Abstract
In Salmonella enterica serovar Typhimurium, the stationary-phase sigma factor σS (RpoS) is required for virulence, stress resistance, biofilm formation, and development of the rdar morphotype. This morphotype is a multicellular behavior characterized by expression of the adhesive extracellular matrix components cellulose and curli fimbriae. The Crl protein of Escherichia coli interacts with σS and activates expression of σS-regulated genes, such as the csgBAC operon encoding the subunit of the curli proteins, by an unknown mechanism. Here, we showed using in vivo and in vitro experiments that the Crl protein of Salmonella serovar Typhimurium is required for development of a typical rdar morphotype and for maximal expression of the csgD, csgB, adrA, and bcsA genes, which are involved in curli and cellulose biosynthesis. In vitro transcription assays and potassium permanganate reactivity experiments with purified His6-Crl showed that Crl directly activated σS-dependent transcription initiation at the csgD and adrA promoters. We observed no effect of Crl on σ70-dependent transcription. Crl protein levels increased during the late exponential and stationary growth phases in Luria-Beratani medium without NaCl at 28°C. We obtained complementation of the crl mutation by increasing σS levels. This suggests that Crl has a major physiological impact at low concentrations of σS.
Bacteria are routinely exposed to limited nutrients and other stressful conditions in their natural habitats. Consequently, they often grow and divide slowly and enter a stationary phase. The stationary-phase physiology, particularly multiple-stress resistance, and even cell morphology are determined by the general stress response. This response is controlled at the molecular level by a sigma subunit of RNA polymerase, σS (RpoS). In enterobacteria, RNA polymerase is composed of a core enzyme, E, with the subunit structure α2ββ′ω, which associates with one of the seven different σ subunits to form the holoenzyme (Eσ). Each σ subunit targets the RNA polymerase to a different set of promoters, modulating the gene expression patterns. The RNA polymerase holoenzyme containing the σ70 subunit is responsible for the transcription of most genes during exponential growth. When an organism enters the stationary phase or when it is under specific stress conditions during the exponential growth phase (high osmolarity, low pH, or high and low temperatures), σS, which is encoded by the rpoS gene, accumulates in the cell, associates with the core enzyme, and directs the transcription of genes essential for the general stress response and for stationary-phase survival (for reviews see references 19 and 21). Regulation of σS occurs during transcription and posttranscription and involves numerous regulators (for a review see reference 19). Recent studies have shown that the crl gene product is a regulator of σS activity in Escherichia coli (8, 35). Crl modulates the expression of σS-regulated genes (poxB, katE, bolA, and csgB) (35). Crl also interacts directly with the σS protein in vitro and associates with EσS at the csgBAC promoter, which governs the synthesis of curli fimbria subunits (8). However, the stimulating effect of Crl could not be reproduced in vitro with purified products, and the mechanism by which Crl influences σS activity is unknown.
To investigate the role of Crl at the molecular and physiological levels, our strategy was to first search for a particular phenotype associated with the crl knockout mutant. The Crl protein was initially implicated in the production of curli fibers in E. coli (1, 31, 32). These thin aggregative surface fimbriae, which are conserved among Enterobacteriaceae, are involved in a number of different processes. These processes include adhesion to soluble extracellular matrices, biofilm formation, interaction with eukaryotic cells, contact with major histocompatibility complex class I molecules, and induction of a proinflammatory response (2-7, 11, 14, 17, 20, 24, 31-33, 36, 42, 46, 47, 51, 54-57). Curli fimbriae and cellulose are the major components of the extracellular matrix involved in the multicellular behavior of Salmonella and some natural isolates of E. coli called the rdar (red dry and rough) morphotype (for a review see reference 40). Cells expressing the rdar morphotype absorb the dye Congo red (CR) from the medium and turn dark red. They have a dry and rough surface and form irregularly shaped, flat colonies that spread on plates (40) (Fig. 1). The rdar colonies also absorb the fluorescent dye calcofluor, which binds cellulose (41, 52, 58) (Fig. 1).
FIG. 1.
rdar morphotypes of Salmonella strains: differential expression of the rdar morphotypes of Salmonella wild-type strain ATCC 14028 and its mutant derivatives ATCCcrl, ATCCrpoS, ATCCbcsA, ATCCcsgB, ATCCcrl bcsA, and ATCCcrl csgB after 5 days of growth at 28°C on CR plates (A) and on LB0 plates supplemented with calcofluor (B). ATCC 14028 and ATCCrpoS have typical rdar and saw morphotypes, respectively (42, 46). The crl mutant has a previously undescribed morphotype (rdarcrl), which was affected by elimination of curli production in the crl csgB double-mutant strain and by elimination of cellulose production in the crl bcsA double-mutant strain.
Expression of the rdar morphotype in Salmonella enterica serovar Typhimurium has been extensively studied. This morphotype occurs during the stationary growth phase under certain environmental conditions (for reviews see references 16 and 30) and is most efficient at low temperatures and low osmolarity. The rdar morphotype is positively regulated by csgD, a gene encoding a putative response regulator belonging to the FixJ family (for a review see reference 16). CsgD is required for expression of csgBAC, the operon encoding the curli subunits. CsgD also stimulates cellulose production via activation of transcription of adrA, which encodes a diguanylate cyclase protein (40, 52, 58). RpoS is required for development of the rdar morphotype and for csgBAC, csgDEFG, and adrA gene expression, although it is not known whether RpoS acts directly or indirectly at these promoters through CsgD or additional regulatory proteins (18, 31, 42, 45, 46, 52, 58).
Our studies indicated that expression of the rdar morphotype and transcription of the genes involved in curli and cellulose biosynthesis are affected in a crl knockout mutant of Salmonella serovar Typhimurium. In vitro transcription experiments with purified Crl protein of Salmonella showed that Crl stimulates σS-dependent transcription at the adrA and csgD promoters. This is the first report showing that Crl has a direct role in transcription initiation and that a crl knockout mutation has a physiological effect in Salmonella.
MATERIALS AND METHODS
Bacterial strains, phages, and growth conditions.
The E. coli and Salmonella serovar Typhimurium strains used in this study are listed in Table 1. Bacteriophage P22HT105/1int was used to transduce mutations between Salmonella strains (49). Green plates, which were used to screen for P22-infected cells or lysogens, were prepared as described previously (53). Strains were routinely cultured at 37°C in Luria-Bertani (LB) medium unless indicated otherwise (48). Antibiotics were used at the following concentrations: carbenicillin, 100 μg ml−1; chloramphenicol, 15 μg ml−1 for the chromosomal resistance gene and 30 μg ml−1 for the plasmid resistance gene; kanamycin, 50 μg ml−1; and tetracycline, 20 μg ml−1.
TABLE 1.
Bacterial strains and plasmids used in this study
| Strain or plasmid | Characteristics | Reference or source |
|---|---|---|
| E. coli strains | ||
| JM109 | recA1 supE44 endA1 hsdR17 gyrA96 relA1 thi Δ(lac-proAB) F′ (traD36 proAB+lacIqlacZΔM15) | 48 |
| MC1061 | hsdR mcrB araD139 Δ(araABC-leu)7679 ΔlacX74 galU galK rpsL thi | 48 |
| Salmonella serovar Typhimurium strains | ||
| ATCC 14028 | Wild type | American Type Culture Collection |
| ATCCrpoS | ATCC 14028 ΔrpoS::cm | This study |
| ATCCcrl | ATCC 14028 Δcrl::cm | This study |
| ATCCcsgD | ATCC 14028 ΔcsgD::cm | This study |
| ATCCcsgB | ATCC 14028 ΔcsgB::cm | This study |
| ADR1a | MAE52 adrA::MudJ | 45 |
| MAE150 | MAE52 bcsA::MudJ | 58 |
| ATCCadrA | ATCC 14028 adrA::MudJ | This study |
| ATCCbcsA | ATCC 14028 bcsA::MudJ | This study |
| ATCCrpoScrl | ATCC 14028 ΔrpoS Δcrl::cm | This study |
| ATCCcsgB-lacZ | ATCC 14028 csgB-lacZY | This study |
| ATCCcsgD-lacZ | ATCC 14028 csgD-lacZY | This study |
| ATCCrpoScsgD | ATCC 14028 ΔrpoS ΔcsgD::cm | This study |
| ATCCrpoSbcsA | ATCC 14028 ΔrpoS::cm bcsA::MudJ | This study |
| ATCCrpoSadrA | ATCC 14028 ΔrpoS::cm adrA::MudJ | This study |
| ATCCrpoScsgD-lacZ | ATCC 14028 ΔrpoS::cm csgD-lacZY | This study |
| ATCCrpoScsgB-lacZ | ATCC 14028 ΔrpoS::cm csgB-lacZY | This study |
| ATCCcrlbcsA | ATCC 14028 Δcrl::cm bcsA::MudJ | This study |
| ATCCcrlcsgB | ATCC 14028 Δcrl ΔcsgB::cm | This study |
| ATCCcrladrA | ATCC 14028 Δcrl::cm adrA::MudJ | This study |
| ATCCcrlcsgD-lacZ | ATCC 14028 Δcrl::cm csgD-lacZY | This study |
| ATCCcrlcsgB-lacZ | ATCC 14028 Δcrl::cm csgB-lacZY | This study |
| ATCCcsgDadrA | ATCC 14028 ΔcsgD::cm adrA::MudJ | This study |
| ATCCcsgDbcsA | ATCC 14028 ΔcsgD::cm bcsA::MudJ | This study |
| ATCCcsgDcsgB-lacZ | ATCC 14028 ΔcsgD::cm csgB-lacZY | This study |
| Plasmids | ||
| pQE30 | Cbr, for expression of His-tagged proteins | QIAGEN |
| pQEcrl | Cbr, pQE30::crl, expresses a His6-Crl protein | This study |
| pACYC184 | Cmr Tetr, cloning vector | 10 |
| pACcrl-1 | Tetr, pACYC184::crl, cat, and crl in the same orientation | This study |
| pACcrl-2 | Tetr, pACYC184::crl, cat, and crl in the opposite orientation | This study |
| pSTK4 | Cmr, pACYC184::rpoS | 23 |
| pJCD01 | Cbr, pUC with transcriptional terminators | 28 |
| pJCDadrA | Cbr, pJCD01::adrA | This study |
| pGO1 | Cbr, pJCD01::lac | 34 |
| pU-2750 | Cbr, pJCD01::csgD | U. Gerstel et al., unpublished |
Phenotypic assays.
For routine monitoring of multicellular behavior, Salmonella strains were grown on LB agar plates without NaCl (LB0) at 28°C. We determined colony morphology and color using LB0 agar supplemented with Congo red (40 μg ml−1) and Coomassie brilliant blue (20 μg ml−1) (CR plates), as described previously (41). We determined cellulose production using LB0 agar supplemented with calcofluor (fluorescent brightener 28; 50 μg ml−1), as described previously (41). We detected calcofluor binding of agar-grown colonies by observing the cells under a 312-nm UV light source.
DNA manipulations.
We used standard molecular biology techniques (39, 48) for DNA manipulations. DNA was sequenced by Genomexpress (Paris, France). Oligonucleotides were obtained from Distribio (France). Strains were constructed either by generalized transduction using phage P22HT105/1int (49) or by linear DNA transformation as described below.
Construction of plasmids.
The plasmids used in this study are listed in Table 1. We determined the nucleotide sequence of a PCR-amplified crl gene from Salmonella strain ATCC 14028. The crl gene in ATCC 14028 was identical to that in Salmonella strain LT2 (http://genomeold.wustl.edu/projects/bacterial/styphimurium/). We constructed pACcrl-1 and pACcrl-2 by using primers specific for the ends of crl (5′-TTCCGAGAATTCTTATGCCGACAGTTTTACCGGCTCG-3′ and 5′-AGGCTCGAATTCTAAAGGAGATCGCAATGACGTTACCGA-3′) to amplify a 0.5-kb promoterless crl gene from ATCC 14028 total DNA by PCR. EcoRI restriction sites (underlined) were incorporated at the 5′ and 3′ ends. After digestion with EcoRI, the fragment was inserted into the EcoRI site of pACYC184 in both orientations to obtain pACcrl-1 and pACcrl-2 (in pACcrl-1 the crl and cat genes are in the same orientation and the crl gene is likely transcribed from the cat promoter). We checked the nucleotide sequence of the crl insert in pACcrl-1 and pACcrl-2 by DNA sequencing. We constructed pQEcrl, which expresses an N-terminal His6 fusion to the crl gene product under the control of the pQE30 isopropyl-β-d-thiogalactopyranoside (IPTG)-inducible promoter, as follows. We used primers specific for the ends of crl (5′-AGGCTCGGATCCATGACGTTACCGAGTGGACACCCGA-3′ and 5′-TTCCGAAAGCTTTTATGCCGACAGTTTTACCGGCTCG-3′) to amplify a 0.45-kb fragment of ATCC 14028 total DNA by PCR. BamHI and HindIII restriction sites (underlined) were incorporated at the 5′ and 3′ ends, respectively. After digestion with BamHI and HindIII, the PCR-amplified fragment was ligated into the BamHI and HindIII sites of pQE30. We checked the nucleotide sequence of the crl insert in pQEcrl by DNA sequencing. The DNA templates for in vitro transcription were prepared as follows. A 679-bp csgD fragment and a 172-bp adrA fragment were amplified by PCR from strain ATCC 14028 using primers 5′-GGGCTGCAGCACATCTGACAGCTGCCCCC and 5′-GGAGGATCCAGATCATAATTTGTCG-3′ for csgD and primers 5′-CGAGAATTCGGCGTAGTGCTATCGG-3′ and 5′-GGCGGATCCGCCTGTTCAACCGC-3′ for adrA. After digestion with BamHI and PstI (underlined sites), the csgD promoter fragment from bp −684 to 121 relative to the transcriptional start site was inserted into BamHI- and PstI-digested pJCD01. The resulting plasmid was sequenced and designated pU-2750. After digestion with EcoRI and BamHI, the adrA promoter fragment from about bp −92 to 73 relative to the major transcriptional start site determined by using the homologous gene yaiC in E. coli (9) was inserted into EcoRI- and BamHI-digested pJCD01 to obtain pJCDadrA.
One-step inactivation of chromosomal genes and construction of chromosomal lac fusions.
We created chromosomal mutations in the rpoS, crl, csgB, and csgD genes of Salmonella strain ATCC 14028 using PCR-generated linear DNA fragments and the λRed recombination method, as described by Datsenko and Wanner (13). Briefly, we used 63- to 66-nucleotide (nt) primers with 43- to 46-nt homology with the gene of interest on the 5′ end of the primer and 20-nt homology with the FLP recognition target-flanked antibiotic resistance cassette of plasmid pKD3 at the 3′ end (for sequences see reference 13). We used primers RpoS-P1 and RpoS-P2 (Table 2) for disruption of the rpoS gene, primers Crl-P1 and Crl-P2 for disruption of the crl gene, primers CsgB-P1 and CsgB-P2 for disruption of the csgB gene, and primers CsgD-P1 and CsgD-P2 for disruption of the csgD gene. ATCC 14028 containing plasmid pKD46, which carries the λ recombination genes gam, bet, and exo under control of the araBAD promoter (13), was grown overnight at 30°C, diluted in LB medium containing carbenicillin and l-arabinose (1 mM), and then grown to an optical density at 600 nm of 0.5. Electrocompetent cells were prepared, transformed with the PCR-generated linear fragments, plated on LB medium containing chloramphenicol (15 μg ml−1), and incubated at 37°C. We then characterized the resulting colonies using a combination of PCRs performed with locus-specific primers and common test primers (13). Finally, isogenic strains were constructed by P22HT/int-mediated transduction of mutations into the appropriate strains. When required, the Cmr resistance cassette was eliminated using temperature-sensitive helper plasmid pCP20, which encodes the FLP recombinase (13). We constructed single-copy csgB-lacZY and csgD-lacZY transcriptional gene fusions from mutants ATCCcsgB::Cm and ATCCcsgD::Cm using conditional plasmids containing promoterless lacZY genes and the FLP recognition target site, as described previously (15). We then used PCR assays to ensure that the plasmids were integrated in the correct location and to determine the presence of multiple plasmid integrants (using common test primers, such as those described by Ellermeier et al. [15]). We also used flanking locus-specific primers to amplify junction fragments that were subsequently analyzed by DNA sequencing. Isogenic strains were constructed by P22HT/int-mediated transduction of the mutations into the appropriate strains.
TABLE 2.
Primers used for gene mutagenesis
| Primer | Sequence | Gene mutated |
|---|---|---|
| RpoS-P1 | GTCAGAATACGCTGAAAGTTCATGATTTAAATGAAGACGCGGAGTGTAGGCTGGAGCTGCTTC | rpoS |
| RpoS-P2 | CTCGCGGAACAGCGCTTCGATATTCAGCCCCTGCGTCTGCAGACATATGAATATCCTCCTTAG | rpoS |
| Crl-P1 | CAGTTGCTTCATTAAAGGAGATCGCAATGACGTTACCGAGTGGGTGTAGGCTGGAGCTGCTTC | crl |
| Crl-P2 | CTGATGGCGCTGCGCCATCAGGCATGGCAGCAGGGTTATGCCGCATATGAATATCCTCCTTAG | crl |
| CsgB-P1 | GATAATTTTCGCTATGTACGACCAGGTCCAGGGTGACAGCATGAAAGTGTAGGCTGGAGCTGCTTC | csgB |
| CsgB-P2 | CCCATCGGATTGATTTAAAAGTCGTAACGGTATTAGCGTTGGGTGCATATGAATATCCTCCTTAG | csgB |
| CsgD-P1 | GCTGTCAGATGTGCGATTAAAAAAAGTGGAGTTTCATCATGTTGTGTAGGCTGGAGCTGCTTC | csgD |
| CsgD-P2 | GCTGCTACAATCCAGGTCAGATAGCGTTTCATGGCCTTACCGCCATATGAATATCCTCCTTAG | csgD |
Antibodies, electrophoresis, and immunoblot analysis of proteins.
Preparation of whole-cell extracts and sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis were carried out as described by Silhavy et al. (50). We determined the amounts of protein in whole-cell lysates using a DC protein assay kit (Bio-Rad). We loaded equal amounts of protein in each slot. The relative molecular weights of the proteins were estimated using molecular weight standards (Euromedex). Denaturing SDS-polyacrylamide gels were stained using Bio-Safe Coomassie blue (Bio-Rad). Antibodies against the Crl protein of Salmonella were produced in rabbits by injecting highly purified His6-Crl protein (P.A.R.I.S Production d'Anticorps, France). Rabbit antibodies against the σS protein of S. enterica serovar Typhimurium were obtained from Coynault et al. (12). Proteins were transferred on Hybond P membranes (Amersham Life Sciences) and incubated with the polyclonal rabbit antibody serum as previously described (12). The bound antibodies were detected using a secondary anti-rabbit antibody linked to peroxidase and an ECL Plus Western blot detection system kit (Amersham Life Sciences).
Electron microscopy.
Electron microscopy was performed with bacteria that had been grown for 5 to 6 days on LB0 plates at 28°C. For scanning electron microscopy, bacteria were fixed in 2.5% (vol/vol) glutaraldehyde in 0.1 M cacodylate buffer (pH 7.2) by incubation for 1 h at room temperature. They were then washed three times for 5 min each time in 0.2 M cacodylate buffer (pH 7.2), postfixed by incubation for 1 h in 1% (wt/vol) osmium tetroxide in 0.2 M cacodylate buffer (pH 7.2), and then rinsed with distilled water. Bacteria were dehydrated by incubation in a graded series of 25, 50, 75, and 95% ethanol solutions for 5 min for each solution. Samples were then dehydrated by incubation for 10 min in 100% ethanol, followed by critical point drying with CO2. Dried specimens were sputter coated with 15 nm of gold-palladium, using a GATAN ion beam coater. Cells were then examined with a JEOL JSM 6700F field emission scanning electron microscope operating at 5 kV, using secondary electron detectors. For transmission electron microscopy, agar pieces containing bacteria were cut, fixed, treated with ruthenium red (25), and embedded in SPURR resin. We used ruthenium red dye for staining bacterial polysaccharide. We avoided disturbing the fragile extracellular material by fixing and staining the bacteria directly on agar, followed by analysis of thin sections. The fixation and processing steps were carried out in a fume cupboard at room temperature in vials on a rotary mixer. Bacteria were fixed by incubation for 2 h in 3% glutaraldehyde containing 0.75% ruthenium red in 0.1 M sodium cacodylate buffer (pH 7.4). The samples were then washed four times in 0.2 M sucrose-0.075 M sodium cacodylate buffer (pH 7.4) by incubation for 90 min and postfixed with 1% osmium tetroxide-0.75% ruthenium red in cacodylate buffer (pH 7.5). Samples were then washed twice in ultrapure water and three times in 0.1 M cacodylate buffer (pH 7.4). Samples were subsequently dehydrated by incubation in a graded series of 30 to 100% ethanol solutions for 15 min per step, followed by incubation for 90 min in pure ethanol and for 15 min in propylene oxide. The samples were then sequentially transferred into 3:1, 1:1, 1:3 propylene oxide—low-vicosity Agar Resin LV (Agar Scientific, Oxford Instrument, France) by incubation for 1 h per step. They were finally incubated in 100% resin overnight. The samples were incubated in fresh resin three times for 3 h, embedded in silicone flat molds, and polymerized by incubation at 60°C for 2 days. Ultrathin transverse sections were cut using a diamond knife and a Leica Ultracut UCT (Leica Microsystems SA, Rueil Malmaison, France). The grid-mounted sections were stained with uranyl acetate and lead citrate. Electron micrographs were recorded using a Mega view charge-coupled device camera (ELOISE SARL, France) mounted on a JEOL 1200 EX II transmission electron microscope (JEOL EUROPE SA) operating at 80 kV.
Enzymatic assay.
β-Galactosidase activity was measured as described by Miller (29) and was expressed in Miller units.
Overproduction and purification of His6-Crl.
A 4-liter culture of JM109 carrying pQEcrl was grown in LB medium containing 100 μg ml−1 ticarpen at 37°C to an optical density of 0.6, and then 1 mM IPTG was added. The cells were harvested after 4 h, washed, resuspended in 60 ml of buffer A (50 mM Na2HPO4 [pH 8], 300 mM NaCl) supplemented with 1 mM 4-(2-aminoethyl)-benzenesulfonyl fluoride, and then lysed by sonication. The crude cell extract was centrifuged at 15,000 × g for 30 min. The supernatant was added to 10 ml of Ni-nitrilotriacetic acid agarose (QIAGEN) and gently mixed for 1 h. The slurry was packed onto an Econo-Pac column (Bio-Rad) and washed with 250 ml of buffer A containing 20 mM imidazole. His6-Crl was eluted with buffer A containing 250 mM imidazole, dialyzed against buffer B (20 mM Tris-HCl [pH 8], 20 mM NaCl), and loaded onto a Hitrap Q anion-exchange column (5 ml; GE Healthcare). After washing with 10 ml of buffer B, His6-Crl was eluted from the column between 0.4 M and 0.6 M NaCl using a linear 0.02 to 1 M NaCl gradient. We pooled only the first fractions, which were dialyzed against buffer C (10 mM Tris-HCl [pH 8], 10 mM MgCl2, 100 mM KCl, 50% glycerol, 1 mM dithiothreitol [DTT]). This resulted in 14 mg of purified His6-Crl, as determined by the Bradford assay.
Runoff transcription assays.
Single-round transcription assays were performed by using pU-2750 and pJCDadrA digested with the AseI restriction enzyme. The 285-bp lacUV5 fragment was synthesized by PCR using primer E5 (5′-CCTGGCAGTTCCCTACTCT-3′) and primer E7 (5′-CTGGCAGATGCGTCTTCCG-3′) with the pGO1 plasmid (34). Reconstituted RNA polymerase (60 nM) (σ/core ratio, 2.5: 1) was incubated in buffer T (40 mM HEPES [pH 8.0], 10 mM MgCl2, 100 mM potassium glutamate, 2 mM DTT) with different concentrations of Crl (0, 0.5, 2, and 8 μM) for 20 min at 30°C. DNA templates were added at a final concentration of 10 nM, and the mixtures were incubated for 12 min at 30°C. Elongation was started by adding 3 μl of a mixture containing 1 mM ATP, 1 mM GTP, 1 mM CTP, 100 μM UTP, 0.5 μCi [α-32P]UTP, and 600 μg ml−1 heparin to the template polymerase mixture. After 10 min, the reactions were stopped by adding formamide containing 10 mM EDTA, xylene cyanol, bromophenol blue, and 1% SDS. After heating to 90°C, the samples were subjected to electrophoresis on a 7.5% polyacrylamide sequencing gel, and the transcripts were quantified using a PhosphorImager (Molecular Dynamics). The transcript sizes were approximately 191 nt for the csgD promoter, 158 nt for the adrA promoter, and 128 nt for the lacUV5 promoter.
Abortive initiation assays.
We used the 172-bp adrA fragment described above and the 191-bp csgD fragment (synthesized by PCR using primers 5′-CTGTGGGTTGAAATAGCCCATTATCC-3′ and 5′-CCCGGATCCCAACTTTACTATCAAATCTAAAC-3′) as templates with the dinucleotide initiators ApA for adrA and ApG for csgD to synthesize ApApUpU and ApGpUpU, respectively. ApApUpU is complementary to the adrA template strand located between nt 8 and 11 downstream from the last T of the Pribnow box. This transcription start site maps close to that identified for the homologous yaiC gene in E. coli (9). A mixture of DNA template (10 nM), dinucleotide (1 mM), and 60 nM [α-32P]UTP was added to an equal volume of the holoenzyme (60 nM) preincubated with buffer or Crl at various concentrations. In the case of the lacUV5 fragment the UTP concentration was 50 μM (27). Aliquots were removed at various times. The aliquots were spotted onto Whatmann 3MM paper prespotted with 100 mM EDTA, and chromatograms were developed as previously described (27).
KMnO4 reactivity experiments.
We generated the 172-bp adrA and 191-bp csgD fragments by PCR using a combination of an unlabeled primer and a primer end labeled with T4 polynucleotide kinase and [γ-32P]ATP. Open complexes were allowed to form in buffer T without DTT by incubation at 30°C for 20 min and were incubated with KMnO4 (3 mM) for 15 s. The reaction was stopped by adding 3 μl of DTT (200 mM) and 200 μl of a solution containing 0.4 M sodium acetate (pH 8.0), 2.5 mM EDTA, and 60 μg ml−1 of salmon sperm DNA. After phenol extraction and precipitation with ethanol, the samples were resuspended in 100 μl of piperidine (1 M), heated at 90°C for 30 min, evaporated to dryness, and evaporated twice with 20 μl water. The samples were resuspended in formamide containing 20 mM EDTA, bromophenol, and xylene cyanol blue and loaded onto an 8% denaturing polyacrylamide gel. The gels were dried and analyzed using a PhosphoImager.
RESULTS
rdar morphotype development is affected by a crl knockout mutation.
We determined the role of crl in the rdar morphotype of Salmonella by plating ATCC 14028 and Δcrl and ΔrpoS mutant derivatives of this strain on LB0 plates containing either CR Fig. 1A) or calcofluor (Fig. 1B) and incubating the plates at 28°C. ATCC 14028 developed a typical rdar morphotype and bound CR and calcofluor (Fig. 1A and B). Curli and cellulose production requires RpoS. Therefore, the rpoS mutant did not bind the dyes and displayed a saw (smooth and white) morphotype (42, 46, 52, 58) (Fig. 1A and B). The crl mutant developed a rough and dry morphotype and bound both CR and calcofluor (Fig. 1A and B). However, the crl colonies were organized differently than the wild type colonies (rdarcrl morphotype). Introducing plasmid pACcrl-1 expressing the Crl protein into the crl mutant restored the typical rdar morphotype (Fig. 2), whereas we observed no complementation with the negative controls pACYC184 and pACcrl-2, in which the crl gene was cloned in the promoterless orientation (Fig. 2). Together, these results showed that expression of a typical rdar morphotype in Salmonella serovar Typhimurium required Crl.
FIG. 2.
rdar morphotypes of Salmonella strains harboring different plasmids, including ATCC 14028 and Δcrl and ΔrpoS mutant derivatives of this strain that harbor the vector pACYC184, the crl gene on pACcrl-1 and pACcrl-2, and the rpoS gene on pSTK4 (Table 1), after 5 days of growth on CR plates at 28°C. In pACcrl-2 the crl gene was cloned in the promoterless orientation. Complementation of the crl mutation was observed with pACcrl-1 and pSTK4. Complementation of the rpoS mutation was observed with pSTK4.
crl mutant still produces curli and cellulose, but the architecture of the colonies is affected.
We determined whether curli and cellulose contribute to the specific morphotype (rdarcrl) of the crl mutant by introducing mutations into the biosynthetic curli (csgB) and cellulose (bcsA) genes in the crl mutant. We examined the morphotypes of the double mutants after 5 days of growth at 28°C on LB0 plates containing CR and calcofluor. The bcsA mutation eliminated calcofluor binding by the crl strain (Fig. 1B). Also, on CR plates, the crl csgB and crl bcsA strains had different morphotypes than the crl strain (Fig. 1A). These results showed that curli and cellulose production contributed to the rdarcrl morphotype of the crl strain.
We investigated the architecture of the colonies of the wild-type and crl strains after 5 days of growth on LB0 at 28°C by microscopic analysis (Fig. 3). Scanning electron microscopy of colonies formed by the wild-type and crl strains revealed a bacterial pellicle that was several cell layers thick in both types of colonies (Fig. 3A, panels a to d). However, the crl colony architecture was different from the wild-type colony architecture. There were extensive folds of the bacterial crl pellicle (Fig. 3A, compare panels a and b). We observed cell-cell interactions and production of an extracellular matrix in both the wild-type and crl strains (Fig. 3A, panels e and f).
FIG. 3.
Electron micrographs of colonies of Salmonella strains. (A) Scanning electron micrographs of wild-type strain ATCC 14028 (panels a, c, and e) and ATCCcrl (b, d, and f) after 5 days of growth on CR plates at 28°C. Panels a and b show one-quarter of a colony (upper surface). Panels c and d show cross fractures of colonies, revealing the inner structure. The arrows indicate the surface (S) and the inner part (I) of the colony. In panel c, bacteria were extruded from the cross fracture, revealing the inner compartmentalization with a highly organized matrix having a honeycomb-like structure. Panels e and f show the upper, air-facing surfaces of colonies, revealing the extracellular matrix and cell-cell interactions. The magnifications are ×25 for panels a and b, ×2,500 for panels c and d, and ×45,000 for panels e and f. (B) Representative transmission electron micrographs of ruthenium red-stained transverse ultrathin sections of wild-type (panels a, c, and e) and crl (panels b, d, and f) colonies of Salmonella after 5 days of growth on LB0 at 28°C. The electron micrographs show the presence of closely packed bacteria and a stained matrix between cells. However, the biofilm formed by the wild-type strain was homogeneously stained, whereas the biofilm formed by the crl strain has a less well-organized structure, and areas have either an extensive extracellular matrix (panel d) or very little matrix (panel f). The magnifications are ×7,500 for panels a and b, ×25,000 for panel c, and ×75,000 for panels d, e, and f.
The extracellular matrix was analyzed further by transmission electron microscopy. Ultrathin sections were stained with ruthenium red, which preferentially stains polysaccharides, such as cellulose (25, 44, 52). In the wild-type colonies, an abundant ruthenium red-positive matrix between the bacteria was observed, whereas in the crl strain the matrix was less abundant and less well organized (Fig. 3B). In the mutant, the pattern was heterogeneous; there were areas of cells enclosed in a ruthenium red-stained matrix (Fig. 3B, panel d), and there were areas of cells with no matrix (Fig. 3B, panel f). The orientations of the cells themselves were also slightly different for the wild-type strain and the crl mutant. Matrix-enclosed wild-type cells were tightly packed and aligned in a honeycomb-like structure, whereas the crl cells appeared to be more randomly distributed (Fig. 3B, panels a, c, and b). We also observed this difference in the organization of the structure in scanning electron micrographs of cross fractures that showed the inner parts of colonies (Fig. 3A, panels c and d).
crl knockout mutation decreases curli and cellulose gene transcription.
RpoS and CsgD are known to positively regulate the production of the extracellular matrix components cellulose and curli (for reviews see references 16 and 40) (Fig. 4). Curli biogenesis requires two divergently transcribed operons, the csgBAC operon, encoding the structural components of curli, and the csgDEFG operon. Activation of the csgBAC operon and the adrA gene, which encodes a protein that mediates production of cyclic di-(3′,5′)-guanylate, requires CsgD. The genes involved in cellulose biosynthesis are organized into two divergently transcribed operons, bcsABZC and bcsEFG. The allosteric activator cyclic di-(3′,5′)-guanylate binds to the BcsB subunit of the cellulose synthase, stimulating cellulose biosynthesis. Expression of the csg operons and the adrA gene depends on rpoS.
FIG. 4.
Schematic representation of gene regulation controlling curli and cellulose expression in Salmonella. Positive regulation by RpoS, Crl, and CsgD is indicated by arrows. Solid lines indicate a direct role for RpoS and Crl in activating transcription initiation at the adrA and csgD promoters. Dashed lines indicate either a direct role or an indirect role for RpoS and Crl, and the thickness of the line indicates the strength of control. Positive regulation by CsgD is indicated by solid lines because putative CsgD binding sites have been found upstream from the csgB and adrA promoters (9). However, so far there is no evidence for CsgD binding. See text for details.
To determine the role of crl in the expression of σS-regulated genes involved in the rdar morphotype, we constructed chromosomal csgD-lacZ, csgB-lacZ, and adrA-lacZ transcriptional fusions at the loci of the wild-type alleles. Expression of the fusions was monitored during growth of wild-type and crl strains (Fig. 5A). Cells were grown in LB0 at 28°C. This allowed maximal expression of these genes (30, 45, 46). The expression of the csgD, csgB, and adrA genes was substantially reduced in the crl mutant compared to the expression in the wild-type strain (Fig. 5A). As expected, csgD, csgB, and adrA expression was considerably reduced in the rpoS mutant (Fig. 5A and B). The effect of the rpoS mutation on csgB and adrA expression was even more drastic than the effect on csgD expression (Fig. 5B). Expression of csgB and expression of adrA were decreased 10-fold and 20-fold, respectively, in the rpoS mutant and only 4-fold and 6-fold, respectively, in the csgD strain (Fig. 5B). We observed no detectable expression of these genes in the csgD rpoS double mutant (Fig. 5B).
FIG. 5.
Expression of curli and cellulose biosynthesis genes in Salmonella wild-type and mutant strains. (A) Expression of csgD-lacZ, csgB-lacZ, adrA-lacZ, and bcsA-lacZ gene fusions as a function of bacterial cell growth. Wild-type strain ATCC 14028 (wt) and the ATCCcrl, ATCCrpoS, and ATCCcsgD mutant strains harboring fusions were grown in LB0 at 28°C. Exponential-phase cultures (optical density at 600 nm [OD600], 0.5) of Salmonella were diluted in LB0 prewarmed at 28°C to prolong the exponential phase. Aliquots were removed at various times, and β-galactosidase activity was measured (solid symbols) by the method of Miller (29). The growth phase was determined by measuring the culture turbidity at an optical density of 600 nm (open symbols and dashed lines). The experiments were repeated twice, and the results of a representative experiment are shown. (B) Expression of csgB-lacZ and adrA-lacZ gene fusions in Salmonella wild-type and rpoS, csgD, and rpoS csgD mutant strains grown to the stationary phase (18 h of growth) in LB0 at 28°C.
The bcs genes are thought to be constitutively transcribed, and their expression is not dependent on CsgD, AdrA, and RpoS (58). Accordingly, we found that expression of a chromosomal bcsA-lacZ gene fusion was not decreased in the csgD mutant (Fig. 5A). However, the levels of expression of the bcsA-lacZ gene fusion in the rpoS and crl strains were lower than the levels of expression in the wild type after 10 h of growth (Fig. 5A). Nevertheless, the pattern of bcsA gene fusion expression was substantially different from the pattern of expression of the adrA, csgB, and csgD gene fusions, as follows: (i) bcsA was transcribed during the exponential growth phase, although its transcription increased in the stationary phase in a σS-dependent manner; (ii) the rpoS mutation affected bcsA gene expression much less than it affected csgD, csgB, and adrA gene expression, and the effect of the rpoS mutation on bcsA gene expression was maximal in the early stationary phase; and (iii) transcription of bcsA probably decreased during the late stationary growth phase.
Consistent with the typical rdar morphotype developed by the crl strain containing pACcrl-1 (Fig. 2), we observed complementation of the crl mutation by pACcrl-1 for csgD, csgB, adrA, and bcsA gene expression (data not shown). Together, these results demonstrated that Crl plays a role in the transcription activation of the csgD, csgB, adrA, and bcsA genes required for developing a typical rdar morphotype in Salmonella.
Crl protein levels increase in the late exponential and stationary growth phases.
Using Western blotting, we examined expression of the Crl protein in cells grown in LB0 at 28°C (Fig. 6). In wild-type strain ATCC 14028, the Crl protein was synthesized during the early exponential growth phase, but Crl protein levels increased during the late exponential and stationary growth phases. This expression pattern is consistent with previous results showing that Crl increases the expression of genes involved in the rdar morphotype when cells enter the stationary phase. The expression pattern of the Crl protein did not seem to be affected in the rpoS strain under the environmental conditions used (Fig. 6). We determined whether the crl mutation affected rpoS expression in LB0 at 28°C by comparing the σS levels in the crl and wild-type strains (Fig. 6). Higher levels of σS were found in the crl mutant. This is consistent with previous results for E. coli which showed that RpoS production is increased by a crl mutation (8, 35). Thus, a higher level of σS in the crl strain seems to be less active for csgD, csgB, adrA, and bcsA gene expression than a lower level of σS in the wild-type strain.
FIG. 6.
Expression of Crl and σS during growth of Salmonella strains in LB0 at 28°C. ATCC 14028 (▴), ATCCcrl (○), and ATCCrpoS (□) were grown in LB0 at 28°C. Exponential-phase cultures (optical density at 600 nm [OD600], 0.5) of Salmonella were diluted in LB0 prewarmed at 28°C to prolong the exponential phase. Aliquots were removed at various times and analyzed by Western blotting with anti-Crl and anti-σS antibodies. Ten micrograms of total protein was loaded in each slot. The growth phase was determined by measuring the culture turbidity at an optical density of 600 nm.
Expression of rpoS from a plasmid can restore a typical rdar morphotype in the crl strain.
The cloned rpoS gene on pSTK4 was able to complement the rpoS mutant for development of the rdar morphotype (Fig. 2). However, the crl mutant containing pSTK4 was also able to develop a typical rdar morphotype (Fig. 2). Accordingly, pSTK4 was able to complement the crl mutation for csgD, csgB, and adrA expression (Fig. 7A). The levels of σS from pSTK4 were higher than the levels from the chromosomal copy of rpoS (Fig. 7B). This increase in RpoS expression may alleviate the need for Crl for expression of the genes involved in the rdar morphotype. In the late exponential growth phase, the level of csgD expression in the presence of pSTK4 was slightly lower in the crl strain than in the wild-type strain, suggesting that, even in the presence of high levels of σS, Crl may still contribute to σS activity when cells enter the stationary phase (Fig. 7A).
FIG. 7.
Complementation of the crl mutation by the cloned rpoS gene. (A) Expression of csgD-lacZ, adrA-lacZ, and csgB-lacZ in the ATCC 14028 and ATCCcrl strains containing the pACYC184 vector and the cloned rpoS gene on pSTK4. Exponential-phase cultures (optical density at 600 nm [OD600], 0.5) of Salmonella were diluted in LB0 prewarmed at 28°C to prolong the exponential phase. Aliquots were removed at various times, and β-galactosidase activity was measured (solid lines) by the method of Miller (29). The growth phase was determined by measuring the culture turbidity at an optical density of 600 nm (dashed lines). (B) RpoS production in the Salmonella cultures shown in panel A. Aliquots were removed during the exponential phase (LOG) (optical density at 600 nm = 0.4) and during the stationary phase (STA) (optical density at 600 nm = 2.5) of growth and were analyzed by Western blotting with anti-σS antibodies. Ten micrograms of total protein was loaded in each slot. Lanes 1 and 3, ATCCcsgD-lacZ with the pACYC184 vector; lanes 2 and 4, ATCCcsgD-lacZ with rpoS on pSTK4 (Table 1).
Crl protein activates in vitro transcription by EσS from the csgD and adrA promoters.
We constructed pQEcrl, which expressed a recombinant Crl protein containing six histidine residues at its N terminus. In pQEcrl, expression of the six-His-tagged protein is controlled by the IPTG-inducible promoter of pQE30. As expected, after induction with IPTG, high levels of a 17-kDa protein were produced in E. coli JM109 carrying pQEcrl (Fig. 8B, lane 2). We determined whether the His6-Crl protein was still active by introducing pQE30 and pQEcrl into the wild-type and crl strains of Salmonella. These derivative strains were spread on LB0 containing CR and grown for 6 days at 28°C. The pQEcrl plasmid was able to restore a typical rdar morphotype in the crl strain, whereas the pQE30 vector could not do this (Fig. 8A). This indicated that the His6-Crl protein was active. The overexpressed recombinant protein was obtained from JM109 carrying pQEcrl and was purified to homogeneity, as judged by electrophoresis in a denaturing polyacrylamide gel (Fig. 8B) and electrospray mass spectrometry (molecular weight, 17,195.4 ± 1.6). It was then used to produce polyclonal antibodies against Crl and in subsequent in vitro studies.
FIG. 8.
In vivo activity and purification of the recombinant His6-Crl protein. (A) Complementation of the crl mutant by His6-Crl for expression of the rdar morphotype. ATCC 14028 and its Δcrl mutant derivative containing the pQE30 vector and pQEcrl expressing the His6-Crl protein were grown on CR plates for 6 days at 28°C. (B) Coomassie blue staining of 12% SDS-polyacrylamide gel. The overexpressed recombinant His6-Crl protein was purified from JM109 carrying pQEcrl using immobilized metal ion chromatography (lane 1) (see Materials and Methods). The protein was purified further on a Hitrap Q anion-exchange column. Fractions that eluted between 400 and 500 mM NaCl contained highly purified His6-Crl (lanes 4 to 6). Lane 1, purified His6-Crl protein on Ni-nitrilotriacetic acid resin (10 μg); lane 2, IPTG-induced JM109 extract containing pQEcrl (10 μg); lane 3, molecular weight standards; lanes 4 to 6, purified His6-Crl fractions eluted from the Hitrap Q anion-exchange column.
The in vivo studies showed that Crl played a role in the transcriptional activation of the σS-regulated genes csgB, csgD, adrA, and bcsA. We determined whether Crl played a direct role by carrying out in vitro transcription assays with the purified His6-Crl protein. As the promoters for the bcsABZC operon have not been identified yet, we focused on the csgB, csgD, and adrA genes. We cloned the three promoters in front of transcriptional terminators (28). In preliminary experiments with csgB, no transcript of the expected size was observed with either EσS or Eσ70. This was consistent with the results of Bougdour et al. (8), who suggested that an unknown factor that activates csgB expression, possibly the CsgD activator protein, was missing in the in vitro assay. By contrast, we observed significant levels of transcripts of the expected sizes in in vitro single-round runoff assays using the EσS holoenzyme for the csgD promoter and for the adrA promoter (Fig. 9A). Addition of Crl increased the levels of transcripts initiating at the csgD and adrA promoters 1.5- and 2.5-fold, respectively, on linear templates (Fig. 9A, lanes 1 to 5). This showed that the Crl protein plays a direct role in transcription initiation by EσS from the csgD and adrA promoters. We detected lower levels of csgD and adrA transcripts when Eσ70 was used instead of EσS under the same conditions (Fig. 9A, lanes 6 and 7). This showed that σS plays a direct role in the transcription of csgD and adrA. However, Crl did not affect the efficiency of σ70-dependent transcription at the csgD and adrA promoters and at the σ70-dependent lacUV5 promoter (Fig. 9A, lanes 6 to 12). Thus, transcriptional activation by Crl appeared to be specific for EσS.
FIG. 9.
In vitro transcription experiments. (A) Single-round in vitro transcription from the csgD and adrA promoters and the σ70-dependent promoter lacUV5 by EσS (lanes 1 to 5) and Eσ70 (lanes 6 to 12) with or without His6-Crl. The holoenzyme was preincubated in the absence of His6-Crl (lanes 1, 5, 6, 8, and 12) or in the presence of His6-Crl (lanes 2 and 9, 250 nM; lanes 3 and 10, 1 μM; lanes 4, 7, and 11, 4 μM). Open complexes were then allowed to form by incubation at 30°C for 12 min before addition of heparin and XTPs. (B) Effect of His6-Crl in kinetic abortive initiation assays. For csgD and adrA, EσS was preincubated in the absence of His6-Crl (⧫) or in the presence of His6-Crl (▪, 250 nM; ▴, 1 μM). For lacUV5, Eσ70 was preincubated in the absence of His6-Crl (⧫) or in the presence of His6-Crl (▪, 250 nM; ▴, 1 μM). After addition of a mixture containing the promoter fragment, the dinucleotide initiator, and [α-32P]UTP, the incorporation of labeled [α-32P]UMP into small abortive transcripts was monitored as a function of time. ApG was used to initiate the synthesis of ApGpUpU at the csgD promoter. ApA was used to initiate the synthesis of ApApUpU at the adrA and lacUV5 promoters.
Crl facilitates open complex formation by EσS at the csgD and adrA promoters.
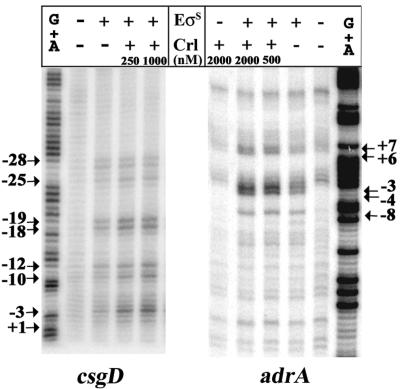
We showed that Crl had a direct effect on open complex formation by performing kinetic abortive initiation experiments (Fig. 9B). This method does not require preincubation of a promoter and RNA polymerase and can directly and accurately monitor the rate of open complex formation. The csgD transcript starts with the sequence pppGpUpUpA, with an A at position −1 on the nontemplate strand (42). Therefore, we used the dinucleotide ApG and [α-32P]UTP to obtain reiterative synthesis of ApG*pU*pU. The reaction was initiated by adding EσS RNA polymerase to a mixture containing ApG, labeled UTP, and the csgD fragment, and we monitored the rate of tetranucleotide formation as a function of time (Fig. 9B). After a short lag time, the synthesis was linear up to 30 min. Crl at concentrations of 250 nM and 1 μM increased the amount of tetranucleotide synthesized 1.7-fold and 2-fold, respectively. We obtained similar results with the adrA promoter using ApA and [α-32P]UTP to synthesize ApA*pU*pU from the template sequence (Fig. 9B). However, the Crl stimulation appeared to be less efficient than that observed in the runoff assays (Fig. 9A and B). We verified that the tetranucleotide synthesis was initiated from the promoters being studied by forming and probing open complexes for single-stranded thymines using KMnO4 (Fig. 10). At the adrA promoter, EσS formed a transcription bubble extending from position −8 to position 7 according to the transcription start determined from the homologous E. coli yaiC promoter (9). At the csgD promoter, the pattern of the single-stranded thymines appeared to be more extended, but the most reactive base mapped at position −3, in agreement with the location of the transcription start site of the Salmonella csgD promoter (42). At each promoter, the reactivity of the transcription bubble thymines was clearly more intense in the presence of Crl, indicating that Crl had increased the amount of open complexes. Together, these results showed that Crl enhanced both the quantity of open complexes and the rate of formation of open complexes by EσS from the csgD and adrA promoters. In contrast, Crl did not affect the rate of formation of open complexes by Eσ70 from the σ70-dependent promoter lacUV5 (Fig. 9B).
FIG. 10.
KMnO4 reactivity patterns of the csgD and adrA open complexes formed with and without His6-Crl. After preincubation of EσS with or without His6-Crl at different concentrations, open complexes were allowed to form by incubation for 20 min at 30°C before the preparations were subjected to KMnO4 attack. The positions of the reactive thymines were calibrated according to the G+A sequencing reaction and are indicated on the left for csgD (template strand) and on the right for adrA (nontemplate strand). The transcription start site of the Salmonella csgDEFG operon was determined previously (42), whereas the adrA start site was only tentatively assigned based on the homologous E. coli yaiC gene (9). Note that despite the conservation of the adrA and yaiC promoter sequences, the adrA start site might be shifted by a few nucleotides downstream from the start site of yaiC. Similar variation in the selection of start sites for Salmonella and E. coli genes was detected at csgD (42).
DISCUSSION
We wanted to determine the physiological impact and the molecular function of Crl, a protein shown to bind σS and to enhance its activity. However, until now, there was no obvious phenotype associated with the crl mutant, although the protein has been identified as a regulator of curli biosynthesis in E. coli. It is noteworthy that the requirement for Crl seemed to vary from strain to strain (32, 37, 38). Curli production resulting in multicellular and aggregative behavior of the cells has been thoroughly investigated in different S. enterica serovars. At 28°C more than 90% of Salmonella serovar Typhimurium and Enteritidis strains develop the rdar morphotype, which is characterized by the formation of an extracellular matrix composed mainly of curli and cellulose (43). Single-cell behavior results from knockout of these two extracellular matrix components. Therefore, we decided to explore the effect of a crl null mutation on the rdar morphotype of Salmonella serovar Typhimurium.
The crl knockout mutant of Salmonella serovar Typhimurium strain ATCC 14028 was able to produce some curli and cellulose, but it developed an atypical rdar morphotype (rdarcrl) (Fig. 1). Scanning electron microscopy and transmission electron microscopy of wild-type colonies expressing the rdar morphotype and crl colonies expressing the rdarcrl morphotype revealed different arrangements of the crl and wild-type cells in the extracellular matrix (Fig. 3). We also observed substantial differences in the quantity and organization of the extracellular matrix in thin sections stained with ruthenium red. Complementation of the crl mutation could be achieved with the cloned crl gene in pACcrl-1, confirming that Crl has a role in the development of the rdar morphotype (Fig. 2).
At the molecular level, the crl mutation decreased expression of lacZ gene fusions in the curli and the cellulose biosynthesis genes csgD, csgB, adrA, and bcsA (Fig. 5). CsgD depends on RpoS and Crl and controls the expression of csgBAC and adrA but not the expression of bscA. This indicates that at least two σS-regulated genes implicated in the formation of the rdar morphotype in Salmonella are affected by crl. The crl null allele effects were not as severe as the effects conferred by an rpoS null mutation. The effects caused by the rpoS mutation alone and the rpoS crl double mutation were similar, and we were not able to obtain complementation of the crl mutation with the cloned crl gene in the absence of RpoS (data not shown). Also, expression of Crl from pACcrl-1 did not modify the “saw” morphotype of the rpoS mutant (Fig. 2). Together, these results indicate that Crl function requires RpoS. Since the level of the RpoS protein did not decrease in the crl null strain but rather increased (Fig. 6) (see below), Crl appears to function together with RpoS, as previously suggested (35).
In vitro studies with purified His6-Crl confirmed this conclusion by demonstrating that Crl has a direct role in σS-dependent transcription initiation at the csgD and adrA promoters (Fig. 9 and 10). In these experiments, we found no noticeable effect of the Crl protein on the basal level of σ70-dependent expression of adrA and csgD and on the σ70-dependent expression of lacUV5. This suggests that Crl-mediated activation is specific for σS-dependent transcription. KMnO4 reactivity experiments and abortive initiation assays clearly showed that Crl activates EσS-dependent transcription by enhancing the rate of open complex formation. However, Crl is not a classical transcription factor since it does not bind DNA (35). In contrast, it associates with σS and likely also with the EσS holoenzyme (8), which allows it to form open complexes more efficiently. Several mechanisms could explain Crl activation of σS-dependent promoters. σS has been shown to bind core in vitro with the weakest affinity among the seven sigma factors of Enterobacteriacae (26). Consistent with the model of sigma factor competition for a limited amount of core available in the cell, Crl could facilitate association of σS with the core enzyme. However, according to the study of Bougdour et al. (8) and our preliminary data, Crl does not play a role in the competition between σS and σ70 for core binding. Therefore, Crl should stimulate EσS recruitment to the promoter. Further assays are needed to determine whether Crl facilitates closed complex formation, helps promoter opening, or affects both mechanisms. Also, our experiments do not exclude the possibility that Crl has additional effects at other steps of the transcription process (for example, facilitating promoter escape by the EσS RNA polymerase). The apparently more efficient stimulation of transcription at the adrA promoter in single-round runoff experiments than in abortive initiation assays supports this idea. Like many transcription factors, Crl may affect different steps of transcription initiation at different promoters in a nonexclusive manner.
However, both in the absence and in the presence of the Crl protein we were unable to obtain noticeable in vitro transcription from the csgB promoter. This result, which is consistent with the results of Bougdour et al. (8), may have been due to the lack of the CsgD protein in the purified system. Indeed, it has been suggested, although it has never been shown, that CsgD binds to conserved 11-bp sequences upstream of the csgB and adrA promoters, activating transcription of these genes (9, 16). Unlike the csgB promoter, the adrA promoter does not necessarily require CsgD since in vitro transcription of adrA was successfully obtained in the absence of CsgD (Fig. 9). Until now, it was not known whether the role of RpoS at adrA and csgBAC is uniquely mediated through csgD or whether RpoS also plays a direct role in transcription initiation at these promoters. The rpoS mutation affected adrA and csgB expression more than a mutation in csgD affected this expression (Fig. 5B). This suggests that, in addition to positively regulating csgD expression, rpoS is also required at some other steps in adrA and csgB gene expression regulation. The results of our in vitro experiments support the hypothesis that σS has a direct role, at least at the adrA promoter. However, this does not exclude the possibility that Eσ70 has an additional role in transcription initiation at the adrA and csgB promoters, and we were able to detect in vitro transcripts by Eσ70 from adrA and also from csgD (Fig. 9). Experiments with purified CsgD are needed to determine whether CsgD can activate transcription by EσS and/or Eσ70 at the adrA and csgB promoters.
In agreement with the study of Zogaj et al. (58), we found that a csgD null mutation did not decrease the expression of the cellulose biosynthetic gene bcsA (Fig. 5A). However, the rpoS mutation affected the transcription of a bcsA-lacZ gene fusion (Fig. 5A). This indicates that at least two genes involved in cellulose biosynthesis are affected by RpoS, the CsgD-dependent gene adrA and the bcsA gene, whose expression is independent of CsgD. In agreement with this, we found that adrA on a plasmid was able to restore calcofluor binding to the adrA and csgD mutant strains but not to the rpoS and bcsA strains (data not shown). We did not expect regulation of bcsA by RpoS, as Zogaj et al. (58) found that a bcsA-lacZ gene fusion (we used the same gene fusion [Table 1]) was indeed partially induced in the stationary phase, but in an RpoS-independent manner. So far, we do not know the reason for this discrepancy. The strain used by Zogaj et al. (58) was a Nalr derivative of ATCC 14028. It is possible that the Nalr mutation interferes with expression of the bcs genes. Further experiments are needed to investigate the regulatory cues of the cellulose biosynthetic genes and their dependence on σS.
The interplay between Salmonella Crl and σS level and activity appears to be intricate. On the one hand, Crl binds to σS, directly increasing its efficiency in transcription initiation both in vivo and in vitro, and as expected, a crl null mutation can be complemented by an increase in RpoS expression (Fig. 2 and 7). On the other hand, the absence of a functional crl gene results in accumulation of increased levels of σS in Salmonella (Fig. 6), as previously shown for E. coli (8, 35). How can these two observations be reconciled?
The former result is fully consistent with the hypothesis that Crl promotes the recruitment of EσS to the σS-dependent promoters, as we would expect an increase in RpoS levels to alleviate the need for Crl. It also seems to be in agreement with the previous finding for E. coli that the requirement of Crl for curli production is different in different strains (31, 32, 37, 38). This difference might be due to σS levels and activities that vary from one strain to another (22, 31, 35). It is likely that the requirement for Crl is more acute in cells expressing low levels of σS. The curli-deficient phenotype of strain HB101 is suppressed by either crl (32) or rpoS (31) on a plasmid. When present in HB101, crl may increase an otherwise low level of RpoS activity, and the increase would then be sufficient for curli production (35).
Altogether, our observations suggest that there is feedback control for Crl-dependent regulation of σS activity. For example, when Crl is missing and σS activity decreases, an increase in σS levels due to a lack of Crl may have a compensatory role. This phenomenon is reminiscent of the autoregulatory loop for the intracellular concentrations of σS and its anti-sigma factor, RssB, in E coli (for a review see reference 19). Phosphorylated RssB that binds σS negatively controls σS levels through ClpXP degradation. This in turn decreases rssB transcription from its σS-dependent promoter, leading to a decrease in the RssB level and consequently to an increase in the σS level. It remains to be determined whether Crl is involved in this autoregulatory control by directly stimulating rssB transcription.
Acknowledgments
We thank Ute Römling for providing bacterial strains and Ulrich Gerstel for constructing plasmid pU-2750.
This work was supported by research funds from the Pasteur Institut and CNRS.
REFERENCES
- 1.Arnqvist, A., A. Olsen, J. Pfeifer, D. G. Russell, and S. Normark. 1992. The Crl protein activates cryptic genes for curli formation and fibronectin binding in Escherichia coli HB101. Mol. Microbiol. 6:2443-2452. [DOI] [PubMed] [Google Scholar]
- 2.Austin, J. W., G. Sanders, W. W. Kay, and S. K. Collinson. 1998. Thin aggregative fimbriae enhance Salmonella enteritidis biofilm formation. FEMS Microbiol. Lett. 162:295-301. [DOI] [PubMed] [Google Scholar]
- 3.Barak, J. D., L. Gorski, P. Naraghi-Arani, and A. O. Charkowski. 2005. Salmonella enterica virulence genes are required for bacterial attachment to plant tissue. Appl. Environ. Microbiol. 71:5685-5691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ben Nasr, A., A. Olsen, U. Sjöbring, W. Müller-Esterl, and L. Björck. 1996. Assembly of human contact phase proteins and release of bradykinin at the surface of curli-expressing Escherichia coli. Mol. Microbiol. 20:927-935. [DOI] [PubMed] [Google Scholar]
- 5.Bian, Z., A. Brauner, Y. Li, and S. Normark. 2000. Expression of and cytokine activation by Escherichia coli curli fibers in human sepsis. J. Infect. Dis. 181:602-612. [DOI] [PubMed] [Google Scholar]
- 6.Bian, Z., Z. Q. Yan, G. K. Hansson, P. Thoren, and S. Normark. 2001. Activation of inducible nitric oxide synthase/nitric oxide by curli fibers leads to a fall in blood pressure during systemic Escherichia coli infection in mice. J. Infect. Dis. 183:612-619. [DOI] [PubMed] [Google Scholar]
- 7.Bokranz, W., X. Wang, H. Tschape, and U. Romling. 2005. Expression of cellulose and curli fimbriae by Escherichia coli isolated from the gastrointestinal tract. J. Med. Microbiol. 54:1171-1182. [DOI] [PubMed] [Google Scholar]
- 8.Bougdour, A., C. Lelong, and J. Geiselmann. 2004. Crl, a low temperature-induced protein in Escherichia coli that binds directly to the stationary phase sigma subunit of RNA polymerase. J. Biol. Chem. 279:19540-19550. [DOI] [PubMed] [Google Scholar]
- 9.Brombacher, E., C. Dorel, A. J. Zehnder, and P. Landini. 2003. The curli biosynthesis regulator CsgD co-ordinates the expression of both positive and negative determinants for biofilm formation in Escherichia coli. Microbiology 149:2847-2857. [DOI] [PubMed] [Google Scholar]
- 10.Chang, A. C., and S. N. Cohen. 1978. Construction and characterization of amplifiable multicopy DNA cloning vehicles derived from the P15A cryptic miniplasmid. J. Bacteriol. 134:1141-1156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Collinson, S. K., S. C. Clouthier, J. L. Doran, P. A. Banser, and W. W. Kay. 1996. Salmonella enteritidis agfBAC operon encoding thin, aggregative fimbriae. J. Bacteriol. 178:662-667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Coynault, C., V. Robbe-Saule, and F. Norel. 1996. Virulence and vaccine potential of Salmonella typhimurium mutants deficient in the expression of the RpoS (sigma S) regulon. Mol. Microbiol. 22:149-160. [DOI] [PubMed] [Google Scholar]
- 13.Datsenko, K. A., and B. L. Wanner. 2000. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97:6640-6645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Dibb-Fuller, M. P., E. Allen-Vercoe, C. J. Thorns, and M. J. Woodward. 1999. Fimbriae- and flagella-mediated association with and invasion of cultured epithelial cells by Salmonella enteritidis. Microbiology 145:1023-1031. [DOI] [PubMed] [Google Scholar]
- 15.Ellermeier, C. D., A. Janakiraman, and J. M. Slauch. 2002. Construction of targeted single copy lac fusions using lambda Red and FLP-mediated site-specific recombination in bacteria. Gene 290:153-161. [DOI] [PubMed] [Google Scholar]
- 16.Gerstel, U., and U. Römling. 2003. The csgD promoter, a control unit for biofilm formation in Salmonella typhimurium. Res. Microbiol. 154:659-667. [DOI] [PubMed] [Google Scholar]
- 17.Gophna, U., M. Barlev, R. Seijffers, T. A. Oelschlager, J. Hacker, and E. Z. Ron. 2001. Curli fibers mediate internalization of Escherichia coli by eukaryotic cells. Infect. Immun. 69:2659-2665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hammar, M., A. Arnqvist, Z. Bian, A. Olsen, and S. Normark. 1995. Expression of two csg operons is required for production of fibronectin- and Congo red-binding curli polymers in Escherichia coli K-12. Mol. Microbiol. 18:661-670. [DOI] [PubMed] [Google Scholar]
- 19.Hengge-Aronis, R. 2002. Signal transduction and regulatory mechanisms involved in control of the sigma(S) (RpoS) subunit of RNA polymerase. Microbiol. Mol. Biol. Rev. 66:373-395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Herwald, H., M. Mörgelin, A. Olsen, M. Rhen, B. Dahlbäck, W. Müller-Esterl, and L. Björck. 1998. Activation of the contact-phase system on bacterial surfaces—a clue to serious complications in infectious diseases. Nat. Med. 4:298-302. [DOI] [PubMed] [Google Scholar]
- 21.Ishihama, A. 2000. Functional modulation of Escherichia coli RNA polymerase. Annu. Rev. Microbiol. 54:499-518. [DOI] [PubMed] [Google Scholar]
- 22.King, T., A. Ishihama, A. Kori, and T. Ferenci. 2004. A regulatory trade-off as a source of strain variation in the species Escherichia coli. J. Bacteriol. 186:5614-5620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kowarz, L., C. Coynault, V. Robbe-Saule, and F. Norel. 1994. The Salmonella typhimurium katF (rpoS) gene: cloning, nucleotide sequence, and regulation of spvR and spvABCD virulence plasmid genes. J. Bacteriol. 176:6852-6860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.La Ragione, R. M., W. A. Cooley, and M. J. Woodward. 2000. The role of fimbriae and flagella in the adherence of avian strains of Escherichia coli O78:K80 to tissue culture cells and tracheal and gut explants. J. Med. Microbiol. 49:327-338. [DOI] [PubMed] [Google Scholar]
- 25.Luft, J. H. 1971. Ruthenium red and violet. I. Chemistry, purification, methods of use for electron microscopy and mechanism of action. Anat. Rec. 171:347-368. [DOI] [PubMed] [Google Scholar]
- 26.Maeda, H., N. Fujita, and A. Ishihama. 2000. Competition among seven Escherichia coli sigma subunits: relative binding affinities to the core RNA polymerase. Nucleic Acids Res. 28:3497-3503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Malan, T. P., A. Kolb, H. Buc, and W. R. McClure. 1984. Mechanism of CRP-cAMP activation of lac operon transcription initiation activation of the P1 promoter. J. Mol. Biol. 180:881-909. [DOI] [PubMed] [Google Scholar]
- 28.Marschall, C., V. Labrousse, M. Kreimer, D. Weichart, A. Kolb, and R. Hengge-Aronis. 1998. Molecular analysis of the regulation of csiD, a carbon starvation-inducible gene in Escherichia coli that is exclusively dependent on sigmaS and requires activation by cAMP-CRP. J. Mol. Biol. 276:339-353. [DOI] [PubMed] [Google Scholar]
- 29.Miller, J. H. 1972. Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
- 30.Olsen, A., A. Arnqvist, M. Hammar, and S. Normark. 1994. Environmental regulation of curli production in Escherichia coli. Infect. Agents Dis. 2:272-274. [PubMed] [Google Scholar]
- 31.Olsen, A., A. Arnqvist, M. Hammar, S. Sukupolvi, and S. Normark. 1993. The RpoS sigma factor relieves H-NS-mediated transcriptional repression of csgA, the subunit gene of fibronectin-binding curli in Escherichia coli. Mol. Microbiol. 7:523-536. [DOI] [PubMed] [Google Scholar]
- 32.Olsen, A., A. Jonsson, and S. Normark. 1989. Fibronectin binding mediated by a novel class of surface organelles on Escherichia coli. Nature 338:652-655. [DOI] [PubMed] [Google Scholar]
- 33.Olsen, A., M. J. Wick, M. Mörgelin, and L. Björck. 1998. Curli, fibrous surface proteins of Escherichia coli, interact with major histocompatibility complex class I molecules. Infect. Immun. 66:944-949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Orsini, G., A. Kolb, and H. Buc. 2001. The Escherichia coli RNA polymerase anti-sigma 70 AsiA complex utilizes alpha-carboxyl-terminal domain upstream promoter contacts to transcribe from a −10/−35 promoter. J. Biol. Chem. 276:19812-19819. [DOI] [PubMed] [Google Scholar]
- 35.Pratt, L. A., and T. J. Silhavy. 1998. Crl stimulates RpoS activity during stationary phase. Mol. Microbiol. 29:1225-1236. [DOI] [PubMed] [Google Scholar]
- 36.Prigent-Combaret, C., G. Prensier, T. T. Le Thi, O. Vidal, P. Lejeune, and C. Dorel. 2000. Developmental pathway for biofilm formation in curli-producing Escherichia coli strains: role of flagella, curli and colanic acid. Environ. Microbiol. 2:450-464. [DOI] [PubMed] [Google Scholar]
- 37.Provence, D. L., and R. Curtiss III. 1994. Isolation and characterization of a gene involved in hemagglutination by an avian pathogenic Escherichia coli strain. Infect. Immun. 62:1369-1380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Provence, D. L., and R. Curtiss III. 1992. Role of crl in avian pathogenic Escherichia coli: a knockout mutation of crl does not affect hemagglutination activity, fibronectin binding, or curli production. Infect. Immun. 60:4460-4467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Robbe-Saule, V., F. Schaeffer, L. Kowarz, and F. Norel. 1997. Relationships between H-NS, sigma S, SpvR and growth phase in the control of spvR, the regulatory gene of the Salmonella plasmid virulence operon. Mol. Gen. Genet. 256:333-347. [DOI] [PubMed] [Google Scholar]
- 40.Römling, U. 2005. Characterization of the rdar morphotype, a multicellular behaviour in Enterobacteriaceae. Cell. Mol. Life Sci. 62:1234-1246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Römling, U. 2001. Genetic and phenotypic analysis of multicellular behavior in Salmonella typhimurium. Methods Enzymol. 336:48-59. [DOI] [PubMed] [Google Scholar]
- 42.Römling, U., Z. Bian, M. Hammar, W. D. Sierralta, and S. Normark. 1998. Curli fibers are highly conserved between Salmonella typhimurium and Escherichia coli with respect to operon structure and regulation. J. Bacteriol. 180:722-731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Römling, U., W. Bokranz, W. Rabsch, X. Zogaj, M. Nimtz, and H. Tschäpe. 2003. Occurrence and regulation of the multicellular morphotype in Salmonella serovars important in human disease. Int. J. Med. Microbiol. 293:273-285. [DOI] [PubMed] [Google Scholar]
- 44.Römling, U., and M. Rohde. 1999. Flagella modulate the multicellular behavior of Salmonella typhimurium on the community level. FEMS Microbiol. Lett. 180:91-102. [DOI] [PubMed] [Google Scholar]
- 45.Romling, U., M. Rohde, A. Olsen, S. Normark, and J. Reinköster. 2000. AgfD, the checkpoint of multicellular and aggregative behaviour in Salmonella typhimurium regulates at least two independent pathways. Mol. Microbiol. 36:10-23. [DOI] [PubMed] [Google Scholar]
- 46.Römling, U., W. D. Sierralta, K. Eriksson, and S. Normark. 1998. Multicellular and aggregative behaviour of Salmonella typhimurium strains is controlled by mutations in the agfD promoter. Mol. Microbiol. 28:249-264. [DOI] [PubMed] [Google Scholar]
- 47.Ryu, J. H., and L. R. Beuchat. 2005. Biofilm formation by Escherichia coli O157:H7 on stainless steel: effect of exopolysaccharide and curli production on its resistance to chlorine. Appl. Environ. Microbiol. 71:247-254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
- 49.Schmieger, H. 1972. Phage P22-mutants with increased or decreased transduction abilities. Mol. Gen. Genet. 119:75-88. [DOI] [PubMed] [Google Scholar]
- 50.Silhavy, T. J., M. L. Bernman, and L. W. Ernqvist. 1984. Experiments with gene fusions. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
- 51.Sjöbring, U., G. Pohl, and A. Olsen. 1994. Plasminogen, absorbed by Escherichia coli expressing curli or by Salmonella enteritidis expressing thin aggregative fimbriae, can be activated by simultaneously captured tissue-type plasminogen activator (t-PA). Mol. Microbiol. 14:443-452. [DOI] [PubMed] [Google Scholar]
- 52.Solano, C., B. Garcia, J. Valle, C. Berasain, J. M. Ghigo, C. Gamazo, and I. Lasa. 2002. Genetic analysis of Salmonella enteritidis biofilm formation: critical role of cellulose. Mol. Microbiol. 43:793-808. [DOI] [PubMed] [Google Scholar]
- 53.Sternberg, N. L., and R. Maurer. 1991. Bacteriophage-mediated generalized transduction in Escherichia coli and Salmonella typhimurium. Methods Enzymol. 204:18-43. [DOI] [PubMed] [Google Scholar]
- 54.Sukupolvi, S., R. G. Lorenz, J. I. Gordon, Z. Bian, J. D. Pfeifer, S. J. Normark, and M. Rhen. 1997. Expression of thin aggregative fimbriae promotes interaction of Salmonella typhimurium SR-11 with mouse small intestinal epithelial cells. Infect. Immun. 65:5320-5325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Tükel, C., M. Raffatellu, A. D. Humphries, R. P. Wilson, H. L. Andrews-Polymenis, T. Gull, J. F. Figueiredo, M. H. Wong, K. S. Michelsen, M. Akcelik, L. G. Adams, and A. J. Bäumler. 2005. CsgA is a pathogen-associated molecular pattern of Salmonella enterica serotype Typhimurium that is recognized by Toll-like receptor 2. Mol. Microbiol. 58:289-304. [DOI] [PubMed] [Google Scholar]
- 56.Van der Velden, A. W., A. J. Baumler, R. M. Tsolis, and F. Heffron. 1998. Multiple fimbrial adhesins are required for full virulence of Salmonella typhimurium in mice. Infect. Immun. 66:2803-2808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Woodward, M. J., M. Sojka, K. A. Sprigings, and T. J. Humphrey. 2000. The role of SEF14 and SEF17 fimbriae in the adherence of Salmonella enterica serotype Enteritidis to inanimate surfaces. J. Med. Microbiol. 49:481-487. [DOI] [PubMed] [Google Scholar]
- 58.Zogaj, X., M. Nimtz, M. Rohde, W. Bokranz, and U. Römling. 2001. The multicellular morphotypes of Salmonella typhimurium and Escherichia coli produce cellulose as the second component of the extracellular matrix. Mol. Microbiol. 39:1452-1463. [DOI] [PubMed] [Google Scholar]