Abstract
The influenza A virus promoter is recognized by the influenza A virus RNA-dependent RNA polymerase, and directs both transcription and replication of the viral RNA genome. Within the sequence of this promoter, flu strains exhibit a natural, unique variation, either a U or a C, at the fourth position from the 3′ end. Promoters that contain a C residue (C4 promoter), which are invariably found in genome segments that encode the three RNA polymerase subunits (PB1, PB2 and PA), down-regulate transcription but activate genome replication. Here, we have determined the structure of the C4 promoter by NMR spectroscopy and compared it with the structure of the U4 promoter, which was determined previously. The structure of the internal loop in the C4 promoter is similar to that of the U4 promoter. However, the terminal stem of the C4 promoter is strikingly different from that of the U4 promoter. These structural data suggest that the internal loop is important for polymerase binding to the promoter, and the terminal stem is crucial for differential regulation of transcription and replication.
INTRODUCTION
The genome of influenza A virus (vRNA) consists of eight single-stranded RNA segments of negative polarity (1). These vRNA segments are transcribed into mRNA and replicated to produce cRNA, a full-length complementary copy of the vRNA (Fig. 1D). Both transcription and replication of the genome are performed by the same viral ribonucleoprotein (RNP) complex, which consists of three RNA-dependent RNA polymerase (RdRp) protein subunits (PB1, PB2 and PA) and nucleoproteins (NP). The mRNA carries a host-derived 7-methyl guanosine cap structure at its 5′ end and a poly(A) tail at its 3′ end, and directs the synthesis of all viral proteins. The cRNA serves as a template for the synthesis of progeny vRNA, which is used in the construction of cRNP complexes.
Figure 1.
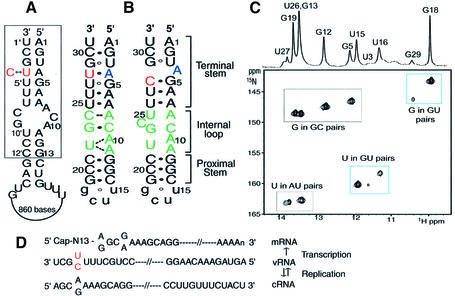
(A) Terminal sequences of vRNA from the influenza A virus. Boxed sequences are converged in all of the influenza A virus variants. Numbering of the 3′ strand is followed by a prime notation (′). The sequence shown is that of vRNA segment 8 of influenza A/PR/8/34. The C4′ residue is a natural variation in vRNA of various strains of influenza A virus. (B) Secondary structures of the U4 promoter, which was determined previously (left), and the C4 promoter, which was determined in this study (right). Bases in red indicate sites of natural sequence variation observed in vRNA segments from influenza A virus. Non-wild-type residues are given as lower case letters. (C) The imino region of the 15N-1H HSQC spectrum at pH 5.5. (D) Graphic of transcription and replication of influenza A virus vRNA.
The influenza A virus RdRp binds specifically to the partial duplex structure found in the viral genes (also referred to as the promoter or panhandle RNA), which consists of the 5′ and 3′ termini of each RNA genome segment (2,3). In this partial duplex, 13 nt at the 5′ end and 12 nt at the 3′ end are highly conserved among most influenza A virus variants (Fig. 1A). All of the regulatory signals for the initiation of transcription and replication reside in these terminal sequences (4–6), and several lines of evidence suggest that the promoter is also involved in gene packaging and polyadenylation (7,8).
Within the conserved sequences of the promoter, there is single, natural sequence variation at position 4 from the 3′ terminus (this position contains either a U or C). According to sequence data, the three polymerase genes (PB1, PB2 and PA) invariably carry a C residue at this position (C4), while most of the other viral genes carry a U residue at this position (U4) (9). Two exceptions are the neuraminidase (NA) and matrix protein (M) genes, in which both nucleotides have been observed at position 4 from the 3′ end. Research using two isogenic A/WSN/33 influenza viruses (a C4 and U4 virus) revealed that the C4 RNA promoter is involved in the down-regulation of transcription (10). Because viral polymerase should be required in only catalytic amounts (11), this down-regulation would be important for utilizing the host cells’ resources effectively. In addition to the effect on transcription, researchers also observed that the C4 promoter activates replication, compared with the U4 promoter, indicating that the C/U variation at position 4 is related to regulation of transcription and replication efficiencies. Recently, it was also reported that a single, natural sequence variation in DNA promoter, which is the recognition site of TATA-binding protein, reveals decreased transcription efficiency by inducing helical bending change (12,13). This implies that the structural change of promoter by a single sequence variation is involved in transcription efficiency.
The structure of the U4 RNA promoter was determined previously in our laboratory by NMR spectroscopy, and revealed the following remarkable features: (i) an A–C mismatch that is stacked into a helix, extending the helical pattern to the end of the internal loop; (ii) an adenosine located at the junction between the internal loop and proximal stem that is displaced toward the minor groove, forming a novel (A-A)-U motif; and (iii) coincidence of the assumed boundary of the open complex with position 4 from the 3′ terminus, at which bending (46 ± 10°) occurs (14).
Even with the structural information that exists, two important questions remain: (i) how are the U4 and C4 nucleotides involved in the differential regulation of transcription and replication, and (ii) what are the structural bases for the specific interactions between either the U4 or C4 RNA promoter and RdRp? In order to address these questions, we have solved the solution structure of the C4 promoter (PDB accession no. 1MFY). It revealed that the structural characters of the internal loop are similar to those of the U4 promoter, but the terminal-stem structure is quite different. The terminal stem of the C4 promoter has a unique adenosine bulge structure and flexible base pairs. These structural differences between the terminal stems of the U4 and C4 promoters raise the possibility that the conformation of the promoter might be involved in the differential transcription and replication.
MATERIALS AND METHODS
RNA sample preparation
The RNA samples were prepared by cleaving substrate RNA with a trans-cleaving hammerhead ribozyme RNA, as was done previously in the preparation of the U4 promoter (14). The use of a hammerhead ribozyme RNA enabled the preparation of an RNA molecule that has an intact sequence in the terminal region (Fig. 1A and B). The substrate RNA was prepared by transcription from DNA templates using T7 RNA polymerase, and was then purified with Prep-cell (Bio-Rad). The cleavage reaction by the hammerhead ribozyme RNA was performed by incubation in 28 mM MgCl2 and 50 mM Tris–HCl (pH 7.9), for 60 min at 55°C, with an RNA/ribozyme ratio of 20:1. The cleaved RNAs were purified by polyacrylamide gel electrophoresis and then electroeluted from the gel (15). The RNAs for the NMR sample were subsequently dialyzed against buffer with either 10% D2O/90% H2O or 99.9% D2O. Unlabeled and 13C,15N-labeled RNAs were prepared at concentrations of ∼1.5 and ∼0.5 mM, respectively, in 0.01 mM EDTA/10 mM phosphate buffer (pH 6.5).
NMR spectroscopy
NMR spectra were acquired at 400, 600 and 800 MHz on Bruker DRX spectrometers and a 600 MHz Varian Unity Inova spectrometer. The spectra from the Bruker spectrometers were processed with XWIN-NMR software, while those from the Varian spectrometer were processed with NMR-Pipe 2.1 and Varian VNMR software. All of the spectra were analyzed with SPARKY 3.95 (University of California, San Francisco). Using an unlabeled sample, two-dimensional (2D) D2O NOESY (nuclear Overhauser effect correlation spectroscopy) with 80, 150 and 250 ms mixing times were obtained at 294 K. Also, 2D H2O NOESY (90% H2O/10% D2O) with 150 and 300 ms mixing times was recorded at 278 K. Two-dimensional DQF-COSY (double quantum filtered correlated spectroscopy) was recorded on unlabeled RNA at 294 K. In order to estimate stability of the base pairs, 1D temperature experiments were performed at 278, 283, 288, 293, 298, 303 and 308 K. The heteronuclear (13C, 15N) experiments used for assignment were 13C-CT-HSQC (heteronuclear single quantum correlation spectroscopy), 15N-1H-HSQC, 2D HCCH-COSY and 2D HCCH-TOCSY. The 3D experiment, HCCH-COSY-TOCSY, was carried out at 294 K (16). The proton-detected 31P-1H heteronuclear correlation (HETCOR) was obtained at 294 K (17). The 13C T1ρ (rotating frame spin-lattice relaxation time) measurements on the adenine C2 were performed on a 13C, 15N-labeled sample with the pulse sequence described previously (18). Relaxation times were extracted by linear fitting of the logarithm of the integrated peak volumes from the 2D spectra.
Spectral analysis and structural calculation
Exchangeable base proton resonances were assigned from the 2D H2O NOESY, and base pairings were identified using 15N-1H HSQC. Most of the sugar protons were assigned in the D2O NOESY and confirmed by DQF-COSY, 2D HCCH-COSY, 2D HCCH-TOCSY, 13C-CT-HSQC and 3D HCCH-COSY-TOCSY. Several of the protons (H3′, H4′ and H5′/H5″) were newly assigned by 31P-1H HETCOR spectra. A total of 445 NOE distance restraints and 196 dihedral restraints were used for the structure calculation. NOE distance restraints from D2O NOESY were characterized as strong (0–3.4 Å), medium (0–4.5 Å), weak (0–6 Å) and very weak (0–7 Å), and the NOE restraints from H2O NOESY were classified as strong (1.6–4.1 Å), medium (2–5 Å) and weak (2–7 Å). Dihedral torsion restraints were obtained from 31P-1H HETCOR (α, β and ε), DQF-COSY (δ) and D2O NOESY (χ) (19). Structures were calculated using a distance geometry simulated annealing (DGSA) protocol with CNS software (20). A total of 100 starting structures were generated by distance geometry. Based on the simulated annealing protocol, the structures were heated to 3000 K for 20 ps and then cooled over 27 ps to 300 K. During this initial round of calculations, the distance force constant was 50 kcal mol–1A–2, and the dihedral angle constant scaled from 10 to 400 kcal mol–1rad–2. A total of 15 final structures were analyzed by Insight II (Biosym Technologies, San Diego) and MOLMOL (Institut für Molekularbiologie und Biophysik, Switzerland).
RESULTS
Overall structure of the C4 promoter
The C4 promoter structure consists of a terminal stem, an internal loop, a proximal stem and a UUCG tetraloop (Fig. 1B). H2O NOESY and 15N –1H HSQC showed that the C4 promoter has a G–U wobble pair, four G–C pairs and two A–U pairs in the terminal and proximal stems. The overall helix has a standard A-form geometry, as evidenced by typical NOEs in D2O NOESY. The terminal stem is rather flexible, because of a lack of stable base pairs and the helix-disrupting A-bulge. The internal loop showed no evidence of base pairing in the NMR spectra, even when obtained at low pH (pH 5.5). In spite of this fact, all of the residues in the 5′ strand of the internal loop are well stacked into the helix. The structure of the UUCG tetraloop was similar to that suggested by a previously determined structure (14,21). A total of 726 restraints derived from NMR data were used for structure calculations to obtain the 15 converged structures (Table 1). The average structure is shown in Figure 2A.
Table 1. Structural statistics for the C4 promoter structure of influenza A virus.
| Total number of restraints | 726 |
| NOE distance restraints | 445 |
| Dihedral restraints (α, β, γ, δ, ε, ζ and χ) | 196 |
| Base planarity restraints | 27 |
| Base pair restraints including hydrogen bonding | 58 |
| Root mean square deviation for all heavy atoms relative to the mean structure (Å) | |
| Terminal stem (residues 1–7, 26–31) | 1.8 ± 0.2 |
| Internal loop (residues 8–11, 22–25) | 1.7 ± 0.5 |
| UUCG tetraloop (residues 14–19) | 0.3 ± 0.1 |
| All nucleotides | 2.4 ± 0.5 |
| NOE violations (Å) | 0 (>0.5 Å) |
| Angle violations (°) | 0 (>5°) |
| Mean deviation from covalent geometry | |
| Bond lengths (Å) | 0.001 |
| Angles (°) | 0.5 |
| Impropers (°) | 0.2 |
Figure 2.
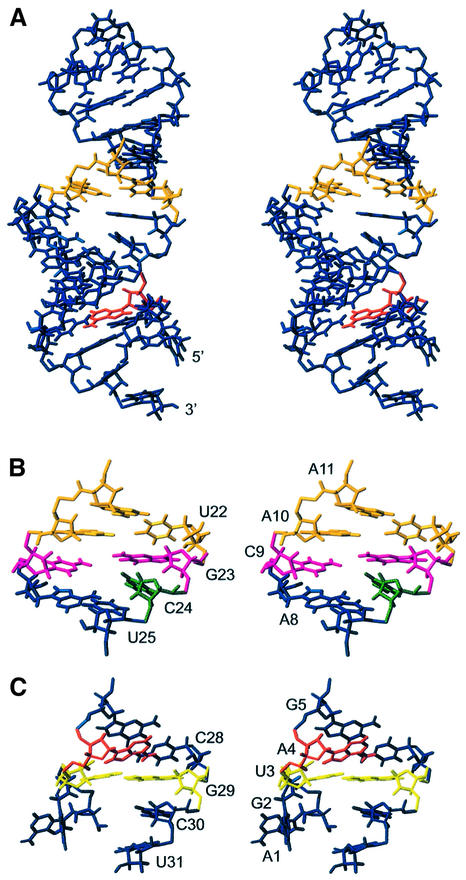
(A) Stereo view of the overall structure of the influenza A virus C4 promoter. Residues in orange are (A-A)-U structures. Residues in red represents the A4-bulge. (B) Stereo view of the internal loop in the C4 promoter. The C9 and G23 (magenta) are placed in nearly the same plane. The C24 (green) has a divergent conformation. (C) Stereo view of the terminal stem in the C4 promoter. Residues in yellow, U3 and G29, form a base pair in the terminal stem.
Terminal stem
The terminal stem is composed of a single base (A4) bulge and two small flanking stems. In the D2O NOESY, sequential NOEs, such as H8/H6i-H1′i–1, H8/H6i-H2′i–1 and H8/H6i-H3′i–1, were identified for most of the residues in the terminal stem, but were not detected for U31 to C30 and A4 to U3. This indicates that the terminal stem forms a typical A-helix, although the sequential connectivity is partially broken. Analysis of the 15N-1H HSQC showed that this terminal stem has one G–C base pair (G5–C28), two A–U base pairs (A6–U27 and A7–U26) and a U–G wobble pair (U3–G29) (Fig. 1C). Three canonical base pairs (G5–C28, A6–U27 and A7–U26) constructed one small stem beside the internal loop. The G5–C28 base pair was newly induced by the natural variation (U to C) at position 4 from the 3′ end. The G5 imino resonance in the 15N-1H HSQC spectra, which is evidence of the G5–C28 base pair, had weaker intensity than did the other G–C pairs in the stem. Also, the G5–C28 base pair melted easily at lower temperature in the variable temperature 1D experiment (data not shown). These results show that the G5–C28 base pair is less stable than the other G–C pairs.
The A1–U31, G2–C30 and U3–G29 base pairs were part of another small stem located near the end of the terminal stem (Fig. 2C). Actually, the first two base pairs (A1–U31 and G2–C30) were not identified in either the H2O NMR spectra or the 15N-1H HSQC spectra, even at low pH (5.5), due to rapid solvent exchange. This observation implies that the small terminal stem is somewhat flexible. In addition to the A1–U31 and G2–C30 base pairs, the U3–G29 wobble pair was also flexible. The peak intensities of U3 and G29 were weaker than those of the other base pairs represented in the 15N-1H HSQC spectra (Fig. 1C), indicating that the small stem in the promoter terminus is completely flexible.
A4-bulge structure
Because of the presence of the G5–C28 and U3–G29 base pairs, the A4 residue is not paired and forms a bulge structure in the terminal stem (Fig. 2C). The weak but evident long-range NOE between A4 and G29 (A4H2–G29H1′), and a sequential NOE (A4H2-G5H1′) in the D2O NOESY (Fig. 3A) indicated that A4 stacks slightly between the G5–C28 and U3–G29 base pair planes. However, although this stacking interaction exists, sequential NOEs from U3 to A4 and A4 to G5 are lacking. These observations indicate that the A4-bulge is not fully stacked in the helical stem. Actually, the A4-bulge is protruded toward the minor groove in the calculated structure. Several previous studies of single adenosine bulges showed that an adenine bulge flanked on one side by a non-Watson–Crick base pair adopts an extra-helical conformation (22,23). In this regard, the partial stacking of the A4-bulge studied here is consistent with previous findings.
Figure 3.
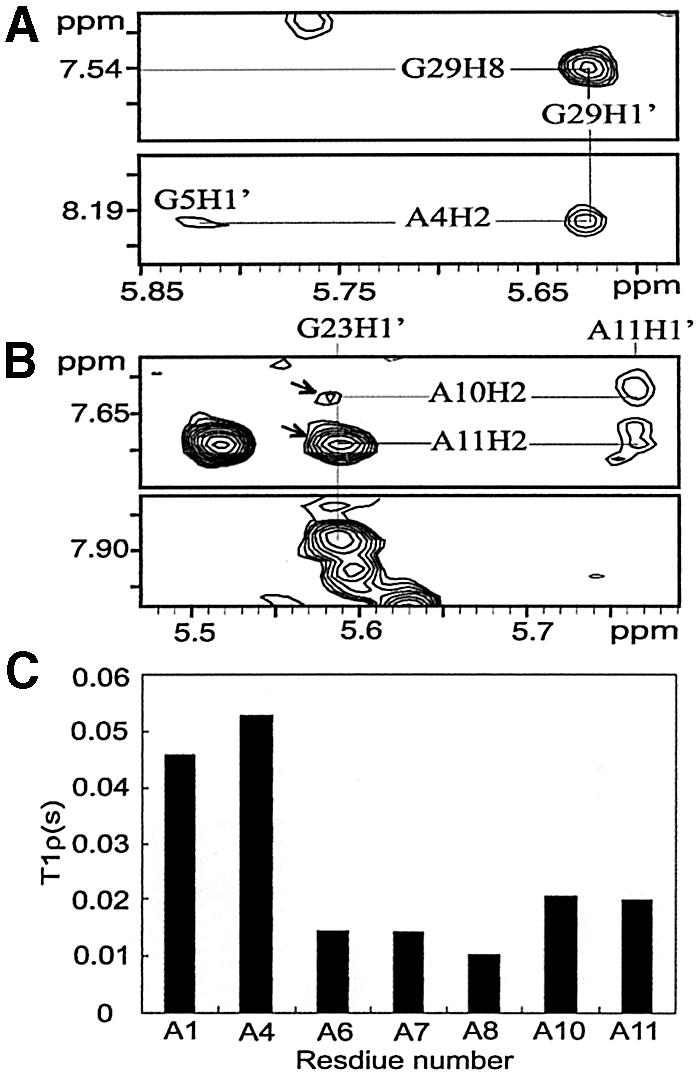
(A) NOEs between A4 and G29, and (B) NOEs between A11/A10 and G23, as observed with D2O NOESY with a 250 ms mixing time. Arrows indicate the cross-peaks of A11H2–G23H1′ and A10H2–G23H1′. (C) Histogram of the T1ρ for adenine C2 resonances in the C4 promoter.
Measurement of 13C T1ρ relaxation times for adenines in the influenza A virus RNA revealed notable dynamic character associated with the A4-bulge. The 13C T1ρ relaxation time for a particular carbon type (C2) is related to the internal motion of the base. Disorder on the pico- to nanosecond time- scale increases the T1ρ value, whereas chemical-exchange processes on the micro- to millisecond time-scale decrease the T1ρ value (24). The T1ρ of A4 residue (52.7 ms) was much larger than those of adenines in the stem (∼14 ms), and was even larger than that of the terminal A1 residue (45 ms) (Fig. 3C). This augmentation of the T1ρ value implies that the A4 base in the stem is highly disordered on the pico- to nanosecond time-scale.
Internal loop
We next assessed the structure of the internal loop of the C4 promoter and found that it is entirely flexible, with no base pairs. In the H2O NOESY, there was no evidence of base pairing, such as an imino–amino cross-peak for G–C pairs or an H2–imino cross-peak, which is associated with A–U pairs. This absence of base pairing in the internal loop resulted in a highly flexible internal loop structure. However, despite the absence of base pairing, all of the residues in the 5′ strand of the internal loop were well stacked into the helix. In D2O NOESY, the sequential connectivity of the A8 to A11 nucleotide in the 5′ strand was continuous. In addition, the A8H2–C9H1′, A10H2–A11H1′ and A11H2–G12H1′ cross-peaks confirmed the presence of a stacked structure. Contrary to the 5′ strand, for most of the residues in the 3′ strand, the sequential NOEs were weak or absent in the NMR spectra. Most notable was the C24 residue, which showed only intra-residue NOEs. Due to this lack of inter-residue NOEs, the internal loop displayed a less-converged structure.
According to our calculated structure, the A8 and U25 residues are rather close to each other, and are nearly co-planar (Fig. 2B). Also, both residues are stacked into the helix, as is evidenced by the NOE connectivity for A7 to C9 and U25 to U26. These observations indicate that A8 and U25 are part of the extended helix of the terminal stem, although we did not observe their base pairing, possibly due to rapid water exchange. This base stacking (for A8 and U25) stabilized the junction between the stem and the internal loop. Like the A8 and U25 residues, base pairing between C9 and G23 was not observed, but these residues were shown to be placed in nearly the same plane in the calculated structure. Also, the C9 residue was well stacked into A8 and A10; thus, its stacking stabilization might result from this base pair-like geometry with G23.
C24 in the 3′ strand had no sequential connectivity in the D2O NOESY. Also, the 3JH1′-H2′ obtained in the DQF-COSY was 6 Hz, indicating that C24 has a dynamic sugar conformation [equilibrium between the C2′ endo (3JH1′-H2′ > 8–9 Hz) and the C3′ endo (3JH1′-H2′ < 2 Hz)]. The C24 base was also shown to protrude towards the major or minor groove, and to display a divergent conformation in the calculated structure. Thus, we suggest that the lack of conformational rigidity associated with the C24 residue might destabilize the C9–G23 base pair and, consequently, the entire internal loop.
The A10 and A11 residues exhibit intra-stand H2i-H1′i+1 NOEs typical of A-form RNA helices. Also, two cross-strand NOEs, A10H2–G23H1′ and A11H2–G23H1′, were observed in the D2O NOESY (Fig. 3B). In the context of the potential base pair between C9 and G23, the A11H2–G23H1′ peak is unusual, because the distance between A11 and G23 is too far for NOEs to be detected in the standard A-form helix geometry. However, this unusual NOE could be explained by the displacement of A11 into the minor groove. The U22 residue was shown not to be base paired with either A10 or A11, and to display divergent conformations. Consequently, the A10, A11 and U22 residues revealed structural and spectral analogies to the previously determined (A10-A11)-U22 motif of the U4 promoter.
Comparison with the U4 promoter
The terminal structures of both promoters are quite different in terms of their secondary structure (Fig. 1B). The A4 residue of the U4 promoter is paired with U28, and displays bending property (46 ± 10°) near this base pair (14). However, the A4 residue of the C4 promoter was found not to be paired, because C28 is paired with G5; thus A4 forms a bulge structure of highly dynamic character. The secondary structure near A4 and G5 is important for transcriptional efficiency (25). Therefore, these differences in the structure of the A4 residue in the C4 promoters may be related to the interaction between vRNA and RdRp during transcription initiation. In addition to the differences in secondary structure, the stabilities of the terminal stems in both promoters are notably divergent. In the C4 promoter, one of the stems flanking the A4-bulge revealed more flexible and unstable base pairs than did the corresponding region of the U4 promoter (Fig. 4B and D). Despite the identical sequences in this region (A1–U3 and G29–U31), the differences in base pair stability suggest that the A4-bulge influences the structure of the terminal stem.
Figure 4.
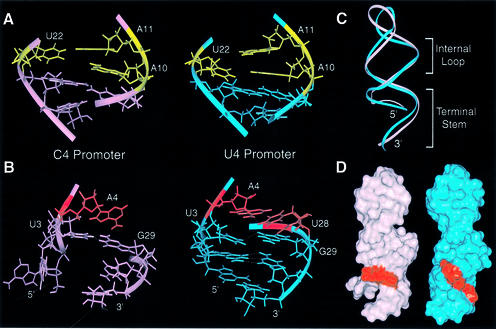
(A) Comparison between the internal loops of the C4 (left) and U4 (right) promoters. In both structures, the (A10-A11)-U22 motif and the potential hydrogen-bonding geometries of the C9–G23 and A8–U25/C24 pairs are similar. (B) Comparison between terminal stems of the C4 (left) and U4 (right) promoters. Residues in red are the A4-bulge (C4) and the A4–U28 pair (U4). The dotted lines indicate hydrogen bonding. (C) Superposition of ribbon representations of the C4 (pink) and the U4 (blue) promoters. (D) Surface representations of the C4 (left) and U4 (right) promoters. Residues in red are the A4-bulge of the C4 promoter (left) and the A4–U28 base pair of the U4 promoter (right).
Although the terminal stem structures of the C4 and U4 promoters are different, both promoters show similar internal loop structures. For example, A10, A11 and U22 in the C4 promoter exhibit singular spectral features such as a long NOE (A11H2–G23H1′) (Fig. 3B). These spectral features are also evident in the (A10-A11)-U22 structure of the U4 promoter. Consequently, the A11 residues of both promoters are displaced into the minor groove, which results in a widening of the major grooves of each helix. It is intriguing that although the C24 residue of the C4 promoter is bulged out and that of the U4 promoter is base paired with the A8 residue, the overall internal loop structures of both promoters closely resemble each other (Fig. 4A and C).
DISCUSSION
DNA-dependent RNA polymerase (DdRp) carries out a series of steps during the transcription reaction: specific recognition of the DNA promoter by DdRp, promoter localization, melting of the DNA to form an active open complex, synthesis of the first phosphodiester bond, abortive RNA synthesis, promoter clearance, transcriptional elongation, and termination (26). Similarly, during the initiation of transcription, RdRp must recognize the promoter sequences and then localize its catalytic core at the promoter in order to form an open complex. Studies of two isogenic influenza viruses, the C4 and U4 viruses, revealed that the C4 promoter down-regulates transcription compared with the U4 promoter (10). Because the C4 promoter activates replication even during transcription, the transcriptional yield is decreased and the replication yield is increased for this promoter. This differential transcription of C4 and U4 promoters might be the result of differential promoter recognition and localization by the RdRp. Based on the solution structures of both promoters, we propose a model for distinct modes of initiation carried out by the C4 and U4 promoters during influenza A virus transcription (Fig. 5).
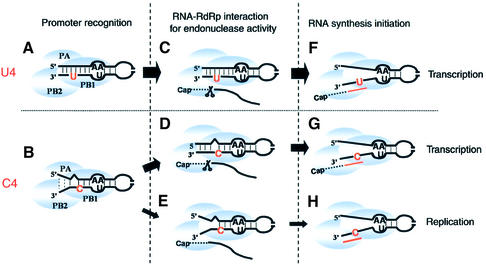
Figure 5.
Transcription-initiation models for the U4 and C4 promoters. Residues in red indicate the naturally variable sites in both promoters. (A and B) Promoter recognition by the influenza RdRp by using the common (A-A)-U structure. (C and D) RNA–RdRp complexes that have endonuclease activity, on both promoters. (E) The C4 promoter–RdRp complex. This complex induces cap-structure binding, but does not display endonuclease activity. (F and G) Transcription initiation at the U4 and C4 promoters using a cap primer. (H) De novo initiation of vRNA replication from the C4 promoter.
In our model, in the first step of transcription initiation, the influenza A virus RdRp specifically recognizes the common structure of both promoters, the (A10-A11)-U22 structure, for the cognate RNA binding (Fig. 5A and B). A modification interference assay identified residues A8 to A11 in the internal loop as the polymerase binding site (27), while photochemical crosslinking assays suggested that the residues C20 to G23 are critical for the transcription activity (28). Also, not the 3′ bulge (5′ A: U–U 3′) but the 5′ bulge (5′ A–A: U 3′) in the internal loop is important for packaging (29). Since the packaging only occurred in vRNA, it is possible that the (A-A)-U structure could be a unique signal of the vRNA for the polymerase binding. Comparison of the C4 and U4 promoters reveals that the internal loop structures of both promoters are essentially the same; in both promoters, A11, which is displaced toward the minor groove, results in the widening of the major groove that contains the (A10-A11)-U22 residues. It is possible that the widening of the major groove in the internal loop reveals points of contact for the RdRp during transcription initiation. Also, because RNA-binding proteins prefer preformed and flexible structures as binding sites (30,31), unique structures with local, dynamic properties, such as the (A-A)-U motif, would be useful for specific recognition by the RdRp.
Continuing with our proposed model, after specific recognition of the promoter, the RdRp localizes the promoter at its catalytic core. Then, before it initiates transcription, the RdRp uses its endonuclease activity to snatch the cap structure from a host mRNA. It is during these steps of the transcription process that the influenza RdRp distinguishes between the C4 and U4 promoters. The conspicuous structural differences between the U4 and C4 promoters are in the terminal stems of both promoters. The C4 promoter has flexible base pairs and an A4-bulge protruding toward the minor groove, while the U4 promoter has the A4–U28 pair with bending properties and other stable base pairs (Fig. 4B). The bent part of the U4 promoter, which exposes the minor groove, has been proposed as an anchor site for the RdRp, as has been observed for the TATA-binding protein (32). Therefore, we suggest that the influenza RdRp binds to the region around the A4 residue, and the structure of the minor groove allows the protein complex to distinguish between the two promoters and initiate differential transcription. The highly dynamic properties of the A4 base in the C4 promoter imply that the RdRp contacts occur at this site.
In addition to the A4 structure, the flexibility of base pairs in the terminal stem might also be critical for differential transcription from the two promoters. The stable duplex structure of the terminal region might induce the endonuclease activity of the RNA–RdRp complex, and this activity is essential for cap-dependent transcription. The endonuclease activity of the RdRp requires both the 5′ and 3′ regions of the influenza promoter (33). Although the single-stranded 5′ end can induce binding of the cap structure, it alone could not effectively trigger endonuclease activity. In the case of the U4 promoter, the terminus is a predominantly stable double-helical structure, thus activating endonuclease activity (Fig. 5C) and the subsequent transcription process. However, for the C4 promoter, the terminal region is in equilibrium between the single- and double-stranded structures, so that a lower proportion of the C4 promoter molecules than that of the U4 promoter molecules induces endonuclease activity (Fig. 5D) and transcription initiation. Actually, it has been shown that single nucleotide mutation (U4 to C4) of the influenza promoter results in slightly lower endonuclease activity (34).
Replication of influenza vRNA is cap-independent and is initiated de novo (35), so that the initiation step does not require the endonuclease activity. Also, de novo initiation generally requires that several nucleotides at the 3′ end of the promoter be unpaired (36,37). In fact, in an NMR study of the cRNA promoter of influenza A virus, which is the template for replication, we found that the terminal region of the cRNA is entirely unpaired (our unpublished data). Thus, it is tempting to speculate that the portion of the C4 promoter with single-stranded character prevents induction of the endonuclease activity, and thus allows replication to be initiated by the RNA–RdRp complex. In other words, endonuclease-positive versions of the RNA–RdRp complex on both promoters (Fig. 5C and D) will initiate transcription by using the cap primer (Fig. 5F and G), but the endonuclease-negative complex (Fig. 5E) on the C4 promoter will begin replication (Fig. 5H) with the assistance of viral NP (38). Consequently, during the same time period, the C4 promoter will give rise to more replication product and less mRNA than will the U4 promoter.
So far, three distinct promoter models have been proposed to explain the initiation of the transcription of the influenza virus RNA: the panhandle (14,39), RNA-fork (40,41) and corkscrew (42,43), where the corkscrew model has been suggested especially for the promoter–polymerase open complex. The differences in these models reside at the very end of the promoter structure, and the debate on the promoter structure has not yet been finished. Although Fodor et al. (40) and Flick et al. (42) assumed that the 5′ and 3′ ends of the vRNA would not form the panhandle structure, we and Baudin et al. (39) have shown that the 5′ and 3′ ends are base paired in the naked state. Thus, we suggest that initially the polymerase recognizes and binds to the proposed bulge region of double-stranded vRNA termini, followed by melting of the termini, generating the RNA fork or the corkscrew structure in the open complex. The importance of the terminal-stem structure was demonstrated in one study where the influenza A virus promoter was cloned into a more internally located position. Even when the native promoter was located in the middle of the double-stranded RNA, while leaving the terminal-stem-disrupting mutant promoter in the original position, the initiation site of the transcription was at an internally displaced location (44). It has been known that the polymerase binding affinity increases significantly when both the 5′ and 3′ strands exist (45). From these results, and our structural data, we infer that the influenza A virus promoter forms a double-stranded terminal structure, and its helical structure has possible regulatory roles in the initiation of RNA synthesis.
Acknowledgments
ACKNOWLEDGEMENTS
We thank Professor Masatsune Kainosho and Dr Chojiro Kosima for assistance with the NMR experiments. This work was supported by the National Creative Research Initiative (grant to B.-S.C.), and by a National Research Laboratory program (grant to C.C.) from the Ministry of Science and Technology, Republic of Korea. M.-K.L., S.-H.B. and C.-J.P. were supported partially by the BK21 project.
PDB no. 1MFY
REFERENCES
- 1.McCauley J.W. and Mahy,B.W. (1983) Structure and function of the influenza virus genome. Biochem. J., 211, 281–294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Desselberger U., Racaniello,V.R., Zazra,J.J. and Palese,P. (1980) The 3′ and 5′-terminal sequences of influenza A, B and C virus RNA segments are highly conserved and show partial inverted complementarity. Gene, 8, 315–328. [DOI] [PubMed] [Google Scholar]
- 3.Hsu M.T., Parvin,J.D., Gupta,S., Krystal,M. and Palese,P. (1987) Genomic RNAs of influenza viruses are held in a circular conformation in virions and in infected cells by a terminal panhandle. Proc. Natl Acad. Sci. USA, 84, 8140–8144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Fodor E., Pritlove,D.C. and Brownlee,G.G. (1994) The influenza virus panhandle is involved in the initiation of transcription. J. Virol., 68, 4092–4096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Pritlove D.C., Fodor,E., Seong,B.L. and Brownlee,G.G. (1995) In vitro transcription and polymerase binding studies of the termini of influenza A virus cRNA: evidence for a cRNA panhandle. J. Gen. Virol., 76, 2205–2213. [DOI] [PubMed] [Google Scholar]
- 6.Cianci C., Tiley,L. and Krystal,M. (1995) Differential activation of the influenza virus polymerase via template RNA binding. J. Virol., 69, 3995–3999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Zheng H., Palese,P. and Garcia-Sastre,A. (1996) Nonconserved nucleotides at the 3′ and 5′ ends of an influenza A virus RNA play an important role in viral RNA replication. Virology, 217, 242–251. [DOI] [PubMed] [Google Scholar]
- 8.Pritlove D.C., Poon,L.L., Fodor,E., Sharps,J. and Brownlee,G.G. (1998) Polyadenylation of influenza virus mRNA transcribed in vitro from model virion RNA templates: requirement for 5′ conserved sequences. J. Virol., 72, 1280–1286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Robertson J.S. (1979) 5′ and 3′ terminal nucleotide sequences of the RNA genome segments of influenza virus. Nucleic Acids Res., 6, 3745–3757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lee K.-H. and Seong,B.L. (1998) The position 4 nucleotide at the 3′ end of the influenza virus neuraminidase vRNA is involved in temporal regulation of transcription and replication of neuraminidase RNAs and affects the repertoire of influenza virus surface antigens. J. Gen. Virol., 79, 1923–1934. [DOI] [PubMed] [Google Scholar]
- 11.Inglis S.C. and Mahy,B.W. (1979) Polypeptides specified by the influenza virus genome. III. Control of synthesis in infected cells. Virology, 95, 154–164. [DOI] [PubMed] [Google Scholar]
- 12.Wu J., Parkhurst,K.M., Powell,R.M., Brenowitz,M.and Parkhurst,L.J. (2001) DNA bends in TATA-binding protein-TATA complexes in solution are DNA sequence-dependent. J. Biol. Chem., 276, 14614–14622. [DOI] [PubMed] [Google Scholar]
- 13.Powell R.M., Parkhurst,K.M. and Parkhurst,L.J. (2002) Comparison of TATA-binding protein recognition of a variant and consensus DNA promoters. J. Biol. Chem., 277, 7776–7784. [DOI] [PubMed] [Google Scholar]
- 14.Bae S.-H., Cheong,H.-K., Lee,J.-H., Cheong,C., Kainosho,M. and Choi,B.-S. (2001) Structural features of an influenza virus promoter and their implications for viral RNA synthesis. Proc. Natl Acad. Sci. USA, 98, 10602–10607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Wyatt J.R., Chastain,M. and Puglish,J.D. (1991) Synthesis and purification of large amounts of RNA oligonucleotides. Biotechniques, 11, 764–769. [PubMed] [Google Scholar]
- 16.Hu W., Kakalis,L.T., Jiang,L., Jiang,F., Ye,X. and Majumdar,A. (1998) 3D HCCH-COSY-TOCSY experiment for the assignment of ribose and amino acid side chains in 13C labeled RNA and protein. J. Biomol. NMR, 12, 559–564. [DOI] [PubMed] [Google Scholar]
- 17.Sklenar V., Miyashiro,H., Zon,G., Miles,H.T. and Bax,A. (1986) Assignment of the 31P and 1H resonances in oligonucleotides by two-dimensional NMR spectroscopy. FEBS Lett., 208, 94–98. [DOI] [PubMed] [Google Scholar]
- 18.Farrow N.A., Muhandiram,R., Singer,A.U., Pascal,S.M., Kay,C.M., Gish,G., Shoelson,S.E., Pawson,T., Forman-Kay,J.D. and Kay,L.E. (1994) Backbone dynamics of a free and phosphopeptide-complexed Src homology 2 domain studied by 15N NMR relaxation. Biochemistry, 33, 5984–6003. [DOI] [PubMed] [Google Scholar]
- 19.Varani G., Aboul-ela,F. and Allain,F.H. (1996) NMR investigation of RNA structure. Prog. Nucl. Mag. Res. Sp., 29, 51–127. [Google Scholar]
- 20.Brünger A.T., Adams,P.D., Clore,G.M., DeLano,W.L., Gros,P., Grosse-Kunstleve,R.W., Jiang,J.S., Kuszewski,J., Nilges,M., Pannu,N.S. et al. (1998) Crystallography and NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D, 54, 905–921. [DOI] [PubMed] [Google Scholar]
- 21.Allain F.H. and Varani,G. (1995) Structure of the P1 helix from group I self-splicing introns. J. Mol. Biol., 250, 333–353. [DOI] [PubMed] [Google Scholar]
- 22.Greenbaum N.L., Radhakrishnan,I., Patel,D.J. and Hirsh,D. (1996) Solution structure of the donor site of a trans-splicing RNA. Structure, 4, 725–733. [DOI] [PubMed] [Google Scholar]
- 23.Kalurachchi K., Uma,K., Zimmermann,R.A. and Nikonowicz,E.P. (1997) Structural features of the binding site for ribosomal protein S8 in Escherichia coli 16S rRNA defined using NMR spectroscopy. Proc. Natl Acad. Sci. USA, 94, 2139–2144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Palmer A.G. III (1997) Probing molecular motion by NMR. Curr. Opin. Struct. Biol., 7, 732–737. [DOI] [PubMed] [Google Scholar]
- 25.Neumann G. and Hobom,G. (1995) Mutational analysis of influenza virus promoter elements in vivo. J. Gen. Virol., 76, 1709–1717. [DOI] [PubMed] [Google Scholar]
- 26.McClure W.R. (1985) Mechanism and control of transcription initiation in prokaryotes. Annu. Rev. Biochem., 54, 171–204. [DOI] [PubMed] [Google Scholar]
- 27.Tiley L.S., Hagen,M., Matthews,J.T. and Krystal,M. (1994) Sequence-specific binding of the influenza virus RNA polymerase to sequences located at the 5′ ends of the viral RNAs. J. Virol., 68, 5108–5116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Fodor E., Seong,B.L. and Brownlee,G.G. (1993) Photochemical cross-linking of influenza A polymerase to its virion RNA promoter defines a polymerase binding site at residues 9 to 12 of the promoter. J. Gen. Virol., 74, 1327–1333. [DOI] [PubMed] [Google Scholar]
- 29.Tchatalbachev S., Flick,R. and Hobom,G. (2001) The packaging signal of influenza viral RNA molecules. RNA, 7, 979–989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Bouvet P., Allain,F.H., Finger,L.D., Dieckmann,T. and Feigon,J. (2001) Recognition of pre-formed and flexible elements of an RNA stem–loop by nucleolin. J. Mol. Biol., 309, 763–775. [DOI] [PubMed] [Google Scholar]
- 31.Mao H., White,S.A. and Williamson,J.R. (1999) A novel loop–loop recognition motif in the yeast ribosomal protein L30 autoregulatory RNA complex. Nature Struct. Biol., 6, 1139–1147. [DOI] [PubMed] [Google Scholar]
- 32.Kim J.L. and Burley,S.K. (1994) 1.9 Å resolution refined structure of TBP recognizing the minor groove of TATAAAAG. Nature Struct. Biol., 1, 638–653. [DOI] [PubMed] [Google Scholar]
- 33.Hagen M., Chung,T.D., Butcher,J.A. and Krystal,M. (1994) Recombinant influenza virus polymerase: requirement of both 5′ and 3′ viral ends for endonuclease activity. J. Virol., 68, 1509–1515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Leahy M.B., Dobbyn,H.C. and Brownlee,G.G. (2001) Hairpin loop structure in the 3′ arm of the influenza A virus virion RNA promoter is required for endonuclease activity. J. Virol., 75, 7042–7049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Young R.J. and Content,J. (1971) 5′-terminus of influenza virus RNA. Nature New. Biol., 230, 140–142. [DOI] [PubMed] [Google Scholar]
- 36.Dreher T.W. (1999) Functions of the 3′-untranslated regions of positive strand RNA viral genomes. Annu. Rev. Phytopathol., 37, 151–174. [DOI] [PubMed] [Google Scholar]
- 37.Chen D. and Patton,J.T. (1998) Rotavirus RNA replication requires a single-stranded 3′ end for efficient minus-strand synthesis. J. Virol., 72, 7387–7396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Portela A. and Digard,P. (2002) The influenza virus nucleoprotein: a multifunctional RNA-binding protein pivotal to virus replication. J. Gen. Virol., 83, 723–734. [DOI] [PubMed] [Google Scholar]
- 39.Baudin F., Bach,C., Cusack,S. and Ruigrok,R.W. (1994) Structure of influenza virus RNP. I. Influenza virus nucleoprotein melts secondary structure in panhandle RNA and exposes the bases to the solvent. EMBO J., 13, 3158–3165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Fodor E., Pritlove,D.C. and Brownlee,G.G. (1995) Characterization of the RNA-fork model of virion RNA in the initiation of transcription in influenza A virus. J. Virol., 69, 4012–4019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kim H.J, Fodor,E., Brownlee,G.G. and Seong,B.L. (1997) Mutational analysis of the RNA-fork model of the influenza A virus vRNA promoter in vivo. J. Gen. Virol., 78, 353–357. [DOI] [PubMed] [Google Scholar]
- 42.Flick R., Neumann,G., Hoffmann,E., Neumeier,E. and Hobom,G. (1996) Promoter elements in the influenza vRNA terminal structure. RNA, 2, 1046–1057. [PMC free article] [PubMed] [Google Scholar]
- 43.Flick R. and Hobom,G. (1999) Interaction of influenza virus polymerase with viral RNA in the ‘corkscrew’ conformation. J. Gen. Virol., 80, 2565–2572. [DOI] [PubMed] [Google Scholar]
- 44.Flick R. and Hobom,G. (1999) Transient bicistronic vRNA segments for indirect selection of recombinant influenza. Virology, 262, 93–103. [DOI] [PubMed] [Google Scholar]
- 45.Gonzalez S. and Ortin,J. (1999) Characterization of influenza virus PB1 protein binding to viral RNA: two separate regions of the protein contribute to the interaction domain. J. Virol., 73, 631–637. [DOI] [PMC free article] [PubMed] [Google Scholar]