Abstract
The expression of genes involved in the pathogenesis of Staphylococcus aureus is known to be controlled by global regulatory loci, including agr, sarA, saeRS, arlRS, and sarA-like genes. As part of our continuing efforts to understand the regulatory mechanisms that involve sarA-like genes, we describe here the characterization of a novel transcriptional regulator called SarX, a member of the SarA protein family. The transcription of sarX was growth phase dependent and was expressed maximally during the stationary phase of growth, which was significantly decreased in the mgrA mutant. MgrA acted as an activator of sarX expression as confirmed by transcriptional fusion and Northern blot analyses. Purified MgrA protein bound to the upstream region of the sarX promoter as demonstrated by gel shift assay. The expression levels of various potential target genes involved in virulence and regulation, specifically those affected by sarA and mgrA, were analyzed with isogenic sarX mutant strains. Our data indicated that SarX acted as a repressor of the agr locus and consequently target genes regulated by the agr system. We propose that SarX is an important regulator in the SarA protein family and may be part of the common pathway by which agr and members of the sarA gene family control virulence in S. aureus.
Staphylococcus aureus is an opportunistic pathogen that causes a broad range of human infections. In both community and hospital settings, S. aureus can cause food poisoning, pneumonia, meningitis, skin infections (e.g., acne, boils, or cellulitis), arthritis, osteomyelitis, endocarditis, and toxic shock syndrome (2, 6, 35). S. aureus is a major cause of hospital-acquired (nosocomial) infections of surgical wounds and infections associated with indwelling medical devices. Despite antimicrobial therapy, the morbidity and mortality associated with S. aureus infections remain high, due, in part, to the organism's ability to develop resistance to virtually all antibiotics, including vancomycin (2, 35). Because of the lack of an effective vaccine and the emergence of multiple antibiotic-resistant strains, S. aureus continues to be a serious public concern.
The primary site of infection for S. aureus is generally the skin or a wound, from which the organism can spread to the bloodstream, proceeding to invade various host tissues. Within the tissue, S. aureus produces a large number of virulence factors, which include surface-associated proteins, enzymes, exotoxins, capsular polysaccharides, and gene products that facilitate tissue colonization, tissue destruction, and immune evasion. The expression of many of these virulence genes is coordinately controlled by regulatory loci, such as two-component regulatory systems (agr, saeRS, srrAB, and arlRS) and global transcriptional regulators (sarA, sigB, sarA paralogs, tcaRA, etc.) (2, 7, 8, 12, 48, 50).
The sarA and sarA paralogs in S. aureus are crucial for the regulation of various genes involved in virulence and the regulation process (2, 8, 12, 42, 48). The sarA locus comprises a major open reading frame, sarA, driven by three distinct promoters, resulting in three overlapping transcripts with a common terminating end (4, 10, 14, 39). This locus up-modulates the expression of selected cell wall proteins and exoproteins (e.g., α- and β-hemolysins). DNA binding studies have revealed that SarA, the major sarA effector molecule, binds to several target gene promoters, including those for regulatory systems (e.g., agr, sarS, and sarV) (13, 15, 41, 51) and virulence genes (e.g., hla, spa, cna, bap, ica, and fnbA), to modulate gene transcription (5, 16, 42, 58, 59), thus implicating both direct and indirect pathways for SarA-mediated regulation. A search of the published S. aureus genomes revealed at least 12 SarA paralogs, which include SarR, SarS (also called SarH1), SarT, Rot, SarU, SarV, and MgrA (also called Rat or NorR) (11, 25, 29, 36-43, 53, 55, 56, 60). Member of the SarA family of proteins share homology (20 to 45% identity and 45 to 65% similarity to SarA protein) with each other as well as with the MarR family of proteins involved in regulating multidrug resistance in gram-negative bacteria (1, 12). Though only partially characterized, the known attributes of eight sarA-like genes suggest that most of these gene products function as repressors or activators and regulate each other as well as target genes either by binding directly to promoter regions or by acting indirectly via other regulators. Structural analysis suggests that SarA proteins exist as dimers or monomers with two homologous domains and are homologous to the winged-helix family of transcription factors, but with unique variations (31, 33, 34). While many of these genes either activate or repress the expression of virulence and regulatory genes, the exact mode of such regulation is not well defined.
The agr locus is a two-component signal transduction system required for exoprotein synthesis during the post-exponential growth phase and for concomitant repression of cell wall protein synthesis (28, 44, 46, 47, 48, 50). Mutants lacking this switching ability exhibit reduced virulence in several animal model systems (13). The agr locus is composed of two divergent transcriptional units, agr RNAII and agr RNAIII. RNAII consists of four genes (agrDBCA), some of which encode the two-component regulatory system, AgrCA. agrD encodes a 46-residue peptide which is processed into a 7- to 9-residue cyclic peptide (known as autoinducing peptide, or AIP) that plays an important role in regulating the agr locus in a cell density-dependent manner. It has recently been shown that either phosphorylated or nonphosphorylated AgrA can bind to specific sites on the agr promoter region to modulate transcription from the agr P2 and P3 promoters (27). RNAIII, a 514-nucleotide (nt) RNA effector molecule of the agr system, is involved in regulating agr-dependent gene expression at both the transcriptional and, to a lesser extent, the translational level (48, 50). Transcription profiling studies demonstrated that an agr mutation could alter expression of at least 138 genes, with 104 genes up-regulated and 34 genes down-regulated (17).
Besides autoactivation of agr by AIP and AgrA, SarA has been shown to bind to the agr promoter to positively modulate RNAII and RNAIII transcription (15, 51). Recently we demonstrated that inactivation of sarR partially affects agr transcription in the early growth phase but that the sarR sarA double mutation has a stronger effect. Binding site analysis of SarA and SarR suggests that both proteins bind to a common region on the agr promoter (42). It has also been reported that an mgrA mutant in various strains of S. aureus (24) is defective in agr transcription, indicating the possibility that MgrA has both direct and indirect effects on agr expression.
In addition to the SarA protein family, other regulators of agr have been described, including RAP-TRAP, two-component regulatory systems such as srrAB and arlSR, and a novel membrane protein, SvrA (3, 18, 19, 20, 48, 49). RAP-TRAP is a regulatory system that appears to activate transcription of RNAII (3). The svrA inactivation also abolishes the expression of the agr locus and is thought to function as a sensor molecule for the RAP-TRAP pathway; however, this has not yet been confirmed experimentally (20). The two-component system srrAB is involved in the down-regulation of agr RNAIII under low-oxygen conditions by binding to the intergenic P2-P3 promoter region (49). While the agr locus is clearly important for the pathogenesis and production of staphylococcal diseases (e.g., bacterial spread and tissue lysis due to proteases and hemolysins), the mechanism by which it is regulated by other loci is not well defined.
In our continuing effort to understand the underlying mechanism by which the SarA protein family functions at various levels, we report here the characterization of another paralog of SarA, a 141-residue polypeptide called SarX (SA0623 of S. aureus N315) (29). The expression of sarX is maximal during the post-exponential phase, and among the SarA family of proteins, only MgrA activates its transcription. Inactivation of sarX did not affect the expression of regulatory genes in the sarA family or saeRS, but it did have a significant negative effect on the transcription of agr RNAII and RNAIII. As the agr locus is involved in the regulation of virulence genes, we also evaluated the expression of specific agr-regulated genes.
MATERIALS AND METHODS
Bacterial strains and growth conditions.
Bacterial strains used in this study are listed in Table 1, and plasmids used are described in the text. Standard Escherichia coli strains were used for manipulation of DNA and overexpression of protein as previously described (41). Phage φ11 was used as a generalized transducing phage for the S. aureus strains. S. aureus strain RN4220, a restriction-deficient derivative of strain 8325-4, was used as the initial recipient for the transformation of plasmid constructs. S. aureus cells were grown at 37°C with aeration in Trypticase soy broth (TSB) or 0.3GL agar supplemented with antibiotics when necessary (41). Luria-Bertani (LB) medium was utilized for growing Escherichia coli. Antibiotics were used at the following concentrations: for S. aureus, erythromycin at 5 μg/ml, kanamycin at 50 μg/ml, tetracycline at 5 μg/ml, and chloramphenicol at 10 μg/ml, and for E. coli, ampicillin at 50 μg/ml, chloramphenicol at 30 μg/ml, erythromycin at 200 μg/ml, and spectinomycin at 75 μg/ml.
TABLE 1.
Strains used
| S. aureus strain | Comment(s) | Reference or source |
|---|---|---|
| RN4220 | Mutant of 8325-4 that accepts foreign DNA | 46 |
| RN6390 | Laboratory strain that maintains its hemolytic pattern when propagated on sheep erythrocyte agar (parental strain) | 46 |
| RN6911 | agr mutant of RN6390 with Δagr::tetM | 46 |
| ALC488 | sarA mutant of RN6390 with ΔsarA::ermC | 39 |
| ALC812 | ALC488 with sarB (nt 1 to 1349 plus 180 bp of upstream sequence) integrated into the geh locus on the chromosome | 39 |
| ALC1713 | sarR mutant of RN6390 with ΔsarR::ermC | 38 |
| ALC1905 | sarT mutant of RN6390 with ΔsarT::ermC | 55 |
| ALC2272 | sarU mutant of RN6390 with ΔsarU::ermC | 40 |
| ALC1893 | sarS mutant of RN6390 with ΔsarS::ermC | 11 |
| ALC2319 | sarV mutant of RN6390 with ΔsarV::ermC | 41 |
| ALC2530 | mgrA mutant of RN6390 with ΔmgrA::ermC | 25 |
| ALC2531 | ALC2530 complemented with the mgrA gene into the geh locus on the chromosome | 25 |
| ALC2535 | sarA mgrA double mutant of RN6390 | 25 |
| ALC4167 | sarX mutant of RN6390 with a deletion of open reading frame of the sarX gene and replacement with an ermC gene | This study |
| ALC4168 | ALC4167 complemented with sarX gene into the geh locus on the chromosome | This study |
| ALC4169 | ALC4167 with pSK236 containing a 1.5-kb DNA fragment containing the sarX gene with a 353-bp upstream sequence at the EcoRI site | This study |
| ALC4170 | RN6390 with pSK236 containing a 1.5-kb DNA fragment containing the sarX gene with a 353-bp upstream sequence at the EcoRI site | This study |
| AM1020 | sarX sarA double mutant (sarA::kan ΔsarX::ermC) | This study |
| AM1021 | sarX agr double mutant (ΔsarX::ermC Δagr::tetM) | This study |
Genetic manipulations in E. coli and S. aureus.
Based on homology with SarA, the sarX gene product was initially identified in the S. aureus N315 genome database (20) (www.tigr.org and www.ncbi.nlm.nih.gov/). In order to construct a sarX mutant, the sarX gene, with flanking sequences on both sides, was amplified by PCR with primers 5′-AGCGACTTAAATTCGATTCGTTA-3′ and 5′-ATTTCAAATCCACATTATGGAATA-3′, using chromosomal DNA from strain RN6390 as the template. The 2.18-kb PCR fragment was cloned into vector pCR2.1 (Invitrogen, CA) in E. coli. An additional SwaI site was created near the putative ribosome-binding site of the sarX gene in the 2.18-kb DNA fragment by using a Muta-Gene mutagenesis kit for site-directed mutagenesis (Bio-Rad, Hercules, CA). A 371-bp fragment comprising the sarX coding region (N-terminal 124 residues out of 141 residues, published) was deleted by restricting with SwaI and then replacing the deleted fragment with an ∼1.2-kb ermC gene. The fragment containing the ermC replacement of the deleted sarX gene was then cloned into the temperature-sensitive shuttle vector pCL52.2 (54). Construction and selection of the putative chromosomally inserted RN6390 sarX mutant (tetracycline-sensitive and erythromycin-resistant colonies) were performed as described previously (38, 41, 42). A phage φ11 lysate of the putative sarX mutant was then prepared to infect a fresh RN6390 strain to reconstruct the sarX mutant (in an attempt to avoid any putative genomic mutations that might have occurred during the temperature shift to promote homologous recombination). The correct mutation was confirmed by PCR and Northern and Southern hybridization with sarX and ermC probes as previously described (38, 41, 42). One clone, designated ALC4167, was selected for further study.
To complement the sarX mutation, a 1.5-kb fragment encompassing the sarX gene, 353 bp upstream of the sarX translation start site, was cloned into shuttle plasmid pSK236 and single-copy integration vector pCL84 (21, 30). The recombinant pSK236 plasmid was electroporated into RN4220, selecting for chloramphenicol-resistant colonies, and the correct transformant was verified by restriction analysis of the recombinant plasmid. The plasmid from RN4220 was then electroporated into parental strain RN6390 and the sarX mutant (ALC4167) to construct trans-complemented strains carrying the sarX gene. To complement with a single copy of the sarX gene, recombinant pCL84 with sarX was electroporated into CYL316, a derivative of the RN4220 strain, and selected for tetracycline-resistant and lipase-negative colonies (30). A phage φ11 lysate of the CYL316 strain containing single-copy integration of the sarX gene onto the geh locus was prepared and transduced into the sarX mutant to construct single-copy complementation of the sarX gene. The authenticity of the complemented strain was determined by lipase-negative phenotype, Southern hybridization with sarX and lipase-specific probes, and also Northern hybridization with a sarX probe as described earlier (38, 41, 42).
Isolation of RNA and Northern blot hybridization.
Total RNA from S. aureus was prepared by using a Trizol isolation kit (Invitrogen, CA) and a reciprocating shaker as previously described (38, 41, 55). The optical densities at 650 nm (OD650) of various cultures were measured with a spectrophotometer (Spectronic 20). The concentration of total RNA was determined by measuring the absorbance at 260 nm. Ten micrograms each of total RNA samples was analyzed by Northern blotting as described earlier (38, 41). The regions containing the agr RNAII, agr RNAIII, sarA, sarR, sarT, sarS, sarU, rot, mgrA, saeR, lytS, lytR, arlR, arlS, spa, fnbA, sspA, sspB, aur, hlb, hla, lytM, lytN, atl, pbp2, and pbp4 genes were either amplified by PCR or excised from the plasmids containing the desired genes with restriction endonucleases. An internal fragment of the 16S rRNA gene (nt 777 to 1500; GenBank accession no. X68417) was used to probe 16S rRNA as a loading control. For detection of specific transcripts, gel-purified DNA probes were radiolabeled with [α-32P]dCTP by use of a random-primed DNA labeling kit (Roche Diagnostics GmbH) and hybridized under aqueous-phase conditions at 65°C. The blots were subsequently washed and autoradiographed.
Transcriptional fusion studies of agr RNAII, agr RNAIII, and hla promoters linked to the gfpuvr reporter gene.
Construction of the plasmids pALC1743, -1742, and -1740, containing agr RNAIII, agr RNAII, and hla promoter fragments, respectively, linked to the gfpuvr reporter gene, were described earlier (26, 38, 40, 41). A 185-bp sarX promoter fragment (positioned from +13 to −171 bp of the sarX promoter) (Fig. 1) was amplified by PCR using chromosomal DNA of S. aureus strain RN6390 and primers with flanking EcoRI or XbaI sites. An EcoRI and XbaI fragment containing the sarX promoter was cloned into shuttle plasmid pALC1484, generating a transcriptional fusion of the sarX promoter to the gfpuvr reporter gene. Restriction analysis and DNA sequencing confirmed the orientation and authenticity of the promoter reporter gene constructs. The recombinant plasmids containing the respective promoter regions were first introduced into S. aureus strain RN4220 by electroporation. Plasmids purified from RN4220 transformants were then electroporated into RN6390, its isogenic mutants ALC4167 (sarX::ermC), ALC488 (sarA::ermC), and ALC2530 (mgrA::ermC), and single-copy-complemented mgrA (ALC2531) and sarA (ALC812) strains as required.
FIG. 1.
Transcription and promoter analysis of the sarX gene in S. aureus. (A) Northern analysis of the sarX transcripts in the wild-type strain RN6390 at the different growth phases. The blot was probed with a 450-bp sarX DNA fragment containing the entire open reading frame of the sarX gene and a 723-bp internal region of 16S rRNA genes, respectively. (B) Primer extension of the sarX transcript by using total RNA isolated from the wild-type strain RN6390 at the post-exponential phase of growth. (C) The nucleotide sequence of the 185-bp promoter region of sarX is shown and marked with putative promoter recognition sites (−35 and −10) and regulatory regions in bold. The transcriptional start site (+1) was identified based on a primer extension experiment. The ribosome-binding sequences (RBS) and translational start codon (TTG) are marked in bold. The predicted translational start codon (ATG) in various S. aureus genomes is underlined.
After overnight incubation, S. aureus strains harboring the recombinant plasmids were diluted 1:100 in TSB containing chloramphenicol (10 μg/ml) and grown at 37°C with shaking at 250 rpm. Aliquots (200 μl) were transferred in triplicate hourly for 10 h to microtiter plates to assay for optical density (OD650) and green fluorescent protein fluorescence with an FL600 fluorescence reader (BioTek Instruments, Winooski, VT). Promoter activation was plotted as mean fluorescence per optical density (OD650) over time, using the average values from triplicate readings.
Purification of MgrA and SarX proteins.
The cloning, purification, and authentication of His6-tagged MgrA protein were described earlier (41). The 360-bp DNA fragment containing the deduced full-length sarX gene was amplified by PCR using chromosomal DNA from S. aureus RN6390 as the template and primers containing flanking restriction sites (NdeI and BamHI) to facilitate in-frame cloning in the expression vector pET14b (Novagen, Madison, WI). The recombinant His6-tagged SarX protein expression and purification were accomplished in a manner similar to that utilized for SarA or MgrA protein as described elsewhere (38, 41, 42). The purified His6-tagged SarX protein was found to be more than 98% pure in a 12% polyacrylamide-sodium dodecyl sulfate gel. The concentration of the purified proteins was determined by the Bradford protein assay (Bio-Rad, Hercules, CA) (6a), using bovine serum albumin as the standard.
Gel shift and primer extension analysis.
In order to determine if the recombinant MgrA protein binds to the sarX promoter region, a 185-bp fragment (positions +13 to −171), representing the sarX promoter region, was end labeled with [γ-32P]ATP by using T4 polynucleotide kinase. The labeled fragment (0.1 ng, or 0.8 fmol) was incubated at room temperature for 20 min with various amounts of purified MgrA protein in 25 μl of binding buffer (25 mM Tris-HCl, pH 7.5, 0.1 mM EDTA, 75 mM NaCl, 1 mM dithiothreitol, and 5% glycerol) containing 0.5 μg of calf thymus DNA (Amersham Pharmacia Biotech). The reaction mixtures were analyzed in an 8.0% nondenaturing polyacrylamide gel. The band shifts were detected by exposing dried gels to X-ray films or scanning with a PhosphorImager scanner (Amersham Biosciences, NJ). Similarly, a 179-bp DNA fragment of the agr intergenic promoter region was used to perform gel shift assays with purified SarX protein. Unlabeled specific (185-bp sarX or 179-bp agr fragment) and nonspecific (200-bp P2 sarA fragment) (4) competitors were used effectively to compete with the labeled DNA substrates.
Mapping of the 5′ end of the sarX transcript by primer extension was performed using the primer 5′-AGACTGCTAAATCATTTAGCGACA-3′, complementary to the sarX coding strand and located from nucleotide positions 85 to 108 downstream from the putative start codon TTG. Reverse transcription was carried out by using total RNA isolated from wild-type RN6390 in stationary-phase culture as described previously (38, 41).
RESULTS
Identification of the sarX gene.
A search for additional SarA paralogs in the published S. aureus N315 genome, using the default setting of 25 as the cutoff for the BLASTP search (29) (www.ncbi.nlm.nih.gov/genomes/), revealed at least 11 proteins homologous to SarA (SA0573, protein gi 13700508). These paralogs, including SarT (55, 56), SarS (also called SarH1) (11, 57), SarU (also called SarH2) (40), SarR (38), Rot (43, 53), MgrA (also called Rat or NorR) (25, 37, 39, 60), and SarV (41), have been shown to play a role in the regulation of virulence genes. The remaining paralogs, which are of unknown function, are SA2174, SA2091, and SA0141 (29). SarX (SA0623) is the fifth most closely related protein that came up in our homology search for SarA paralogs, with SarT, SarS, SarU, and SarR being the first to fourth hits (29). SarX is a 141-residue polypeptide that shares 51, 46, 51, 49, 54, and 47% homology with SarV, SarA, SarU, SarR, SarS, and Rot, respectively. SarX of S. aureus strain RN6390 is highly conserved among the six published Staphylococcus genomes of methicillin-susceptible S. aureus 476, (100% homology), methicillin-resistant S. aureus N315, MW2, Mu50 (100% homology), COL (99% homology), and 252 (99% similarity and 98% identity). There exists a significant degree of homology of S. aureus SarX to the hypothetical protein (SE0436) of S. epidermidis ATCC 12228 (122 residues, E value of 1e−30, with 68% identity and 89% similarity) and the hypothetical protein (SERP0321) of S. epidermidis RP62A (119 residues, E value of 1e−29, with 66% identity and 88% similarity). Based on sequence alignment, it was observed that specific residues of SarX were conserved within the SarA protein family (see Fig. S1S in the supplemental material). A number of putative transcriptional regulators with unknown function and low homology were found upon conducting a BLASTP comparison of SarX against other microbial genomes with the same default setting. These data indicate that SarX is likely to be regulatory in nature and is highly conserved in staphylococcal species.
Analyses of sarX expression during various growth phases were performed using Northern hybridization. Two transcripts of the sarX gene were found, one weakly expressed at ∼1,500 nt (P2) during the post-exponential phase and another highly (and maximally) expressed at ∼500 nt (P1) during the post-exponential phase of growth, thus indicating a strong growth phase dependency for sarX transcription (Fig. 1A). A similar transcription pattern was observed with S. aureus strains 8325-4 and SH1000 (rsbU+, a sigB optimal expression derivative of RN6390) and other prototypic S. aureus strains, such as Newman (methicillin-susceptible S. aureus) and COL (methicillin-resistant S. aureus) (data not shown). Northern hybridization with open reading frames (SA0622 and SA0624) adjacent to SarX suggested that the larger transcript may have originated from a cryptic promoter within the SA0622 (717-residue) coding region (data not shown).
Transcriptional start site and promoter structure of the sarX gene.
In order to determine the transcriptional start site and the promoter sequence of the major sarX transcript, primer extension was performed with total RNA isolated from the wild-type strain RN6390 (Fig. 1B). The transcriptional start site was mapped to an “A” which is located 29 nt downstream from the predicted SarX initiation codon ATG in the S. aureus genome (Fig. 1C). Based upon the mapped transcriptional start site, the predicted putative promoter region is AAGACA(−35)-N17-GATAAT(−10), which is closely homologous to the −10 and −35 consensus sequences of σA-dependent promoters (Fig. 1C). The computer-generated published translational start codon (ATG) for SarX was located at the −29-nt position from the mapped transcriptional start site (+1) and displays a lack of any defined ribosome-binding sequences (RBS). However, a strong RBS, TGGGGG, is located at the +24-nt position from the transcriptional start site. Based on the mapped transcriptional start site and the presence of a strong putative RBS, we suggest that TTG, located 9 nt downstream of the RBS, is the likely SarX translational start codon. Analysis of the downstream sequence of the sarX gene revealed the presence of a potential rho-independent transcriptional terminator sequence from a position 11 to 42 bp immediately after the stop codon (TAA). Within this region, a 12-bp inverted repeat sequence from positions 11 to 22 and 30 to 42 after the stop codon of the sarX gene could form a potential hairpin structure with a 12-nt-long heteroduplex. Thus, primer extension suggests that SarX is a 119-residue polypeptide starting with a TTG codon and not a 141-residue polypeptide as predicted from various S. aureus genomes. Interestingly, the predicted SarX proteins in S. epidermidis ATCC 12228 and RP62A are 122 and 119 residues, respectively, more closely aligned to the deduced 119-residue size of their S. aureus counterpart. The alignment of the 119-residue SarX with members of the SarA protein family is much more ordered and of higher quality than the alignment of the 141-residue counterpart with these members (e.g., leaving fewer gaps) (see Fig. S1S in the supplemental material).
Analysis of sarX transcription in RN6390 and an assortment of isogenic mutants.
To determine the phenotypic effects of sarX, we constructed a sarX mutant in RN6390 by allelic replacement, essentially replacing the sarX gene with an ermC cassette (see Materials and Methods). Northern blot assays were performed with a sarX probe (450 bp within the coding region) with the various regulatory mutants in our collection to determine if the transcription of the sarX gene is affected by inactivation of other regulatory systems, including those in the sarA family of genes. Our results disclosed that the level of sarX transcript was undetectable only with the mgrA mutant, whereas no significant changes were observed with other mutants tested (Fig. 2A). Upon introduction of a single copy of the mgrA gene into the lipase locus (geh) of the mgrA mutant, the expression of sarX transcript returned to near-parental level (Fig. 2B), suggesting that the expression of sarX is truly activated by the mgrA gene product. As a control, the sarA mutant and its complemented strain were also assessed for sarX transcript, and no significant alteration was observed. Interestingly, no transcription of sarX was detected with the sarA mgrA double mutant, indicating that only mgrA is involved in the regulation of the sarX gene. This result was also confirmed by promoter fusion studies (sarX P1 promoter-gfpuvr) with the various isogenic strains. We found a substantial reduction in mean green fluorescent protein fluorescence for the mgrA mutant ALC2530 (73 ± 25 U per OD650) compared to that for the wild-type strain RN6390 (2,054 ± 200 U per OD650) and the single-copy-complemented mgrA mutant ALC2531 (1,809 ± 160 U per OD650). We consistently observed that there was some degree of decreased or increased sarX transcript level in some of the mutants, particularly for sarR and sae, which we speculate might be due to an indirect effect. As expected, single-copy or multicopy complementation of the sarX mutant in a plasmid carrying the sarX gene restored or increased the level of the sarX transcript. There are no significant growth differences between the sarX mutant and the wild type and various complemented strains (data not shown). Growth phase variation did not differ significantly from the data presented above for the expression of sarX transcript when Northern blot hybridization was performed with total RNA isolated in the early phase of growth of various mutants.
FIG. 2.
Northern analysis of the sarX transcript in the wild-type (wt) strain, various isogenic mutants, and complemented strains at the post-exponential phase (OD650 of ∼1.7) of growth (A) and at both the exponential and post-exponential phases (B). A 450-bp DNA fragment containing the sarX gene was used for hybridization. cp s and cp m denote complementation in single copy and multicopy, respectively, with the indicated genes. The 723-bp internal fragment of 16S rRNA was used for hybridization as a loading control (A), or the region of 23S and 16S rRNA of the ethidium bromide-stained gel was used for the hybridization (B).
Binding of MgrA protein to the sarX promoter region.
As the level of sarX transcript decreased in an mgrA mutant and was restored to the wild-type level in a single-copy-complemented mgrA strain, we speculated that MgrA might bind to the sarX promoter region to activate sarX transcription. We employed a 185-bp sarX promoter fragment (positioned from +13 to −171 nt) (Fig. 1C) for DNA binding assays to test this supposition. The γ-32P-radiolabeled DNA fragment was used in gel shift assays with various amounts of purified MgrA protein (Fig. 3). Retarded DNA-protein complex was detected with as little as 0.1 to 0.2 μg of MgrA (2.6 to 5.2 nM, assuming that MgrA is a dimeric protein like SarR or SarA). As the concentration of protein increased, the retarded protein-DNA complex became the predominant band, with complete conversion at ∼0.5 μg of MgrA protein. The dissociation constant of MgrA was calculated, based on the amount of protein (∼200 ng) needed to shift 50% of the labeled probe, to be 2.06 × 10−4 M (52). Unlabeled promoter fragment effectively competed with labeled fragment, as shown in Fig. 3, suggesting that the DNA-protein complex was specific. Hence, MgrA protein can bind to the sarX promoter region, presumably acting as an activator of sarX transcription.
FIG. 3.
Autoradiogram of an 8.0% polyacrylamide gel showing gel shifts for purified MgrA protein with a γ-32P-labeled, 185-bp sarX promoter fragment. Mobility of the band was noted in the presence of increasing amounts of MgrA protein, as indicated on the top. Unlabeled, specific 185-bp sarX fragment and nonspecific 200-bp PsarA fragment (nt 1 to 200; GenBank accession no. U46541) competitors with 50-fold excess (molar ratio), as indicated, contained 0.5 μg of the purified MgrA protein.
Previously, we mapped the DNA binding site of the MgrA protein upstream of the sarV promoter region (41). DNA sequence analysis upstream of the sarX promoter region revealed the presence of the sequence TGTTGGN8ACAACG, which is consistent with our earlier observation of the MgrA binding site on the sarV promoter (Fig. 1C) (41). It is possible that the sequence-specific binding of MgrA to upstream of the promoter region determines the function of MgrA, whether it will activate or repress the target gene. We are in the process of investigating whether the sequence-specific binding is one of the underlying mechanisms of gene regulation by the SarA family of proteins.
Transcription and transcriptional fusion assays for regulatory genes in a sarX mutant.
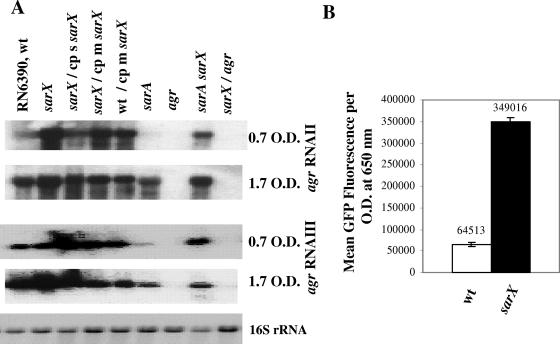
To determine whether inactivation of sarX affects other regulatory genes involved in virulence, we performed Northern blot analysis with probes for 12 different regulatory loci (agr RNAII, agr RNAIII, sarA, sarR, sarS, sarT, sarU, mgrA, rot, sarV, saeR, lytS, lytR, arlR, and arlS). The functions of many SarA paralogs have been described previously, including SarR as a regulator of SarA and agr expression, SarS as an activator of protein A expression, SarT as a repressor of α-hemolysin expression, and MgrA as an activator of agr expression and repressor of sarV transcription (11, 24, 37, 38, 41, 42, 56, 57). The lytSR locus is a two-component regulatory system that negatively controls extracellular murein hydrolase activity via positive regulation of the lrgAB operon (9, 23). Another two-component regulatory mutant (arlRS) up-regulates the expression of autolysis genes while augmenting transcription of agr and other genes, including those for extracellular serine protease and the multidrug resistance transporter gene norA (18, 19). The saeRS locus encodes another two-component regulatory system that positively regulates the expression of α- and β-hemolysins, DNase I, coagulase, and protein A (22). As shown in Fig. 4A, there was a significant elevation in RNAII and RNAIII transcription during the exponential and post-exponential phases of growth in the sarX mutant. The wild-type levels were restored in the mutant complemented with a single-copy sarX gene in the lipase (geh) locus. A significant reduction of transcript levels of agr RNAII and RNAIII in the post-exponential phase was observed with the sarX mutant or wild-type strain carrying the sarX gene on a multicopy plasmid. This is consistent with a dose-dependent effect of sarX on the transcription of the agr locus. An increase in agr transcription was verified using agr RNAII and agr RNAIII promoter fusions with the gfpuvr reporter gene in the wild type and an isogenic sarX mutant (data for agr RNAII not shown), thus suggesting that the increase in agr promoter activity in the sarX mutant is likely due to inactivation of sarX. Northern and transcriptional fusion studies suggest that the sarX gene product possibly acts as a repressor of the expression of agr transcripts.
FIG. 4.
Analysis of expression of agr transcripts in a sarX mutant and various isogenic mutant strains. (A) Northern blots of agr RNAII and RNAIII and 16S rRNA transcripts (a loading control) for the wild-type (wt), sarX mutant, single-copy-complemented (cp s), and trans-complemented (cp m) strains and isogenic sarA, agr, and double mutant strains from the exponential (OD650 of ∼0.7) and post-exponential (OD650 of ∼1.7) phases of growth. The blots were probed with 3.0-kb agr RNAII, 0.5-kb agr RNAIII, and 0.723-kb internal fragments of 16S rRNA genes (as a loading control). (B) gfpuvr expression as driven by agr RNAIII in the wild type and the sarX mutant at the 8-h time point (post-exponential phase). To minimize variations in fluorescence attributable to cell density, the data are presented as the averages of triplicate samples in fluorescence unit per absorbance at 650 nm. GFP, green fluorescent protein.
Interestingly, expression of both agr transcripts was reduced significantly in the sarA sarX double mutant compared with levels for the single sarX mutant; we speculate, therefore, that there is some sort of cross talk between sarA and sarX genes in their regulation of the agr locus. In contrast, the transcription of other sarA family genes, arlRS, lytSR, and saeR, was not significantly altered in the sarX mutant (data not shown). It should be stressed, however, that expression of sarS, sarT, sarU, and sarV was difficult to detect even in the parental strain under routine laboratory culture conditions due to their repression by other SarA paralogs and the agr system.
Northern blot analysis for the expression of target genes.
The data presented above clearly indicated that the expression of sarX, normally expressed in the post-exponential phase in the parental strain, was decreased in the mgrA mutant. MgrA, a member of the SarA protein family, binds to the sarX promoter region, presumably to activate sarX transcription. Analyses of various regulatory genes suggested that the expression of agr transcripts is significantly affected by the inactivation of sarX. We thus measured the expression of a few virulence genes known to be regulated by agr-dependent or -independent mechanisms in the sarX mutant. Among those genes tested were cell surface-associated proteins (e.g., spa and fnbA), hemolysins (α-hemolysins [hla] and β-hemolysins [hlb]), proteases (a V8 protease, or sspA; a zinc metalloprotease, or aur; and a cysteine protease, or sspB), and genes involved in the cell wall integrity (autolysin, or atl; penicillin binding proteins, or Pbp2 and Pbp4; an endopeptidase, or lytM; and a cell wall hydrolase, or lytN) (41). Significantly increased transcript levels of hemolysins and proteases were observed for the sarX mutant, while transcript levels of these genes returned to near-normal levels in the mutant complemented in single copy or trans-complemented by sarX in multicopy (Fig. 5A). Transcriptional fusion studies of hla-gfpuvr, hlb-gfpuvr, and sspA-gfpuvr supported the Northern blot data; a representation of hla-gfpuvr fusion data is shown in Fig. 5B at 8 h of growth for the sarX mutant and the parental strain. No detectable change in expression of the protein A (spa) or fibrinogen binding protein (fnbA) transcription was observed for the sarX mutant compared to the parental strain at the exponential phase, which is consistent with the agr repression of the cell wall-associated proteins. There was no significant effect on the transcription of those genes involved in cell wall integrity (data not shown). Analysis of the sarA sarX and sarX agr double mutants (Fig. 5A) suggests that the expression of hemolysins and protease (sspA) transcription is agr dependent. We also believed that sarX might be involved directly or indirectly in a sarA-dependent manner in regulation of the sspA transcription, as retention of the sspA transcript was found in the sarA sarX and sarX agr double mutants compared to the sarA or agr single mutant (Fig. 5A). Whether this effect is direct or indirect and whether it is physiologically relevant remain to be determined.
FIG. 5.
Analysis of expression of target gene transcripts in a sarX mutant and its various isogenic strains. (A) Northern blots of α- and β-hemolysins (hla and hlb), V8 protease (sspA), and 16S rRNA transcripts (a loading control) for the wild-type (wt), sarX mutant, single-copy-complemented (cp s), and multicopy-complemented (cp m) sarX mutant strains and isogenic sarA, agr, and double mutant strains from the various growth phases as indicated. (B) Promoter activation of the 235-bp hla promoter fused to a gfpuvr reporter gene in the wild type and the sarX mutant at the 8-h time point. To minimize variations in fluorescence attributable to cell density, the data are presented as the averages of triplicate samples in fluorescence unit per absorbance at 650 nm. GFP, green fluorescent protein.
Phenotypic effects of the sarX gene.
Alteration in agr transcription and subsequently hemolysin and protease transcription in the sarX mutant, coupled with the activation effect of sarX by mgrA, suggests that sarX might play an important role in controlling genes mediated by mgrA and agr in S. aureus. Because it has previously been shown that mgrA and agr independently positively regulate autolysis in strain RN6390 (25), we tested the sarX mutant for susceptibility to lysis in the presence of a detergent and an antibiotic. Interestingly, the sarX mutant had no effect on Triton X-100- or penicillin G-mediated cell lysis (data not shown), which is consistent with our Northern hybridization finding that genes involved in cell wall integrity are not affected by sarX inactivation. Phenotypic assays for hemolysins on sheep blood agar plates or zymogram analyses of proteases in casein or gelatin gels indicated that levels of hemolysins and proteases are increased in the sarX mutant and decreased in complemented strains. These data suggest that the sarX gene product represses expression of agr and many of the agr-dependent target genes. As the agr locus is known to activate transcription of many exoprotein and protease genes, the effect of sarX on the target genes could be either direct (agr independent) or indirect (agr dependent).
Effects on the expression of genes surrounding sarX (SA0623), particularly SA0622, were analyzed. The SA0622 gene, also known as rbf (regulator for biofilm formation), is involved in regulation of biofilm formation under high-salt and -glucose concentrations (32). Northern blot analysis with total RNA isolated from various isogenic sarX strains (e.g., wild-type, sarX mutant, and complemented sarX strains) under different concentrations of glucose (0.25%, 0.68%, and 2.7%) and NaCl (0.5 M and 1 M) with sarX and SA0622 (2-kb) gene probes yielded no detectable change in SA0622 (rbf) transcription in the sarX mutant. Similarly, results of microtiter plate assays for biofilm formation utilizing various concentrations of glucose and NaCl in TSB demonstrated no significant effect on biofilm formation in the sarX mutant compared to the wild type (data not shown). However, the sarX mutant aggregated significantly on the side of culture glass tubes and tended to lyse when the cells were grown for more than 12 h in TSB. These phenotypes can be eliminated by adding DNase I to the culture media; therefore, we speculate that production of factors associated with aggregation of cells is probably increased due to the inactivation of the sarX gene. Further work will be required to elucidate the reason for such aggregation in the sarX mutant.
Binding of SarX protein to the intergenic promoter region of agr locus.
Since agr transcription during both phases of growth is increased in the sarX mutant and restored to the parental level in the single-copy-complemented sarX mutant, we speculate that SarX binds directly to the agr P2-P3 promoter region. We purified the SarX (119-residue) protein as a His-tag fusion and used it for the gel shift assay with a 179-bp labeled DNA fragment containing the intergenic agr P2-P3 promoters, with various amounts of purified fusion SarX protein (Fig. 6). Retarded protein-DNA complex was detected with as little as 25 ng of SarX protein (0.78 nM). As the concentration of SarX increased, the retarded protein-DNA complex became the predominant band, with a complete shift at ∼0.5 μg of SarX protein. The dissociation constant of SarX was calculated, based on the amount of protein (∼100 ng) needed to shift 50% of the labeled probe, to be 1.25 × 10−4 M (52). Unlabeled competitor DNA fragment effectively eliminated retarded complex, whereas unlabeled nonspecific competitor DNA fragment did not. The SarX-DNA complex displayed an unusual laddering pattern with the agr promoter fragment. The same type of laddering pattern was observed when gel shift assays were performed with some of the other SarA paralog proteins, particularly SarR and SarT, on their corresponding target promoter regions (38, 40, 42, 56). One plausible explanation for this phenomenon is that the agr promoter region perhaps contains multiple SarX binding sites; alternatively, SarX could bind in multimeric form to a common site or multiple sites within the agr fragment. Interestingly, the same type of laddering was reported with the agr promoter fragment either with a crude lysate of S. aureus (45) or with purified SarA (51). Under our experimental conditions, we observed the laddering pattern with SarR but not with SarA within the same 179-bp agr promoter region, which is consistent with earlier observations of various promoter fragments (15, 34, 41, 58, 59). The specificity and mode of action of the SarX protein along with other SarA paralogs, particularly MgrA, to the agr promoter region will be investigated. We surmise that the 119-residue SarX protein can bind to the agr promoter region and perhaps function to repress transcription.
FIG. 6.
Autoradiogram of an 8.0% polyacrylamide gel showing gel shifts for purified His fusion SarX protein (119 residue) with a γ-32P-labeled, 179-bp agr promoter fragment. Mobility of the band was noted in the presence of increasing amounts of SarX protein, as indicated on the top. Cold competition lanes with 50-fold excess (molar ratio) of the unlabeled 179-bp agr fragment and 200 bp P2 sarA DNA fragment (as indicated) contained 0.5 μg of the purified SarX protein.
DISCUSSION
S. aureus exhibits complex patterns of protein expression in response to different environmental conditions (2, 8, 12, 13, 35, 48), consistent with the notion that large numbers of complex regulatory networks coordinate expression of most virulence factors. In prior studies, we and others have characterized eight SarA-like proteins, SarA, SarR, SarS, Rot, SarT, SarU, SarV, and MgrA, that are involved in regulation of sarA family genes and virulence genes in S. aureus. Members of the SarA protein family in S. aureus are similar both in sequence and in structure and are also homologous to members of the MarR protein family of gram-negative bacteria (1, 12).
The levels of transcription of sarA-like genes are highly variable in strain RN6390. For instance, sarA, sarR, and mgrA transcripts are highly expressed in the parental strain, whereas the transcripts of sarU, sarT, sarV, and sarS are not readily detectable under normal laboratory growth conditions. We found the sarX gene to be poorly transcribed in the early and late exponential phases of growth, whereas transcription was significantly higher in the post-exponential phase in the parental strain. Northern analysis of sarX expression in mutants with mutations in agr, sae, and sigB and seven sarA family genes revealed that transcription of sarX remains unaltered in these mutants, with the exception of mgrA. SarX expression is decreased in the mgrA mutant and is restored upon complementation. Similar results were observed with a sarX promoter-gfpuvr fusion.
Based on the transcriptional start site found in the primer extension experiment and the presence of a strong ribosome-binding site, we propose TTG to be the translational start codon of the sarX gene. Use of TTG as a start codon is not unprecedented in S. aureus. The spa (protein A), agrB, and blaZ genes of S. aureus all have TTG as their translational start codon (27). In the future, we plan to determine the size of SarX by Western analysis of cell extracts from the post-exponential growth phase. When SarX (119 residues) was aligned with other SarA paralogs along with MarR of E. coli, we found that the alignment was more significant than that with the published 141-residue SarX (see Fig. S1S in the supplemental material). Interestingly, many basic residues are conserved within members of the SarA protein family, consistent with the notion that the DNA binding domains might be highly conserved in this protein family. Indeed, detailed mutational analysis of SarR and SarA indicates that many of the conserved basic residues (e.g., K52, K80, R82, R88, etc., of SarR) are involved in DNA binding (14, 34, 42).
By use of gel shift studies, we found that purified MgrA binds to the sarX promoter region with high affinity. Two MgrA consensus-binding sequences were found within the −51- to −75-bp sarX promoter region located 18 bp upstream of the −35 region of the sarX promoter. Interestingly, the organization of these two consensus-binding sequences is distinct from that in the sarV promoter region, where MgrA acts as a repressor (41). In the case of the sarV promoter region, two out of three consensus-binding sequences are organized in the same manner as in the sarX promoter region, but additional consensus-binding sequences are located adjacent to the second binding site on the opposite strand of DNA. Therefore, we speculate that one of the determining factors for whether SarA paralogous proteins function as repressors or activators may be the organization of sequence-specific binding sites in their cognate target DNA.
It has been shown by transcriptional profiling that agr, sarA, mgrA, and rot regulate the expression of more than 100 genes involved in regulation, virulence, autolysis, and metabolic processes in S. aureus (17, 37, 53). Inactivation of the sarX gene had a positive effect on the expression of agr, while the effects on sae, lyt, sarA, and sarA paralogs were minimal. sarX is the only regulator in the SarA protein family known to negatively regulate agr transcription. The regulation of secreted proteins by sarX entails up-regulation of hemolysins (e.g., hla and hlb) and proteases (e.g., sspA), concordant with the notion that sarX regulates agr.
We have demonstrated that the SarX protein binds to the agr intergenic promoter region with high affinity, which implies direct regulation of agr by SarX, but the exact binding region is yet to be determined. Although it is known that mgrA is involved in positive regulation of agr transcription in various strains of S. aureus (24), no consensus MgrA binding site was found in the agr P2-P3 promoter region, leading us to speculate that SarX may interact with MgrA to modulate the expression of the agr transcription.
Recently, we have shown that the SarA-SarR pathway may be involved in positive regulation of agr transcription in the exponential phase of growth (42) and that post-exponential control of agr transcription may be governed by some other regulatory systems. A SarX-MgrA pathway may play that role. This hypothesis is further supported by results of the promoter fusion studies (sarX P1 promoter-gfpuvr) with the various isogenic strains. We found that the values of sarX promoter fusion were significantly decreased for a sarX mutant compared with values for the wild-type and the complemented sarX mutant strains (unpublished data). Therefore, it is reasonable to suggest that SarX itself is involved in regulating its own expression, probably in association with MgrA. Based on published and unpublished observations, we propose a simple working model for the regulation of the agr system by the SarA protein family in Fig. 7.
FIG. 7.
Proposed model for the regulation of the expression of agr transcription in S. aureus. There are mainly two types of pathways involved in agr regulation: autoactivation by the AgrA and SarA family proteins. Among SarA paralogs, two pathways are involved: the SarA-SarR pathway operates in the exponential phase of growth, whereas the SarX-MgrA pathway operates mostly in the post-exponential phase of growth, when SarX is expressed. Other regulatory systems may be involved directly or indirectly. We also speculate that these regulators interact with each other to ensure optimal expression of agr transcription. The letters B, D, C, and A appearing in boxes are the agrB, agrD, agrC, and agrA genes, respectively, encoded within the agr RNAII transcript.
To further define the precise function of the sarX gene product, we are in the process of analyzing various knockouts of sarX with other regulatory genes, particularly with sarA and mgrA, and investigating the molecular interaction of SarX with the upstream region of its target genes. Analysis of a sarX knockout in combination with mutations in sarA, mgrA, and other regulatory genes, coupled with our studies of the structure-function relationship of these proteins, may provide a clearer understanding of the mechanism of gene regulation by SarX and other SarA paralogs. Increasing our understanding of the precise mechanisms by which the SarA protein family controls the expression of various target genes will perhaps open up new perspectives for antimicrobial chemotherapy utilizing key inhibitors of these systems.
Supplementary Material
Acknowledgments
We thank Leonard Copertino, MaryBeth Maloney, and Binata Ray for their technical help. We thank Keith Weaver for critical reading and helpful comments on the manuscript.
This work was supported in part by NIH grant AI37142 to A.L.C. and 2010-initiative start-up fund to A.C.M.
Footnotes
Supplemental material for this article may be found at http://jb.asm.org/.
REFERENCES
- 1.Alekshun, M. N., S. B. Levy, T. R. Mealy, B. A. Seaton, and J. F. Head. 2001. The crystal structure of MarR, a regulator of multiple antibiotic resistance, at 2.3 A resolution. Nat. Struct. Biol. 8:710-714. [DOI] [PubMed] [Google Scholar]
- 2.Arvidson, S., and K. Tegmark. 2001. Regulation of virulence determinants in Staphylococcus aureus. Int. J. Med. Microbiol. 291:159-170. [DOI] [PubMed] [Google Scholar]
- 3.Balaban, N., T. Goldkorn, Y. Gov, M. Hirshberg, N. Koyfman, H. R. Mathews, R. T. Nhan, B. Singh, and O. Uziel. 2001. Regulation of Staphylococcus aureus pathogenesis via target of RNAIII-activating protein (TRAP). J. Biol. Chem. 276:2658-2667. [DOI] [PubMed] [Google Scholar]
- 4.Bayer, M. G., J. H. Heinrichs, and A. L. Cheung. 1996. The molecular architecture of the sar locus in Staphylococcus aureus. J. Bacteriol. 178:4563-4570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Blevins, J. S., A. F. Gillaspy, T. M. Rechtin, B. K. Hurburt, and M. S. Smeltzer. 1999. The staphylococcal accessory regulator (sar) represses transcription of the S. aureus collagen adhesin gene (cna) in an agr-independent manner. Mol. Microbiol. 33:317-326. [DOI] [PubMed] [Google Scholar]
- 6.Boyce, J. M. 1997. Epidemiology and prevention of nosocomial infections, p. 309-329. In K. B. Crossley and G. L. Archer (ed.), The staphylococci in human diseases, vol. 12. Churchill Livingston, New York, N.Y. [Google Scholar]
- 6a.Bradford, M. M. 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72:248-254. [DOI] [PubMed] [Google Scholar]
- 7.Brandenberger, M., M. Tschierske, P. Giachino, A. Wada, and B. Berger-Bachi. 2000. Inactivation of a novel three-cistronic operon tcaR-tcaA-tcaB increases teicoplanin resistance in Staphylococcus aureus. Biochim. Biophys. Acta 1523:135-139. [DOI] [PubMed] [Google Scholar]
- 8.Bronner, S., H. Monteil, and G. Prevost. 2004. Regulation of virulence determinants in Staphylococcus aureus. FEMS Microbiol. Rev. 28:485-492. [DOI] [PubMed] [Google Scholar]
- 9.Brunskill, E. W., and K. W. Bayles. 1996. Identification of LytSR-regulated genes from Staphylococcus aureus. J. Bacteriol. 178:5810-5812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cheung, A. L., J. M. Koomey, C. A. Butler, S. J. Projan, and V. A. Fischetti. 1992. Regulation of exoprotein expression in Staphylococcus aureus by a locus (sar) distinct from agr. Proc. Natl. Acad. Sci. USA 89:6462-6466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cheung, A. L., K. A. Schmidt, B. Bateman, and A. C. Manna. 2001. SarS, a SarA homolog repressible by agr, is an activator of protein A synthesis in Staphylococcus aureus. Infect. Immun. 69:2448-2455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cheung, A. L., and G. Zhang. 2002. Regulation of virulence determinants in Staphylococcus aureus by the SarA protein family. Front. Biosci. 7:D1825-D1842. [DOI] [PubMed] [Google Scholar]
- 13.Cheung, A. L., A. S. Bayer, G. Zhang, H. Gresham, and Y.-Q. Xiong. 2004. Regulation of virulence determinants in vitro and in vivo in Staphylococcus aureus. FEMS Microbiol. Lett. 1649:1-9. [DOI] [PubMed] [Google Scholar]
- 14.Cheung, A. L., and A. C. Manna. 2005. Role of the distal sarA promoters in SarA expression in Staphylococcus aureus. Infect. Immun. 73:4391-4394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Chien, Y.-T., and A. L. Cheung. 1998. Molecular interactions between two global regulators, sar and agr, in Staphylococcus aureus. J. Biol. Chem. 237:2645-2652. [DOI] [PubMed] [Google Scholar]
- 16.Chien, Y.-T., A. C. Manna, S. J. Projan, and A. L. Cheung. 1999. SarA, a global regulator of virulence determinants in Staphylococcus aureus, binds to a conserved motif essential for sar-dependent gene regulation. J. Biol. Chem. 274:37169-37176. [DOI] [PubMed] [Google Scholar]
- 17.Dunman, P. M., E. Murphy, S. Haney, D. Palacios, G. Tucker-Kellogg, S. Wu, E. L. Brown, R. J. Zagursky, D. Shlaes, and S. J. Projan. 2001. Transcriptional profiling-based identification of Staphylococcus aureus genes regulated by agr and/or sarA loci. J. Bacteriol. 183:7341-7353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Fournier, B., and D. C. Hooper. 2000. A new two-component regulatory system involved in adhesion, autolysis, and extracellular proteolytic activity of Staphylococcus aureus. J. Bacteriol. 182:3955-3964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Fournier, B., A. Klier, and G. Rapoport. 2001. The two-component system ArlS-ArlR is a regulator of virulence gene expression in Staphylococcus aureus. Mol. Microbiol. 41:247-261. [DOI] [PubMed] [Google Scholar]
- 20.Garvis, S., J. M. Mei, J. Ruiz-Albert, and D. W. Holden. 2002. Staphylococcus aureus svrA: a gene required for virulence and expression of the agr locus. Microbiology 148:3235-3243. [DOI] [PubMed] [Google Scholar]
- 21.Gaskill, M. E., and S. A. Khan. 1988. Regulation of the enterotoxin B gene in S. aureus. J. Biol. Chem. 263:6276-6280. [PubMed] [Google Scholar]
- 22.Giraudo, A. L., A. L. Cheung, and R. Nagel. 1997. The sae locus of Staphylococcus aureus controls exoprotein synthesis at the transcriptional level. Arch. Microbiol. 168:53-58. [DOI] [PubMed] [Google Scholar]
- 23.Groicher, K. H., B. A. Firek, D. F. Fujimoto, and K. W. Bayles. 2000. The Staphylococcus aureus lrgAB operon modulates murein hydrolase activity and penicillin tolerance. J. Bacteriol. 182:1794-1801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ingavale, S., W. van Wamel, T. T. Luong, C. Y. Lee, and A. L. Cheung. 2005. Rat/MgrA, a regulator of autolysis, is a regulator of virulence genes in Staphylococcus aureus. Infect. Immun. 73:1423-1431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ingavale, S. S., W. van Wamel, and A. L. Cheung. 2003. Characterization of RAT, an autolysis regulator in Staphylococcus aureus. Mol. Microbiol. 48:1451-1466. [DOI] [PubMed] [Google Scholar]
- 26.Kahl, B. C., M. Goulian, W. van Wamel, M. Herrmann, S. M. Simon, G. Kaplan, G. Peters, and A. L. Cheung. 2000. Staphylococcus aureus RN6390 replicates and induces apoptosis in a pulmonary epithelial cell line. Infect. Immun. 68:5385-5392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Koenig, R. L., J. L. Ray, S. J. Maleki, M. S. Smeltzer, and B. K. Hurlburt. 2004. Staphylococcus aureus AgrA binding to the RNAIII-agr regulatory region. J. Bacteriol. 186:7549-7555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kornblum, J., B. Kreiswirth, S. J. Projan, H. Ross, and R. P. Novick. 1990. Agr: a polycistronic locus regulating exoprotein synthesis in Staphylococcus aureus, p. 403-420. In R. P. Novick (ed.), Molecular biology of the staphylococci. VCH Publishers, New York, N.Y.
- 29.Kuroda, M., T. Ohta, I. Uchiyama, T. Baba, H. Yuzawa, I. Kobayashi, L. Cui, A. Oguchi, K.-I. Aoki, Y. Nagai, J. Lian, T. Ito, M. Kanamori, H. Matsumaru, A. Maruyama, H. Murakami, A. Hosoyama, Y. Mizutani-Ui, N. K. Takahashi, T. Sawano, R.-I. Inoue, C. Kaito, K. Sekimizu, H. Hirakawa, S. Kuhara, S. Goto, J. Yabuzaki, M. Kanehisa, A. Yamashita, K. Oshima, K. Furuya, C. Yoshino, T. Shiba, M. Hattori, N. Ogasawara, H. Hayashi, and K. Hiramatsu. 2001. Whole genome sequencing of methicillin-resistant Staphylococcus aureus. Lancet 357:1225-1240. [DOI] [PubMed] [Google Scholar]
- 30.Lee, C. Y., S. L. Buranen, and Z. H. Ye. 1991. Construction of single copy integration vectors for Staphylococcus aureus. Gene 103:101-105. [DOI] [PubMed] [Google Scholar]
- 31.Li, R., A. C. Manna, A. L. Cheung, and G. Zhang. 2003. Crystal structure of the SarS protein from Staphylococcus aureus. J. Bacteriol. 185:4219-4225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lim, Y., M. Jana, T. T. Luong, and C. Y. Lee. 2004. Control of glucose and NaCl-induced biofilm formation by rbf in Staphylococcus aureus. J. Bacteriol. 186:722-729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Liu, Y., A. Manna, R. Li, W. E. Martin, R. C. Murphy, A. L. Cheung, and G. Zhang. 2001. Crystal structure of the SarR protein from Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 98:6877-6882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Liu, Y., A. C. Manna, C.-H. Pan, I. A. Kriksunov, D. J. Thiel, A. L. Cheung, and G. Zhang. 2006. Structural and functional analyses of the global regulatory protein SarA from Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 103:2392-2397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lowy, F. 1998. Staphylococcus aureus infections. N. Engl. J. Med. 339:520-532. [DOI] [PubMed] [Google Scholar]
- 36.Luong, T. T., S. W. Newell, and C. Y. Lee. 2003. Mgr, a novel global regulator in Staphylococcus aureus. J. Bacteriol. 185:3703-3710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Luong, T. T., P. M. Dunman, E. Murphy, S. J. Projan, and C. Y. Lee. 2006. Transcription profiling of the mgrA regulon in Staphylococcus aureus. J. Bacteriol. 188:1899-1910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Manna, A., and A. L. Cheung. 2001. Characterization of sarR, a modulator of sar expression in Staphylococcus aureus. Infect. Immun. 69:885-896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Manna, A. C., M. G. Bayer, and A. L. Cheung. 1998. Transcriptional analysis of different promoters in the sar locus in Staphylococcus aureus. J. Bacteriol. 180:3828-3836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Manna, A. C., and A. L. Cheung. 2003. sarU, a sarA homolog, is repressed by SarT and regulates virulence genes in Staphylococcus aureus. Infect. Immun. 71:343-353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Manna, A. C., S. S. Ingavale, M. Maloney, W. van Wamel, and A. L. Cheung. 2004. Identification of sarV (SA2062), a new transcriptional regulator, is repressed by SarA and MgrA (SA0641) and involved in the regulation of autolysis in Staphylococcus aureus. J. Bacteriol. 186:5267-5280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Manna, A. C., and A. L. Cheung. Transcriptional regulation of the agr locus and the identification of DNA binding residues of the global regulatory protein SarR in Staphylococcus aureus. Mol. Microbiol., in press. [DOI] [PubMed]
- 43.McNamara, P. J., K. C. Milligan-Monroe, S. Khalili, and R. A. Proctor. 2000. Identification, cloning, and initial characterization of rot, a locus encoding a regulator of virulence factor expression in Staphylococcus aureus. J. Bacteriol. 182:3197-3203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Morfeldt, E., D. Taylor, A. von Gabain, and S. Arvidson. 1995. Activation of alpha-toxin translation in Staphylococcus aureus by the trans-encoded antisense RNA, RNAIII. EMBO J. 14:4569-4577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Morfeldt, E., K. Tegmark, and S. Arvidson. 1996. Transcriptional control of the agr-dependent virulence gene regulator, RNAIII, in Staphylococcus aureus. Mol. Microbiol. 21:1227-1237. [DOI] [PubMed] [Google Scholar]
- 46.Novick, R. P., H. F. Ross, S. J. Projan, J. Kornblum, B. Kreiswirth, and S. Moghazeh. 1993. Synthesis of staphylococcal virulence factors is controlled by a regulatory RNA molecule. EMBO J. 12:3967-3977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Novick, R. P., S. J. Projan, J. Kornblum, H. F. Ross, G. Ji, B. Kreiswirth, F. Vandenesch, and S. Moghazeh. 1995. The agr P2 operon: an autocatalytic sensory transduction system in Staphylococcus aureus. Mol. Gen. Genet. 248:446-458. [DOI] [PubMed] [Google Scholar]
- 48.Novick, R. P. 2003. Autoinduction and signal transduction in the regulation of staphylococcal virulence. Mol. Microbiol. 48:1429-1449. [DOI] [PubMed] [Google Scholar]
- 49.Pragman, A. A., J. M. Yarwood, T. J. Tripp, and P. M. Schlievert. 2004. Characterization of virulence factor regulation by SrrAB, a two-component system in Staphylococcus aureus. J. Bacteriol. 186:2430-2438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Projan, S. J., and R. P. Novick. 1997. The molecular basis of pathogenicity, p. 55-81. In K. B. Crossley and G. L. Archer (ed.), The staphylococci in human diseases. Churchill Livingston, New York, N.Y.
- 51.Rechtin, T. M., A. F. Gillaspy, M. A. Schumacher, R. G. Brennan, M. S. Smeltzer, and B. K. Hurlburt. 1999. Characterization of the SarA virulence gene regulator of Staphylococcus aureus. Mol. Microbiol. 33:307-316. [DOI] [PubMed] [Google Scholar]
- 52.Riggs, A. D., H. Suzuki, and S. Bourgeois. 1970. lac repressor-operator interaction: equilibrium studies. J. Mol. Biol. 48:67-83. [DOI] [PubMed] [Google Scholar]
- 53.Said-Salim, B., P. M. Dunman, F. M. McAleese, D. Macapagal, E. Murphy, P. J. McNamara, S. Arvidson, T. J. Foster, S. J. Projan, and B. N. Kreiswirth. 2003. Global regulation of Staphylococcus aureus genes by Rot. J. Bacteriol. 185:610-619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Sau, S., L. Sun, and C. Y. Lee. 1997. Molecular characterization and transcriptional analysis of type 8 capsule genes in Staphylococcus aureus. J. Bacteriol. 179:1614-1621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Schmidt, K. A., A. C. Manna, S. Gill, and A. L. Cheung. 2001. SarT, a repressor of α-hemolysin in Staphylococcus aureus. Infect. Immun. 69:4749-4758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Schmidt, K. A., A. C. Manna, and A. L. Cheung. 2003. SarT influences sarS expression in Staphylococcus aureus. Infect. Immun. 71:5139-5148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Tegmark, K., A. Karlsson, and S. Arvidson. 2000. Identification and characterization of SarH1, a new global regulator of virulence gene expression in Staphylococcus aureus. Mol. Microbiol. 37:398-409. [DOI] [PubMed] [Google Scholar]
- 58.Tormo, M. A., M. Marti, J. Valle, A. C. Manna, A. L. Cheung, I. Lasa, and J. R. Penades. 2005. SarA is an essential positive regulator of Staphylococcus epidermidis biofilm development. J. Bacteriol. 187:2348-2356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Trotonda, M. P., A. C. Manna, A. L. Cheung, I. Lasa, and J. R. Penadés. 2005. SarA positively controls Bap-dependent biofilm formation in Staphylococcus aureus. J. Bacteriol. 187:5790-5798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Truong-Bolduc, Q. C., X. Zhang, and D. C. Hooper. 2003. Characterization of NorR protein, a multifunctional regulator of norA expression in Staphylococcus aureus. J. Bacteriol. 185:3127-3138. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.