and groups compatible with the 50% majority-rule consensus were included in the tree. The SplitsTree graph is based on the concatenated C and cox nucleotide sequences. Characters within genes, as well as the genes themselves, point at alternative phylogenetic relationships. Thus, two immunity groups can be related in more than one way, i.e., connected by more than one node in the tree. The topology of the tree suggests that recombinational events in the past were followed by differentiation due to other mechanisms.
TABLE 3.
Pairwise amino acid sequence identities (bl2seq) of C (lower matrix) and Cox (upper matrix)
| Phage | % Identity
|
||||||
|---|---|---|---|---|---|---|---|
| P2 | P2 Hy dis | WΦ | HK109 | ΦD160 | ΦD145 | P2-EC4 | |
| P2 | 83 | 44 | 44 | 35 | 35 | 47 | |
| P2 Hy dis | 64 | 47 | 47 | 38 | 38 | 50 | |
| WΦ | 46 | 51 | 81 | 30 | 31 | 85 | |
| HK109 | 43 | 43 | 55 | 32 | 34 | 86 | |
| ΦD160 | 50 | 59 | 46 | 45 | 82 | 37 | |
| ΦD145 | 46 | 50 | 42 | 38 | 78 | 37 | |
| P2-EC4 | 45 | 47 | 69 | 65 | 41 | 41 | |
The immunity repressor C.
The inferred amino acid sequence of the C proteins fell into seven groups (Fig. 3) that corresponded largely to previously identified immunity groups, which were identified by the plating efficiencies of phage isolates on different lysogens (9, 14). We have assigned these different classes with roman numerals, depending on what type of immunity they belong to; i.e., class I constitutes the isolates with the same immunity as phage P2 (P2, PK, ΦD266, HK240, HK241, 299, P2-EC7, P2-EC30, P2-EC31, P2-EC45, P2-EC48, and P2-EC58), class II constitutes the isolates with phage P2 Hy dis immunity (P2 Hy dis, ΦD124, ΦD252, HK111, HK114, P2-EC5, P2-EC10, and P2-EC67), class III has the same immunity as WΦ (WΦ, P2-EC44, and phage 18), class IV has the HK109 immunity (HK109, HK113, HK239, L-413C, P2-EC53, P2-EC59, P2-EC61, P2-EC62, and P2-EC64), class V has the ΦD160 immunity (ΦD160 and P2-EC12), class VI has the immunity of phage ΦD145 (ΦD145, P2-EC15, and P2-EC46), and class VII consists of only one single prophage, P2-EC4.
FIG. 3.
Alignment of the deduced amino acid sequences for the different C proteins. The full amino acid sequence of the type phage of each class is noted. The sequence of variants within a class is written out only where it differs from the type phage, and identical amino acids are indicated by a dot. The classes contain the following phages: class I, P2, PK, ΦD266, HK240, HK241, 299, P2-EC7, P2-EC30, P2-EC31, P2-EC45, P2-EC48, and P2-EC58; class II, P2 Hy dis, ΦD124, ΦD252, HK111, HK114, P2-EC5, P2-EC10, and P2-EC67; class III, WΦ, P2-EC44, and phage 18; class IV, HK109, HK113, HK239, L-413C, P2-EC53, P2-EC59, P2-EC61, P2-EC62, and P2-EC64; class V, ΦD160 and P2-EC12; class VI, ΦD145, P2-EC15, and P2-EC46; class VII, P2-EC4. The α-helices predicted by JPRED, indicated below the alignments, and the presumed DNA binding HTH motif are shaded in gray. The consensus sequences are indicated at the bottom. The preferred integration sites are noted on the right side, where nd denotes not determined.
Most of these immunity classes consisted of well-defined discrete groups, but as mentioned above, three groups showed either an uncertain or a different position in the phylogenetic analysis. The result was also in some cases not in accordance with previous phenotypical investigations of their immunity. Immunity class VI (ΦD145), for instance, contains three members that show a high identity to class V (78%), but they have previously been classified as belonging to different immunity groups, since ΦD145 was able to plate on a ΦD160 lysogen and vice versa (9). However, we found that the plating efficiency of ΦD145 on a ΦD160 lysogen was 10-fold reduced compared to a nonlysogen, indicating that phage ΦD145 has some sensitivity to the ΦD160 repressor. Phage ΦD145 also grows well on the class II HK114 lysogen but not at all on an HK111 lysogen, which belongs to the same immunity class as HK114. The reciprocal spot tests make the situation even more confusing, since HK111 produces plaques on bacteria lysogenic for ΦD145, with reduced plating efficiency, while HK114 does not grow at all on a ΦD145 lysogen. HK111 and HK114 belong to the same immunity class and cannot grow on each other's lysogens. The C protein of class VI consists of 103 or 104 amino acid residues, thus making it the longest C protein in this study. The C proteins are identical except for a short region starting at codon 23, where ΦD145 has a sequence of six amino acids, while P2-EC15 and P2-EC46 have a different sequence of five amino acids.
Another immunity class, the previously unreported class VII, consisted of only one member, P2-EC4. The slightly weaker bootstrap support for this position (87%) motivated a closer investigation of its integrity. The C protein of P2-EC4 has a high identity to class III and class IV, 69% and 65%, respectively. The three-dimensional structures of the C proteins are not known, but secondary structure predictions using JPRED (version 2; School of Life Sciences, University of Dundee, Scotland, United Kingdom [http://www.compbio.dundee.ac.uk/∼www-jpred/]) indicate that they contain at least three α-helices, where helix 2 and helix 3 most likely constitute a DNA binding helix-turn-helix (HTH) motif since they are separated by a conserved glycine residue. In this case, α-helix 3 should be the DNA recognition helix, which is supported by the fact that only the last two amino acids in this helix are conserved. Since the C proteins of P2-EC4 and WΦ differ in only one out of the eight amino acids in α-helix 3 (Fig. 3), the P2-EC4 C gene was cloned (pEE2050), and the capacity of WΦ to plate on bacteria containing plasmid pEE2050 or plasmid pEE900 (containing the WΦ C repressor) or on the nonlysogenic strain C-1a was determined. WΦ plated with the same efficiency on C-1a, and C-1a containing pEE2050, but formed no plaques on C-1a containing pEE900. The same pattern was shown when the plating efficiency of HK109 was assayed. HK109 formed plaques at the same frequency on the nonlysogenic strain C-1a and the construct with the pEE2050 plasmid but was unable to form any plaques on a C-1a strain lysogenic for HK109 (strain TD204). This implies that the C repressor of P2-EC4 is unable to block transcription from the Pe promoter of WΦ as well as HK109.
Localization of presumptive early promoters and C operators.
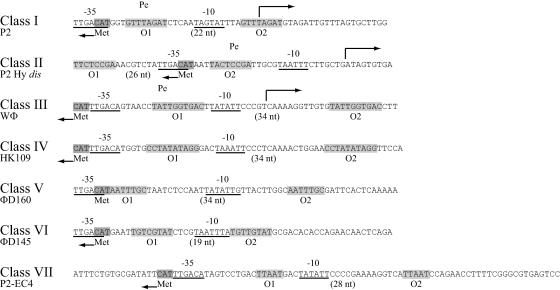
In phages P2, P2 Hy dis, and WΦ, the strong early Pe promoters and the C operators have been located (22, 30) (Fig. 4). In these phages, the initiation codon of the C protein and the −35 region of Pe are either overlapping, as in the case for P2 and P2 Hy dis, or located back to back. The C operators have been shown to consist of two directly repeated sequences located on either side of the −10 region (P2 and WΦ) or of the −35 region (P2 Hy dis). Since the early promoters are expected to be strong, we have searched the equivalent regions of class IV to class VII, and presumptive promoters can also be found at similar positions in these classes (Fig. 4). A search for direct repeats spanning the −10 or −35 region of Pe revealed directly repeated sequences of 10, 8, 9, and 5 nucleotides (nt) spanning the −10 regions in class IV, V, VI, and VII, respectively. As expected, the sequences of these presumptive operators differ between the classes.
FIG. 4.
Comparison of the Pe promoter and operator regions. The location of the Pe promoters, transcriptional start sites, and C operators of P2, Hy dis, and WΦ have been determined previously. The locations of the presumed Pe promoters of the other P2-like type phages are indicated by the −35 and −10 regions, and direct repeats presumed to constitute operator sequences are shaded in light gray. The initiation codon of the C genes is shaded in dark gray.
The Cox protein.
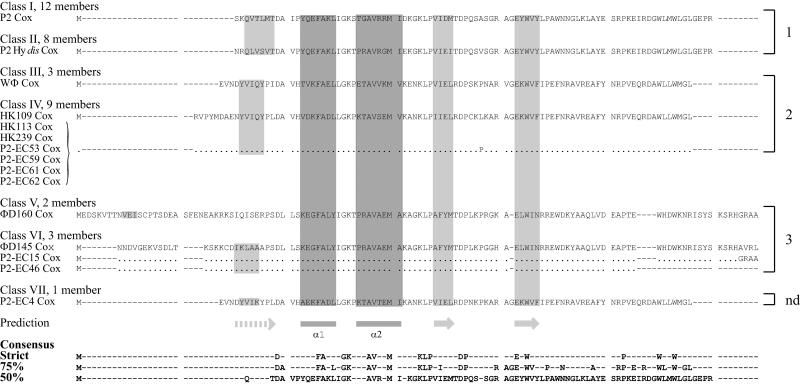
The P2 Cox protein links the developmental switch with site-specific recombination, since it acts as a repressor of Pc, which controls expression of the C repressor and the integrase, and as a directionality factor by inhibiting the integration of the phage genome into the host chromosome but promoting the excision. P2 Cox is a small, slightly basic protein of 91 amino acids. The N-terminal end has been predicted to contain an HTH motif presumed to be involved in DNA binding (32). Since the HTH motif, according to the secondary structure prediction of the Cox proteins using JPRED, is preceded by a β-strand and followed by two β-strands, the Cox proteins most likely belong to the winged-helix family of prokaryotic excisionases (15, 31, 33) and the DNA binding domain of the Mu repressor (19). The identity within different immunity repressor classes was mirrored by the variation of the Cox protein. Generally, phages that had identical C proteins also had identical Cox proteins, and the same nodes were uncertain in the phylogenetic analysis. The variation between classes was, however, lower than that for the C protein. Class I and class II differed at 51 positions in the C protein but only at 22 positions for the approximately same-sized Cox protein (Fig. 5). Thus, the identity between class I and class II was high (83%) (Table 3), which fits with the finding that the Cox proteins of P2 and P2 Hy dis are functionally interchangeable (30).
FIG. 5.
Alignment of the deduced amino acid sequences for the different Cox proteins. The full amino acid sequence of the type phage of each class is noted. The sequence of variants within a class is written out only where it differs from the type phage, and identical amino acids are indicated by a dot. The classes contain the following phages: class I, P2, PK, ΦD266, HK240, HK241, 299, P2-EC7, P2-EC30, P2-EC31, P2-EC45, P2-EC48, and P2-EC58; class II, P2 Hy dis, ΦD124, ΦD252, HK111, HK114, P2-EC5, P2-EC10, and P2-EC67; class III, WΦ, P2-EC44, and phage 18; class IV, HK109, HK113, HK239, L-413C, P2-EC53, P2-EC59, P2-EC61, P2-EC62, and P2-EC64; class V, ΦD160 and P2-EC12; class VI, ΦD145, P2-EC15, and P2-EC46; class VII, P2-EC4. Secondary structures predicted by JPRED are indicated below the alignments. The presumptive DNA binding HTH motif (dark gray) is preceded by a β-strand and followed by two β-strands (light gray), which together form a winged-helix structure. The consensus sequences are indicated at the bottom. The preferred integration sites are noted on the right side, where nd denotes not determined.
Recombination and selection.
The phylogenetic analyses of the amino acid sequences revealed two things. First, bootstrap analyses showed that some clusters had weak support (bootstrap percentages under 70%) (Fig. 2), especially in the Cox tree, where some characters indicated an alternative phylogenetic tree. Many characters contradicting a tree (i.e., homoplastic characters) may be a sign of homologous recombination. Second, it was also evident from the analyses that the positions of class V, class VI, and class VII differed between the two trees (Fig. 2), a fact that by itself indicated a recombinational event between C and Cox. The subsequent investigation of the causes of the homoplastic characters that contradicted some of the groups resulted in high probabilities for homologous recombination as the main evolutionary mechanism. In addition, a 100% bootstrap support does not mean that a cluster is free from homoplasies, and the analysis showed several cases of recombination between such clearly separated groups.
The investigation of the distribution of informative characters between pairs of immunity classes showed several cases of a long run of homologous characters intervened by sections consisting almost exclusively of homoplastic characters (data not shown).
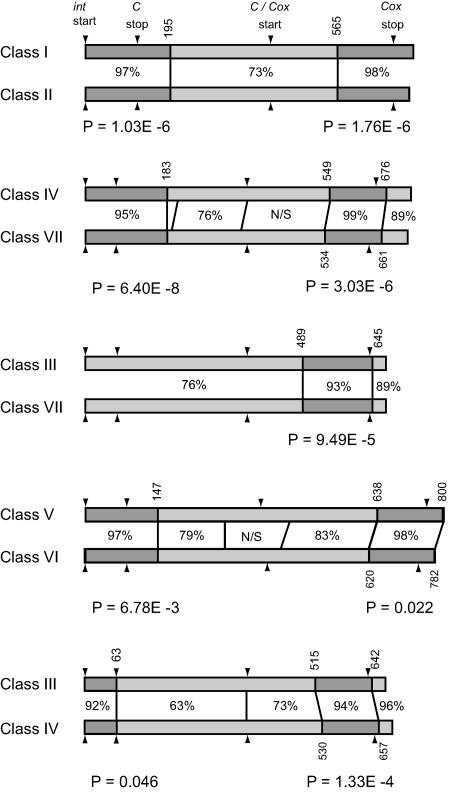
The statistical analyses for detecting and estimating the extent of recombination, using the Geneconv program, showed that none of the immunity classes studied had been unaffected by recombination. Five of the 21 pairwise tests resulted in statistically significant apparent recombinational events. All of the classes seemed to have experienced at least one instance of recombination, and the detected breakpoints for recombination were all within coding regions (Fig. 6). The program did not give any statistical support for recombination events with genes other than those contained in the data set, i.e., recombination described as resulting in “outer fragments” in Geneconv.
FIG. 6.
Location of recombination breakpoints in the genes C and cox for the five pairs of immunity groups that resulted in statistically significant recombinant fragments. Dark gray bars indicate regions of high nucleotide similarity between classes, and light gray bars indicate regions of lower similarity. Similarities are written between the nucleotide bars, and breakpoint coordinates, relative to the start codon of int, are written vertically above and below the bars. The P values below each pair are global probabilities for recombinational events reported by the Geneconv program. N/S designates a nonsignificant fragment. Arrowheads mark the start and stop codons for the genes given at the top. The intergenic region between C and cox was removed from all sequences prior to the analyses.
The recombination between the C and cox genes was also visualized by the analysis of the concatenated nucleotide sequences with the SplitsTree program (1). In the figure generated by the program, all of the immunity classes were connected to a net structure in the center, from which they radiated (Fig. 2). Most groups in the tree were related in more than one way, which is illustrated as multiple nodes leading to an immunity group. The topology of the tree indicated a complex pattern of recombination among the different immunity classes, taking place during the formation of the groups, followed by a period of positive selection in order to fine-tune the different immunities.
To detect whether any selective forces were acting on the C and Cox proteins, the ratio of synonymous substitutions per synonymous site (Ks) to nonsynonymous substitutions per nonsynonymous site (Ka) of the nucleotide sequences was calculated. The Ka/Ks ratio for C was calculated to be 0.34 (standard deviation, 0.13), and the Ka/Ks ratio for cox was calculated to be 0.45 (standard deviation, 0.30). The predominance of synonymous substitutions could be interpreted either as purifying selection leaving only neutral changes or that the directional selection took place such a long time ago that synonymous substitutions have had the time to accumulate and that the signal is no longer distinguishable.
The regulatory intergenic region.
The length of the DNA sequences between C and cox genes in the different immunity classes varies from 92 nucleotides (ΦD160) to 162 nucleotides (HK109), and the sequences differ to such a high degree that alignments are impossible. Comparison of the intergenic regions within immunity classes is possible, though, and generally show a high identity, with the lowest score being 98%.
Since the Pc promoters in P2, P2 Hy dis, and WΦ are weak, with a low identity to the consensus E. coli promoter (22, 30, 32), Pc promoters of phages from other immunity groups may also have weak promoters. A search of the intergenic region for possible Pc promoters gave no strong indications of possible promoters. Thus, their location remains to be determined.
The Cox protein of the P2-like phages analyzed couples the control of the transcriptional switch with site-specific recombination, since it acts as a repressor of Pc and as a directionality factor during site-specific recombination. Thus, Cox should bind in the vicinity of Pc and within attP. In the case of P2 and P2 Hy dis, which integrate into the same attachment site, the Cox proteins are interchangeable, even though α-helix 2, believed to be involved in DNA recognition, differs in three out of eight amino acids (22) (Fig. 5). WΦ has the same integration site as HK109, and a direct repeat (CCTAGAA[A/G]GGAC), located upstream of the Pc promoter, has been implicated as the recognition sequence of Cox. This sequence cannot be found in the intergenic region of HK109; only a subset of it, AGAA, can be found. Instead, a repeat of 6 nt (TTTGAG) is located in this region. Thus, it is possible that the Cox proteins of WΦ and HK109 are noninterchangeable.
ΦD160 and ΦD145 integrate at the same attachment site (see below), and, as can be seen in Fig. 5, they have identical amino acid sequences in α-helix 2. Thus, their Cox proteins are expected to recognize the same DNA sequence. A directly repeated sequence of 10 nt can also be found in the intergenic regions of both phages (data not shown) and in the attP region of ΦD145 (Fig. 7A) with the consensus sequence G4G5C6T6C5T4A4G5T4T6, taking all six repeats into account.
FIG. 7.
Identification of the integration site of ΦD145. (A) DNA sequence of the ΦD145 attP region. The stop codons of the ogr and int genes are shaded in dark gray. The presumed arm-binding sites P1 and P2 and P′1 and P′2 are underlined. The hypothetical IHF binding site is indicated in italics and shaded in light gray. The inverted repeats in the presumed core are underlined. Arrows indicate the presumptive Cox recognition sequences. (B) The core sequence of the attP region is capitalized and shaded in light gray. The nucleotides in the host attachment site, attB, and the left (attL) or right (attR) junctions of the prophage that are identical to the core of attP are capitalized. The imperfect inverted repeat, assumed to be recognized by the integrase, is underlined. The phage sequences are indicated in light gray, and nucleotides common to all four sequences are shaded in dark gray.
Identification of integration sites.
The integration sites for phages WΦ and P2 have previously been identified in E. coli (2, 22). In the sequenced K-12 strain MG1655, P2 integrates at 2,165.2 kb (46.6 min), and WΦ integrates at 4,104.4 kb (88.6 min). In the case of P2, the preferred integration site, locI, has been identified in E. coli strain C since this site is occupied by a defective prophage in K-12, leading to integration at several secondary sites. Using primers located on either side of locI, and primers within the phage genomes, we could show by PCR that all phages belonging to immunity class I and class II were integrated at locI. None of the others were integrated at this location, named site 1 in Fig. 3. PCR amplification with primers on either side of the integration site of WΦ, and with primers within the phage genomes, showed that all phages belonging to class III and class IV integrate at this site, denoted site 2 in Fig. 3. This left six phages and prophages with unknown integration sites, namely, those of class V, class VI, and class VII.
In order to identify these chromosomal integration sites, the DNA sequence of the attP region of phage ΦD145 was determined after PCR amplification using primers designed from well-conserved regions of the completely sequenced genomes of P2, WΦ, and L-413C. The primers were placed in the ogr gene, located to the left of attP, and in orf78, located downstream of cox, i.e., on the right side of attP (Fig. 1). The sequence between the ends of ogr and int are shown in Fig. 7A. The P2 integrase is heterobivalent. It has two DNA binding motifs that recognize different DNA sequences, the arm-binding sites, which are present as two direct repeats on either side of the core, and the core binding motif that recognizes an imperfect inverted repeat in the core. The arm-binding motif is located at the N terminus of the integrase (16), and the WΦ integrase has a similar arm-binding motif, since they recognize similar arm sequences (22). Similar repeats were also found in the presumed attP region of ΦD145, which indicated that they constitute the arm-binding sites (P1 and P2, P′1 and P′2). A comparison between the 12 direct repeats in the arm sequences of P2, WΦ, and ΦD145 showed that they were highly similar and that the consensus sequence was T12G10T12G12G12A10C12A8.
Furthermore, an IHF binding site is located to the left of the core in the P2 attP site, and since a potential IHF binding site can be found to the left of an imperfect repeat in ΦD145, it may constitute the core sequence (Fig. 7A). A GenBank search for a bacterial sequence with similarity to this hypothetical ΦD145 core gave no hits, but a search for similar integrases potentially integrating within the same core sequence resulted in a candidate integrase. The integrase of the Erwinia carotovora subsp. atroseptica prophage ΦECA29 (3) was found to be 73% identical to the integrase of ΦD145, and there was a high degree of similarity between the putative ΦD145 and ΦECA29 core regions.
PCR amplifications of three different E. coli ΦD145 lysogen genomes with primers designed from the E. carotovora DNA flanking the ΦECA29 prophage and primers from within phage ΦD145 DNA all generated identical PCR fragments. Sequence analyses of the prophage junctions confirmed that ΦD145 integrates into E. coli at the equivalent location as ΦECA29 in Erwinia, namely, into the very beginning of the pflA gene. In fact, the attB sequence contains the start codon of pflA, and the integration of ΦD145 disrupts the gene. The pflA gene codes for an enzyme, pyruvate formate-lyase-activating enzyme, that is involved in the fermentation of pyruvate in microaerobic conditions (40).
The attB sequence, corresponding to ΦD145 attP of the nonlysogenic strain E. coli C-1a, was determined after PCR amplification using primers on either side of the presumed attB sequence. Analysis of the sequence showed that this region is identical to E. coli K-12 strain MG1655 (11). As shown in Fig. 7B, the ΦD145 core sequence and the attB sequence share an identical region of 13 nt, but the sequence similarity can be extended if mismatches are tolerated. The identical region should correspond to the region where branch migration occurs during recombination, while the imperfect inverted repeats should constitute the integrase recognition sequence. As can be seen in Fig. 7B, the imperfect repeat makes it clear that the recombination has occurred within the 13-nt region of identity.
The center of the attB region lies between positions 950293 and 950294 in the MG1655 map, but the region should contain at least 20 nt centered around this position if the inverted repeats are recognized by the integrase.
ΦD145 integrates at the 5′ end of the pflA gene, and the crossing over occurs between positions +3 and +15 from the start codon of the gene. This creates a truncated gene at attR that codes for a peptide of eight amino acids. After that, there are two inverted repeats; one of them is the ogr/int terminator that prevents transcription from the bacteria to continue into the prophage or vice versa. In the attL region, the start codon of pflA is substituted by an AAG codon that codes for lysine. In the same frame, but 60 nt upstream of the lysine codon, there is a start codon. This ATG codon is preceded by a potential ribosomal binding site. In the equivalent region of ΦECA29 and Erwinia, there is also an in-frame start codon at the same distance, but the N-terminal additions to PflA show low similarity between ΦD145 and ΦECA29. This implies that the pflA gene is on the same transcript as the int gene and that the 20 additional amino acids at the N terminus seem not to affect its biological activity.
By using the same set of primers as those used for analyzing the integration site of ΦD145, ΦD160, P2-EC12, P2-EC15, and P2-EC46 were found to use the same integration site as ΦD145. However, this site is empty in ECOR4, and the integration site of P2-EC4 thus remains unknown.
DISCUSSION
In this work, we have analyzed the developmental switches of 38 P2-like coliphages or prophages that can be classified into seven discrete immunity classes. The analysis further indicates that homologous recombination has played an important role in generating the different classes. Interestingly, among all these phages, we found only four different host attachment sites, and we found no case where phages of the same immunity class had different host attachment sites. The majority of genes in P2-like phages from various γ-proteobacteria seem to have the same phylogenetic relationship. This phylogeny resembles the tree of their different γ-proteobacterial hosts, which indicates that P2-like phages coevolve with their hosts (Nilsson, unpublished). What then is the biological significance of having different immunities and integration sites?
Since P2-like phages are commonly encountered as prophages in E. coli, superinfections of a lysogen, or possibly mixed infections, can be expected to be frequent events in nature. The outcome of such infections will depend on whether the phages have the same or different immunities and whether they share the same or have different host attachment sites. If a P2 lysogen is superinfected with another P2-like phage with the same immunity and attachment site, it will be repressed and must integrate to be stably maintained in the cell. However, P2 tandem double prophages are unstable due to int expression (7, 8), so the superinfecting phage will either integrate at secondary sites, which could affect its capacity to excise upon induction, or replace the resident prophage. Consequently, gaining a different immunity should be an advantage under these circumstances, since the superinfecting phage would be able to enter the lytic cycle. This is also in accordance with our observation that the immunities have diverged within the different integration site classes. When the regular integration site is occupied, there ought to be strong selection for a new immunity. But this is in conflict with the fact that there must be plenty of phages with different immunities around, even though they integrate at other sites, and there may be no need for developing a new immunity.
We have observed that immunity seems to be complete between members of the same immunity class, but we have also observed that it is sometimes incomplete or nonreciprocal between phages from different groups, like the immunity of the class VI phage ΦD145. There are several possible explanations for this. In uniform and infinitely large populations of phages and hosts, immunity would be expected to evolve through frequency-dependent selection and to result in a multitude of phages with different immunities. However, phages form classes containing many phages with the same immunity, and the number of immunity classes is limited. Consequently, an unknown spatial clonal variation of bacterial hosts, each harboring phages from all immunity classes and integrating at all possible locations, might exist. According to this hypothesis, phage ΦD145 should be found in host populations that do not contain ΦD160 or HK111, which are able to block transcription from their Pe promoters. It is also difficult to explain the coexistence of ΦD145 and HK114. Phage ΦD145 should be able to outnumber and possibly eradicate HK114 since it can use that lysogen for its lytic growth, and HK114 cannot grow on a ΦD145 lysogen. One explanation would be that they never meet. Another hypothesis is that the fitness of the phage is not dependent on immunity only. The number of possible hosts, lysogenic or not, that a phage with a specific type of immunity can use may be limited, but this could be balanced by lysogenic conversion genes that increase the fitness of the host and, indirectly, the phage as well. A third hypothesis is that immunity is a characteristic that is under such evolutionary constraint that the rate of change is slow. Expression of immunity involves both the C protein and the operators to which it binds, and there are many possible outcomes of a change in either of these. The ΦD145 and ΦD160 C proteins show 78% identity at the amino acid level (Table 3). They differ by only two amino acids in the third α-helix, which is believed to be the part that recognizes the operators. It is not surprising that this small difference affects immunity. Some phages may express an immunity that is “under construction” but sufficient under the actual circumstances.
The disagreement of the phylogenetic analyses of C and Cox together with the results of the analyses for detecting recombination performed in this work support homologous recombination as the causative agent for the generation of new immunity classes. P2 Hy dis is such an example. P2 Hy dis was obtained after P2 infection of E. coli B, containing a cryptic prophage with dis immunity (6, 13). Later, three nonhomologous regions between P2 and P2 Hy dis were identified by electron microscope heteroduplex mapping (12). Whether P2 Hy dis was generated by one or several recombination events is not known, since the cryptic prophage has not been sequenced. However, a comparison of the DNA sequence of the int gene regions of phages P2 and ΦD266, belonging to immunity class I, with those of ΦD252 and P2 Hy dis, belonging to immunity class II, shows a high level of identity. The variation is only between 0.5 and 1% at the nucleotide level (unpublished data). Thus, homologous recombination between the int genes and gene B between phages of immunity groups I and II would generate the same heteroduplex loop in this region as that obtained with P2 and P2 Hy dis.
Considering the clonal evolution of P2-like coliphages, the question is from where the different immunities originated. One possible scenario is that a phage with a different host preference occasionally infects E. coli, allowing recombination between homologous regions with a resident P2-like prophage that will gain a new immunity and higher survival fitness when competing with other P2-like coliphages. Since a survey of immunities of P2-like phages in other enterobacteria has not been performed, this hypothesis cannot be validated.
The evolution of new phage attachment sites is more complex due to the complex structure of attP. The heterobivalent Int protein binds not only to the core sequence but also to the arm sites located on each side of the core, excluding recombination between an incoming phage and a prophage as a possibility for gaining new site preferences. In addition, the Cox binding sites located between the core and one of the arm sites must be compatible with the binding sites in the transcriptional switch. The generation of new site preferences may therefore occur by two routes. P2 saf is a P2 mutant with an altered site preference that has a single-base-pair substitution within the core sequence (37, 38). P2 saf was isolated from a P2 prophage located at secondary integration site II (not to be confused with regular integration site 2 in this paper) and is an example of how new attachment sites may evolve after integration and excision from secondary attachment sites. Under these conditions, the recombinant phage should have the same immunity as before, but it should have a new site preference. Alternatively, new attachment sites could be generated by homologous recombination between two phage genomes present simultaneously in the same cell, where one phage originates from a different bacterial host. Since the Cox protein has a dual function, the recombination event must include the whole attP, int, C, and cox region, possibly leading to a change in attachment site and immunity at the same time. A possible example of such a recombination event is the Erwinia prophage ΦECA29, which integrates at the same location as the P2-like coliphages of class V and VI, but since then, new recombination has occurred, giving different immunities. It is also interesting that ΦD145 integrates into the coding part of a gene. It integrates into pflA and provides the 5′ sequence that changes the start codon to an AAG and restores the 5′ end of the pflA gene, as opposed to most genetic elements, which restore the 3′ end of the disrupted gene. The core region and attB of ΦD145 have some mismatches, in contrast to P2 and WΦ, where their att sites are identical. As with the immunity, the integration of ΦD145 indicates differences that could be interpreted as imperfect and evolution on the way.
The fact that we have not found any P2-like phages belonging to the same immunity group that have different attachment sites is intriguing, since under laboratory conditions, P2 triple lysogens can be established where P2 is integrated at secondary sites (23). It is possible that phages having that same immunity but different attachment sites are not as competitive and are therefore removed by selection.
Acknowledgments
We thank Joe Bertani for sharing unpublished results, for critical reading of a draft version of the paper, and for many valuable comments. We also thank Sara Renberg Eriksson for sequencing the attP region of phage ΦD160 and Georgina Ibrahim Isak for providing unpublished observations.
This work was supported by grants from the Swedish Research Council and the Sven and Lilly Lawski Foundation.
REFERENCES
- 1.Bandelt, H.-J., and A. W. M. Dress. 1992. Split decomposition: a new and useful approach for the phylogenetic analysis of distance data. Mol. Phylogenet. Evol. 1:242-252. [DOI] [PubMed] [Google Scholar]
- 2.Barreiro, V., and E. Haggård-Ljungquist. 1992. Attachment sites for bacteriophage P2 on the Escherichia coli chromosome: DNA sequences, localization on the physical map, and detection of a P2-like remnant in E. coli K-12 derivatives. J. Bacteriol. 174:4086-4093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bell, K. S., M. Sebaihia, L. Pritchard, M. T. Holden, L. J. Hyman, M. C. Holeva, N. R. Thomson, S. D. Bentley, L. J. Churcher, K. Mungall, R. Atkin, N. Bason, K. Brooks, T. Chillingworth, K. Clark, J. Doggett, A. Fraser, Z. Hance, H. Hauser, K. Jagels, S. Moule, H. Norbertczak, D. Ormond, C. Price, M. A. Quail, M. Sanders, D. Walker, S. Whitehead, G. P. Salmond, P. R. Birch, J. Parkhill, and I. K. Toth. 2004. Genome sequence of the enterobacterial phytopathogen Erwinia carotovora subsp. atroseptica and characterization of virulence factors. Proc. Natl. Acad. Sci. USA 101:11105-11110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bertani, G. 1957. A disimmune mutant of temperate bacteriophage P2. Bacteriol. Proc., p.38.
- 5.Bertani, G. 1951. Studies on lysogenesis. I. The mode of phage liberation by lysogenic Escherichia coli. J. Bacteriol. 62:293-300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bertani, L. E. 1957. The effect of the inhibition of protein synthesis on the establishment of lysogeny. Virology 4:53-71. [DOI] [PubMed] [Google Scholar]
- 7.Bertani, L. E. 1970. Split-operon control of a prophage gene. Proc. Natl. Acad. Sci. USA 65:331-336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bertani, L. E. 1971. Stabilization of P2 tandem double lysogens by int mutations in the prophage. Virology 46:426-436. [DOI] [PubMed] [Google Scholar]
- 9.Bertani, L. E., and G. Bertani. 1971. Genetics of P2 and related phages. Adv. Genet. 16:199-237. [DOI] [PubMed] [Google Scholar]
- 10.Bertani, L. E., and E. W. Six. 1988. The P2-like phages and their parasite, P4, p. 73-143. In R. Calendar (ed.), The bacteriophages 2, vol. 2. Plenum Publishing Corporation, New York, N.Y. [Google Scholar]
- 11.Blattner, F. R., G. Plunkett III, C. A. Bloch, N. T. Perna, V. Burland, M. Riley, J. Collado-Vides, J. D. Glasner, C. K. Rode, G. F. Mayhew, J. Gregor, N. W. Davis, H. A. Kirkpatrick, M. A. Goeden, D. J. Rose, B. Mau, and Y. Shao. 1997. The complete genome sequence of Escherichia coli K-12. Science 277:1453-1474. [DOI] [PubMed] [Google Scholar]
- 12.Chattoraj, D. K., and R. B. Inman. 1973. Electron microscope heteroduplex mapping of P2 Hy dis bacteriophage DNA. Virology 55:174-182. [DOI] [PubMed] [Google Scholar]
- 13.Cohen, D. 1959. A variant of phage P2 originating in Escherichia coli, strain B. Virology 7:112-126. [DOI] [PubMed] [Google Scholar]
- 14.Dhillon, E. K. S., T. S. Dhillon, Y. Y. Lam, and A. H. C. Tsang. 1980. Temperate coliphages: classification and correlation with habitats. Appl. Environ. Microbiol. 39:1046-1053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.ElAntak, L., M. Ansaldi, F. Guerlesquin, V. Méjean, and X. Morelli. 2005. Structural and genetic analyses reveal a key role in prophage excision for the TorI response regulator inhibitor. J. Biol. Chem. 280:36802-36808. [DOI] [PubMed] [Google Scholar]
- 16.Frumerie, C., L. Sylwan, A. Ahlgren-Berg, and E. Haggard-Ljungquist. 2005. Cooperative interactions between bacteriophage P2 integrase and its accessory factors IHF and Cox. Virology 332:284-294. [DOI] [PubMed] [Google Scholar]
- 17.Geisselsoder, J., and M. Mandel. 1970. Physical properties of phage 299. Mol. Gen. Genet. 108:158-166. [PubMed] [Google Scholar]
- 18.Glover, S. W., and G. Kerszman. 1967. The properties of a temperate bacteriophage WΦ isolated from Escherichia coli strain W. Genet. Res. Camb. 9:135-139. [Google Scholar]
- 19.Ilangovan, U., J. M. Wojciak, K. M. Connolly, and R. T. Clubb. 1999. NMR structure and functional studies of the Mu repressor DNA-binding domain. Biochemistry 38:8367-8376. [DOI] [PubMed] [Google Scholar]
- 20.Jacob, F., and E. L. Wollman. 1956. Sur les processus de conjugaison et de recombinaison chez Escherichia coli. I. L'induction par conjugaison ou induction zygotique. Ann. Inst. Pasteur (Paris) 91:486-510. [PubMed] [Google Scholar]
- 21.Jesaitis, M. A., and J. J. Hutton. 1963. Properties of a bacteriophage derived from Escherichia coli K235. J. Exp. Med. 117:285-302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Liu, T., and E. Haggård-Ljungquist. 1999. The transcriptional switch of bacteriophage WΦ, a P2-related but heteroimmune coliphage. J. Virol. 73:9816-9826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lundqvist, B., and G. Bertani. 1984. Immunity repressor of bacteriophage P2. Identification and DNA-binding activity. J. Mol. Biol. 178:629-651. [DOI] [PubMed] [Google Scholar]
- 24.Maynard Smith, J., and N. H. Smith. 1998. Detecting recombination from gene trees. Mol. Biol. Evol. 15:590-599. [DOI] [PubMed] [Google Scholar]
- 25.Nilsson, A. S., and E. Haggård-Ljungquist. 2001. Detection of homologous recombination among bacteriophage P2 relatives. Mol. Phylogenet. Evol. 21:259-269. [DOI] [PubMed] [Google Scholar]
- 26.Nilsson, A. S., and E. Haggård-Ljungquist. 2006. The P2-like bacteriophages, p. 365-390. In R. Calendar (ed.), The bacteriophages. ASM Press, Washington, D.C.
- 27.Nilsson, A. S., J. L. Karlsson, and E. Haggård-Ljungquist. 2004. Site-specific recombination links the evolution of P2-like coliphages and pathogenic enterobacteria. Mol. Biol. Evol. 21:1-13. [DOI] [PubMed] [Google Scholar]
- 28.Ochman, H., and R. K. Selander. 1984. Standard reference strains of Escherichia coli from natural populations. J. Bacteriol. 157:690-693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Pizer, L. I., H. S. Smith, M. Miovic, and L. Pylkas. 1968. Effect of prophage W on the propagation of bacteriophage T2 and T4. J. Virol. 2:1339-1345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Renberg-Eriksson, S. K., A. Ahlgren-Berg, J. DeGrooth, and E. Haggård-Ljungquist. 2001. Characterization of the developmental switch region of bacteriophage P2 Hy dis. Virology 290:199-210. [DOI] [PubMed] [Google Scholar]
- 31.Rogov, V. V., C. Lucke, L. Muresanu, H. Wienk, I. Kleinhaus, K. Werner, F. Lohr, P. Pristovsek, and H. Ruterjans. 2003. Solution structure and stability of the full-length excisionase from bacteriophage HK022. Eur. J. Biochem. 270:4846-4858. [DOI] [PubMed] [Google Scholar]
- 32.Saha, S., E. Haggård-Ljungquist, and K. Nordström. 1987. The Cox protein of bacteriophage P2 inhibits the formation of the repressor protein and autoregulates the early operon. EMBO J. 6:3191-3199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sam, M. D., D. Cascio, R. C. Johnson, and R. T. Clubb. 2004. Crystal structure of the excisionase-DNA complex from bacteriophage lambda. J. Mol. Biol. 338:229-240. [DOI] [PubMed] [Google Scholar]
- 34.Sambrook, J., and D. W. Russell. 2001. Molecular cloning: a laboratory manual, 3rd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 35.Sasaki, I., and G. Bertani. 1965. Growth abnormalities in Hfr derivatives of Escherichia coli strain C. Virology 40:365-376. [DOI] [PubMed] [Google Scholar]
- 36.Sawyer, S. 1989. Statistical tests for detecting gene conversion. Mol. Biol. Evol. 6:526-538. [DOI] [PubMed] [Google Scholar]
- 37.Six, E. 1966. Specificity of P2 for prophage site I on the chromosome of Escherichia coli strain C. Virology 29:106-125. [DOI] [PubMed] [Google Scholar]
- 38.Yu, A., L. E. Bertani, and E. Haggård-Ljungquist. 1989. Control of prophage integration and excision in bacteriophage P2: nucleotide sequences of the int gene and att sites. Gene 80:1-12. [DOI] [PubMed] [Google Scholar]
- 39.Yu, A., and E. Haggård-Ljungquist. 1993. The Cox protein is a modulator of directionality in bacteriophage P2 site-specific recombination. J. Bacteriol. 175:7848-7855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zhu, J., and K. Shimizu. 2004. The effect of pfl gene knockout on the metabolism for optically pure D-lactate production by Escherichia coli. Appl. Microbiol. Biotechnol. 64:367-375. [DOI] [PubMed] [Google Scholar]