Abstract
Numerous prior studies have indicated that viable rotavirus reassortants containing structural proteins of heterologous parental origin may express unexpected phenotypes, such as changes in infectivity and immunogenicity. To provide a structural basis for alterations in phenotypic expression, a three-dimensional structural analysis of these reassortants was conducted. The structures of the reassortants show that while VP4 generally maintains the parental structure when moved to a heterologous protein background, in certain reassortants, there are subtle alterations in the conformation of VP4. The alterations in VP4 conformation correlated with expression of unexpected VP4-associated phenotypes. Interactions between heterologous VP4 and VP7 in reassortants expressing unexpected phenotypes appeared to induce the conformational alterations seen in VP4.
Rotavirus (family Reoviridae) is responsible for many cases of infantile gastroenteritis, resulting in approximately 600,000 deaths annually worldwide (8). The genome of rotavirus contains 11 segments of double-stranded RNA (7). The capsid structure consists of four major structural proteins (VP2, VP4, VP6, and VP7) and two enzymes (VP1 and VP3) arranged in three capsid layers (15). The outer capsid layer, with a T=13 icosahedral symmetry, consists of 780 molecules of VP7 and 120 molecules of the spike protein VP4. The intermediate capsid, with the same icosahedral symmetry, is made of 780 molecules of VP6. The inner capsid layer, with a T=1 symmetry composed of 120 VP2 molecules, surrounds the genome and VP1 and VP3, which are attached internally to this layer at the fivefold vertices. VP4 functions in cell binding, entry, hemagglutination, and neutralization (7). VP4 is also the target for trypsin-like proteases that cleave it into VP5* and VP8*, a cleavage that increases virus infectivity. VP4 is present on the capsid surface as 60 dimeric spikes and is the determinant of P neutralization serology. Recent X-ray crystallographic studies of VP8* indicate that the distal domain of the VP4 spike is composed of VP8* (6). VP7, a glycosylated protein, is also located on the virion surface as a relatively smooth lattice of trimers. VP7 is also a neutralizing antigen and determines G neutralization serology. VP6 of the intermediate capsid is the major protein component of virions and forms a lattice of trimers surrounding the inner capsid. In the virion, the outer capsid proteins VP4 and VP7 are known to interact with the underlying VP6 protein (13, 16, 19).
Several studies have shown that VP4-VP7 interactions in reassortants affect the expression of important viral phenotypes, such as sensitivity of VP4 to protease cleavage and plaque size (1, 17), neutralization with specific monoclonal antibodies (3, 7), receptor binding (11), and virion stability (3). Reassortment brings rotavirus structural proteins from diverse origins into a context in which a protein must interact with an interaction partner different from the one with which the protein evolved and adapted for functional interaction. Reassortant viruses are isolated with many different constellations of genes encoding proteins of heterologous parental origin. These viruses are quite viable, allowing their isolation, and grow to high titers in cultured cells. However, some of these reassortants display unexpected phenotypes. In some cases, a phenotype from virus A can be transferred to virus B by segregation of a single genome segment (protein), whereas transfer of the same phenotype of virus A to a different second parent (virus C) requires segregation of two viral genes. This implies that the genetic background of the recipient virus can affect phenotypic expression (1, 2, 3). Phenotype expression is regulated by protein-protein interactions within the virion in these reassortants (2). For example, in recombinants derived from SA11-4F and B223, expression of the phenotype of neutralization by VP4-specific monoclonal antibody (MAb) 2G4 segregated in unexpected patterns. This pattern was determined by whether VP4 interacted with its homologous VP7 protein or a heterologous VP7 protein. That the unexpected phenotypic expression required interaction of heterologous combinations of VP7 and VP4 was demonstrated by showing that the soluble VP4 and VP7 proteins displayed expected phenotypes, while assembly of VP4 and VP7 into virus particles resulted in an unexpected phenotype (2). Several important phenotypes were displayed in unexpected patterns when proteins of heterologous parental origin interacted in the virus particle: trypsin cleavage of VP4 (1), expression of neutralizing epitopes (1, 2, 5), receptor binding (11), and stability of viral infectivity during purification (3) (see Table 1).
TABLE 1.
Genotypes and phenotypes of parental and reassortant viruses used in this study
| Virus | Parental origin of indicated genome segment (protein)a
|
Trypsin cleavage of VP4 (EOP)b | Neutralization by α-VP4 MAb 2G4c | Virion stability during purification (%)d | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 (VP1) | 2 (VP2) | 3 (VP3) | 4 (VP4) | 5 (NSP1) | 6 (VP6) | 7 (NSP3) | 8 (NSP2) | 9 (VP7) | 10 (NSP4) | 11 (NSP5) | ||||
| SA11-4F | F | F | F | F | F | F | F | F | F | F | F | 2.7 × 10−3 | Yes | 93.3 |
| SA11-C13 | S | S | S | S | S | S | S | S | S | S | S | <6.3 × 10−6 | Yes | 10.6 |
| B223 | B | B | B | B | B | B | B | B | B | B | B | <1.2 × 10−5 | No | 0.5 |
| R-A29 | S | S | S | F | S | S | S | S | S | S | S | 1.4 × 10−2 | Yes | 60.6 |
| R-924 | F | F | F | S | F | F | F | F | F | F | F | NDf | Yes | 6.9 |
| R-197 | B | B | B | F | B | B | B | B | B | B | B | <8.0 × 10−6e | No | 8.2 |
| R-202 | F | F | F | F | F | B | F | F | F | F | F | ND | ND | 65.5 |
| R-600 | B | B | B | B | B | F | B | B | B | B | B | ND | ND | 1.2 |
| R-004 | F | F | F | B | F | F | F | F | F | F | F | ND | Yes | 0.5 |
Origin of segments: B, B223; F, SA11-4F; S, SA11-C13. Bold type indicates differing segment.
EOP, titer in the absence of trypsin/titer in the presence of trypsin (from reference 2).
Neutralization phenotype using VP4-specific MAB 2G4 (from reference 3).
Percent initial titer of virus grown in the presence of trypsin recovered by a standard CsCl purification procedure (from reference 4).
Unexpected phenotypic expression is shown in bold.
ND, not determined.
Here, we determined the structures of parental and reassortant rotaviruses that displayed expected and unexpected phenotypes in the studies described above (1-3, 16) to verify that altered protein interactions result in visible conformational changes. Changes in the structure of the VP4 spike were documented in some of the reassortants, confirming the previous suggestion that heterologous interactions between VP4 and VP7 can affect the conformation of one or both of these proteins and lead to unexpected expression of phenotypes such as virion stability, infectivity, growth to high titer, neutralization, and receptor binding.
Cells and virus.
MA104 African green monkey kidney cells were propagated in medium 199 supplemented with 5% fetal bovine serum, 0.15% NaHCO3, 0.03% glutamine, 25 U of penicillin per ml, and 250 μg of streptomycin per ml. The cell medium was changed to medium 199 without serum but containing the other supplements 18 to 24 h prior to infection. All virus stocks were propagated in MA104 cells grown in 150-cm2 flasks with medium 199 lacking fetal bovine serum. The reassortant viruses used were isolated and characterized in previous studies (1-3). For this study, each reassortant was plaque purified by three sequential plaque-to-plaque passages. The final plaque was passaged to a high titer twice on MA104 cells, and the genotype of the second-passage virus was confirmed by polyacrylamide gel electrophoresis as previously described (1). The reassortant genotypes and phenotypes are summarized in Table 1. The titers of the second-passage stocks were as follows: SA11-4F, 1.10 × 108 PFU/ml; SA11-Cl3, 1.30 × 108 PFU/ml; B223, 3.80 × 107 PFU/ml; R-004, 1.40 × 107 PFU/ml; R-197, 6.30 × 107 PFU/ml; R-600, 4.15 × 107 PFU/ml; R-202, 1.85 × 107 PFU/ml; R-924, 1.80 × 108 PFU/ml; R-A29, 1.35 × 108 PFU/ml. For each infection, the second-passage virus was activated in the presence of 5 μg of trypsin per ml for 30 min at 30°C. MA104 cells in 150 cm2 flasks were infected at a multiplicity of infection of 10 PFU/cell. After adsorption, medium 199 without serum was added to each flask so that the virus was propagated in the presence of 1 μg of trypsin per ml. When cytopathic effects were confluent, the infected-cell lysates were pooled and subjected to three freeze-thaw cycles. The lysates were then clarified by centrifugation (30 min at 12,000 rpm in a Beckman JA14 rotor) to remove cell debris. The virus in the clarified supernatant was pelleted (60 min at 25,000 rpm in a Beckman SW28 rotor) and resuspended in TNC buffer (10 mM Tris, 150 mM NaCl, 10 mM CaCl2, pH 7.5). Resuspended virus was layered on top of a preformed CsCl gradient (density, 30 to 60%), and the virus was banded by centrifugation (3.5 h at 33,000 rpm in a Beckman SW41 rotor). After density gradient centrifugation, the double-layer particle (DLP) and triple-layer particle (TLP) bands were collected, diluted with TNC buffer, and pelleted (45 min at 35,000 rpm in a Beckman SW41 rotor). The purified DLPs and TLPs were resuspended in an appropriate volume of TNC buffer, and the concentration and purity of the virus were determined by absorbance and negative-stain electron microscopy.
cryo-EM.
Specimen preparation for electron cryomicroscopy (cryo-EM) was carried out by following standard procedures (12). The sample, embedded in a thin layer of vitreous ice, was transferred and loaded on a GATAN cold-stage cryoholder. Electron micrographs of the frozen-hydrated specimens were recorded on a JEOL 1200 transmission electron microscope operating at 100 kV with a magnification of ×30,000 using an electron dose of ∼5 e−/Å2. From each specimen area in the grid, a focal pair with intended defocus values of ∼1 and ∼2 μm was recorded. The images were taken with a 1-s exposure to Kodak S0-163 EM film (Eastman Kodak Co., Rochester, N.Y.) and developed for 12 min in Kodak D-19 developer, followed by fixation for 10 min in Kodak fixer.
Three-dimensional reconstructions.
Micrographs were selected for correct defocus, ice quality, contrast, and particle concentration and scanned by using a Zeiss SCAI microdensitometer (Carl Zeiss, Inc., Englewood, Colo.), with a scanning interval of 14 μm, corresponding to 4.67 Å in the object. Reconstructions were carried out by using the data from closer-to-focus images to a resolution within the first zero of the contrast transfer function with appropriate corrections with defocus values of −1.45 μm (SA11-4F), −1.33 μm (SA11-Cl3), −1.52 μm (R-A29), −1.40 μm (R-924), −1.45 μm (R-197), and −1.40 μm (R-202). Determination of the defocus values, orientation determination and refinement, and three-dimensional reconstruction were conducted by using the ICOS Toolkit software (9). After the final refinement, the average phase residual between the images and their corresponding projections was less than 45°. Threshold values (contour levels) for the reconstructions were calculated to account for 780 molecules of VP6 between radii of 250 and 350 Å in all reconstructions.
Characterization of reassortants.
The reassortants examined here were described previously (1). However, since they were re-plaque-purified for this work, the parental origin of each genome segment was confirmed by labeling cells infected with each reassortant with [32P]orthophosphate, extracting the labeled double-stranded RNA, and determining the parental origin of the reassortant genome segments by polyacrylamide gel electrophoresis and comparison of mobility to that of parental segments. All reassortants contained the constellations of genome segments described previously (data not shown; Table 1 contains a summary). Previously, unusual presentation of viral phenotypes was described for some of these reassortants (1-3); these are also shown in Table 1.
Not all purified reassortant stocks yielded virus suitable for cryo-EM analysis. Most of the reassortants containing a genetic background of simian SA11 proteins (R-924, R-A29, and R-202) and one reassortant containing a B223 (bovine) background (R-197) were stable and purified as TLPs. Reassortants R-004 and R-600, containing a B223 genetic background, were unsuitable for structural analysis because they either yielded only DLPs (lacking VP4 and/or VP7) or were unstable during purification (16).
The TLP structures of the parental strains (SA11-Cl3 and SA11-4F) and the reassortants (R-924, R-A29, R-202, and R-197) were computed to a resolution of approximately 23 Å. Micrographs of either the native or the reassortant particles are not shown because of the morphological similarity of the TLPs in cryo-EM. Differences between the structures are visible only on three-dimensional reconstructions. These differences, particularly in the vicinity of the spikes, were examined by computationally extracting a conical section containing one VP4 spike, as shown in Fig. 1, from each of the reconstructions. The differences elsewhere, if any, were minimal and insignificant.
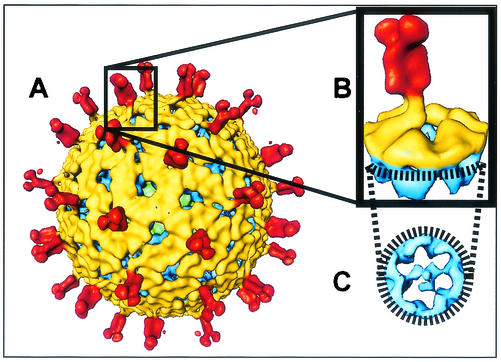
FIG. 1.
TLP structure and extraction of an individual VP4 spike. (A) Typical rotavirus TLP structure (SA11-4F strain) viewed along the icosahedral threefold axis. (B) Conical section extracted from the TLP structure, which contains an individual spike along with the surrounding VP7 and VP6 mass densities. In this view, the overall shape of the VP4 spike and its connections with VP7 can be seen. The region of the map used for the conical section is indicated by a rectangle. (C) Slice of the conical section illustrating the connections made between the internal globular domain of VP4 and VP6. The view is looking down on the conical section from the outer surface toward the center of the capsid. The TLP structure is colored radially. The spike structure between radii of 385 and 500 Å is red; the region between radii of 350 and 385 Å, representing the VP7 layer, is yellow; and the mass density below a radius of 350 Å, representing the VP6 layer, is blue. The green regions in the TLP structure represent the VP2 layer. The same color scheme is used in Fig. 2.
Interactions of VP4 with VP6 and VP7 in parental viruses.
Reconstructions of parental viruses SA11-Cl3 and SA11-4F were necessary to provide information on the conformation of the proteins when they interact with their native interaction partners. These two strains are thought to differ only in the VP4 protein. SA11-Cl3 contains all genes of simian origin, whereas SA11-4F is thought to be a reassortant that contains a set of genes homologous with SA11-Cl3, except for VP4, which shows high homology to VP4 of bovine origin (10).
The structure of SA11-Cl3 grown in the presence of trypsin and suspended in TNC buffer, pH 7.5, was determined to a resolution of ∼22 Å by using 144 particles (Fig. 2A, top). The structure of SA11-4F was determined to a similar resolution under identical conditions by using 322 particles (Fig. 2B, top). The structure of B223, another parental strain, could not be determined because of instability of the virus during purification, consistent with previous observations (3).
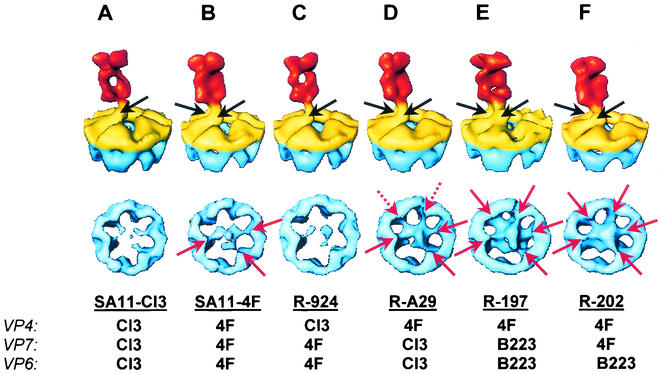
FIG. 2.
VP4 spike structures from the parental and reassortant strains. (Top) Conical sections, as illustrated in Fig. 1B, from the structures of parental strains (A, SA11-Cl3; B, SA11-4F) and reassortant strains (C, R-924; D, R-A29; E, R-197; F, R202). (Bottom) Corresponding cross-sectional views of these structures, as illustrated in Fig. 1C, showing the internal globular domain and its interactions with VP6. The black arrows indicate connections between VP4 and VP7. The red arrows indicate points of contact between the globular domain of VP4 and VP6. The dashed red arrows denote additional connections present in R-A29 that are not seen in SA11-4F.
The reconstructions of SA11-Cl3 and SA11-4F were adjusted for proper contour to display protein densities at equal levels. The spikes of SA11-Cl3 (Fig. 2A, top) have a lower mass density than those of SA11-4F (Fig. 2B, top), as evidenced by the reduction in density at the center of the SA11-Cl3 spike. In addition, the SA11-Cl3 spike makes one main connection to VP7, visualized as a slender stalk (arrow in Fig. 2A, top), compared to two connections to VP7 for the SA11-4F spike (arrows in Fig. 2B, top). One of the stalk domains of the SA11-4F spike, shown in the forefront of Fig. 2B (top), is large and robust.
Previous studies of SA11-4F documented the presence of a large internal globular domain at the base of the spike, which is below the VP7 layer and interacts with VP6 (16, 19). When viewed down from the outer surface of the capsid, the SA11-Cl3 spike had a very low mass density because of the internal globular domain that was difficult to see even at low contour levels (Fig. 2A, bottom). In contrast, the SA11-4F spike had a higher mass density because of the internal globular domain, which became more pronounced at lower threshold values, connecting it with the VP6 layer at three locations (arrows in Fig. 2B, bottom).
Apart from the differences associated with the VP4 spikes, no other significant differences in the structures of VP7 or VP6 were noted between SA11-4F and SA11-Cl3, an observation fitting with the homology of these gene products. The lower mass density of the VP4 spike in SA11-Cl3 could be due to decreased stoichiometry and/or increased conformational flexibility of the VP4 spikes. Comparative gel analysis of the proteins in these two strains indicates noticeably lower stoichiometry of VP4 in SA11-Cl3 (data not shown). The mass density due to the lower domain is reproducibly absent in several independent reconstructions of SA11-Cl3, perhaps indicating that VP4 spikes in this virus are not stably associated with the capsid. This may also give rise to conformational flexibility of the spikes. These observations, taken together, support the notion that the differences between these viruses in the trypsin cleavage of VP4 and the stability of virion infectivity during purification (Table 1) result from the different origins of VP4 that are manifested as the difference in spike structure between the two viruses. No purified TLPs could be isolated from the third parental virus, B223. This correlates with the instability of B223 during CsCl purification and the rapid decay of B223 infectivity during storage of virus grown in the presence of trypsin (3).
Interactions among VP4, VP7, and VP6 in simian reassortants.
To determine whether differences between the spikes of SA11-Cl3 and SA11-4F are associated with VP4, VP6, or VP7, reassortants with reciprocal parental origins of these proteins were imaged and reconstructed. Reassortant R-924 contains VP4 from SA11-Cl3 on a background of all other proteins from SA11-4F. The structure of R-924 was determined from 126 particles to a resolution of ∼23 Å (Fig. 2C). Reassortant R-A29 has the reciprocal constellation of genome segments and has VP4 of SA11-4F on a background of all other proteins from SA11-Cl3. The structure of R-A29 was determined from 234 particles and reconstructed to a resolution of ∼24 Å (Fig. 2D). The SA11-Cl3 spike on an SA11-4F background of reassortant R-924 had the same general features as in the native virus, where it interacted with SA11-Cl3 proteins. It has the low mass density of the SA11-Cl3 spike, and it makes a relatively thinner connection to VP7 of SA11-4F origin (arrow in Fig. 2C, top). The internal globular domain of R-924 was not well defined, just as in the case of the native SA11-Cl3 structure (Fig. 2C, bottom). Likewise, the SA11-4F spike on an SA11-Cl3 background in the reassortant R-A29 has the same robustness seen in the structure of native SA11-4F. It has two connections to VP7 (arrows in Fig. 2D, top) and has a very high mass density because of the lower globular domain that makes five contacts with VP6 (arrows in Fig. 2D, bottom).
These results indicate that the robust nature of the SA11-4F spike and the large globular domain segregate with the segment 4 product (VP4) and not with the product of segment 6 (VP6) or 9 (VP7). This result is not surprising, since SA11-4F and SA11-Cl3 are thought to share all genes, except segment 4, which is of bovine origin in SA11-4F (10). Movement of the SA11-Cl3 VP4 spike protein onto a background of SA11-4F proteins (R-924) resulted in no significant change in the conformation of the external spike or the globular domain (Fig. 2D). The lack of conformational change in the spike between the respective native and reassortant virions correlated with the conservation of the trypsin cleavage sensitivity of VP4, the neutralization by MAb 2G4, and the stability of the virion with the parental origin of VP4 in the reassortants (Table 1). Thus, the expected phenotypes were expressed and were accompanied by the parental conformation of the VP4 spike.
Interactions among VP4, VP7, and VP6 in simian-bovine reassortants.
To determine the effect of placing the SA11-4F (bovine origin) spike on a background of the other proteins of bovine origin, SA11-4F-B223 reassortants that segregated VP4 were examined. Reassortant R-197 contains the VP4 spike from SA11-4F on a background of all other proteins derived from B223. During purification, most of the particles produced from the infection of cells with R-197 were DLPs, as was the case exclusively with B223, R-600, and R-004. However, in the case of R-197, a faint band of TLPs could be isolated with a very low particle concentration. The structure of R-197 was determined from 32 particles to a resolution of ∼23 Å (Fig. 2E). Reassortant R-004 containing the B223 VP4 spike on a background of all other proteins from SA11-4F could not be purified as intact TLPs. This result was not surprising since R-004 was previously shown to spontaneously lose its spikes (16).
The SA11-4F origin spike of R-197 has structural features of both the SA11-Cl3 and SA11-4F spikes (Fig. 2E), although it is more akin to SA11-4F. The distal portion of the spike has a low central mass density, like that of native SA11-Cl3 (albeit not as much), but has two connections to VP7 like that of the native SA11-4F spike (compare Fig. 2E to Fig. 2A and B, top). The R-197 spike does retain a high mass density because of the internal globular domain, which makes five contacts with VP6, much like that seen in native SA11-4F (compare Fig. 2E and B, bottom). The gel analysis indicates lowered VP4 stoichiometry for R-197 compared to that seen in SA11-4F (data not shown). In the structure of R-197, even with the inclusion of only 37 particles, the mass density due to the internal globular domain is clearly visible. It is possible that the VP4 spike in R-197 is conformationally more rigid than the spike in SA11-Cl3, in which the mass density due to the lower domain was significantly lower. However, the conformation of the VP4 spike in R-197, particularly in the distal domains, appears to be altered, perhaps because of the heterologous interactions of the SA11-4F spikes with bovine VP7 and VP6.
These observations on R-197 spike structure reinforce the conclusion that the strong globular domain is a property of the SA11-4F VP4 spike. The alteration of the conformation of the spike in R-197 is consistent with other observations, such as its low sensitivity to trypsin cleavage, and absence of neutralization by MAb 2G4 (Table 1), whereas these phenotypes were the opposite in native SA11-4F. It is indeed surprising that despite its lower stability, comparable to that of SA11-Cl3 (Table 1), the reconstructed spike in R-197 is robust, with a well-defined internal globular domain. This suggests that SA11-4F VP4 is intrinsically conformationally rigid.
The site of trypsin cleavage on the spike structure is unknown, but the unexpected trypsin resistance of R-197 suggests that the cleavage site is altered in the reassortant spike. It is, however, unlikely that the cleavage site is completely hidden in R-197 since trypsin cleavage stabilizes the spike structure, allowing the spike to change to a conformation stable enough to be imaged and reconstructed (4). MAb 2G4 binds the VP4 spike at the sides of the saddle between the two distal lobes (14, 18). The conformation of this region appears to be altered in R-197 (Fig. 2E, top), which may account for the failure of the virus to express the 2G4 epitope normally present on SA11-4F VP4. The stability of virion infectivity during purification and storage was previously shown to correlate mainly with the parental origin of VP4 (3). In the case of R-197, we note that the stability is about 10-fold less than that of the SA11-4F parent but about 10-fold greater than that of the B223 parent. This represents the significantly increased stability conferred by SA11-4F VP4 on a background of all other genes from extremely unstable B223, consistent with prior observations. Unfortunately, the reciprocal reassortant (R-004) could not be purified for this study. Prior work with this reassortant showed that the VP4 spikes were so unstable that they spontaneously fell off the virions (16). Thus, it was not possible to demonstrate an altered conformation of the B223 spike on an SA11-4F background, although the unexpected phenotypes of the virus strongly suggest altered interactions between VP4 and VP7 and/or VP6 in R-004 (Table 1).
VP6 from B223 does not disrupt an SA11-4F-based reassortant.
To determine if an intermediate VP6 capsid of B223 origin would disrupt or alter the structure of a TLP with all other proteins from SA11-4F, the structure of reassortant R-202 was determined. The R-202 structure was determined from 121 particles and reconstructed to a resolution of 22.8 Å (Fig. 2F). The SA11-4F-derived spike was like that of native SA11-4F, having a structure with robust distal density and two connections to VP7 (Fig. 2F, top) and a high mass density due to a lower globular domain that made five contacts with VP6 (Fig. 2F, bottom). The structure of the reciprocal reassortant, R-600, could not be determined because only DLPs could be purified from this reassortant.
Our structural studies on R-202 indicate that heterologous interactions between VP4 and VP6 do not affect the structure of the spike and strongly suggest that the altered spike structure seen in reassortant R-197 is due to altered interactions between VP4 and VP7. The importance of the VP4-VP7 interaction was emphasized by the fact that some reassortants expressed unexpected phenotypes when VP4 and VP7 were of heterologous parental origin, but the same VP4 protein expressed the expected phenotypes when moved to a heterologous background in the presence of the homologous VP7 protein (2). The reciprocal reassortant (R-600), which moved SA11-4F VP6 onto a background of B223 proteins, could not be purified. This was not surprising, given the instability of this reassortant during CsCl purification and storage as crude lysate (3). In this case, R-600 was slightly more stable than its parent, B223, contributing all of its proteins except VP6, which suggests that the instability of the reassortant was not due to the heterology between VP4 and VP7 and/or VP6 (3).
In summary, some rotavirus reassortants were previously shown to display unexpected phenotypes and these phenotypes were hypothesized to result from heterologous interactions between the proteins of the outer capsid that result in altered conformations (2-4). Here, evidence is provided that shows that this is indeed the case. Reassortants (R-A29 and R-924) in which VP4 interacted with VP7 and VP6 of heterologous parental origin but which expressed the expected VP4-associated phenotypes showed no alterations in the structures of any of the interacting proteins. In contrast, in a reassortant (R-197) in which VP4 interacted with heterologous VP7 and VP6 and which showed unexpected expression of VP4-associated phenotypes, an altered conformation of the VP4 spike was observed. Retention of the parental VP4 spike structure in a reassortant in which only VP6 was of heterologous parental origin showed the importance of VP4-VP7 heterology in unexpected phenotypic expression. These limited structural studies suggest that conformational alterations induced by suboptimal heterologous protein interactions can result in altered viral phenotype expression and that heterology between VP4 and VP7 is the determining factor in the phenotypes examined here. Unexpected phenotypic expression in reassortants should not be surprising, and important reassortants, such as those for vaccines, should be characterized for expression of the expected phenotypes.
Acknowledgments
This work was supported by National Institutes of Health grants AI-16687 (R.F.R.) and AI-36040 (B.V.V.P.). J.B.P. acknowledges the support of the Keck Center for Computational Biology through National Science Foundation training grant BIR-9256580.
REFERENCES
- 1.Chen, D., J. W. Burns, M. K. Estes, and R. F. Ramig. 1989. Phenotypes of rotavirus reassortants depend upon the recipient genetic background. Proc. Natl. Acad. Sci. USA 86:3743-3747. [DOI] [PMC free article] [PubMed]
- 2.Chen, D. Y., M. K. Estes, and R. F. Ramig. 1992. Specific interactions between rotavirus outer capsid proteins VP4 and VP7 determine expression of a cross-reactive, neutralizing VP4-specific epitope. J. Virol. 66:432-439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chen, D. Y., and R. F. Ramig. 1992. Determinants of rotavirus stability and density during CsCl purification. Virology 186:228-237. [DOI] [PubMed] [Google Scholar]
- 4.Crawford, S. E., S. K. Mukherjee, M. K. Estes, J. A. Lawton, A. L. Shaw, R. F. Ramig, and B. V. V. Prasad. 2001. Trypsin cleavage stabilizes the rotavirus VP4 spike. J. Virol. 75:6052-6061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Dormitzer, P. R., and H. B. Greenberg. 1992. Calcium chelation induces a conformational change in recombinant herpes simplex virus-1-expressed rotavirus VP7. Virology 189:828-832. [DOI] [PubMed] [Google Scholar]
- 6.Dormitzer, P. R., Z. Y. Sun, G. Wagner, and S. C. Harrison. 2002. The rhesus rotavirus VP4 sialic acid binding domain has a galectin fold with a novel carbohydrate binding site. EMBO J. 21:885-897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Estes, M. K. 2001. Rotaviruses and their replication, p. 1747-1785. In D. M. Knipe, P. M. Howley, D. E. Griffin, R. A. Lamb, M. A. Martin, B. Roizman, and S. E. Straus (ed.), Fields virology, 4th ed., Lippincott Williams and Wilkins, Philadelphia, Pa.
- 8.Kapikian, A. Z. 2002. Ecological studies, rotavirus vaccination, and intussusception. Lancet 359:1065-1066. [DOI] [PubMed] [Google Scholar]
- 9.Lawton, J. A., and B. V. V. Prasad. 1996. Automated software package for icosahedral virus reconstruction. J. Struct. Biol. 116:209-215. [DOI] [PubMed] [Google Scholar]
- 10.Mattion, N. M., and M. K. Estes. 1991. Sequence of a rotavirus gene 4 associated with unique biologic properties. Arch. Virol. 120:109-113. [DOI] [PubMed] [Google Scholar]
- 11.Mendez, E., C. F. Arias, and S. Lopez. 1996. Interactions between the two surface proteins of rotavirus may alter the receptor-binding specificity of the virus. J. Virol. 70:1218-1222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Pesavento, J. B., J. A. Lawton, M. E. Estes, and B. V. V. Prasad. 2001. The reversible condensation and expansion of the rotavirus genome. Proc. Natl. Acad. Sci. USA 98:1381-1386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Prasad, B. V., G. J. Wang, J. P. Clerx, and W. Chiu. 1988. Three-dimensional structure of rotavirus. J. Mol. Biol. 199:269-275. [DOI] [PubMed] [Google Scholar]
- 14.Prasad, B. V. V., J. W. Burns, E. Marietta, M. K. Estes, and W. Chiu. 1990. Localization of VP4 neutralization sites in rotavirus by three-dimensional cryo-electron microscopy. Nature 343:476-479. [DOI] [PubMed] [Google Scholar]
- 15.Prasad, B. V. V., and M. E. Estes. 2000. Electron cryomicroscopy and computer image processing techniques: use in structure-function studies of rotavirus, p. 9-32. In U. Desselberger (ed.), Rotaviruses—methods and protocols, vol. 34. Humana Press, Inc., Totowa, N.J. [DOI] [PubMed]
- 16.Shaw, A. L., R. Rothnagel, D. Chen, R. F. Ramig, W. Chiu, and B. V. V. Prasad. 1993. Three-dimensional visualization of the rotavirus hemagglutinin structure. Cell 74:693-701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Taniguchi, K., K. Nishikawa, N. Kobayashi, T. Urasawa, H. Wu, M. Gorziglia, and S. Urasawa. 1994. Differences in plaque size and VP4 sequence found in SA11 virus clones having simian authentic VP4. Virology 198:325-330. [DOI] [PubMed] [Google Scholar]
- 18.Tihova, M., K. A. Dryden, A. R. Bellamy, H. B. Greenberg, and M. Yeager. 2001. Localization of membrane permeabilization and receptor binding sites on the VP4 hemagglutinin of rotavirus: implications for cell entry. J. Mol. Biol. 314:985-992. [DOI] [PubMed] [Google Scholar]
- 19.Yeager, M., J. A. Berriman, T. S. Baker, and A. R. Bellamy. 1994. Three-dimensional structure of the rotavirus haemagglutinin VP4 by cryo-electron microscopy and difference map analysis. EMBO J. 13:1011-1018. [DOI] [PMC free article] [PubMed] [Google Scholar]