Abstract
Va14Ja18 natural T (NKT) cells play an immunoregulatory role, which is controlled by a self glycolipid(s) presented by CD1d. Although the synthetic antigen α-D-galactosylceramide (α-D-GalCer) stimulates all Va14Ja18 NKT cells, α-anomeric D-glycosylceramides are currently unknown in mammals. We have used β-D-GalCer-deficient mice and β-D-glucosylceramide (β-D-GlcCer)-deficient cells to define the chemical nature of a natural NKT cell antigen. β-D-GalCer-deficient mice exhibit normal NKT cell development and function, and cells from these animals potently stimulate NKT hybridomas. In striking contrast, the same hybridomas fail to react to CD1d1 expressed by a β-D-GlcCer-deficient cell line. Importantly, human β-D-GlcCer synthase cDNA transfer, and hence the biosynthesis of β-D-GlcCer, restores the recognition of mutant cells expressing CD1d1 by the Va14Ja18 NKT hybridomas. Additionally, suppression of β-D-GlcCer synthesis inhibits antigen presentation to Va14Ja18 NKT cells. The possibility that β-D-GlcCer itself is the natural NKT cell antigen was excluded because it was unable to activate NKT hybridomas in a cell-free antigen-presentation assay. These findings suggest that β-D-GlcCer may play an important role in generating and/or loading a natural Va14Ja18 NKT antigen.
Natural T (NKT) cells represent a unique cellular component of the innate immune system because they utilize a rearranged antigen-specific T cell receptor (Tcr) typical of the adaptive immune system to initiate their effector function. A major subset of mouse NKT cells expresses an invariant Va14Ja18 Tcr α-chain that pairs predominantly with Vb8.2 Tcr β-chain. Va14Ja18 NKT cells are thought to play a key role in immunity to pathogens and tumors as well as to regulate autoreactive and alloreactive responses. These functions of NKT cells are accomplished by their capacity to rapidly elicit a robust effector response upon Tcr engagement in vivo. The rapid NKT cell response to antigen facilitates communications between components of the innate and adaptive immune systems (reviewed in ref. 1).
Cell-to-cell communication through the dynamic formation of the immunological synapse is fundamental to the initiation of a cellular immune response. Central to the synapse is the interaction between the presented antigen and its cognate receptor (2). Thus, knowing the structure of the natural antigen recognized by the Tcr is critically important in understanding the molecular events that occur at the immune synapse. Although much has been learned regarding the interactions between peptide antigens and their cognate Tcr (reviewed in ref. 3), the natural Va14Ja18 NKT cell antigen remains unknown.
Current evidence suggests that Va14Ja18 NKT cells recognize a CD1d1-restricted self-glycolipid antigen(s) that resides within the late endosomes/lysosomes (4–12). Several approaches have been utilized to determine the nature of the Va14Ja18 NKT cell antigen(s). In one approach, a synthetic glycolipid, α-D-galactosylceramide (α-D-GalCer), which displays anti-tumor activity, was identified as a potent in vivo stimulant of all Va14Ja18 NKT lymphocytes (5, 12–19). α-D-Glucosylceramide (α-D-GlcCer) was identified as a second synthetic antigen of Va14Ja18 NKT cells (12, 16). A second approach used an in vitro antigen reconstitution assay to identify extracted cellular phospholipids as antigens of a few Va14Ja18 NKT hybridomas (20). Herein, we report a third approach, which utilizes cells deficient in the enzymes that synthesize β-anomeric forms of D-GalCer and D-GlcCer. The data indicate that β-D-GlcCer may play an important role in generating and/or loading a natural Va14Ja18 NKT cell antigen.
Materials and Methods
Mice.
β-D-GalCer synthase-deficient mice, B6.129-CGT+/0 mice (21), were obtained from B. Popko (University of Chicago, Chicago). Homozygotes were obtained by interbreeding B6.129-CGT+/0 heterozygotes and ascertained by PCR-based genotyping of tail DNA, as described (21). Heterozygous and homozygous wild-type mice served as controls. B6.129-CD1d10/0 mice have been described (22). All mice were bred and maintained in compliance with the regulations of the Vanderbilt University Institutional Animal Care and Use Committee.
Cell Lines and Hybridomas.
All cell lines (23–25) and hybridomas (4, 23, 26, 27) used in this study have been reported previously; they were maintained as described.
CD1d1 Expression.
B16, GM95, and CG1 were infected with recombinant vaccinia virus rVV-CD1d1, rVV-Qa2, or rVV-H2Kb, containing the cDNA encoding CD1d1, H2Q7b, or H2Kb, respectively, at a multiplicity of infection of 10, as described (28). To determine CD1d1 expression, 5 × 105 cells were reacted with 0.5 μg of biotinylated 1B1, a CD1d1-reactive mAb, biotinylated-1-1-2, a Qa2-specific mAb, or biotinylated AF6, an H2Kb-reactive mAb, and detected with streptavidin-CyChrome by using a FACScan or FACSCalibur flow cytometer (Becton Dickinson). All reagents were from PharMingen.
Generation of CD1d-α-d-GalCer Tetramers.
Tetramers of mouse CD1 were produced by streptavidin-mediated tetramerization of folded CD1d1–β2-microglobulin (β2m) heterodimers. A BirA substrate peptide tag was added to the C terminus of CD1d1 by PCR (forward 5′-AGATCTCAATCTGAAGCCCAGCAA-3′ and reverse 5′-GATATCTGCAGAATTGCCCTTGTCCCGCAGCTCCATCTTCATTGCCTCAAAGATTCCTCCAAGTCCACCTCCTTGCCTGGCATCCCAGTAGAGGATGAT-3′ primers). Human β2m was also PCR cloned (forward 5′-GATCTATCCAGCGTACTCCAAAGATT-3′ and reverse 5′-CATGTCTCGATCCCACTT-3′ primers). The products were cloned separately into the BglII–XhoI-digested pMT/BiP/V5-His expression vector (Invitrogen). Drosophila S2 (Invitrogen) cells secreting folded CD1d1–β2m were obtained by cotransfection of the above plasmids together with pCoHygro selection vector (Invitrogen), selection in 0.3 mg/ml hygromycin (Invitrogen), and induction of protein expression with 1 mM CuSO4. Culture supernatant was concentrated by tangential flow filtration (Pall) against a membrane with a molecular weight cut-off of 30,000. CD1d1–β2m complexes were purified by Ni chromatography (HisTrap chelating column; Amersham Pharmacia). CD1d1–β2m complexes were biotinylated by using biotin protein ligase (Avidity) (29) and dialyzed, and biotinylation efficiency was ascertained by using avidin peroxidase (Sigma) to detect plate-bound, biotinylated CD1. CD1d1 monomers (3 μM) were loaded with 9 μM α-D-GalCer (Kirin Brewery, Gunma, Japan) for 12–16 h at room temperature (18). Tetramers were prepared by mixing monomers with streptavidin-allophycocyanin (Molecular Probes) at a molar ratio of 4:1. Va14Ja18 NKT lymphocytes were detected as described previously (30).
NKT Cell Stimulation and ELISA.
About 5 × 104 hybridoma cells per well were mixed with 5 × 105 stimulator thymocytes and cocultured in triplicate for 18–20 h at 37°C. Similarly, equal numbers (5 × 104) of hybridoma cells and virally transduced cell lines were cocultured as above. In some experiments, β-D-GlcCer synthase activity was inhibited in B16 by using 2.5 μM dl-threo-1-phenyl-2-palmitoylamino-3-morpholino-1-propanol (PPMP) plus 10 μM (±)-threo-1-phenyl-2-decanoylamino-3-morpholino-1-propanol hydrochloride (PDMP) (both from Sigma) for the indicated time periods (31). Cells were then virally transduced and used to stimulate NKT hybridomas. Finally, experiments were performed wherein B16 and GM95 cells were treated with 60 μM β-D-glucosylsphingosine (β-D-GlcSph) or 20 μM β-D-GlcCer for 2 days as described (32). Control and treated cells were virally transduced and cocultured with hybridomas as above. IL-2 secreted upon activation was monitored by ELISA as described (23).
Cell-Free Antigen Reconstitution and Inhibition Assay.
Soluble mouse CD1d1 (sCD1d1) was Ni-affinity (Amersham Pharmacia) purified as described (33) and bound to ELISA plates at a concentration of 5 μg/ml. After binding at 4°C for 18 h and blocking of unbound sites with 2% fetal bovine serum (FBS; Life Technologies), plate-bound sCD1d1 was loaded with lipids at the concentrations indicated for 4 h at 37°C. After removing excess lipids, ≈5 × 104 hybridoma cells were added to each well. Controls included wells bound with 2 μg/ml anti-CD3ɛ (positive) or with 5 μg/ml BSA (negative) loaded with 1 μg/ml α-D-GalCer. IL-2 secreted upon activation was monitored by ELISA. Inhibition of α-D-GalCer presentation was assessed by adding increasing concentrations of phosphatidylinositol (PI), β-D-GlcCer (Matrya), or p99 peptide (gift from J. Yewdell, National Institutes of Health, Bethesda) (34) to a solution of sCD1d1 previously loaded with 0.0567 μM stimulatory ligand. After displacement, sCD1d1 was bound to ELISA plates and incubated with hybridomas; IL-2 secreted upon activation was monitored by ELISA. The data are presented as percent inhibition in comparison to maximal stimulation by 0.0567 μM α-D-GalCer.
Thin-Layer Chromatography (TLC).
Total cellular lipids from 5 × 106 cell pellets were extracted by the Bligh–Dyer method (35) and separated by TLC and visualized by charring with orcinol-containing sulfuric acid buffer at 80°C (24).
CD1d1 Internalization and Recycling Assays.
These assays were performed as described (36). Cleavable sulfo-NHS-SS-biotin (Pierce) was used to label surface molecules. To measure internalization, labeled cells were washed and incubated for the indicated times at 37°C, and then surface biotin was stripped off with 0.005 μM reduced glutathione in 0.075 mM NaCl solution containing 0.001 mM EDTA, 0.075 mM NaOH, and 10% FBS. To quantitate recycling, labeled cells were incubated at 37°C for 2 h and then stripped of surface biotin. Cells were incubated at 37°C for the indicated times and surface biotin was stripped a second time. After detergent lysis, the postnuclear lysate was precleared by immune precipitation with normal mouse serum. CD1d1 (1H6), transferrin receptor (C2F2, PharMingen), and H2b class Ia (Y3+B22–249) were captured onto microtiter plates with the indicated mAb. Biotinylated molecules captured by specific mAb were detected by avidin-peroxidase ELISA.
Intracellular Staining and Confocal Microscopy.
Cells (5 × 106) were plated in glass-bottom microwells (MatTek) and incubated overnight at 37°C to allow the cells to adhere to the wells. The cells were then infected with rVV-CD1d1 or rVV-CD1d1Δcyt (cytoplasmic tail-deleted CD1d1) (11) at a multiplicity of infection of 3 for 8 h (28). Infected cells were permeabilized with 0.05% saponin, fixed with 1% vol/vol paraformaldehyde, and stained with 1H6 (11) and 1D4B, a CD1d1- and a LAMP-1 (CD107a)-specific mAb. Specific mAb binding was detected with anti-mouse Ig-FITC (for 1H6) and anti-rat Ig-phycoerythrin (for 1D4B). Labeled cells were visualized by using an MRC 1024 confocal laser-scanning microscope equipped with a krypton–argon laser (Bio-Rad). Dual emission fluorescent images of 0.5-μm optical slices were collected sequentially and processed by using CONFOCAL ASSISTANT (version 4.02, Todd Clark Brelji).
Results and Discussion
β-d-GalCer Deficiency Does Not Alter Recognition by or Antigen Presentation to Va14Ja18 NKT Cells.
All Va14Ja18 NKT cells are potently activated by α-D-GalCer (5, 12–19). Therefore, an α-anomeric glycosphingolipid is most likely the natural NKT cell antigen. The sugar donor for the synthesis of β-D-GalCer is UDP-α-D-galactose. Mammalian cells are not known to synthesize UDP-β-D-galactose. Therefore, we reasoned that if an α-anomeric glycolipid is indeed the natural NKT cell antigen, then β-D-GalCer could serve as the sugar donor for the synthesis of the glycolipid antigen in vivo. Thus, we tested the ability of cells from β-D-GalCer synthase-deficient mice (B6.129-Cgt0/0; ref. 21) to stimulate Va14Ja18 NKT hybridomas and the capacity of these mice to develop Va14Ja18 NKT cells.
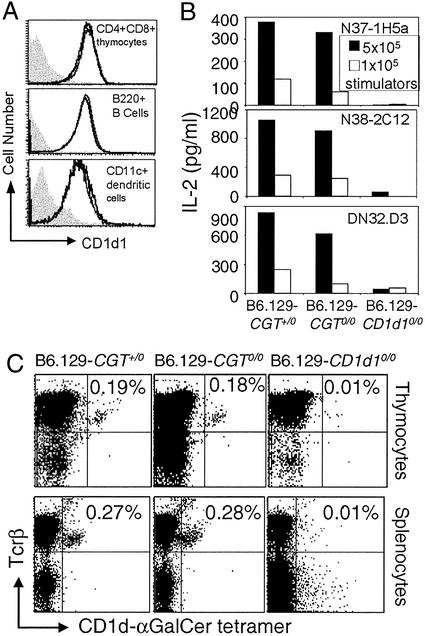
CD4+8+ thymocytes, splenic B220+ B lymphocytes, and CD11c+ dendritic cells from B6.129-Cgt0/0 mice express similar levels of CD1d1 as the control wild-type heterozygotes (Fig. 1A). Further, Va14Ja18 NKT hybridomas recognize CD1d1 expressed by thymocytes of B6.129-Cgt0/0 mice as efficiently as those from B6.129-Cgt0/+ mice (Fig. 1B). As expected, the negative control thymocytes from B6.129-CD1d10/0 mice did not activate the Va14Ja18 NKT hybridomas (Fig. 1B). Thus, B6.129-Cgt0/0 mice express functional CD1d1.
Figure 1.
Expression of functional CD1d1 and development of Va14Ja18 NKT cells in β-D-GalCer synthase-null mice. (A) CD1d1 expression in CD4+8+ thymocytes, B220+ B lymphocytes, and CD11c+ dendritic cells was monitored by flow cytometry after staining with biotinylated CD1-specific mAb 1B1. Note that cells from B6.129-Cgt+/0 and B6.129-Cgt0/0 mice express equal levels of CD1d1, and hence the two histograms overlap. The filled gray histogram represents CD1d1 expression in B6.129-CD1d10/0-deficient cells. (B) The functional status of CD1d1 expressed by β-D-GalCer-containing and -deficient cells was determined in the stimulator–responder coculture system. IL-2 secreted (an indicator of T cell activation) from 5 × 104 hybridoma cells stimulated with 1–5 × 105 wild-type and mutant thymocytes was monitored by ELISA. (C) Dot plots demonstrating the percentage of Va14Ja18 NKT lymphocytes within the thymus and spleen of B6.129-Cgt0/+, B6.129-Cgt0/0, and B6.129-CD1d10/0 mice. Mononuclear cells were stained with Tcrβ-specific H57–597-phycoerythrin mAb and CD1d1-α-D-GalCer tetramer-allophycocyanin. Results are representative of at least three experiments involving at least six mice in each group.
The ligand(s) that positively selects Va14Ja18 NKT cells during development is thought to be similar to the natural antigen (7). Therefore, to confirm the functional nature of CD1d1 in the absence of β-D-GalCer, we determined whether B6.129-Cgt0/0 mice develop this subset of T lymphocytes. For this purpose, CD1d1-α-D-GalCer tetramer was generated and used to detect Va14Ja18 NKT cells. The data revealed the development and/or maintenance of equal numbers of Va14Ja18 NKT cells in B6.129-Cgt+/0 and B6.129-Cgt0/0 (Fig. 1C) as well as in B6.129-Cgt+/+ (data not shown) littermate thymi and spleens (Fig. 1C). Hepatic NKT cells could not be studied because of degeneration of the B6.129-Cgt0/0 liver (data not shown). Because B6.129-Cgt0/0 mice die between 21 and 28 days of age (data not shown and ref. 21), the numbers of Va14Ja18 NKT cells of mice at this young age are lower than those in adult 6- to 8-week-old animals (37). Thus, NKT cell development is normal in β-D-GalCer synthase-null mice, and hence β-D-GalCer is neither the natural antigen nor the precursor of the natural antigen.
β-d-GlcCer Is Essential for Antigen Recognition by Va14Ja18 NKT Cells.
Because α-D-GalCer and α-D-GlcCer but not α-D-mannosylceramide activate NKT cells (12, 16), we reasoned that α-anomeric D-GlcCer might be the natural Va14Ja18 NKT cell antigen. Using the same rationale as discussed above, we conducted a search for β-D-GlcCer synthase-deficient mice and cell lines. β-D-GlcCer synthase-deficient mice are embryonic lethal (38) and hence could not be studied. Instead, GM95, derived from B16 melanoma but deficient in β-D-GlcCer synthase (24), and CG1, GM95 reconstituted with human β-D-GlcCer synthase cDNA (25), were studied.
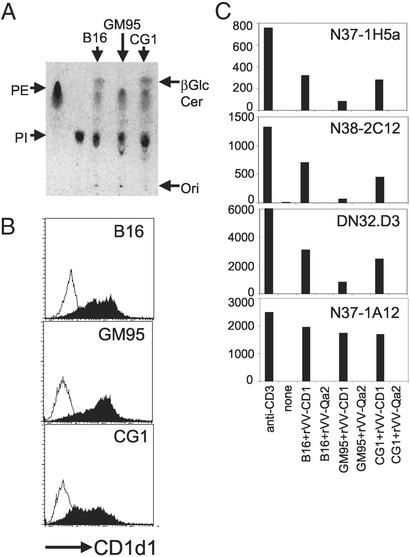
First, we confirmed that B16 and CG1 do indeed synthesize β-D-GlcCer and that GM95 is defective by TLC analysis of their total cellular lipids (Fig. 2A). Second, because none of these cell lines express CD1d1 (data not shown), they were transduced with rVV-CD1d1, resulting in similar levels of cell-surface CD1 expression (Fig. 2B). rVV-Qa2 and rVV-H2Kb were used as negative controls (data not shown). Thus, the β-D-GlcCer deficiency in GM95 does not alter cell-surface expression of CD1d1.
Figure 2.
Defective presentation of the natural Va14Ja18 NKT cell antigen by CD1d1 expressed by β-D-GlcCer synthase-deficient GM95 cells. (A) Total cellular lipids were extracted from B16, GM95, and CG1 cells, separated by TLC, and visualized by charring. Commercial β-D-GlcCer was separated alongside as a standard. PE, phosphatidylethanolamine. (B) The expression of CD1d1 after infection with rVV-CD1d1 (filled histogram) or rVV-Qa2 (open histogram) was monitored after staining with biotinylated-1B1 as in Fig. 1A. (C) Mouse CD1 was expressed in the three cell lines by recent infection with rVV-CD1d1. The ability of Va14Ja18-positive (N37–1H5a, N38–2C12, and DN32.D3) and -negative (N37–1A12) NKT cell hybridomas to recognize CD1d1 expressed by β-D-GlcCer-containing and -deficient cells was determined in the stimulator–responder coculture system described for Fig. 1B. Results are representative of at least three experiments.
Third, to determine whether the CD1d1 expressed by the β-D-GlcCer-deficient GM95 cell line was functional, rVV-CD1d1-infected GM95 cells were cocultured with a panel of NKT hybridomas. Va14Ja18-positive and -negative hybridomas recognize CD1d1 expressed by B16, which synthesizes β-D-GlcCer (Fig. 2C). Remarkably, the same Va14Ja18 NKT hybridomas did not recognize CD1d1 expressed by the β-D-GlcCer-deficient GM95 (Fig. 2C). Recognition by the three hybridomas was restored in the β-D-GlcCer synthase-reconstituted CG1 infected with rVV-CD1d1 (Fig. 2C). Furthermore, GM95 transduced with rVV-CD1d1 was fully capable of presenting α-D-GalCer to Va14Ja18 NKT hybridomas (Fig. 3D), indicating functional expression of CD1 in the mutant.
Figure 3.
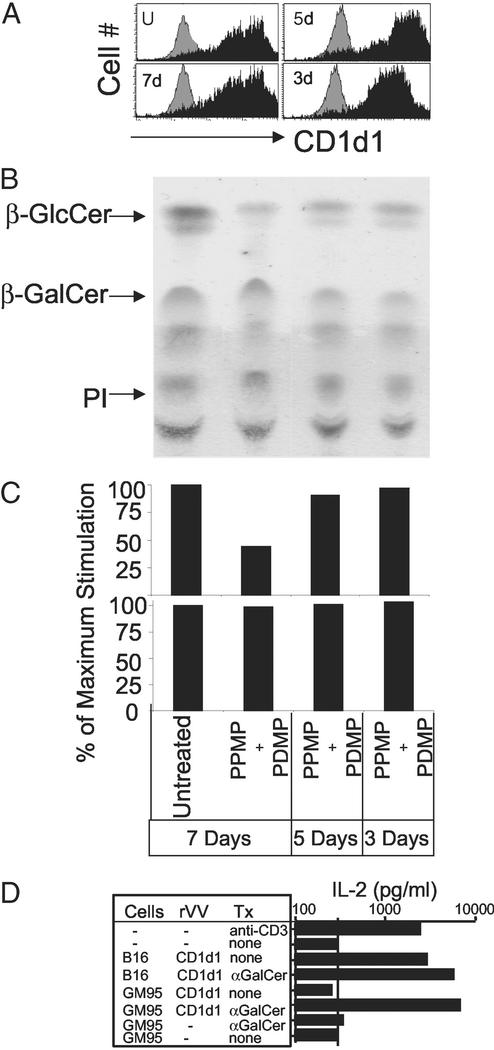
CD1d1 molecules expressed by β-D-GlcCer synthase-deficient cells preserve structural and functional integrity. (A) CD1d levels on uninfected or rVV-CD1d1-infected B16 cells treated for the indicated times with 2.5 μM PPMP and 10 μM PDMP were determined as in Fig. 1A. (B) β-D-GlcCer levels of PPMP+PDMP-treated B16 cells were evaluated by TLC. (C) The ability of Va14Ja18 (Upper, N37–1H5a; N38–2C12 and DN32-D3 not shown) and non-Va14 (Lower, N37–1A12) NKT cell hybridomas to recognize CD1d1 expressed by PPMP+PDMP-treated B16 cells was determined in the stimulator–responder coculture system. (D) Uninfected or recent rVV-CD1d1-infected B16 and GM95 were pulsed with vehicle or 0.5 μM α-D-GalCer for 4 h and washed extensively to remove unbound antigen. The ability of these cells to stimulate N38–2C12 Va14Ja18 NKT hybridoma was tested as described for Fig. 1B. Tx, treatment. Results are representative of three experiments.
As an independent test, β-D-GlcCer biosynthesis was inhibited in B16 (Fig. 3A) with PPMP and PDMP, potent inhibitors of β-D-GlcCer synthase (31). The expression of CD1d1 was not affected by the loss of β-D-GlcCer in this cell line (Fig. 3A) but inhibited antigen recognition by Va14Ja18 NKT hybridomas in a β-D-GlcCer concentration-dependent manner (Fig. 3C). Thus, β-D-GlcCer is essential for antigen recognition by Va14Ja18 NKT hybridomas.
Surprisingly, in striking contract to the lack of recognition of rVV-CD1d1-infected mutant cells by Va14Ja18 NKT hybridomas, these T cells efficiently recognize GM95 stably expressing CD1d1 (data not shown). This recognition might be caused by the leaky phenotype of the β-D-GlcCer synthase-deficient GM95 (24, 25). Alternatively, it may be attributable to the oligoclonal nature of this cell line. However, none of the five single-cell clones derived from GM95 by limiting dilution transduced to express CD1d1 upon rVV-CD1d1 infection activated any of the Va14Ja18 NKT hybridomas (data not shown). Thus, we predict that small amounts of antigen synthesized in GM95 assemble with CD1d1, which over time, because of the long half-life (>1 day) and multiple rounds of recycling of CD1d1 (7), become as efficient in antigen presentation as the wild-type cells.
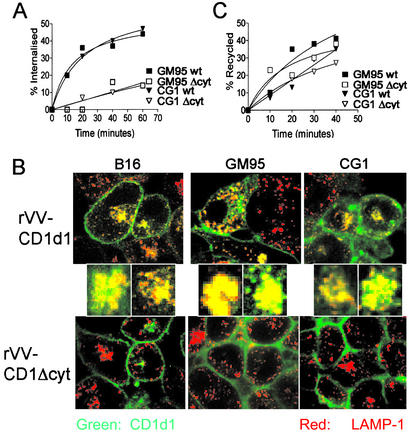
Finally, a recent study revealed that β-D-GlcCer deficiency in GM95 retains tyrosinase, a melanosome-resident protein, in the Golgi apparatus because of a short (17-aa residues) transmembrane domain (32). Thus, CD1d1 targeting to the late endosome/lysosome, albeit less likely because of its 30-aa-residue transmembrane domain, could be altered in GM95. Therefore, first, we determined the rates of CD1d1 internalization and recycling in B16 and GM95, and second, we compared CD1d1 colocalization with LAMP-1 (a marker for late endosomes/lysosomes) in B16, GM95, and CG1 infected with rVV-CD1d1 or with control rVV-CD1d1Δcyt (a mutant defective in negotiating the low-pH compartment). The data revealed that the rates of internalization and recycling of CD1d1 were similar in GM95 and CG1 (Fig. 4 A and C). The internalized wild type, but not the mutant CD1d1, significantly colocalizes with LAMP-1 in all cell lines tested, including GM95 (Fig. 4B). A previous study demonstrated that exogenous addition of β-D-GlcSph, but not β-D-GlcCer, significantly rescues melanosomal targeting of tyrosinase in GM95 cells (32). To independently ascertain that CD1d1 trafficking in GM95 was intact, β-D-GlcSph was added to GM95 cells; after 2 days they were infected with rVV-CD1d1. CD1d1 so expressed by β-D-GlcSph-treated GM95 was not recognized by Va14Ja18 NKT hybridomas (data not shown). Previous studies demonstrated that efficient α-D-GalCer presentation requires CD1d1 internalization and recycling (7, 12, 16). Thus, normal α-D-GalCer presentation by CD1d1 (Fig. 3D) and colocalization of CD1 with LAMP-1 (Fig. 4B) suggest intact intracellular trafficking through the natural NKT cell antigen-containing compartment in β-D-GlcCer-deficient GM95.
Figure 4.
Normal CD1d1 trafficking through late endosomes/lysosomes in β-D-GlcCer synthase-deficient cells. (A) Internalization of wild-type CD1d1 and CD1d1Δcyt expressed by GM95 and CG1 was monitored by using biotin labeling of cell-surface molecules as described in Materials and Methods. H2b class I and transferrin receptor were used as negative and positive controls (data not shown). (B) The traffic of CD1d1 through the low-pH endosomal/lysosomal compartment was monitored by the colocalization of mouse CD1 with LAMP-1 by using confocal microscopy. Wild-type CD1d1 or CD1d1Δcyt was expressed by infection of B16, GM95, and CG1 with the corresponding rVV. CD1d1 was detected 8 h after viral transduction with a specific mAb 1H6 (green) and LAMP-1 with 1D4B (red). (C) Recycling of wild-type CD1d1 or CD1d1Δcyt expressed by GM95 and CG1 was monitored after biotin labeling of cell-surface molecules as described in Materials and Methods. Results are representative of three (A and C) or four (B) independent experiments.
Cell-Free CD1d1 Presents α-d-GalCer but Not β-d-GlcCer to Va14Ja18 NKT Hybridomas.
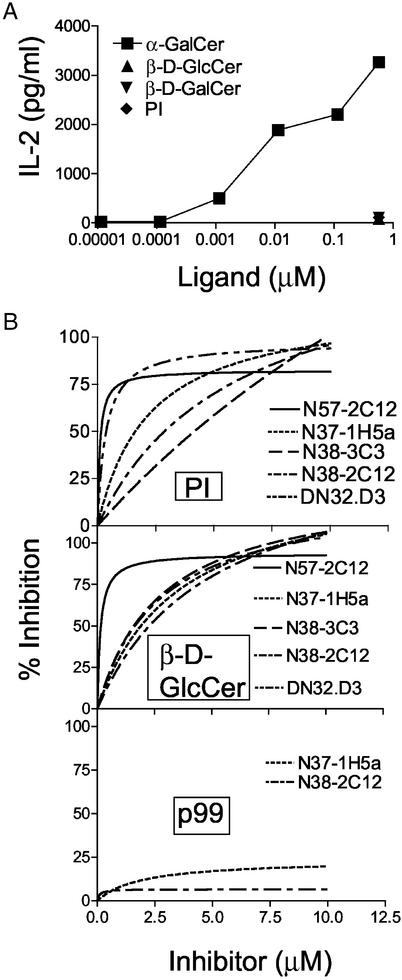
To determine whether mouse CD1 directly presents β-D-GlcCer to Va14Ja18 NKT cells, a cell-free functional CD1d1 reconstitution assay (described in Materials and Methods) was established. PI, β-D-GlcCer, β-D-GalCer, and α-D-GalCer were used as ligands to reconstitute stimulatory function in otherwise inert plate-bound soluble CD1d1 (sCD1d1). As expected (20, 39), the data revealed that α-D-GalCer reconstituted functional sCD1d1 because the complex activated all of the Va14Ja18-positive but not the Va14-negative NKT cell hybridomas tested (Fig. 5A). Neither the Va14Ja18-positive nor the Va14-negative NKT hybridomas were activated by any of the other CD1d1 ligands (Fig. 5A).
Figure 5.
Specific cell-free recognition of CD1d1-α-D-GalCer by Va14Ja18 NKT hybridomas. (A) Microtiter wells were coated with 5 μg/ml sCD1d1, blocked with fetal bovine serum, and pulsed with the indicated lipids. Increasing concentrations of α-D-GalCer or a fixed concentration of PI, β-D-GalCer, or β-D-GlcCer was used. Cell-free presentation of lipids to NKT hybridomas by plate-bound sCD1d1 was monitored. (B) Plate-bound sCD1d1 was first pulsed with 0.0567 μM α-D-GalCer, and then increasing concentrations of PI or β-d-GlcCer were added. The ability of PI, β-D-GlcCer, and p99, a hydrophobic peptide known to bind CD1d1, to inhibit the presentation of α-D-GalCer was tested by using Va14Ja18 NKT hybridomas. In all assays, IL-2 secretion indicated the recognition of antigen by the hybridomas. Results are representative of at least three experiments.
To confirm that PI and β-D-GlcCer indeed bind plate-bound sCD1d1, we established a competition assay. Briefly, plate-bound sCD1d1 was loaded with an optimal stimulatory dose (0.0567 μM) of α-D-GalCer, and the ability of increasing concentrations of PI and β-D-GlcCer to inhibit the presentation of the antigen was determined. A peptide, p99, that binds CD1d1 (34) but does not compete with α-D-GalCer for CD1 binding (39), was used as a negative control for the inhibition assay. PI and β-D-GlcCer inhibit the presentation of α-D-GalCer by plate-bound sCD1d1 to Va14Ja18 NKT hybridomas (Fig. 4B). As expected from a previous report (39), p99 did not inhibit the presentation of α-D-GalCer to Va14Ja18 NKT cell hybridomas (Fig. 5B). Thus, PI and β-D-GlcCer bind plate-bound sCD1d1 but do not directly activate Va14Ja18 NKT cells.
Implications for the Chemical Basis of a Natural Va14Ja18 NKT Cell Antigen.
Our data reveal that β-D-GlcCer binds CD1d1 but is not directly recognized by Va14Ja18 NKT cells. However, cells deficient in β-D-GlcCer are unable to present the natural CD1d1-restricted antigen to Va14Ja18 NKT cells. Therefore, because α-D-GlcCer stimulates Va14Ja18 NKT cells in vitro (12, 16), we predict that β-D-GlcCer may be a precursor for the biosynthesis of a natural antigen. If this is indeed true, then α-D-GalCer is a close mimic of a natural Va14Ja18 NKT cell antigen.
β-D-GlcCer synthase activity localizes to the cytosolic face of the Golgi apparatus, and hence β-D-GlcCer biosynthesis occurs at this intracellular site (40, 41). β-D-GlcCer then actively flips to the luminal face of the Golgi apparatus, where it subserves a precursor function for glucosphingolipid biosynthesis (40, 41). If β-D-GlcCer is a precursor of the Va14Ja18 NKT cell antigen, then the biosynthesis of the antigen occurs either in the Golgi apparatus or in a late intracellular vesicle, but distal to the site of biosynthetic assembly of CD1d1. This conclusion is consistent with prior reports demonstrating that CD1d1 assembles with the Va14Ja18 NKT cell antigen in late endosomes/lysosomes (6, 7, 11, 12, 23).
An alternative explanation that may account for defective antigen presentation in β-D-GlcCer-deficient cells is that β-D-GlcCer may be essential for the function(s) of an accessory protein(s) that loads Va14Ja18 NKT cell antigen onto CD1d1 in late endosomes/lysosomes. Alterations in function(s) could occur either because of improper intracellular targeting of such proteins to the site of CD1d1-antigen assembly or purely because of protein dysfunction in the absence of β-D-GlcCer. Identification of the accessory proteins that facilitate CD1d1-antigen assembly and their characterization in GM95 will distinguish between these two possibilities that account for defective CD1d1 antigen presentation in β-D-GlcCer-deficient cells.
Because humans develop Va24Ja18/Vb11 T lymphocytes, an exact homologue of mouse Va14Ja18/Vb8.2 NKT cells, the findings reported herein have implications for antigen recognition by human NKT cells as well. Finally, solving the molecular mechanism(s) for defective CD1d1-restricted antigen presentation in β-D-GlcCer-deficient GM95 has the potential to yield significant new insights into glycolipid antigen presentation and recognition within the immune system.
Acknowledgments
We thank the Kirin Brewery for generous supplies of synthetic α-d-GalCer, B. Popko for B6.129-CGT0/+ mice, A. Bendelac for the DN32.D3 hybridoma, O. Naidenko and M. Kronenberg for help in preparing CD1d1 tetramers, Gerrit van Meer for insightful discussions, and members of the Joyce laboratory for technical assistance and helpful discussions. This work was supported by National Institutes of Health Grants AI46455 (to R.R.B.) and AI42284 (to S.J.), the Juvenile Diabetes Research Foundation, and the Human Frontiers in Science Program (to S.J.).
Abbreviations
- α-d-GalCer
α-d-galactosylceramide
- β-d-GalCer
β-d-galactosylceramide
- β-d-GlcCer
β-d-glucosylceramide
- β-d-GlcSph
β-d-glucosylsphingosine
- β2m
β2-microglobulin
- NKT
natural T
- PDMP
(±)-threo-1-phenyl-2-decanoylamino-3-morpholino- 1-propanol hydrochloride
- PPMP
dl-threo-1-phenyl-2-palmitoylamino-3-morpholino-1-propanol
- PI
phosphatidylinositol
- sCD1d1
soluble CD1d1
- Tcr
T cell receptor
Footnotes
This paper was submitted directly (Track II) to the PNAS office.
References
- 1.Joyce S. Cell Mol Life Sci. 2001;58:442–469. doi: 10.1007/PL00000869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Grakoui A, Bromley S K, Sumen C, Davis M M, Shaw A S, Allen P M, Dustin M L. Science. 1999;285:221–227. [PubMed] [Google Scholar]
- 3.Krummel M, Wulfing C, Sumen C, Davis M M. Philos Trans R Soc London B. 2000;355:1071–1076. doi: 10.1098/rstb.2000.0644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bendelac A, Lantz O, Quimby M E, Yewdell J W, Bennink J R, Brutkiewicz R R. Science. 1995;268:863–865. doi: 10.1126/science.7538697. [DOI] [PubMed] [Google Scholar]
- 5.Brossay L, Chioda M, Burdin N, Koezuka Y, Casorati G, Dellabona P, Kronenberg M. J Exp Med. 1998;188:1521–1528. doi: 10.1084/jem.188.8.1521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Chiu Y H, Jayawardena J, Weiss A, Lee D, Park S H, Dautry-Varsat A, Bendelac A. J Exp Med. 1999;189:103–110. doi: 10.1084/jem.189.1.103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chiu Y H, Park S H, Benlagha K, Forestier C, Jayawardena-Wolf J, Savage P B, Teyton L, Bendelac A. Nat Immunol. 2002;3:55–60. doi: 10.1038/ni740. [DOI] [PubMed] [Google Scholar]
- 8.Exley M, Garcia G, Balk S P, Porcelli S. J Exp Med. 1997;186:109–120. doi: 10.1084/jem.186.1.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Molano A, Park S H, Chiu Y H, Nosseir S, Bendelac A, Tsuji M. J Immunol. 2000;164:5005–5009. doi: 10.4049/jimmunol.164.10.5005. [DOI] [PubMed] [Google Scholar]
- 10.Park S-H, Roark J H, Bendelac A. J Immunol. 1998;160:3128–3134. [PubMed] [Google Scholar]
- 11.Roberts T J, Sriram V, Spence P M, Gui M, Hayakawa K, Bacik I, Bennink J R, Yewdell J W, Brutkiewicz R R. J Immunol. 2002;168:5409–5414. doi: 10.4049/jimmunol.168.11.5409. [DOI] [PubMed] [Google Scholar]
- 12.Spada F M, Koezuka Y, Porcelli S A. J Exp Med. 1998;188:1529–1534. doi: 10.1084/jem.188.8.1529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Benlagha K, Weiss A, Beavis A, Teyton L, Bendelac A. J Exp Med. 2000;191:1895–1904. doi: 10.1084/jem.191.11.1895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Burdin N, Brossay L, Koezuka Y, Smiley S T, Grusby M J, Gui M, Taniguchi M, Hayakawa K, Kronenberg M. J Immunol. 1998;161:3271–3281. [PubMed] [Google Scholar]
- 15.Carnaud C, Lee D, Donnars O, Park S-H, Beavis A, Koezuka Y, Bendelac A. J Immunol. 1999;163:4647–4650. [PubMed] [Google Scholar]
- 16.Kawano T, Cui J, Koezuka Y, Toura I, Kaneko Y, Motoki K, Ueno H, Nakagawa R, Sato H, Kondo E, et al. Science. 1997;278:1626–1629. doi: 10.1126/science.278.5343.1626. [DOI] [PubMed] [Google Scholar]
- 17.Kobayashi E, Motoki K, Uchida T, Fukushima H, Koezuka Y. Oncol Res. 1995;7:529–534. [PubMed] [Google Scholar]
- 18.Matsuda J L, Naidenko O V, Gapin L, Nakayama T, Taniguchi M, Wang C-R, Koezuka Y, Kronenberg M. J Exp Med. 2000;192:741–753. doi: 10.1084/jem.192.5.741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Singh N, Hong S, Scherer D C, Serizawa I, Burdin N, Kronenberg M, Koezuka Y, Van Kaer L. J Immunol. 1999;163:2373–2377. [PubMed] [Google Scholar]
- 20.Gumperz J E, Roy C, Makowska A, Lum D, Sugita M, Podrebarac T, Koezuka Y, Porcelli S A, Cardell S, Brenner M B, Behar S M. Immunity. 2000;12:211–221. doi: 10.1016/s1074-7613(00)80174-0. [DOI] [PubMed] [Google Scholar]
- 21.Coetzee T, Fujita N, Dupree J, Shi R, Blight A, Suzuki K, Popko B. Cell. 1996;86:209–219. doi: 10.1016/s0092-8674(00)80093-8. [DOI] [PubMed] [Google Scholar]
- 22.Mendiratta S K, Martin W D, Hong S, Boesteanu A, Joyce S, Van Kaer L. Immunity. 1997;6:469–477. doi: 10.1016/s1074-7613(00)80290-3. [DOI] [PubMed] [Google Scholar]
- 23.De Silva A D, Park J-J, Matsuki N, Stanic A K, Brutkiewicz R R, Medof M E, Joyce S. J Immunol. 2002;168:723–733. doi: 10.4049/jimmunol.168.2.723. [DOI] [PubMed] [Google Scholar]
- 24.Ichikawa S, Nakajo N, Sakiyama H, Hirabayashi Y. Proc Natl Acad Sci USA. 1994;91:2703–2707. doi: 10.1073/pnas.91.7.2703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ichikawa S, Sakiyama H, Suzuki G, Hidari K I-P J, Hirabayashi Y. Proc Natl Acad Sci USA. 1996;93:4638–4643. doi: 10.1073/pnas.93.10.4638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Brossay L, Tangri S, Bix M, Cardell S, Locksley R, Kronenberg M. J Immunol. 1998;160:3681–3688. [PubMed] [Google Scholar]
- 27.Gui M, Li J, Wen L J, Hardy R R, Hayakawa K. J Immunol. 2001;167:6239–6246. doi: 10.4049/jimmunol.167.11.6239. [DOI] [PubMed] [Google Scholar]
- 28.Brutkiewicz R R, Bennink J R, Yewdell J W, Bendelac A. J Exp Med. 1995;182:1913–1919. doi: 10.1084/jem.182.6.1913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Altman J D, Moss P, Goulder P, Barouch D H, McHeyzer W M, Bell J I, McMichael A J, Davis M M. Science. 1997;274:94–96. doi: 10.1126/science.274.5284.94. [DOI] [PubMed] [Google Scholar]
- 30.Boesteanu A, De Silva A D, Nakajima H, Leonard W J, Peschon J J, Joyce S. J Exp Med. 1997;186:331–336. doi: 10.1084/jem.186.2.331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sandvig K, Garred O, van Helvoort A, van Meer G, van Deurs B. Mol Biol Cell. 1996;7:1391–1404. doi: 10.1091/mbc.7.9.1391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sprong H, Degroote S, Claessens T, van Drunen J, Oorschot V, Westerink B H, Hirabayashi Y, Klumperman J, van der Sluijs P, van Meer G. J Cell Biol. 2001;155:369–380. doi: 10.1083/jcb.200106104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Joyce S, Woods A S, Yewdell J W, Bennink J R, De Silva A D, Boesteanu A, Balk S P, Cotter R J, Brutkiewciz R R. Science. 1998;279:1541–1544. doi: 10.1126/science.279.5356.1541. [DOI] [PubMed] [Google Scholar]
- 34.Castano A R, Tangri S, Miller J E W, Holcombe H R, Jackson M R, Huse W D, Kronenberg M, Peterson P A. Science. 1995;269:223–226. doi: 10.1126/science.7542403. [DOI] [PubMed] [Google Scholar]
- 35.Bligh E G, Dyer W J. Can J Biochem Physiol. 1959;37:911–917. doi: 10.1139/o59-099. [DOI] [PubMed] [Google Scholar]
- 36.Turvy D N, Blum J S. J Immunol Methods. 1998;212:9–18. doi: 10.1016/s0022-1759(97)00206-8. [DOI] [PubMed] [Google Scholar]
- 37.Bendelac A, Killeen N, Litman D R, Schwartz R H. Science. 1994;263:1774–1778. doi: 10.1126/science.7907820. [DOI] [PubMed] [Google Scholar]
- 38.Yamashita T, Wada R, Sasaki T, Deng C, Bierfreund U, Sandhoff K, Proia R L. Proc Natl Acad Sci USA. 1999;96:9142–9147. doi: 10.1073/pnas.96.16.9142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Naidenko O V, Maher J K, Ernst W A, Sakai T, Modlin R L, Kronenberg M. J Exp Med. 1999;190:1069–1080. doi: 10.1084/jem.190.8.1069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Burger K N J, van der Bijl P, van Meer G. J Cell Biol. 1996;133:15–28. doi: 10.1083/jcb.133.1.15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Marks D L, Wu K, Paul P, Kaminska R, Pagano R E. J Biol Chem. 1999;274:451–456. doi: 10.1074/jbc.274.1.451. [DOI] [PubMed] [Google Scholar]