Abstract
The Rho family of GTP-binding proteins plays a critical role in a variety of cellular processes, including cytoskeletal reorganization and activation of kinases such as p38 and C-jun N-terminal kinase (JNK) MAPKs. We report here that dominant negative forms of Rac1 and Cdc42Hs inhibit the expression of the muscle-specific genes myogenin, troponin T, and myosin heavy chain in L6 and C2 myoblasts. Such inhibition correlates with decreased p38 activity. Active RhoA, RhoG, Rac1, and Cdc42Hs also prevent myoblast-to-myotube transition but affect distinct stages: RhoG, Rac1, and Cdc42Hs inhibit the expression of all muscle-specific genes analyzed, whereas active RhoA potentiates their expression but prevents the myoblast fusion process. We further show by two different approaches that the inhibitory effects of active Rac1 and Cdc42Hs are independent of their morphogenic activities. Rather, myogenesis inhibition is mediated by the JNK pathway, which also leads to a cytoplasmic redistribution of Myf5. We propose that although Rho proteins are required for the commitment of myogenesis, they differentially influence this process, positively for RhoA and Rac1/Cdc42Hs through the activation of the SRF and p38 pathways, respectively, and negatively for Rac1/Cdc42Hs through the activation of the JNK pathway.
INTRODUCTION
The development of skeletal muscle is a multistep process in which pluripotent mesodermal cells give rise to myoblasts that subsequently withdraw from the cell cycle and differentiate into plurinucleated myotubes. The earliest known markers for the skeletal muscle lineage are the four myogenic basic helix-loop-helix (bHLH) factors MyoD, Myf5, myogenin, and MRF4 (Molkentin and Olson, 1996; Arnold and Winter, 1998). During differentiation, myogenic bHLH factors activate muscle-specific genes (such as myosin light and heavy chains, troponin T, etc.) and coordinate withdrawal from the cell cycle (Lassar et al., 1994). In culture, differentiation of myocytes is induced when cells are exposed to medium containing a low concentration of mitogens (differentiation medium). These myogenic bHLH factors are targets of growth factor–signaling pathways that either negatively or positively regulate myogenic differentiation. In vivo experiments in mice have shown that sonic hedgehog and Wnt proteins coming from the dorsal neural tube control Myf5 expression, whereas MyoD is controlled by factors from the dorsal ectoderm (Munsterberg et al., 1995; Cossu et al., 1996; Stern et al., 1997). In addition, negative regulation by proteins such as BMP4, released by the lateral mesoderm, or proteins of the TGFβ and FGF families plays an important role in this process (Pourquie et al., 1996).
The Rho family of Ras-like GTPases are clustered in two distinct subgroups: the Rac/Cdc42 subgroup, which includes Rac1, Rac2, and Rac3, RhoG, Cdc42Hs, TC10, chp, and RhoH, and the Rho subgroup, in which RhoA, -B and -C, RhoD, RhoL, and Rnd1, Rnd2, and Rnd3 are found. Specific substitutions based on Ras studies result in the expression of proteins that are either in a constitutively active GTP-bound (e.g., G12V and Q61L) or dominant negative GDP-bound (e.g., T17N) conformation. Rho GTPases regulate a variety of cytoskeleton-dependent cell functions (Hall, 1998) and have been implicated in various cellular events (Tapon and Hall, 1997; Fort, 1999; Michiels and Collard, 1999). RhoA, Rac1, and Cdc42Hs control signaling pathways that regulate gene transcription. They are required for the serum-induced activation of SRF (serum response factor) (Hill et al., 1995), a transcription factor that binds the CArG box sequence that is found in the promoters of many muscle-specific genes and is essential for their expression (Mohun et al., 1987; Vandromme et al., 1992; Gauthier et al., 1996). Rac1 and Cdc42Hs, but not RhoA (except in 293T cells) (Teramoto et al., 1996), regulate the activation of members of the MAPK group, the C-jun N-terminal kinases (JNKs)/stress-activated protein kinases and p38 kinases (Coso et al., 1995; Minden et al., 1995). Recent studies show that p38 activity is required for C2C12 myoblast differentiation (Cuenda and Cohen, 1999; Zetser et al., 1999).
The role of Rho GTPases, and particularly the Rac/Cdc42Hs subgroup, in skeletal muscle differentiation is not well understood. Recent reports have revealed that the RhoA/SRF signaling cassette controls MyoD expression and skeletal muscle differentiation without affecting Myf5 expression (Carnac et al., 1998; Takano et al., 1998; Wei et al., 1998). In contrast, constitutively activated Rac1 does not appear to affect MyoD activity (Ramocki et al., 1997). Because Northern blot analysis revealed strong expression of Rac1 and Cdc42Hs mRNAs in muscle, we studied their role in the skeletal muscle differentiation of rat L6 and mouse C2 myoblasts. First, we show that the activity of Rac1 and Cdc42Hs GTPases is required for myogenesis, because expression of dominant negative forms of these GTPases resulted in total inhibition of the expression of the myogenic factors myogenin, troponin T, and myosin heavy chain (MHC) and led to strong inhibition of p38 activity. Second, we demonstrate that active Rac1 and Cdc42Hs inhibit the expression of myogenin, troponin T, and MHC, whereas expression of active RhoA potentiates SRF-dependent myogenin and troponin T expression. We further show by two different approaches that inhibition of skeletal muscle differentiation by active Rac1 and Cdc42Hs is mediated by the JNK pathway, the activation of which also leads to a cytoplasmic distribution of Myf5.
MATERIALS AND METHODS
Northern Blot Analysis
Membranes containing poly(A)+ RNAs isolated from various human tissues (Multiple Tissue Northern, Clontech, Palo Alto, CA) were hybridized for 12–24 h in a mixture containing 50% formamide, 5× Denhardt's solution, 10 mM PO4 buffer, pH 7, 0.75 M NaCl, 0.1% SDS, 10% dextran sulfate, and 100 μg/ml denatured salmon sperm DNA at 42°C with the use of 32P-labeled nick-translated probes (106 cpm/ml). Filters were washed twice in 2× SSC–0.1% SDS (1× SSC is 0.15 M NaCl plus 0.015 M sodium citrate) at room temperature and once in 0.2× SSC–0.1% SDS at 65°C.
Establishment of Stable Cell Lines
G418-resistant GP+E-86 clones expressing constitutively activated forms of Rac1, RhoG, Cdc42Hs, or RhoA were grown to collect retrovirus-containing cell-free supernatants (Roux et al., 1997). Infection was performed on exponentially growing L6 or C2.7 myoblasts (5 × 105 cells per 60-mm dish) with the use of 5 ml of viral supernatant (105 to 5 × 105 colony-forming units/ml). Twenty-four hours later, cells were split and cultured in medium containing G418 (1 mg/ml). After 10 d of selection, resistant colonies were collected, tested for GTPase expression, and used for experiments. Cells were grown continuously in G418.
Cell Culture and Drug Treatment
L6 rat myoblasts (Vandromme et al., 1992) were grown in DMEM/HAM F-12 (ratio 1:1) supplemented with 10% FCS (BioMedia, Boussens, France). To induce differentiation, growth medium was replaced with differentiation medium consisting of DMEM/HAM F-12 supplemented with 2% FCS. Stable cell lines derived from L6 myoblasts were cultured under the same conditions in medium supplemented with 1 mg/ml G418. C2.7 and C2C12 mouse myoblasts were grown in DMEM supplemented with 10% FCS. Differentiation was triggered by replacing growth medium with DMEM containing 2% FCS. Anisomycin was used at a concentration of 5 nM, and SB 203580 was used at a concentration at 10 μM. Control cells were incubated with the same volume of DMSO. Drug-containing medium was replaced every 12 h.
Plasmids and Transient Transfection
Cells were transfected with GFP-Rac1V12, GFP-RhoGV12, GFP-Cdc42HsV12, GFP-RhoAV14 (Gauthier et al., 1998), Myc-Rac1L61Y40, Myc-Rac1L61F37, Myc-Cdc42HsL61Y40, Myc-Cdc42HsL61F37 (provided by A. Hall, NRC Laboratory for Molecular Cell Biology, London, UK) or FLAG-MKK7 (provided by R. Davis, University of Massachusetts, Worcester, MA) and HA-MKK3 (provided by B. Derijard, Centre National de la Recherche Scientifique, Nice, France) with the use of the lipofectamine method (1 μg of plasmid DNA per 35-mm dish) as described by the supplier (Life Technologies, Gaithersburg, MD). Four hours after transfection, the medium was replaced with DMEM/HAM F-12 supplemented with 10% FCS for 4 h. Growth medium was then replaced with differentiation medium, and cells were fixed at different times thereafter and processed for immunohistochemistry.
Immunofluorescence
Cells growing in 35-mm dishes were fixed in 3.7% formaldehyde in PBS followed by 5 min of permeabilization in 0.1% Triton X-100 in PBS and incubated in PBS containing 0.1% BSA. Myogenin expression was visualized with a 60-min incubation with a rabbit polyclonal anti-myogenin antibody (1:30 dilution; Tebu) followed by incubation with a fluorescein-conjugated goat anti-rabbit antibody (1:50 dilution; Cappel-ICN, Costa Mesa, CA). MHC expression was detected with the use of a mouse monoclonal anti-MHC antibody (1:100 dilution; Sigma Chemical, St. Louis, MO), followed by incubation with fluorescein-conjugated anti-mouse antibody (1:50 dilution; Cappel-ICN). Expression of Green Fluorescent Protein (GFP)-tagged proteins was visualized directly. Expression of Myc epitope–tagged Rac1 and Cdc42Hs mutants was detected with a 60-min incubation with the 9E10 anti-Myc antibody (a gift from C. Lambert and D. Mathieu, Centre National de la Recherche Scientifique, Montpellier, France), expression of hemagglutinin (HA)-tagged proteins was monitored with a mouse monoclonal anti-HA antibody (12CA5), and expression of FLAG-tagged proteins was assessed with a mouse anti-FLAG antibody (1:200 dilution; Sigma) followed by incubation with a TRITC-conjugated anti-mouse antibody (1:20 dilution) (Cappel-ICN). Myf5 protein was detected with the C-terminal polyclonal rabbit anti-Myf5 antibody (1:200 dilution; Yablonka-Reuveni et al., 1999) followed by biotin-conjugated anti-rabbit antibody (1:200 dilution; Amersham, Arlington Heights, IL) and streptavidin–Texas red (1:200 dilution; Amersham). Actin filaments were stained with rhodamine-conjugated phalloidin (0.5 U/ml; Sigma). Nuclei were stained with Hoechst stain (0.1 μg/ml; Sigma) applied just before cells were mounted in Mowviol (Aldrich, Milwaukee, WI) and observed under a DMR Leica microscope (Rueil-Nalmaison, France) with the use of 40× or 63× planapochromat lenses. Images thus obtained were captured with an ORCA 100 (B/W) 10-bit cooled charge-coupled device camera (C mount 1×), a C 4742-95 controller, and an HIPIC controller program run on a personal computer (Hamamatsu, Hamamatsu City, Japan). Images were saved in TIFF format (8 bits) for processing in Adobe (Mountain View, CA) Photoshop and assembled in Adobe Illustrator.
Scanning Electron Microscopy
Parental L6 myoblasts and stable L6 RhoGV12, L6 Rac1V12, L6 Cdc42HsV12, and L6 RhoAV14 were fixed and processed as described previously for scanning electron microscopy (SEM) analysis (Gauthier et al., 1998). Cells were observed with the use of a Hitachi (Tokyo, Japan) S4000 scanning microscope.
Gel Electrophoresis and Immunoblotting
Cells cultured in 60-mm dishes were rinsed in cold PBS and lysed in Laemmli sample buffer (40 mM Tris-HCl, pH 6.8, 5 mM DTT, 1% SDS, 7.5% glycerol, 0.01% bromphenol blue) added to the dish. After scraping and boiling, samples (100 μg of protein) were loaded on a 12.5% polyacrylamide gel and transferred onto nitrocellulose. Membranes were saturated in 5% milk in PBS containing 0.1% Tween-20 for 1 h and subsequently incubated with different primary antibodies. The following antibodies were used: mouse monoclonal anti-Rac1 antibody (dilution 1:50; Upstate Biotechnology, Euromedex, Mundolsheim, France); anti-Myc antibody from mouse ascites (dilution 1:1000; B. Rouault, Centre National de la Recherche Scientifique, Montpellier, France); mouse 12CA5 anti-HA antibodies (dilution 1:10); mouse monoclonal anti-troponin T antibody (dilution 1:100; Sigma); mouse monoclonal anti-α-tubulin antibody (dilution 1:10; P. Mangeat, Centre National de la Racherche Scientifique, Montpellier, France); and rabbit polyclonal anti-Myf5 antibody (dilution 1:500; Yablonka-Reuveni et al., 1999). Membranes were washed and incubated with peroxidase-conjugated anti-mouse or anti-rabbit secondary antibodies (dilutions of 1:2000 or 1:5000, respectively; Amersham). After washing, membranes were incubated with chemiluminescence reagents (Renaissance, New England Nuclear, Boston, MA). Autoradiographs were scanned, and images obtained in Adobe Photoshop were assembled in Adobe Illustrator.
Kinase Assay
Parental L6, L6 RhoGV12, L6 Rac1V12, L6 Cdc42HsV12, and L6 RhoAV14 cells cultured in 60-mm dishes were transfected or cotransfected with 2 μg of HA-JNK, HA-p38 (provided by B. Derijard), GFP-RhoGV12, GFP-Rac1V12, GFP-Cdc42HsV12, GFP-Rac1L61, GFP-Rac1L61F37A, GFP-Rac1L61Y40C, GFP-Cdc42HsL61, GFP-Cdc42HsL61F37A, GFP-Cdc42HsL61Y40C, GFP-RhoAV14, FLAG-MKK7, or HA-MKK3 plasmids with the use of the lipofectamine method as described by the supplier (Life Technologies). Four hours after the transfection, the medium was replaced with DMEM/HAM F-12 supplemented with 10% FCS. Twenty-four hours after transfection, cells were washed twice in cold PBS and then solubilized in a Triton lysis buffer containing 20 mM Tris (pH 7.5), 137 mM NaCl, 2 mM EDTA, 1% Triton X-100, 25 mM β-glycerophosphate, 1 mM sodium orthovanadate, 2 mM NaPPi, 10% glycerol, 1 mM PMSF, and 10 μg/ml leupeptin. Soluble extracts were obtained by centrifugation at 12,000 rpm for 30 min at 4°C. Cell extracts were then split into two fractions: the first to test HA-JNK, HA-p38, or GTPase expression, and the second to assay for JNK or p38 kinase activities as follows. The extracts were immunoprecipitated by incubation with the anti-HA antibody 12CA5 (dilution 1:50) and 20 μl of protein G–Sepharose (Pharmacia Biotech, Piscataway, NJ) for 2 h at 4°C. Immunoprecipitates were then washed twice with the Triton lysis buffer and twice with a kinase buffer containing 25 mM HEPES (pH 7.5), 25 mM MgCl2, 25 mM β-glycerophosphate, 2 mM DTT, and 0.1 mM sodium orthovanadate. The kinase assay was performed by incubating the immunoprecipitates for 20 min at 30°C in the presence of 50 μM ATP and 2 μCi of [γ32P]ATP with 3 μg of GST–c-JUN for the JNK assay and 3 μg of GST–ATF2 for the p38 assay. The reactions were stopped by adding 5× Laemmli buffer. Samples were loaded onto a 15% polyacrylamide gel, transferred onto nitrocellulose, analyzed with a Phosphoimager (Molecular Dynamics, Sunnyvale, CA), and normalized as a function of the expression of the various proteins.
RESULTS
The Activities of Rac1 and Cdc42Hs GTPases Are Required for Skeletal Muscle Differentiation
As a prerequisite, we first analyzed the level of Rac1 and Cdc42Hs GTPase mRNA expression in several human organs. Membranes containing poly(A)+ RNAs isolated from various human tissues (Multiple Tissue Northern, Clontech) were probed for Rac1 and Cdc42Hs mRNAs (Figure 1). As described previously (Didsbury et al., 1989), two Rac1 mRNA species of 1.1 and 2.4 kilobases were detected and two Cdc42Hs mRNAs of 1.9 and 2.5 kilobases were detected. Membranes were also probed for S26 mRNA to estimate the amounts of loaded RNAs (Vincent et al., 1993). Although ubiquitously expressed, a high level of both Rac1 and Cdc42Hs mRNAs was found in skeletal and cardiac muscle.
Figure 1.
Rac1 and Cdc42Hs mRNA distribution. Northern blot membrane containing 2 μg of poly(A)+ RNA from various human tissues was probed for Rac1 and Cdc42Hs GTPases. RNA marker sizes (in kilobases) are indicated.
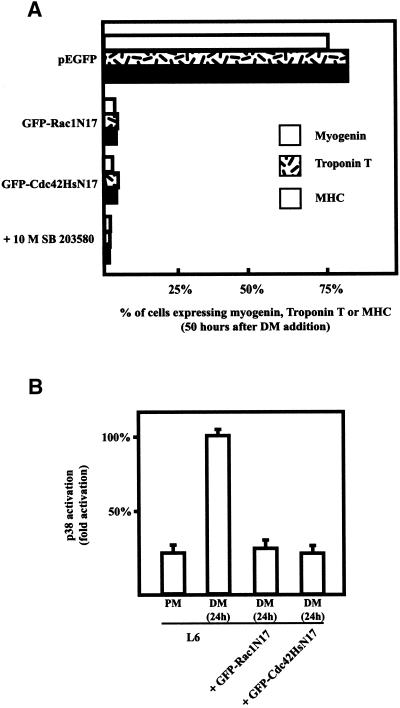
To examine whether theses GTPases are involved in the process of skeletal muscle differentiation, we transfected L6 and C2 myoblasts with GFP-tagged constructs expressing the dominant negative (DN) T17N mutants of Rac1 and Cdc42Hs GTPases. Cells were induced to differentiate by replacing the growth medium (GM) with differentiation medium (DM), fixed, and analyzed for expression of myogenin, troponin T, and MHC, three muscle-specific proteins (Figure 2A). Myogenin is a transcription factor expressed before the establishment of the postmitotic state that has been shown to be one of the earliest known markers for myoblasts committed to myogenesis (Wright et al., 1989; Andres and Walsh, 1996). Troponin T and MHC are major components of the contractile apparatus in muscle fibers and are expressed only several hours after the onset of differentiation. Fifty hours after DM addition, myogenin, troponin T, and MHC proteins were expressed in ∼75% of parental L6 cells and control L6 cells transfected with plasmid encoding GFP alone. In contrast, although they exhibited unaltered morphology, more than 95% of cells transfected with the DN Rac1 and Cdc42Hs mutants failed to express myogenin, troponin T, and MHC. Similar results were obtained with transfected C2 cells (our unpublished results). Expression of DN Rac1 and Cdc42Hs did not affect α-tubulin expression, discounting a deleterious effect of these GTPase mutants (our unpublished results). These data indicate that Rho-dependent pathways are required during the early phases of myogenic differentiation. In addition to the control of cytoskeletal structures, Rho GTPases have been shown to regulate the JNK and p38 MAPK pathways (Coso et al., 1995; Minden et al., 1995). Interestingly, p38 activity was recently shown to be essential for C2C12 myoblast differentiation (Cuenda and Cohen, 1999; Zetser et al., 1999). We first assessed whether p38 inhibition had a similar effect on L6 cells. As shown in Figure 2A, the specific inhibition of p38 by SB 203580 totally blocked L6 myoblast differentiation. We next analyzed the effects of DN Rac1 and Cdc42Hs mutants on p38 activity (Figure 2B). L6 cells were transiently cotransfected with GFP-tagged expression plasmids for Rac1N17, Cdc42HsN17, and HA-tagged p38, and anti-HA immunoprecipitates were assayed for kinase activity. As reported previously (Cuenda and Cohen, 1999), we observed a threefold increase in p38 activity 24 h after culture in differentiating conditions. In contrast, no changes in p38 activity were detected under identical conditions when Rac1N17 and Cdc42HsN17 were expressed. Together, these data show that the inhibition of Rac1 and Cdc42Hs GTPase activities is associated with a down-regulation of p38 kinase activity and a block in skeletal muscle differentiation.
Figure 2.
Rac1 and Cdc42Hs activity and myogenesis. (A) L6 myoblasts were transfected with constructs expressing GFP-tagged Rac1N17 and Cdc42HsN17 and shifted to DM for 50 h. Cells were then fixed and stained for myogenin, troponin T, and MHC expression. L6 myoblasts were also switched to DM containing 10 μM SB 203580. The histogram represents the percentages of myogenin-, troponin T-, or MHC-positive cells and summarizes the data from five independent sets of experiments; 40–50 cells were analyzed in each experiment. (B) L6 myoblasts were cotransfected with HA-p38 and constructs expressing GFP alone as a control, GFP-tagged Rac1N17, or GFP-tagged Cdc42HsN17. p38 activity in the cell lysates was measured as described in MATERIALS AND METHODS from L6 cells in proliferation medium (PM) or L6 cells expressing or not expressing GFP-Rac1N17 or GFP-Cdc42HsN17 in differentiation medium (DM) for 24 h. The results are presented as averages from three independent experiments.
Establishment of Stable L6 Myoblast Cell Lines Expressing Constitutively Active Forms of RhoG, Rac1, Cdc42Hs, and RhoA GTPases
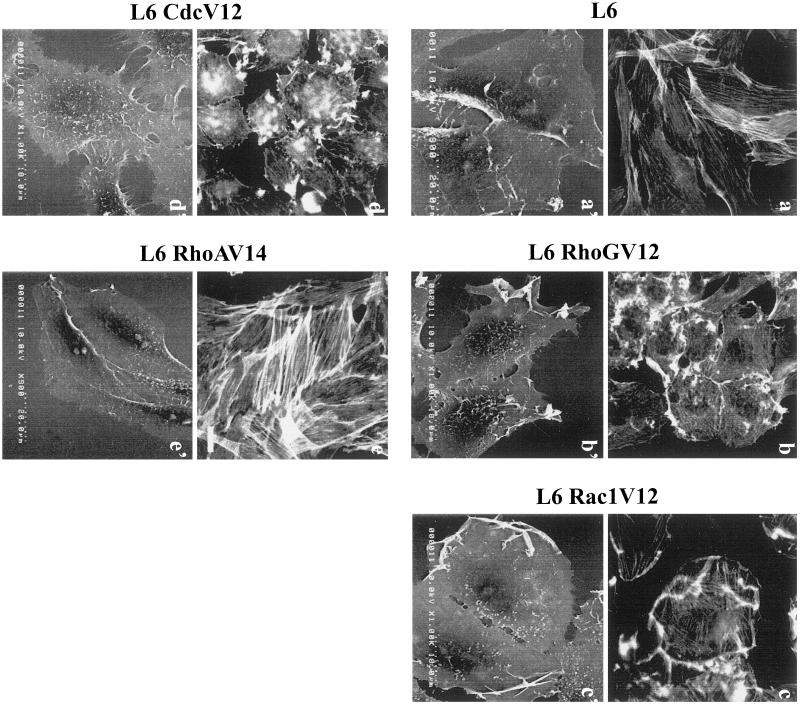
Because the inhibition of Rac1 and Cdc42Hs activities impaired myogenesis of L6 cells, we next investigated the effects of constitutively active Rac1 and Cdc42Hs proteins. L6 myoblastic cells were stably infected with retroviral vectors encoding either HA-tagged Rac1V12 or Myc-tagged Cdc42HsV12. As controls, we also established L6 cells stably expressing RhoGV12 or RhoAV14. RhoG has been shown to act upstream of Rac1 and Cdc42Hs (Roux et al., 1997; Gauthier et al., 1998), whereas RhoA controls distinct signaling pathways (Tapon and Hall, 1997; Sahai et al., 1998). The introduction of the G12V (G14V for RhoA) or Q61L substitution has been described extensively as leading to constitutively active GTP-bound proteins (for review, see Hall, 1998). Expression of activated Rho proteins was monitored by examining the modifications in F-actin–containing structures controlled by each of these GTPases, with the use of rhodamine-labeled phalloidin staining, and in cell plasma membranes, with the use of SEM. F-actin staining and SEM analysis of growing parental L6 cells (i.e., noninfected cells or cells infected with wild-type virions) are shown in Figure 3, a and a′. RhoGV12-expressing cells (Figure 3, b and b′) presented the expected modifications in polymerized actin-containing structures and cell surface morphology, i.e., formation of ruffles, lamellipodia, filopodia, microvilli, and partial disassembly of actin stress fibers (Gauthier et al., 1998). Similarly, Rac1V12-expressing cells (Figure 3, c and c′) displayed ruffles and lamellipodia and partial disassembly of actin stress fibers (Ridley et al., 1992), whereas Cdc42- HsV12-expressing cells (Figure 3, d and d′) showed radial extensions called filopodia, microvilli, and partial disassembly of actin stress fibers (Kozma et al., 1995; Nobes and Hall, 1995b). Although RhoAV14-expressing cells had increased actin stress fibers (Figure 3e) compared with parental L6 cells (Figure 3a), no significant modifications of cell surface morphology were detected by SEM (Figure 3, compare a′ and e′) (Ridley and Hall, 1992).
Figure 3.
Analysis of RhoGV12, Rac1V12, Cdc42HsV12, and RhoAV14 GTPase expression in stable L6 cell lines. Proliferative parental L6 (a and a′), L6 RhoGV12 (b and b′), L6 Rac1V12 (c and c′), L6 Cdc42HsV12 (d and d′), and L6 RhoAV14 (e and e′) cells were fixed and then stained for filamentous actin distribution with rhodamine-labeled phalloidin (a, b, c, d, and e) or processed for SEM (a′, b′, c′, d′, and e′). For each panel, cells shown are representative of more than 100 observed cells. Bar, 10 μm.
Morphological Analysis of Active Rho-expressing Cells during the Course of Differentiation
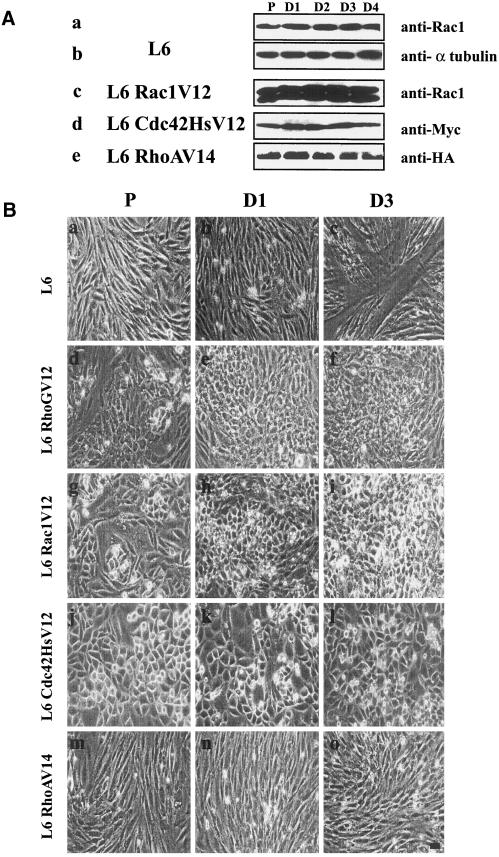
Parental L6, L6 RhoGV12, L6 Rac1V12, L6 Cdc42HsV12, and L6 RhoAV14 myoblasts were next induced to differentiate by replacing the GM with the appropriate DM. Expression of the various GTPases and morphological differentiation were monitored at different times after the shift to DM. Figure 4A shows an immunoblot analysis of endogenous Rac1 in parental L6 cells during the course of differentiation (row a) as well as both endogenous (row c, lower bands) and HA-tagged (row c, upper bands) Rac1 proteins in L6 Rac1V12 cells. The Rac1 protein level remained constant during the course of differentiation, as did the expression of Myc-Cdc42HsV12 (row d) and HA-RhoAV14 (row e) proteins. In this experiment, α-tubulin was monitored as an internal loading control (row b).
Figure 4.
Effect of RhoGV12, Rac1V12, Cdc42HsV12, and RhoAV14 expression on myogenesis. (A) Protein extracts from parental L6 (a and b), L6 Rac1V12 (c), L6 Cdc42HsV12 (d), and L6 RhoAV14 (e) (100 μg/well) collected at the indicated periods (P, proliferative; D1, D2, D3, and D4, 1 to 4 d after addition of differentiation medium) were immunoblotted with anti-Rac1 (a and c), anti-α-tubulin (b), anti-Myc (d), or anti-HA (e) antibody as described in MATERIALS AND METHODS. (B) Proliferating parental L6 and L6 RhoGV12, L6 Rac1V12, and L6 Cdc42HsV12 myoblasts were grown to ∼80% and then induced to differentiate in DM. Shown are phase-contrast images of parental L6 myoblasts (a–c), L6 RhoGV12 (d–f), L6 Rac1V12 (g–i), L6 Cdc42HsV12 (j–l), and L6 RhoAV14 cells (m–o). Panels a, d, g, j, and m, cells in proliferating medium; panels b, e, h, k, and n, 1 d after addition of DM; panels c, f, i, l, and o, 3 d after addition of DM. For each panel, cells shown are representative of more than 100 observed cells. Bar, 10 μm.
The morphological analysis of these cell lines shown in Figure 4B demonstrates that 1 d after addition of DM, L6 parental cells were regularly aligned (panel b) and myotube formation was observed after 3 d (panel c). In sharp contrast, L6 RhoGV12, L6 Rac1V12, and L6 Cdc42HsV12 cells in either GM (panels d, g, and j, respectively) or DM (panels e, h, and k, respectively) were densely packed and adopted a rounded shape. No myotube formation was detected 3 d after DM addition (panels f, i, and l, respectively). Similar inhibition of myotube formation was observed in C2 myogenic cells transfected with RhoGV12, Rac1V12, and Cdc42V12 vectors (our unpublished results). No myotubes were observed even after 7 d in DM (our unpublished results), indicating that myoblast-to-myotube transition is efficiently blocked and not simply delayed. Although L6 RhoAV14 cells have a morphology similar to that of parental L6 cells in GM (panel m) or DM (panel n) conditions, no myotubes were detected after 3 d (panel o).
Active RhoG, Rac1, and Cdc42Hs GTPases Inhibit the Expression of Muscle-specific Proteins
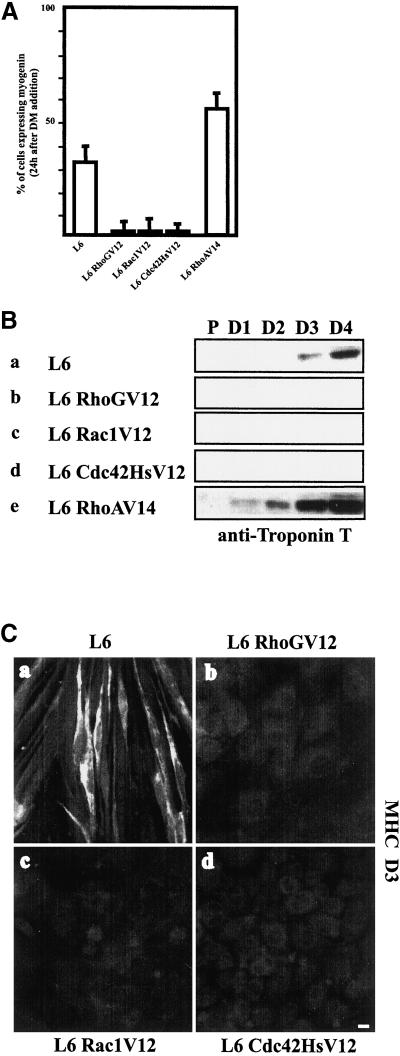
We next examined whether expression of Rho GTPases could affect the regulation of myogenin, troponin T, and MHC (Figure 5). Nuclear myogenin was detected in ∼30–40% of L6 cells after 1 d in DM but in less than 1% of L6 RhoGV12, L6 Rac1V12, and L6 Cdc42HsV12 cells (Figure 5A). After 7 d in DM, myogenin expression was still not detectable in L6 RhoGV12, L6 Rac1V12, and L6 Cdc42HsV12 cells (our unpublished results). In contrast, myogenin expression was increased in L6 RhoAV14 cells (50–60% of myogenin-expressing cells versus 30–40% in parental L6 cells after 1 d in DM) by an SRF-dependent pathway (our unpublished results). The effects of RhoGV12, Rac1V12, and Cdc42HsV12 on myogenin expression were also monitored by transient transfection of L6 or C2 cells, and as in stable cell lines, expression of these GTPases prevented myogenin expression (our unpublished results). In contrast, active Rho GTPases did not affect α-tubulin expression level (our unpublished results). Troponin T expression was monitored by Western blot analysis during the course of differentiation (proliferative myoblasts and after 1–4 d in DM) of parental L6 and RhoGV12-, Rac1V12-, Cdc42HsV12-, and RhoAV14-expressing cells (Figure 5B). As expected, troponin T protein was not detected in proliferative L6 myoblasts and was increased 3–4 d after differentiation (Figure 5B, row a). In contrast, in L6 RhoGV12, L6 Rac1V12, and L6 Cdc42HsV12 cells, no troponin T expression was detected even after 4 d in DM (Figure 5B, rows b, c, and d). This lack of troponin T expression was also observed for all three GTPases by immunocytochemistry on either stable or transiently L6- or C2-transfected cells (our unpublished results). In contrast, troponin T expression was accelerated in RhoAV14-transfected cells, in which it was readily detected after only 1 d in DM (Figure 5B, row e). Similar changes in MHC expression were observed (Figure 5C). MHC was detected in differentiating myoblasts (Figure 5C, panel a) but was not expressed in L6 RhoGV12 (Figure 5C, panel b), L6 Rac1V12 (Figure 5C, panel c), or L6 Cdc42HsV12 (Figure 5C, panel d) cells even after 7 d in DM (our unpublished results). As with Troponin T, MHC appeared earlier in L6 RhoAV14 cells (our unpublished results).
Figure 5.
Effect of RhoGV12, Rac1V12, Cdc42HsV12, and RhoAV14 on muscle-specific protein expression. (A) Proliferating parental L6 and L6 RhoGV12, L6 Rac1V12, L6 Cdc42HsV12, and L6 RhoAV14 myoblasts were grown to ∼80% and shifted to DM for 1 d. Cells were then fixed and stained for myogenin expression and nuclei with Hoechst. The histogram represents the percentage of myogenin-positive cells and summarizes the data from five independent sets of experiments; 40–50 cells were analyzed in each experiment. (B) Protein extracts from parental L6, L6 RhoGV12, L6 Rac1V12, L6 Cdc42HsV12, and L6 RhoAV14 (100 μg/well) collected at the indicated periods (P, proliferative; D1, D2, D3, D4, 1–4 d after addition of differentiation medium) were immunoblotted with an anti-troponin T antibody as described in MATERIALS AND METHODS. Shown are Western blots indicating the expression of troponin T in parental L6 (a), L6 RhoGV12 (b), L6 Rac1V12 (c), L6 Cdc42HsV12 (d), and L6 RhoAV14 (e). (C) Proliferating parental L6 (a) and L6 RhoGV12 (b), L6 Rac1V12 (c), and L6 Cdc42HsV12 (d) myoblasts were grown to ∼80% and shifted to DM for 3 d. Cells were then fixed and stained for MHC expression. For each panel, cells shown are representative of five independent experiments with more than 100 observed cells. Bar, 10 μm.
Together, these data indicate that, as observed previously for the DN mutants, the activation of Rac1 and Cdc42Hs inhibits the early stages of myogenic differentiation. In contrast, the activation of RhoA potentiates the differentiation process at the transcriptional level.
Inhibition of Myogenesis by RhoG, Rac1, and Cdc42Hs Correlates with JNK Activation
We next investigated which pathways controlled by Rac1 and Cdc42Hs might be responsible for myogenesis inhibition. Having already demonstrated that these GTPases induce the expected morphogenic modifications in L6 myoblasts (see Figure 3), we next examined the levels of JNK and p38 MAPK activity (Figure 6A). Parental L6 as well as L6 RhoGV12, L6 Rac1V12, L6 Cdc42HsV12, and L6 RhoAV14 were transfected with HA-tagged JNK or HA-tagged p38, and anti-HA immunoprecipitates were assayed for kinase activity. In agreement with previously published data, expression of RhoG, Rac1, and Cdc42Hs led to a threefold to fourfold increase in both JNK (Figure 6A) and p38 activity (our unpublished results), whereas RhoAV14 had no effect (Coso et al., 1995; Minden et al., 1995; Roux et al., 1997).
Figure 6.
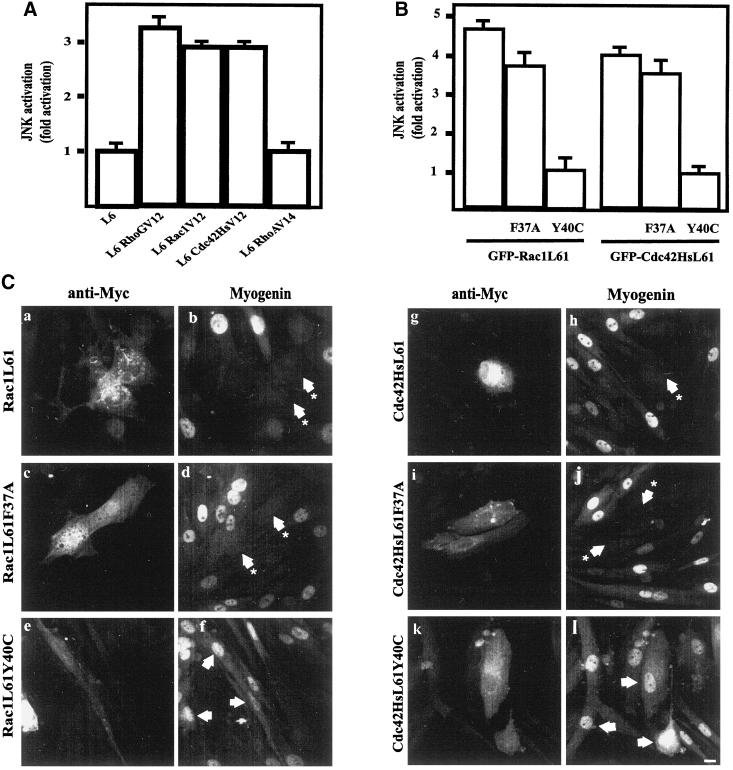
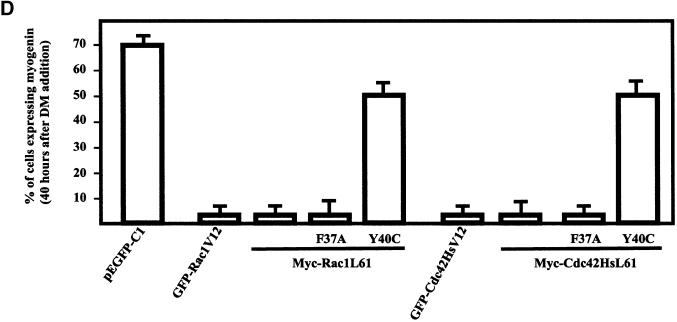
Effect of Rac1 and Cdc42Hs effector loop mutants on myogenin expression. (A) Parental L6, L6 RhoGV12, L6 Rac1V12, L6 Cdc42HsV12, and L6 RhoAV14 myoblasts were transfected with HA-tagged JNK. As a control, cells were transfected with empty vector (pCDNA3). JNK activities in the cell lysates were measured by immunocomplex kinase activity with the use of GST–c-Jun as a substrate, as described in MATERIALS AND METHODS. The results are presented as averages for three independent experiments. (B) L6 myoblasts were cotransfected with HA-JNK and GFP-Rac1L61, GFP-Rac1L61F37A, GFP-Rac1L61Y40C, GFP-Cdc42HsL61, GFP-Cdc42HsL61F37A, or GFP-Cdc42HsL61Y40C. JNK activity in the cell lysates was measured as described in MATERIALS AND METHODS. The results are presented as averages for three independent experiments. (C) L6 myoblasts were transfected with constructs expressing Myc-Rac1L61 (a and b), Myc-Rac1L61F37A (c and d), Myc-RacL61Y40C (e and f), Myc-Cdc42HsL61 (g and h), Myc-Cdc42HsL61F37A (i and j), or Myc-Cdc42HsL61Y40C (k and l). After transfection, cells were induced to differentiate by the addition of DM, fixed 1–2 d later, and processed for Myc epitope detection (a, c, e, g, i, and k) and myogenin expression (b, d, f, h, j, and l). For each panel, cells shown are representative of five independent experiments with more than 100 observed cells. Bar, 10 μm. (D) Summary of the quantification for myogenin expression in cells transfected with empty pEGFP-C1, GFP-tagged Rac1V12 or Cdc42HsV12, or Myc-tagged Rac1L61, Rac1L61F37A, Rac1L61Y40C, Cdc42HsL61, Cdc42HsL61F37A, or Cdc42HsL61Y40C.
To discriminate between morphogenic activities and MAPK activation, we next used effector loop mutants of Rac1 and Cdc42Hs GTPases, which have been shown to differentially bind to and activate downstream effectors (Lamarche et al., 1996). The Y40C mutants of Rac1 and Cdc42Hs have lost their ability to interact with some CRIB (Cdc42Hs- or Rac-interacting Binding) motif–containing proteins, and although they are less efficient in activating PAK (p21 activated kinase) and JNK activity in COS-7 cells, they still induce membrane ruffling and filopodia formation in Swiss 3T3 cells, respectively. Conversely, the F37A mutants of Rac1 and Cdc42Hs still bind CRIB motif–containing proteins and activate PAK and JNK but are less efficient at inducing membrane ruffling and filopodia formation, respectively. We first investigated whether the double Rac1 and Cdc42Hs mutants produced the same differential effects in our cell system (Figure 6B). L6 cells were transiently cotransfected with expression plasmids for Rac1L61, Rac1L61F37A, Rac1L61Y40C, Cdc42HsL61, Cdc42HsL61F37A, Cdc42HsL61Y40C, and HA-tagged JNK, and anti-HA immunoprecipitates were assayed for kinase activity. Whereas the L61F37A mutants of Rac1 and Cdc42Hs activated JNK as efficiently as the single L61 mutant, no JNK activation was detected after expression of L61Y40C mutants in L6 cells, in agreement with previous observations in COS-7 cells (Joneson et al., 1996; Lamarche et al., 1996; Westwick et al., 1997). We next investigated the effects of the double mutants on myogenic differentiation. L6 cells were transiently transfected and induced to differentiate by the addition of DM. Cells were fixed 1–2 d thereafter and analyzed for myogenin expression (Figure 6, C and D). In cells expressing the constitutively active Myc-tagged Rac1L61 (Figure 6, panel a) or Rac1L61F37A (panel c), myogenin expression was detected in <2% of cells (panels b and d), whereas myogenin was expressed in 70% of parental cells or cells transfected with empty pEGFP-C1. Under the same conditions, expression of Myc-tagged Rac1L61Y40C (panel e) did not significantly impair myogenin expression, which occurred in 50% of transfected cells (panel f). Similar to that of Rac1 mutants, expression of Myc-Cdc42HsL61 (panel g) or Myc-Cdc42HsL61F37A (panel i) impaired myogenin expression (panels h and j), whereas expression of Myc-Cdc42HsL61Y40C had no pronounced inhibitory effect (panels k and l).
These data indicate that changes in cell morphology are not responsible for the early inhibition of skeletal muscle differentiation and, furthermore, suggest a role of JNK activation by Rac1 and Cdc42Hs GTPases in this process.
Specific JNK Activation by the MAPK MKK7 Impairs Myogenesis
To demonstrate the direct involvement of the JNK pathway in the inhibition of skeletal muscle differentiation induced by Rac1 and Cdc42Hs, we first treated parental L6 myoblasts with anisomycin, a drug that activates both JNK and p38 (Figure 7A) (Cano et al., 1994). Anisomycin was added 12 h before the addition of DM and included in the culture medium for 1–3 d. Cells were then fixed and analyzed for myogenin expression (Figure 7B). Whereas myogenin expression was detected in almost 40% of cells after 1 d in DM (Figure 7B, panel a), anisomycin treatment completely suppressed myogenin expression (panel c). DNA staining with Hoechst is shown in panels b and d. Similar inhibition of myogenin expression was obtained in C2 myoblasts after anisomycin treatment (our unpublished results).
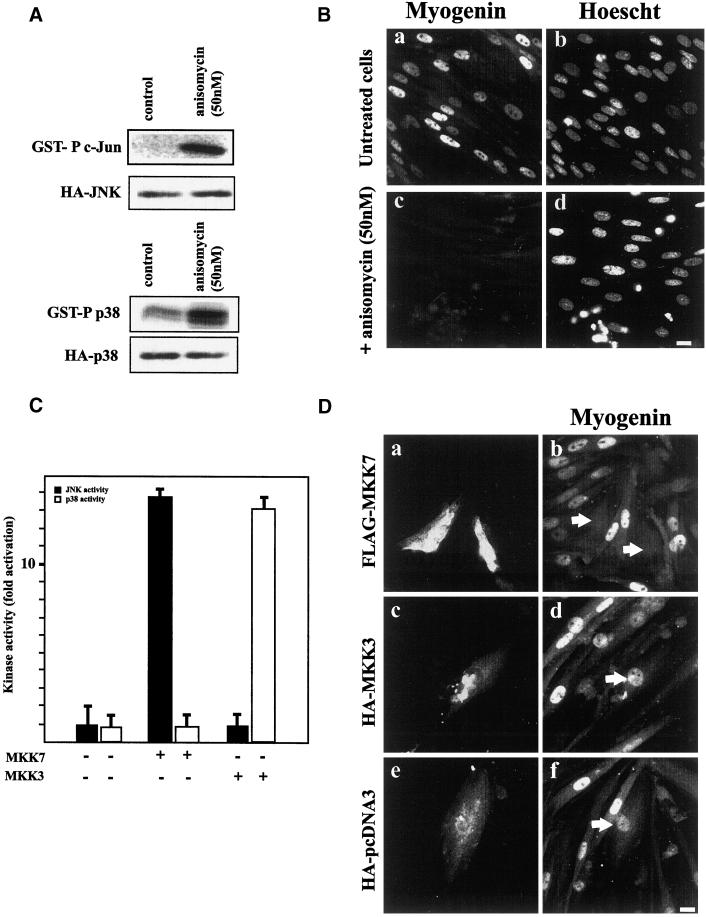
Figure 7.
JNK activation impairs myogenin expression. (A) L6 myoblasts were transfected with HA-tagged JNK or HA-tagged p38. Four hours after transfection, cells were treated with 50 ng of anisomycin for 1 d. JNK or p38 activities in the cell lysate were measured by immunocomplex kinase activity with the use of GST–c-jun or GST–ATF2 as a substrate, respectively. HA-JNK and HA-p38 expression were controlled by Western blot analysis with the use of an anti-HA antibody. (B) L6 myoblasts were treated with anisomycin (50 ng) and then induced to differentiate by the addition of DM. One day later, cells were fixed and stained for myogenin expression (a and c) and for DNA with Hoechst dye (b and d). Bar, 10 μm. (C) L6 myoblasts were cotransfected with FLAG-tagged MKK7 or FLAG-tagged MKK3 and either HA-JNK or HA-p38. JNK and p38 activities in the cell lysate were measured by immunocomplex kinase activity with the use of GST–c-jun or GST–ATF2 as a substrate. The results are presented as averages of three independent experiments. (D) L6 myoblasts were transfected with constructs expressing FLAG-MKK7 (a and b), HA-MKK3 (c and d), or empty HA-tagged pcDNA3 (e and f). After transfection, cells were induced to differentiate by the addition of DM, fixed 1.5 d later, and processed for FLAG (a) or HA epitope (c and e) detection and myogenin expression (b, d, and f). For each panel, cells are representative of three independent experiments. Bar, 10 μm.
We next expressed the MAPK kinase MKK7, which specifically activates JNK but not p38 (Yao et al., 1997). Cotransfection of L6 myoblasts with different combinations of plasmids encoding MKK7, MKK3, HA-tagged JNK, or HA-tagged p38 confirmed the specific JNK activation by MKK7 without affecting p38 activity and that of p38 by MKK3 without affecting JNK activity (Figure 7C). L6 cells were thus transfected with constructs expressing FLAG epitope–tagged MKK7, HA epitope–tagged MKK3, or empty HA–tagged pcDNA3 as a control. After transfection, cells were induced to differentiate by the addition of DM, fixed 1–2 d thereafter, and analyzed for FLAG-tagged MKK7 or HA-tagged MKK3 and pcDNA3 and myogenin expression (Figure 7D). Transient expression of MKK7 (Figure 7D, panel a) strongly inhibited myogenin expression (panel b), whereas expression of MKK3 (panel c) or empty pcDNA3 (panel e) had no effect on myogenin expression (panels d and f, respectively).
In conclusion, these data demonstrate that JNK activation impairs myogenin expression, the earliest known marker for myoblasts committed to the differentiation pathway.
RhoG, Rac1, and Cdc42Hs GTPases Impair Nuclear Myf5 Accumulation
The muscle-determining factor Myf5 has been shown to be expressed in proliferating myoblasts and down-regulated in cells undergoing differentiation (Lindon et al., 1998). The early inhibition of myogenesis observed after RhoGV12, Rac1V12, Cdc42HsV12, and JNK activation led us to investigate their effects on Myf5 expression. Myf5 expression was first analyzed by Western blot analysis in parental L6 cells and RhoG-, Rac1-, and Cdc42Hs-expressing cells, which showed no significant modification of Myf5 protein level between parental and stably transfected cell lines (Figure 8A). We then analyzed Myf5 expression by indirect immunofluorescence with the use of a Myf5 polyclonal antibody (Yablonka-Reuveni et al., 1999) (Figure 8B). As expected, Myf5 was found to be highly expressed in the nucleus of proliferating L6 myoblasts (Figure 8B, panel a) and absent in differentiated cells (panel b). No modification of Myf5 expression or distribution was detected in L6 cells expressing RhoAV14 (panel c). In contrast, Myf5 remained undetectable in the nuclei of RhoGV12-, Rac1V12-, and Cdc42HsV12-expressing stable cell lines cultured under either proliferation (panels d, e, and f) or differentiation conditions (our unpublished results). Nevertheless, there was increased cytoplasmic staining of Myf5 in these transfectants. A similar loss of nuclear Myf5 staining was observed in L6 cells transiently transfected with GFP-tagged RhoGV12, Rac1V12, or Cdc42HsV12 constructs, whereas expression of empty pEGFP-C1 had no effect (our unpublished results). In contrast, expression of DN Rac1 and Cdc42Hs mutants did not affect Myf5 nuclear localization (our unpublished results). Because the data presented above showed that myogenesis inhibition by RhoG, Rac1, and Cdc42Hs GTPase expression correlated with JNK activation, we next examined whether this was also the case for Myf5 localization. L6 myoblasts were treated with anisomycin for 24 h and then fixed and analyzed for Myf5 localization (Figure 8C). Anisomycin treatment of L6 cells led to a loss of Myf5 nuclear staining associated with increased cytoplasmic staining (compare Figure 8B, panel a, with Figure 8C, panel a). Similar results were obtained in anisomycin-treated C2 myogenic cells (our unpublished results). Anisomycin treatment of L6 RhoAV14 myoblasts also resulted in Myf5 nuclear delocalization (compare Figure 8B, panel c, with Figure 8C, panel b), suggesting a dominant effect of JNK activation. To specifically activate JNK, we then transfected L6 myoblasts with FLAG epitope–tagged MKK7. As a control, cells were transfected with constructs expressing HA epitope–tagged MKK3 or empty HA–tagged pcDNA3. Twenty-four hours after transfection, cells were fixed and analyzed for FLAG-tagged MKK7, HA-tagged MKK3, and Myf5 localization. Transient expression of MKK7 (Figure 8C, inset in panel c) led to a loss of Myf5 nuclear staining associated with increased cytoplasmic staining (panel c), whereas expression of MKK3 (inset in panel d) or empty pcDNA3 (our unpublished results) had no effect on Myf5 localization (panel d).
Figure 8.
Myf5 translocation to a cytoplasmic compartment in RhoG-, Rac1-, and Cdc42Hs-expressing cells and after JNK activation. (A) Protein extracts from parental L6, L6 RhoGV12, L6 Rac1V12, and L6 Cdc42HsV12 (100 μg/well) were immunoblotted with anti-Myf5 antibody as described in MATERIALS AND METHODS. (B) Parental L6 (a and b), L6 RhoAV14 (c), L6 RhoGV12 (d), L6 Rac1V12 (e), and L6 Cdc42HsV12 (f) were fixed and stained for Myf5 expression. Panels a, c, d, e, and f, cells in proliferating medium; panel b, 1.5 d after the addition of DM. For each panel, cells shown are representative of five independent experiments with more than 100 observed cells. Bar, 10 μm. (C) Parental L6 (a) and L6 RhoAV14 (b) were treated with anisomycin (50 nM) for 24 h and then fixed and stained for Myf5 localization. L6 myoblasts were transfected with constructs expressing FLAG-tagged MKK7 or HA-tagged MKK3. Twenty-four hours after transfection, cells were fixed and processed for FLAG (c, inset) or HA epitope detection (d, inset) and for Myf5 localization (c and d). For each panel, cells are representative of three independent experiments.
These data demonstrate that in contrast to myogenin, troponin T, and MHC, the level of Myf5 expression remains unaffected upon expression of active RhoG, Rac1, and Cdc42Hs GTPases. However, Myf5 protein is excluded from the nucleus after expression of active RhoG, Rac1, and Cdc42Hs GTPases but not RhoA GTPase as well as after JNK activation.
DISCUSSION
To gain insight into the role of Rho GTPases in skeletal muscle differentiation, we expressed DN T17N versions of Rac1 and Cdc42Hs that have decreased affinity for GTP and act as inhibitors of the endogenous proteins. Expression of DN Rac1 or Cdc42Hs mutants impaired myogenin, troponin T, and MHC expression, showing that the activity of either GTPase is required for myogenesis. In addition, we report that expression of DN mutants of Rac1 or Cdc42Hs also leads to a strong decrease in p38 activity, which is essential for myogenesis.
We have also established stable cell lines expressing constitutively active forms of RhoG, Rac1, Cdc42Hs, and RhoA. These proteins, all of which contain a Gly-to-Val amino acid substitution at position 12 (or position 14 for RhoA), are preferentially bound to GTP and permanently activate their downstream effectors. We found that expression of active RhoG, Rac1, and Cdc42Hs proteins prevents the formation of myotubes and inhibits the expression of myogenin, troponin T, and MHC. We further demonstrated that RhoA activation potentiates the expression of myogenin, MHC, and troponin T during muscle differentiation but still prevents myoblast fusion. Similar findings were obtained with the use of stably infected L6 cell lines or transiently transfected C2 myoblasts. Using effector loop mutants of Rac1 and Cdc42Hs, we also showed that myogenesis inhibition is mediated through the binding to CRIB motif–containing proteins, associated with JNK activation, whereas Rho-dependent cytoskeletal reorganization has no effect on the early stages of myogenesis. The direct implication of the JNK pathway in myogenesis inhibition was demonstrated by expressing MKK7, a specific upstream activator of JNK. We further showed that, although expressed at the same level in normal and infected cells, Myf5 accumulates in the nucleus of normal and RhoA-expressing cells but is detected exclusively in the cytoplasm of RhoG-, Rac1-, and Cdc42Hs-expressing myoblasts as well as in L6 cells in which the JNK pathway has been activated. These data suggest that Rho GTPase activity of the Rac/Cdc42 subgroup might control myogenic differentiation through a differential effect on p38 and JNK MAPK.
Dual Regulation of p38 and JNK by Rac1 and Cdc42Hs GTPases
Our data demonstrate that either the activation or the inhibition of Rac1 and Cdc42Hs prevents the establishment of early phases of myogenic differentiation. This suggested that both GTPases control two pathways, one positively acting on and the other down-regulating the onset of myogenesis. Although Rac1 and Cdc42Hs have been reported to control distinct cytoskeletal structures, they both lead to p38 and JNK MAPK activation (Coso et al., 1995; Minden et al., 1995). Interestingly, p38 activation has been found to be necessary for the myogenic differentiation of C2C12 and C3H10T1/2 cells (Cuenda and Cohen, 1999; Zetser et al., 1999) as well as L6 cells (this paper). Whereas expression of DN Rac1 and Cdc42Hs mutants led to a decrease of p38 activity, overexpression of a specific p38 activator cannot reverse the DN Rac1- and Cdc42Hs-dependent inhibition of myogenesis (our unpublished results). Thus, inhibiting the p38 pathway is not the only cascade modified by the expression of these mutants. Indeed, expression of these DN Rac1 and Cdc42Hs mutants strongly suppresses the morphogenic functions of these GTPases, which might also contribute to their inhibitory effects on myogenesis. In particular, Rho GTPases might regulate the function of cadherins or integrins, adhesion proteins linked to the actin-based cytoskeleton that are implicated in cell–cell as well as cell–extracellular matrix interactions (Ridley and Hall, 1992; Nobes and Hall, 1995a; Braga et al., 1997). Cadherins and integrins have been shown to play a key role both in the early stages of myogenic induction and in later stages of myoblast interaction and fusion (Menko and Boettiger, 1987; Redfield et al., 1997; Goichberg and Geiger, 1998). On the other hand, we show that the inhibitory effect of the constitutively active Rac1 and Cdc42Hs proteins on myogenesis is directly linked to the activation of the JNK pathway. Indeed, we observed a perfect correlation between the inhibitory activity of Rac1 and Cdc42Hs F37A and Y40C double mutants and their ability to activate the JNK pathway. A recent report also showed the inhibition of myogenesis in primary avian myoblasts expressing constitutively active Rac1 and Cdc42Hs (Gallo et al., 1999). However, the authors of that paper were unable to correlate this inhibition to JNK activation with the use of effector mutants of Rac1 and Cdc42Hs, likely because the F37A Rac1 and Cdc42Hs mutants that they used do not activate JNK in these cells. Regardless, our own data with MKK7 expression constructs unambiguously demonstrate that the direct activation of JNK inhibits myogenic differentiation. An attempt to determine whether blocking the JNK pathway reverses myogenesis inhibition was unsuccessful, because inhibiting this pathway was highly toxic in myoblasts. Our data, therefore, suggest that the level of GTP-bound Rac1 and Cdc42Hs might influence the early stages of myogenesis either positively through the p38 pathway or negatively through the JNK pathway. In addition, myogenesis inhibition in Rac1- and Cdc42Hs-expressing cells or in cells treated with anisomycin, in which both JNK and p38 are activated, suggests either a dominant or an earlier inhibitory effect of JNK activation. Although our data point to a critical role of the JNK pathway in the early steps of myogenesis, Rho GTPases might also affect later stages in myogenesis. In particular, RhoAV14-expressing L6 cells failed to produce multinucleated myotubes, suggesting an inhibition of the fusion process.
Opposing Roles for RhoG/Rac1/Cdc42Hs and RhoA in Regulating Myogenesis
Several lines of evidence have suggested a functional antagonism between the Rac1/Cdc42 and RhoA–C subfamilies in controlling cytoskeletal structures (Kozma et al., 1997; Leeuwen et al., 1997; Moorman et al., 1999). As far as myogenesis is concerned, a similar balance between RhoG/Rac1/Cdc42Hs and RhoA activities emerges in the light of the present study. We found that expression of activated RhoA potentiates early steps of myogenesis, because it increases the expression of myogenin, troponin T, and MHC. These data are in agreement with recent studies showing that RhoA but not Rac1/Cdc42 signaling is an integral part of the skeletal muscle differentiation pathway (Carnac et al., 1998; Wei et al., 1998). Nevertheless, expression of activated RhoG, Rac1, and Cdc42Hs fully inhibits myogenesis of rat L6 myoblasts, preventing the expression of myogenin, troponin T, and MHC. Furthermore, the activation of JNK in RhoA-expressing cells still inhibits myogenin expression, suggesting a dominant effect of the Rac/Cdc42 subgroup over RhoA. Our results on the inhibitory effect of Rac1/Cdc42 are in disagreement with previous results showing that the myogenin promoter is activated by constitutively active Rac1, Cdc42Hs, or RhoA (Takano et al., 1998). This discrepancy may be due to the fact that our experiments relied on the measurement of endogenous myogenin levels in stably infected cells expressing moderate levels of GTPases, whereas Takano et al. used a reporter gene assay in transiently transfected cells.
Our data also establish a differential effect of Rho GTPases on Myf5, a muscle-determining factor shown to be expressed in proliferating myoblasts and down-regulated in cells undergoing differentiation (Lindon et al., 1998). Here, we present the first evidence that the nuclear localization of Myf5 is negatively controlled by RhoG/Rac1 and Cdc42Hs, likely through the activation of the JNK pathway. In contrast to MyoD, the myogenic factor Myf5 is not regulated by the RhoA/SRF pathway in C2C12 cells (Carnac et al., 1998), in agreement with our own results with L6 cells. Furthermore, expression of the DN Rac1 and Cdc42Hs mutants does not affect Myf5 localization, suggesting that DN and active Rac1 and Cdc42Hs mutants inhibit myogenesis through distinct signaling pathways.
JNK Activation and Inhibition of Myogenesis
Several but not exclusive targets might be proposed for the inhibitory effect of JNK activation on myogenesis. First, the inhibition might be mediated by Jun proteins. Indeed, the JNK signaling pathway has been reported to activate Jun transcription factor members (Gupta et al., 1996), leading to the formation of activated AP1 complexes known to inhibit myogenic differentiation (Lassar et al., 1989; Li et al., 1992; Park et al., 1992). Myogenesis inhibition might also be mediated through the phosphorylation of Elk-1, a member of the Ternary Complex Factor (TCF) proteins that is involved in gene transcription in cooperation with SRF and that has been shown to be phosphorylated by JNK (Gupta et al., 1996). Changes in TCF phosphorylation, therefore, might inhibit the SRF-dependent activation of muscle-specific genes. Along this line, a new TCF family member acting as a transcriptional repressor was recently characterized in muscle cells (Goldman et al., 1998). A third target might be the nuclear factor κB, the activity of which has been reported to be regulated by GTPases of the Rho family (Sulciner et al., 1996; Perona et al., 1997), through a process controlled by the JNK kinase pathway (Montaner et al., 1998). Recently, nuclear factor κB was shown to activate cyclin D1, leading to inhibition of C2C12 myogenic differentiation (Guttridge et al., 1999). Similar results were obtained in L6 cells expressing RhoG, Rac, and Cdc42Hs, in which cyclin D1 remained expressed at high levels even 5 d after the shift to differentiation medium (M. Meriane, unpublished data).
Although a direct effect of JNK on Myf5 phosphorylation remains to be established, analysis of the primary amino acid sequence of Myf5 reveals the presence of five conserved putative JNK phosphorylation sites. Interestingly, a similar negative correlation has been observed between JNK activation and the phosphorylation and nuclear translocation of NFAT4, a transcription factor of the rel domain family (Chow et al., 1997). Moreover, control of the cytonuclear localization has been reported for the Xenopus MyoD protein, which moves from the cytoplasm to the nucleus during the midblastula transition (Rupp et al., 1994).
Rho GTPases and Myogenesis
It has been reported that Rac1 and Cdc42Hs regulate cell motility and migration in various cell systems (Nobes and Hall, 1995a). In the developing embryo, part of the progenitor population migrates away from the somite to form peripheral body muscle and the muscle of the limb. During the migration, cells are maintained in an undifferentiated state, and myogenic factors are not expressed until they have reached the correct position in the embryo. An attractive function for the Rac1/Cdc42Hs subgroup could be to coordinate cell migration and inhibition of myogenesis, and for RhoA, to inhibit cell migration and favor myogenesis. This scheme is supported by the fact that Rho proteins are associated with the control of morphogenesis in the worm and the fly (Settleman, 1999). Additionally, expression of mutant Rac1 and Cdc42Hs proteins in Drosophila muscle precursor cells leads to aberrant myoblast fusion and migration, respectively (Luo et al., 1994). These GTPases might thus participate in inhibiting cell differentiation and stimulating cell movement and migration in response to signals derived from adjacent tissues, such as TGFβ, whose signaling requires pathways controlled by Rho proteins (Atfi et al., 1997). In this regard, it is interesting to note that JNK activity is negatively regulated by cell density (Lallemand et al., 1998). Functional studies will be necessary to further correlate the activity of Rho GTPases with the level of expression and/or the subcellular localization of myogenic markers during development.
ACKNOWLEDGMENTS
We thank A. Hall for Rac1 and Cdc42Hs mutant cDNAs, J.M. Bellanger for GST–c-Jun and GST–ATF2 proteins, Marie Vandromme for L6 cells, Pierre Travo for constructive microscopy support, Bob Hipskind for helpful discussions, and Naomi Taylor for critical reading of the manuscript. We also thank Anne Blangy, Sophie Mary, and Emmanuel Vignal for continuous support. Electron scanning microscopy was performed at the Center Regional d'Imagerie Cellulaire de Montpellier. We thank the Association Française contre les Myopathies, the Association pour la Recherche contre le Cancer (contract number 9759), the Ligue Nationale contre le Cancer, and the Fondation pour la Recherche Médicale for financial support.
REFERENCES
- Andres V, Walsh K. Myogenin expression, cell cycle withdrawal, and phenotypic differentiation are temporally separable events that precede cell fusion upon myogenesis. J Cell Biol. 1996;132:657–666. doi: 10.1083/jcb.132.4.657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arnold HH, Winter B. Muscle differentiation: more complexity to the network of myogenic regulators. Curr Opin Genet Dev. 1998;8:539–544. doi: 10.1016/s0959-437x(98)80008-7. [DOI] [PubMed] [Google Scholar]
- Atfi A, Djelloul S, Chastre E, Davis R, Gespach C. Evidence for a role of Rho-like GTPases and stress-activated protein kinase/c-Jun N-terminal kinase (SAPK/JNK) in transforming growth factor beta-mediated signaling. J Biol Chem. 1997;272:1429–1432. doi: 10.1074/jbc.272.3.1429. [DOI] [PubMed] [Google Scholar]
- Braga VM, Machesky LM, Hall A, Hotchin NA. The small GTPases Rho and Rac are required for the establishment of cadherin-dependent cell-cell contacts. J Cell Biol. 1997;137:1421–1431. doi: 10.1083/jcb.137.6.1421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cano E, Hazzalin CA, Mahadevan LC. Anisomycin-activated protein kinases p45 and p55 but not mitogen-activated protein kinases ERK-1 and -2 are implicated in the induction of c-fos and c-jun. Mol Cell Biol. 1994;14:7352–7362. doi: 10.1128/mcb.14.11.7352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carnac G, Primig M, Kitzmann M, Chafey P, Tuil D, Lamb N, Fernandez A. RhoA GTPase and serum response factor control selectively the expression of MyoD without affecting myf5 in mouse myoblasts. Mol Biol Cell. 1998;9:1891–1902. doi: 10.1091/mbc.9.7.1891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chow CW, Rincon M, Cavanagh J, Dickens M, Davis RJ. Nuclear accumulation of NFAT4 opposed by the JNK signal transduction pathway. Science. 1997;278:1638–1641. doi: 10.1126/science.278.5343.1638. [DOI] [PubMed] [Google Scholar]
- Coso OA, Chiariello M, Yu JC, Teramoto H, Crespo P, Xu N, Miki T, Gutkind JS. The small GTP-binding proteins Rac1 and Cdc42 regulate the activity of the JNK/SAPK signaling pathway. Cell. 1995;81:1137–1146. doi: 10.1016/s0092-8674(05)80018-2. [DOI] [PubMed] [Google Scholar]
- Cossu G, Kelly R, Tajbakhsh S, Di Donna S, Vivarelli E, Buckingham M. Activation of different myogenic pathways: myf-5 is induced by the neural tube and MyoD by the dorsal ectoderm in mouse paraxial mesoderm. Development. 1996;122:429–437. doi: 10.1242/dev.122.2.429. [DOI] [PubMed] [Google Scholar]
- Cuenda A, Cohen P. Stress-activated protein kinase-2/p38 and a rapamycin-sensitive pathway are required for C2C12 myogenesis. J Biol Chem. 1999;274:4341–4346. doi: 10.1074/jbc.274.7.4341. [DOI] [PubMed] [Google Scholar]
- Didsbury J, Weber RF, Bokoch GM, Evans T, Snyderman R. rac, a novel ras-related family of proteins that are botulinum toxin substrates. J Biol Chem. 1989;264:16378–16382. [PubMed] [Google Scholar]
- Fort P. Small GTPases of the Rho family and cell transformation. Prog Mol Subcell Biol. 1999;22:159–181. doi: 10.1007/978-3-642-58591-3_8. [DOI] [PubMed] [Google Scholar]
- Gallo R, Serafini M, Castellani L, Falcone G, Alema S. Distinct effects of rac1 on differentiation of primary avian myoblasts. Mol Biol Cell. 1999;10:3137–3150. doi: 10.1091/mbc.10.10.3137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gauthier RC, Vandromme M, Tuil D, Lautredou N, Morris M, Soulez M, Kahn A, Fernandez A, Lamb N. Expression and activity of serum response factor is required for expression of the muscle-determining factor MyoD in both dividing and differentiating mouse C2C12 myoblasts. Mol Biol Cell. 1996;7:719–729. doi: 10.1091/mbc.7.5.719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gauthier RC, Vignal E, Meriane M, Roux P, Montcourier P, Fort P. RhoG GTPase controls a pathway that independently activates Rac1 and Cdc42Hs. Mol Biol Cell. 1998;9:1379–1394. doi: 10.1091/mbc.9.6.1379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goichberg P, Geiger B. Direct involvement of N-cadherin–mediated signaling in muscle differentiation. Mol Biol Cell. 1998;9:3119–3131. doi: 10.1091/mbc.9.11.3119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldman D, Sapru MK, Stewart S, Plotkin J, Libermann TA, Wasylyk B, Guan K. Cloning and characterization of GETS-1, a goldfish Ets family member that functions as a transcriptional repressor in muscle. Biochem J. 1998;335:267–275. doi: 10.1042/bj3350267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gupta S, Barrett T, Whitmarsh AJ, Cavanagh J, Sluss HK, Derijard B, Davis RJ. Selective interaction of JNK protein kinase isoforms with transcription factors. EMBO J. 1996;15:2760–2770. [PMC free article] [PubMed] [Google Scholar]
- Guttridge DC, Albanese C, Reuther JY, Pestell RG, Baldwin AS., Jr NF-kappaB controls cell growth and differentiation through transcriptional regulation of cyclin D1. Mol Cell Biol. 1999;19:5785–5799. doi: 10.1128/mcb.19.8.5785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall A. Rho GTPases and the actin cytoskeleton. Science. 1998;279:509–514. doi: 10.1126/science.279.5350.509. [DOI] [PubMed] [Google Scholar]
- Hill CS, Wynne J, Treisman R. The Rho family GTPases RhoA, Rac1, and CDC42Hs regulate transcriptional activation by SRF. Cell. 1995;81:1159–1170. doi: 10.1016/s0092-8674(05)80020-0. [DOI] [PubMed] [Google Scholar]
- Joneson T, McDonough M, Bar-Sagi D, Van Aelst L. RAC regulation of actin polymerization and proliferation by a pathway distinct from Jun kinase. Science. 1996;274:1374–1376. doi: 10.1126/science.274.5291.1374. [DOI] [PubMed] [Google Scholar]
- Kozma R, Ahmed S, Best A, Lim L. The Ras-related protein Cdc42Hs and bradykinin promote formation of peripheral actin microspikes and filopodia in Swiss 3T3 fibroblasts. Mol Cell Biol. 1995;15:1942–1952. doi: 10.1128/mcb.15.4.1942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozma R, Sarner S, Ahmed S, Lim L. Rho family GTPases and neuronal growth cone remodelling: relationship between increased complexity induced by Cdc42Hs, Rac1, and acetylcholine and collapse induced by RhoA and lysophosphatidic acid. Mol Cell Biol. 1997;17:1201–1211. doi: 10.1128/mcb.17.3.1201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lallemand D, Ham J, Garbay S, Bakiri L, Traincard F, Jeannequin O, Pfarr CM, Yaniv M. Stress-activated protein kinases are negatively regulated by cell density. EMBO J. 1998;17:5615–5626. doi: 10.1093/emboj/17.19.5615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamarche N, Tapon N, Stowers L, Burbelo PD, Aspenstrom P, Bridges T, Chant J, Hall A. Rac and Cdc42 induce actin polymerization and G1 cell cycle progression independently of p65PAK and the JNK/SAPK MAP kinase cascade. Cell. 1996;87:519–529. doi: 10.1016/s0092-8674(00)81371-9. [DOI] [PubMed] [Google Scholar]
- Lassar AB, Skapek SX, Novitch B. Regulatory mechanisms that coordinate skeletal muscle differentiation and cell cycle withdrawal. Curr Opin Cell Biol. 1994;6:788–794. doi: 10.1016/0955-0674(94)90046-9. [DOI] [PubMed] [Google Scholar]
- Lassar AB, Thayer MJ, Overell RW, Weintraub H. Transformation by activated ras or fos prevents myogenesis by inhibiting expression of MyoD1. Cell. 1989;58:659–667. doi: 10.1016/0092-8674(89)90101-3. [DOI] [PubMed] [Google Scholar]
- Leeuwen FN, Kain HE, Kammen RA, Michiels F, Kranenburg OW, Collard JG. The guanine nucleotide exchange factor Tiam1 affects neuronal morphology: opposing roles for the small GTPases Rac and Rho. J Cell Biol. 1997;139:797–807. doi: 10.1083/jcb.139.3.797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li L, Chambard JC, Karin M, Olson EN. Fos and Jun repress transcriptional activation by myogenin and MyoD: the amino terminus of Jun can mediate repression. Genes Dev. 1992;6:676–689. doi: 10.1101/gad.6.4.676. [DOI] [PubMed] [Google Scholar]
- Lindon C, Montarras D, Pinset C. Cell cycle-regulated expression of the muscle determination factor Myf5 in proliferating myoblasts. J Cell Biol. 1998;140:111–118. doi: 10.1083/jcb.140.1.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo L, Liao YJ, Jan LY, Jan YN. Distinct morphogenetic functions of similar small GTPases: Drosophila Drac1 is involved in axonal outgrowth and myoblast fusion. Genes Dev. 1994;8:1787–1802. doi: 10.1101/gad.8.15.1787. [DOI] [PubMed] [Google Scholar]
- Menko AS, Boettiger D. Occupation of the extracellular matrix receptor, integrin, is a control point for myogenic differentiation. Cell. 1987;51:51–57. doi: 10.1016/0092-8674(87)90009-2. [DOI] [PubMed] [Google Scholar]
- Michiels F, Collard JG. Rho-like GTPases: their role in cell adhesion and invasion. Biochem Soc Symp. 1999;65:125–146. [PubMed] [Google Scholar]
- Minden A, Lin A, Claret FX, Abo A, Karin M. Selective activation of the JNK signaling cascade and c-Jun transcriptional activity by the small GTPases Rac and Cdc42Hs. Cell. 1995;81:1147–1157. doi: 10.1016/s0092-8674(05)80019-4. [DOI] [PubMed] [Google Scholar]
- Mohun T, Garrett N, Treisman R. Xenopus cytoskeletal actin and human c-fos gene promoters share a conserved protein-binding site. EMBO J. 1987;6:667–673. doi: 10.1002/j.1460-2075.1987.tb04806.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molkentin JD, Olson EN. Defining the regulatory networks for muscle development. Curr Opin Genet Dev. 1996;6:445–453. doi: 10.1016/s0959-437x(96)80066-9. [DOI] [PubMed] [Google Scholar]
- Montaner S, Perona R, Saniger L, Lacal JC. Multiple signaling pathways lead to the activation of the nuclear factor kappaB by the Rho family of GTPases. J Biol Chem. 1998;273:12779–12785. doi: 10.1074/jbc.273.21.12779. [DOI] [PubMed] [Google Scholar]
- Moorman JP, Luu D, Wickham J, Bobak DA, Hahn CS. A balance of signaling by Rho family small GTPases RhoA, Rac1 and Cdc42 coordinates cytoskeletal morphology but not cell survival. Oncogene. 1999;18:47–57. doi: 10.1038/sj.onc.1202262. [DOI] [PubMed] [Google Scholar]
- Munsterberg AE, Kitajewski J, Bumcrot DA, McMahon AP, Lassar AB. Combinatorial signaling by sonic hedgehog and Wnt family members induces myogenic bHLH gene expression in the somite. Genes Dev. 1995;9:2911–2922. doi: 10.1101/gad.9.23.2911. [DOI] [PubMed] [Google Scholar]
- Nobes CD, Hall A. Rho, rac and cdc42 GTPases: regulators of actin structures, cell adhesion and motility. Biochem Soc Trans. 1995a;23:456–459. doi: 10.1042/bst0230456. [DOI] [PubMed] [Google Scholar]
- Nobes CD, Hall A. Rho, rac, and cdc42 GTPases regulate the assembly of multimolecular focal complexes associated with actin stress fibers, lamellipodia, and filopodia. Cell. 1995b;81:53–62. doi: 10.1016/0092-8674(95)90370-4. [DOI] [PubMed] [Google Scholar]
- Park K, Chung M, Kim SJ. Inhibition of myogenesis by okadaic acid, an inhibitor of protein phosphatases, 1 and 2A, correlates with the induction of AP1. J Biol Chem. 1992;267:10810–10815. [PubMed] [Google Scholar]
- Perona R, Montaner S, Saniger L, Sanchez-Perez I, Bravo R, Lacal JC. Activation of the nuclear factor-kappaB by Rho, CDC42, and Rac-1 proteins. Genes Dev. 1997;11:463–475. doi: 10.1101/gad.11.4.463. [DOI] [PubMed] [Google Scholar]
- Pourquie O, Fan CM, Coltey M, Hirsinger E, Watanabe Y, Breant C, Francis-West P, Brickell P, Tessier-Lavigne M, Le Douarin NM. Lateral and axial signals involved in avian somite patterning: a role for BMP4. Cell. 1996;84:461–471. doi: 10.1016/s0092-8674(00)81291-x. [DOI] [PubMed] [Google Scholar]
- Ramocki MB, Johnson SE, White MA, Ashendel CL, Konieczny SF, Taparowsky EJ. Signaling through mitogen-activated protein kinase and Rac/Rho does not duplicate the effects of activated Ras on skeletal myogenesis. Mol Cell Biol. 1997;17:3547–3555. doi: 10.1128/mcb.17.7.3547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Redfield A, Nieman MT, Knudsen KA. Cadherins promote skeletal muscle differentiation in three-dimensional cultures. J Cell Biol. 1997;138:1323–1331. doi: 10.1083/jcb.138.6.1323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ridley AJ, Hall A. The small GTP-binding protein rho regulates the assembly of focal adhesions and actin stress fibers in response to growth factors. Cell. 1992;70:389–399. doi: 10.1016/0092-8674(92)90163-7. [DOI] [PubMed] [Google Scholar]
- Ridley AJ, Paterson HF, Johnston CL, Diekmann D, Hall A. The small GTP-binding protein rac regulates growth factor-induced membrane ruffling. Cell. 1992;70:401–410. doi: 10.1016/0092-8674(92)90164-8. [DOI] [PubMed] [Google Scholar]
- Roux P, Gauthier RC, Doucet BS, Fort P. The small GTPases Cdc42Hs, Rac1 and RhoG delineate Raf-independent pathways that cooperate to transform NIH3T3 cells. Curr Biol. 1997;7:629–637. doi: 10.1016/s0960-9822(06)00289-2. [DOI] [PubMed] [Google Scholar]
- Rupp RA, Snider L, Weintraub H. Xenopus embryos regulate the nuclear localization of XMyoD. Genes Dev. 1994;8:1311–1323. doi: 10.1101/gad.8.11.1311. [DOI] [PubMed] [Google Scholar]
- Sahai E, Alberts AS, Treisman R. RhoA effector mutants reveal distinct effector pathways for cytoskeletal reorganization, SRF activation and transformation. EMBO J. 1998;17:1350–1361. doi: 10.1093/emboj/17.5.1350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Settleman J. Rho GTPases in development. Prog Mol Subcell Biol. 1999;22:201–229. doi: 10.1007/978-3-642-58591-3_10. [DOI] [PubMed] [Google Scholar]
- Stern HM, Lin-Jones J, Hauschka SD. Synergistic interactions between bFGF and a TGF-beta family member may mediate myogenic signals from the neural tube. Development. 1997;124:3511–3523. doi: 10.1242/dev.124.18.3511. [DOI] [PubMed] [Google Scholar]
- Sulciner DJ, Irani K, Yu ZX, Ferrans VJ, Goldschmidt-Clermont P, Finkel T. rac1 regulates a cytokine-stimulated, redox-dependent pathway necessary for NF-kappaB activation. Mol Cell Biol. 1996;16:7115–7121. doi: 10.1128/mcb.16.12.7115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takano H, Komuro I, Oka T, Shiojima I, Hiroi Y, Mizuno T, Yazaki Y. The Rho family G proteins play a critical role in muscle differentiation. Mol Cell Biol. 1998;18:1580–1589. doi: 10.1128/mcb.18.3.1580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tapon N, Hall A. Rho, Rac, and Cdc42 GTPases regulate the organization of the actin cytoskeleton. Curr Opin Cell Biol. 1997;9:86–92. doi: 10.1016/s0955-0674(97)80156-1. [DOI] [PubMed] [Google Scholar]
- Teramoto H, Crespo P, Coso OA, Igishi T, Xu N, Gutkind JS. The small GTP-binding protein rho activates c-Jun N-terminal kinases/stress-activated protein kinases in human kidney 293T cells: evidence for a Pak-independent signaling pathway. J Biol Chem. 1996;271:25731–25734. doi: 10.1074/jbc.271.42.25731. [DOI] [PubMed] [Google Scholar]
- Vandromme M, Gauthier RC, Carnac G, Lamb N, Fernandez A. Serum response factor p67SRF is expressed and required during myogenic differentiation of both mouse C2 and rat L6 muscle cell lines. J Cell Biol. 1992;118:1489–1500. doi: 10.1083/jcb.118.6.1489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vincent S, Marty L, Fort P. S26 ribosomal protein RNA: an invariant control for gene regulation experiments in eucaryotic cells and tissues. Nucleic Acids Res. 1993;21:1498. doi: 10.1093/nar/21.6.1498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei L, Zhou W, Croissant JD, Johansen FE, Prywes R, Balasubramanyam A, Schwartz RJ. RhoA signaling via serum response factor plays an obligatory role in myogenic differentiation. J Biol Chem. 1998;273:30287–30294. doi: 10.1074/jbc.273.46.30287. [DOI] [PubMed] [Google Scholar]
- Westwick JK, Lambert QT, Clark GJ, Symons M, Van Aelst L, Pestell RG, Der CJ. Rac regulation of transformation, gene expression, and actin organization by multiple, PAK-independent pathways. Mol Cell Biol. 1997;17:1324–1335. doi: 10.1128/mcb.17.3.1324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wright WE, Sassoon DA, Lin VK. Myogenin, a factor regulating myogenesis, has a domain homologous to MyoD. Cell. 1989;56:607–617. doi: 10.1016/0092-8674(89)90583-7. [DOI] [PubMed] [Google Scholar]
- Yablonka-Reuveni Z, Rudnicki MA, Rivera AJ, Primig M, Anderson JE, Natanson P. The transition from proliferation to differentiation is delayed in satellite cells from mice lacking MyoD. Dev Biol. 1999;210:440–455. doi: 10.1006/dbio.1999.9284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yao Z, et al. Activation of stress-activated protein kinases/c-Jun N-terminal protein kinases (SAPKs/JNKs) by a novel mitogen-activated protein kinase kinase. J Biol Chem. 1997;272:32378–32383. doi: 10.1074/jbc.272.51.32378. [DOI] [PubMed] [Google Scholar]
- Zetser A, Gredinger E, Bengal E. p38 mitogen-activated protein kinase pathway promotes skeletal muscle differentiation: participation of the Mef2c transcription factor. J Biol Chem. 1999;274:5193–5200. doi: 10.1074/jbc.274.8.5193. [DOI] [PubMed] [Google Scholar]