Abstract
AtPep1 is a 23-aa endogenous peptide elicitor from Arabidopsis leaves that signals the activation of components of the innate immune response against pathogens. Here, we report the isolation of an AtPep1 receptor from the surface of Arabidopsis suspension-cultured cells. An 125I-labeled AtPep1 analog interacted with suspension-cultured Arabidopsis with a Kd of 0.25 nM, and an 125I-labeled azido-Cys-AtPep1 photoaffinity analog specifically labeled a membrane-associated protein of ≈170 kDa. The labeled protein was purified to homogeneity, and its tryptic peptides were identified as gene At1g73080, which encodes a leucine-rich repeat receptor kinase, here called PEPR1. Verification of the binding protein as the receptor for AtPep1 was established by demonstrating the loss of function of microsomal membranes of two SALK insertional mutants and by a gain in function of the alkalinization response to AtPep1 by tobacco suspension-cultured cells expressing the At1g73080 transgene. Synthetic homologs of AtPep1, deduced from the C termini of six known paralogs of PROPEP1, were biologically active and were competitors of the interaction of an AtPep1 radiolabeled analog with the receptor. The data are consistent with a role for PEPR1 as the receptor for AtPep1 to amplify innate immunity in response to pathogen attacks.
Keywords: leucine, rich repeat receptor kinase, pathogen defense, photoaffinity labeling, microsomal membranes
A 23-aa peptide recently isolated from Arabidopsis called AtPep1, when supplied to Arabidopsis leaves at nanomolar concentrations, activates expression of the AtPep1 precursor gene PROPEP1 and defensin, PDF1.2, and induces the production of H2O2 (1). Arabidopsis plants transformed with the AtPep1 precursor gene PROPEP1 overexpress defense genes, producing a phenotype with enhanced resistance toward Pythium irregulare, a root pathogen (1).
The innate immune response in animals and plants is the first line of defense against microbial pathogens and is more ancient than adaptive immunity (2–6). Innate immunity in plants exhibits similarities with those of Drosophila and higher animals, and it appears that these responses have ancient evolutionary relationships (2). The immune responses are initiated by the interaction of conserved molecules derived from the pathogens. In animals, these molecules are called pathogen-associated molecular patterns, and in plants, they are called elicitors. Pathogen-associated molecular patterns and elicitors are recognized at the cell surface by receptors that are typically composed of an extracellular leucine-rich repeat (LRR) domain flanked by disulfide pairs, a transmembrane domain, and an intracellular signaling domain. Although endogenous peptides from plants have been shown to activate defenses against herbivores (7–9) and pathogen-derived peptides have been shown to activate defense responses against pathogens (10–14), AtPep1 is a unique example of an endogenous peptide signal in plants that activates defense genes targeted specifically against pathogens.
Here, we report the isolation of an AtPep1-binding protein from Arabidopsis suspension-cultured cells. Characterization of tryptic digests of the purified receptor protein has led to the identification of the gene in the Arabidopsis Information Resource as At1g73080, a LRR receptor kinase. The identity of this gene as the AtPep1 receptor gene was confirmed by using two SALK insertional mutants of At1g73080 that lost their ability to be photoaffinity-labeled and by a gain of AtPep1 binding to the cell surface of tobacco cells transformed with At1g73080. The receptor appears to be a key component of AtPep1 signaling for the amplification of the innate immune response in Arabidopsis in response to pathogen attacks.
Results and Discussion
Biological Activity of Tyr-AtPep1 Analogs.
To radiolabel AtPep1 to quantify binding to Arabidopsis suspension-cultured cells, a tyrosine residue was attached to the N terminus of synthetic AtPep1 for modification with125I (Fig. 1A). This addition of a single tyrosine to the N terminus of the peptide did not affect its medium alkalinization activity (Fig. 1B). Tyr-AtPep1 was iodinated, and the separated monoiodinated and diiodinated products, I1-Tyr-AtPep1 and I2-Tyr-AtPep1, were found to be as fully active as AtPep1 in causing the alkalinization of Arabidopsis suspension-cultured cells (1) (Fig. 1B). A half-maximal alkalinization response occurred at <2.5 nM, which is within the concentration range for half-maximal activity of AtPep1 and Tyr-AtPep1. Tyr-AtPep1 was iodinated with Na125I as above, and monoiodinated 125I1-Tyr-AtPep1 was recovered as 95% of the total label with a specific activity of ≈2 mCi/nmol (1 Ci = 37 GBq). This species was used for the following binding studies.
Fig. 1.
Kinetic analysis of 125I1-Tyr-AtPep1 binding to Arabidopsis cells. (A) Tyr-AtPep1 was synthesized and iodinated to produce mono- and diiodinated TyrAtPep1 (I1-Tyr-AtPep1 and I2-Tyr-AtPep1, respectively). (B) Biological activity of AtPep1 and analogs from A were assayed in the medium alkalinization assay by using Arabidopsis suspension-cultured cells. (C) Time course of binding of 125I1-Tyr-AtPep1 to Arabidopsis cells. Specific binding was calculated by subtracting nonspecific binding (binding in the presence of 250-fold nonradioactive AtPep1) from total binding in the absence of AtPep1. Error bars indicate standard deviation. (D) Saturation analysis of 125I1-Tyr-AtPep1 binding to Arabidopsis cells. A representative plot from four repetitions is shown. (E) Scatchard analysis of the data from D.
Kinetic Analysis of 125I1-Tyr-AtPep1 Binding.
Arabidopsis suspension-cultured cells were incubated with 0.25 nM125I1-Tyr-AtPep1 and washed with cold Murashige and Skoog medium containing 3% sucrose, and the total bound radioactivity was quantified. To estimate the contribution of nonspecific binding, a 250-fold excess of unlabeled AtPep1 was added to the cells along with 125I1-Tyr-AtPep1, and the remaining bound radioactivity was determined. Specific binding was calculated by subtracting nonspecific binding from total binding. Binding of 125I1-Tyr-AtPep1 to the cells could be detected within 1 min, and the initial rapid binding was complete within ≈4 min (Fig. 1C). The specific binding of 125I1-Tyr-AtPep1 was saturated at a level of ≈0.6 nM (Fig. 1D). From the data, a Kd of 0.25 nM for the binding of 125I1-Tyr-AtPep1 was estimated (Fig. 1E), which is in agreement with the concentration of AtPep1 required for half-maximal medium alkalinization (Fig. 1B).
Photoaffinity Labeling of an AtPep1-Binding Protein.
An azido-Cys-AtPep1 analog was constructed (see Materials and Methods) by using the photoaffinity cross-linker N-(4-[p-azidosalicylamido]butyl)-3′-(2′-pyridyldithio)propionamide, which was coupled to the peptide through a cysteine residue that had been attached to the N terminus of the synthetic AtPep1 peptide (Fig. 2A). This analog was as active as native AtPep1 in alkalinating the medium of Arabidopsis suspension-cultured cells when incubated with the cells under red light to prevent cross-linking (Fig. 2B). Azido-Cys-AtPep1 was iodinated with Na125I, and the purified product, 125I-azido-Cys-AtPep1, was incubated with Arabidopsis cells for 5 min under red light and then irradiated with UV-B for 10 min to activate the azido group to effect cross-linking. Separation of the labeled proteins of the cells by SDS/PAGE revealed the radiolabeling of a 170-kDa protein (Fig. 2C), and the labeling of this protein could be fully competed with a 10-fold excess of unlabeled AtPep1 but not by unlabeled tomato systemin (Fig. 2C), which does not cause alkalinization of the medium of Arabidopsis cells. Photoaffinity labeling of the 170-kDa protein was inhibited by 0.1 mM suramin (Fig. 2D), a heterocyclic, polysulfonated inhibitor of ligand–receptor interactions in both animals and plants (see references in ref. 15). These results further strengthened the proposed role of the 170-kDa AtPep1-binding protein as the AtPep1 receptor. Treatment of the 170-kDa protein with peptide-N-glycosidase F decreased its mass by ≈20 kDa, indicating that the protein was glycosylated (Fig. 3E).
Fig. 2.
Photoaffinity labeling of an AtPep1-binding protein on the surface of Arabidopsis suspension-cultured cells. (A) Construction of the photoaffinity analog azido-Cys-AtPep1. AtPep1 was lengthened at its N terminus with a Cys residue, and the analog was azido-labeled under red light by using the photoaffinity cross-linker N-(4-[p-azidosalicylamido]butyl)-3′-(2′-pyridyldithio)propionamide through the formation of a disulfide bond. (B) Concentration dependence of the alkalinization activity of azido-Cys-AtPep1. (C) Specific binding of 0.25 nM 125I-azido-Cys-AtPep1 to Arabidopsis cells under red light in the presence of increasing concentrations of unlabeled AtPep1 (Left) or tomato systemin (Right). Cells were incubated with 125I-azido-Cys-AtPep1 for 10 min, washed with water, and irradiated with UV-B for 10 min, and the proteins were separated by SDS/PAGE. The labeled proteins in the gels were identified by exposure to x-ray film. (D) Suramin inhibits 125I-azido-Cys-AtPep1-labeling of the 170-kDa binding protein of Arabidopsis suspension-cultured cells. (E) Removal of N-linked glycans with the endoglycosidase peptide-N-glycosidase F (PNGaseF) caused a shift in mobility on SDS/PAGE of ≈12% of the mass of the 170-kDa labeled protein.
Fig. 3.
Purification and identification of an AtPep1-binding protein. (A) (Left) SDS/PAGE analysis of photoaffinity-labeled proteins during purification (described in Materials and Methods). Protein staining is as follows: lane 1, crude extract; lane 2, microsomal fraction; lane 3, the eluate from preparative SDS/PAGE; lane 4, eluate from Con A Sepharose. (Right) Radioautography of gels from Left. (B) Mass spectral analysis of peptides resulting from tryptic digest of the purified, radiolabeled AtPep1-binding protein from lane 4 in A (probability-based molecular weight search score = 609; P = <0.05). Peptide fragments marked with an asterisk matched predicted amino acid sequences of Arabidopsis gene At1g73080. (C) The phylogenetic relationships are based on comparison of the amino acid sequence of PEPR1 with the gene products of other members of the LRR XI subfamily of Arabidopsis LRR receptor-like protein kinases.
Purification of AtPep1-Binding Protein.
The photoaffinity labeling protocol was scaled up to a volume of 3 liters of Arabidopsis suspension-cultured cells, as described in ref. 16 for the isolation of the systemin receptor. One liter of cells was photoaffinity-labeled with125I-azido-Cys-AtPep1, and the remaining 2 liters of cells was treated with nonradioactive I-azido-Cys-AtPep1. Microsomal membrane fractions prepared from both labeled and unlabeled cells were combined to provide enough labeled protein for isolation. The protein was purified by using preparative SDS/PAGE and Con A affinity Sepharose (Fig. 3A). The photoaffinity-labeled protein bound so tightly to the Con A Sepharose that it could not be eluted by 0.5 M α-methyl-d-glucoside. However, the proteins were solubilized from the Sepharose by boiling in 5% SDS and separated by SDS/PAGE. After electrophoresis, the section of the gel containing the radiolabeled protein was treated with trypsin, and the peptide fragments were identified by mass spectroscopy (Fig. 3B) as being derived from the gene At1g73080. Tryptic digests of the deglycosylated protein confirmed the identity of the protein.
At1g73080, which we call PEPR1, encodes a protein of 1,124 aa that includes a signal peptide, 26 LRR motifs with flanking cysteine pairs, a transmembrane region, and a kinase domain. Based on phylogenetic analysis of all Arabidopsis receptor-like kinases (17), At1g73080 belongs to a 28-member subfamily of LRR XI genes that includes HAESA (18), CLV1 (19), and three CLV1-related LRR receptor-like kinase genes, BAM1, BAM2, and BAM3 (20) (Fig. 3C). These putative receptors are the only members of this 28-member subfamily that have been identified as having signaling roles in Arabidopsis, and only CLV1 and PEPR1 have known ligands, both being peptides.
Functionality of PEPRI as the AtPep1 Receptor Gene.
Total RNA was isolated from leaves of each of two SALK insertional mutant lines of PEPR1 (At1g73080), SALK_059281 and SALK_014538 (Fig. 4A Upper), and assayed by RT-PCR (Fig. 4A Lower), establishing that the gene was not expressed in either line. Microsomal membranes prepared from the two SALK mutant lines were subjected to photoaffinity labeling with125I-azido-Cys-AtPep1 and compared with labeling of membranes from wild-type plants and from the SALK insertional line SALK_064539, which encodes a phosphodiesterase gene that is unrelated to PEPRI. A protein of 170 kDa and a lower molecular mass protein were labeled in membranes from the wild type and SALK_064539 but not in membranes of the two mutant lines (Fig. 4B). This labeling suggested that the 170-kDa protein was the same protein that was identified and isolated from Arabidopsis cells, with the lower molecular mass species likely being a degradation product of the 170-kDa species because no labeling of this protein was found in either of the two insertional mutants. Possible loss of resistance against pathogens and changes in the ability to activate defensin and other defense genes in response to wounding and elicitors in the two insertional mutants have yet to be studied.
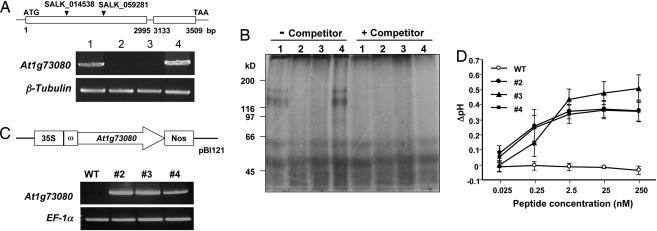
Fig. 4.
Confirmation of the functionality of At1g73080 as coding for the AtPep1 receptor PEPR1. (A) (Upper) The location of the T-DNA in each insertional line, SALK_059281 and SALK_014538. (Lower) RT-PCR expression analyses of At1g73080 mRNA in leaves of the following: lane 1, wild-type plants; lane 2, SALK_014538 line; lane 3, SALK_059281 line; lane 4, SALK_064539 line, which is a control line unrelated to At1g73080 in which the glycerophosphoryl diester phosphodiesterase (At5g55480) gene is disrupted by T-DNA. Expression of the β-tubulin gene was used as a control. (B) Photoaffinity labeling under red light of microsomal fractions prepared from 5-week-old plants. Microsomal proteins were photoaffinity-labeled in the presence or absence of 250-fold unlabeled AtPep1 as a competitor. Lane 1, wild-type plants; lane 2, SALK_014538 line; lane 3, SALK_059281 line; lane 4, SALK_064539 line. (C) (Upper) Composition of the vector construct that was used to transform tobacco suspension-cultured cells to express At1g73080. (Lower) RT-PCR analyses of At1g73080 mRNA in wild-type cells and in three transgenic cell lines (#2, #3, and #4). Expression of the elongation factor 1α (EF-1α) gene was used as a control. (D) Alkalinization responses of wild-type and transgenic tobacco suspension-cultured cells to increasing concentrations of AtPep1. Only the transgenic cells responded to AtPep1 by alkalinating the culture medium.
To further investigate the role of PEPR1 as the AtPep1 receptor, gain-of-function experiments were carried out with tobacco suspension-cultured cells that do not respond to AtPep1 by alkalinating the cell medium. Tobacco cells were transformed with a CaMV-35S-PEPR1 gene (Fig. 4C Upper) to determine whether the foreign receptor protein was targeted to the cell surface, where, in response to AtPep1, it would activate intracellular signaling to effect the alkalinization of the medium of the transgenic tobacco cells. Expression of the PEPR1 gene was confirmed by RT-PCR analysis of RNA from three independently transformed tobacco cell lines (Fig. 4C Lower). All three of the tobacco lines responded to the addition of nanomolar concentrations of AtPep1 by producing a strong, reproducible alkalinization of the cell culture medium, whereas no alkalinization was detected in the culture medium of wild-type cells challenged with AtPep1 (Fig. 4D). These results not only confirm the identity of PEPR1 as the AtPep1 receptor, but they indicate that the receptor-mediated intracellular signaling pathway in tobacco cells that regulates alkalinization in response to peptide ligands can accommodate signaling by PEPR1 and its peptide ligand. This type of accommodation by suspension cultured cells to a foreign receptor and its peptide ligand has been reported for the tomato systemin receptor in tobacco cells (21), the tomato xylanase receptor in tobacco cells (22), the Arabidopsis flagellin receptor in tomato cells (23), and the receptor EFR in Nicotiana benthmiana cells (24). Kinase domains of receptors are known to possess highly conserved regions, and it may be that these regions in foreign receptors are recognized by interacting signaling components, perhaps through phosphorylation events. It will be of interest to determine the phylogenetic distance from Arabidopsis where the PEPR1 gene can still initiate intracellular signaling in response to AtPep1 when transformed into foreign plant species.
PEPR1 may also be the receptor for peptides related to AtPep1 generated from PROPEP1 paralogs (1). The putative peptides encoded by all seven known PROPEP paralogs were synthesized based on their homology to AtPep1 (see Fig. 5 and ref. 1). All of the synthetic peptides were assayed for their alkalinization activities with Arabidopsis suspension-cultured cells. Of the seven paralogs, four were as fully active as AtPep1 (Fig. 5A). The seven peptides were then assayed for their ability to compete with 125I-Tyr-AtPep1 for labeling of Arabidopsis suspension-cultured cells. The five peptides that were most active in the alkalinization assay were potent competitors, whereas the two peptides with weak alkalinization activities were weak competitors of photoaffinity labeling (Fig. 5B). These data support the possibility that PEPR1 may be the receptor for at least five of the peptides that might be generated during pathogen attacks.
Fig. 5.
Comparisons of the alkalinization activities of 23-aa peptides synthesized from the C-terminal sequences of the proteins encoded by the seven PROPEP paralogs with their abilities to compete with 125I1-Tyr-AtPep1 for binding on the cell surface of suspension-cultured Arabidopsis cells. (A) Medium alkalinization of suspension-cultured Arabidopsis cells by the synthetic AtPep peptides deduced from each of the seven paralogs, assayed at the four concentrations shown. (B) Assays using each synthetic AtPep analog in competition with 125I1-Tyr-AtPep1 for binding to Arabidopsis suspension-cultured cells (for details, see Materials and Methods). Each peptide was incubated with cells at the concentrations indicated just before adding 0.25 nM 125I1-Tyr-AtPep1. After 2 min, the cells were recovered and washed, and the bound radioactivity was quantified.
In summary, we report the isolation of the AtPep1-binding protein PEPR1 and present evidence to support its identity as a functional LRR receptor for AtPep1, including kinetic analyses, photoaffinity labeling, mutant analyses, and gain-of-function experiments. The role of the receptor AtPepR1 to amplify defense gene signaling for the innate immune response provides a useful ligand–receptor pair to further investigate defense peptide signaling, both in Arabidopsis and in species throughout the plant kingdom.
Materials and Methods
Alkalinization Assay.
Medium alkalinization assays were carried out by using 1 ml of Arabidopsis suspension-cultured cells as described in ref. 1. Azido-Cys-AtPep1 was mixed with cells in darkness for increasing times before the pH of the culture medium was measured.
Binding Assays.
Radioligand binding assays were performed as described in ref. 22 with minor modifications. Suspension-cultured Arabidopsis cells were used for assays 4–5 days after subculturing. The cells were separated from the medium by using miracloth, washed once with 40 ml of culture medium, and adjusted to a fresh weight of 0.2 mg/ml with fresh medium. A 2-ml aliquot of cells was pipeted into a well of a 12-well culture plate and allowed to equilibrate for 1 h at room temperature while agitated on an orbital shaker (160 rpm). 125I1-Tyr-AtPep1 was added to the medium to give a final concentration of 0.02–1.0 nM. To assay binding, 500 μl of cells was removed and filtered through a 2.5-cm Type A/E Glass Fiber Filter (Pall) by using a 12-well vacuum filtration manifold (Millipore). The cells were washed three times with 5 ml of cold Murashige and Skoog (MS) medium containing 3% sucrose, suspended in 1 ml of MS medium containing 3% sucrose, transferred to a test tube, and then analyzed for total radioactivity in a gamma-ray counter (Isodata 2020; Isodata, Palatine, IL). Specific binding was calculated by subtracting nonspecific binding (binding in the presence of 250-fold native AtPep1) from total binding.
AtPep1 Analogs.
AtPep1, Cys-AtPep1, Tyr-AtPep1, and AtPep1 homologs were synthesized by using solid-phase chemistry with an Applied Biosystems model 431 synthesizer. Cys-AtPep1 was coupled in darkness to the azido-photoaffinity cross-linker, N-(4-[p-azidosalicylamido]butyl)-3′-(2′-pyridyldithio)propionamide (Pierce), and azido-Cys-AtPep1 was purified by HPLC as described in ref. 25. Iodination of Tyr-AtPep1 (50 nmol) was achieved by using IODO-GEN Pre-Coated Iodination Tubes (Pierce) by the indirect method as described by the manufacturer. Iodinated forms of Tyr-AtPep1 were purified by HPLC as described in ref. 25 and quantified by using the bicinchoninic acid method (Pierce) with AtPep1 as a standard. All analogs were analyzed with an LCQ ion trap mass spectrometer (Finnigan, San Jose, CA).125I iodination was performed by using 2 mCi of Na125I with 12.5 nmol of azido-Cys-AtPep1 and Tyr-AtPep1, followed by purification by HPLC as described in ref. 25. The specific radiolabel of each analog was calculated to be ≈2 mCi/nmol.
Photoaffinity Labeling and Purification of AtPep1-Binding Protein.
The procedures for photoaffinity labeling were identical to those described in ref. 25 except that125I-azido-Cys-AtPep1 was added under red light to a final concentration of 0.25 nM (1.4 × 105 cpm). The plate containing the cells was maintained on ice while it was irradiated with a UV-B lamp (F15T8.UV-B, 15 W) for 10 min. The cells were recovered from the well, washed at room temperature with 1 ml of Murashige and Skoog medium, and sedimented by centrifugation at 12,000 × g, and the pellet was mixed with 400 μl of 5% SDS while in the microfuge tube. The cells were disrupted by placing the tube in a boiling water bath for 30 min, and the cell debris was removed by centrifugation in a microfuge at 12,000 × g. Proteins in the supernatant were precipitated by addition of 400 μl of methanol and 100 μl of chloroform and pelleted in a microfuge. The pellet was dissolved in 100 μl of Laemmli sample buffer containing 5% SDS at 65°C, and 10 μl was separated by 8% SDS/PAGE. The gels were dried and exposed to x-ray film for 50 h.
Large-scale photoaffinity labeling of cell surface AtPep1-binding proteins was performed according to the method of Scheer and Ryan (16). Briefly, from 3 liters of Arabidopsis cells, 6 days after subculturing, the 1-liter and 2-liter sets of cells were placed in separate large, shallow plastic dishes (33 cm in diameter). Both sets of cells were incubated under red light while shaking on an orbital shaker (90 rpm) for 10 min. To the 1-liter set of cells was added 125I-azido-Cys-AtPep1 (to a final concentration of 0.25 nM), and to the 2-liter set of cells was added nonradiolabeled I-azido-Cys-AtPep1 (also to a final concentration of 0.25 nM). Both sets of cells were exposed to UV irradiation for 10 min while shaking on an orbital shaker, and each set of cells was collected by filtration as described in ref. 16. Each set of cells was dispersed in 300 ml of extraction buffer, the cells were disrupted in a Parr Cell Disruption Bomb (Parr Instruments, Moline, IL), and microsomal fractions were prepared by differential centrifugation (16). Radiolabeled and unlabeled microsomal proteins (25 and 55 mg, respectively), were mixed together, and the proteins were introduced into single horizontal wells of two 7.5% SDS/PAGE gels (0.6 × 14 × 9 cm) and separated during 16 h at a constant 37 V at room temperature. The gel was sliced horizontally into 5-mm widths, and the radioactivity was monitored with a gamma counter. The single gel piece that contained the most radioactivity from each gel was macerated to allow for protein extraction by using a 20-ml syringe (26). The protein was extracted with 25 ml of Con A binding buffer (20 mM Tris·HCl, pH 7.5/0.5 M NaCl) by gently shaking at room temperature for 30 min. The gel material was centrifuged at 2,000 × g for 10 min, and the supernatant was recovered. The extraction was repeated twice, and the supernatants were pooled (50 ml total). A 75-μl aliquot of Con A Sepharose 4B (Amersham Pharmacia Bioscience), previously equilibrated in the Con A binding buffer, was added to the extracted proteins and incubated at room temperature with constant mixing for 6 h on an orbital shaker (100 rpm). Con A Sepharose was recovered by centrifugation at 500 × g for 10 min at 4°C and washed three times with 50 ml of 0.5 M α-methyl-d-glucoside, followed by three washes with distilled water. The proteins were released from the Con A Sepharose by boiling in 500 μl of 5% SDS, centrifuging as above, and repeating the boiling procedure. The proteins in the supernatants were pooled and separated into 500-μl samples, and each was mixed with 500 μl of methanol and 100 μl of chloroform to precipitate proteins. One sample was subjected to 7.5% SDS/PAGE (0.15 × 14 × 8.5 cm) at a constant 100 V for 8 h at room temperature. The gel was sliced horizontally into 1-mm widths, and the radioactivity was monitored with a gamma counter. One gel piece containing 59% of the radioactivity added to the gel was treated with modified trypsin (Promega) (27). The second sample was treated with peptide-N-glycosidase F (Prozyme, San Leandro, CA), separated by 7.5% SDS/PAGE, and digested with trypsin as above. The peptides from both trypsin-digested proteins were analyzed by MALDI-TOF mass spectrometry (Voyager-DE RP Biospectrometry Workstation; Applied Biosystems). The gene encoding the AtPep1-binding protein was identified by searching in the National Center for Biotechnology Information database with mascot.
Analysis of Transfer DNA (T-DNA) Insertional Lines.
Arabidopsis thaliana T-DNA insertional lines, SALK_014538, SALK_059281, and SALK_064539, were obtained from the Arabidopsis Biological Resource Center (Columbus, OH) (28). Homogeneous lines were screened by two sets of PCR analyses, one using the gene-specific primer pair and the other using a gene-specific primer and the T-DNA left border of the vector as a primer. Microsomal proteins were prepared from 1-month-old wild-type and SALK mutant plants by differential centrifugation as described in ref. 16. The microsomal proteins were adjusted to 1 μg/μl in binding buffer (10 mM Hepes-KOH, pH 7.4/100 mM NaCl/100 mM sucrose/5 mM MgCl2), and 250 μl was added under red light to ice-cold 0.25 nM 125I-azido-Cys-AtPep1 with or without 250-fold unlabeled AtPep1 for 60 min on ice. After incubation, the reaction mixture was irradiated with UV-B (lamp F15T8.UV-B, 15 W) for 15 min. The microsomal proteins were precipitated by centrifugation at 12,000 × g for 5 min after mixing with 250 μl of methanol and 50 μl of chloroform, dissolved in 20 μl of Laemmli sample buffer containing 5% SDS at 65°C, and then separated by SDS/PAGE.
Transgenic Tobacco Cells.
The β-glucuronidase gene (excised between its XbaI site and its SacI site) in a pBI121 vector was replaced with a PCR fragment consisting of an XbaI site, a ω-prime leader sequence (29), and KpnI, SmaI, BamHI, and SacI sites to enhance the translation and further construction. PCR primers for this construction were as follows: ω-XbaI, 5′-CCG TCT AGA TTT TTA CAA CAA TTA CCA ACA ACA ACA AAC AAC AAA CAA CAT TAC AAT TA-3′; ω-BamHI, 5′-CCG AGC TCG GAT CCC GGG GTA CCA TGG TAA TTG TAA ATA GTA ATT GTA ATG TTG TTT-3′. A 3,604-bp fragment consisting of a KpnI site, the coding region of At1g73080 including an intron, and a BamHI site was amplified by PCR using 73080-KpnI (5′-CGG GGT ACC TCG ATC TTT AAA CTC AGA TGA AGA A-3′) and 73080-BamHI (5′-CGC GGA TCC TAC CGG TAA GCC CAG TTA CG-3′) primers. This PCR fragment was ligated into KpnI and BamHI sites of pBI121 vector containing ω-prime leader sequence and then introduced into tobacco suspension-cultured cells by using Agrobacterium tumefaciens strain GV3850 by the method of Nakayama et al. (30). Transgenic cells were assayed in the alkalinization assay as described above.
RT-PCR Analyses.
Total RNA from leaves of T-DNA insertion lines and transgenic tobacco cells were extracted by using TRIzol total RNA isolation reagent (Invitrogen) according to the manufacturer’s instructions. Reverse transcription was performed by using Superscript III Reverse Transcriptase (Invitrogen) and oligo(dT) primer according to the manufacturer’s instructions. To amplify At1g73080, 73080-KpnI and 73080-BamHI primers were used. Arabidopsis β-tubulin gene and the tobacco elongation factor 1α (EF-1α) gene were amplified as internal controls by using β-Tub1 (5′-CAA CGC TAC TCT GTC TGT CC-3′), β-Tub2 (5′-TCT GTG AAT TCC ATC TCG TC-3′), EF-1α-1 (5′-TTT GGC CCT ACT GGT TTG AC-3′), and EF-1α-2 (5′-CTC ATG TCC CTC ACA GCA AA-3′).
Phylogenic Analysis.
Full-length amino acid sequences of all members in subfamily LRR XI (18) were analyzed by using clustal w (31), and a phylogenic tree was produced by using treeview (32).
Acknowledgments
We thank Julia Gothard and Sue Vogtmann for growing our plants, Guido Barona for maintaining the cell cultures, Dr. Jerry Reeves and Dr. David Deavila (Washington State University) for help and advice in radiolabeling, Dr. William Siems for advice and assistance in obtaining mass spectrometric analyses, and Dr. Gerhard Munske (Washington State University) for peptide synthesis. This work was supported by the Toyobo Biotechnology Research Foundation of Osaka, Japan (Y.Y.), National Science Foundation Grant IBN 0090766, the Charlotte Y. Martin Foundation, and the Washington State University College of Agriculture, Human, and Natural Resources Sciences.
Abbreviations
- LRR
leucine-rich repeat
- T-DNA
transfer DNA.
Footnotes
Conflict of interest statement: No conflicts declared.
References
- 1.Huffaker A., Pearce G., Ryan C. A. Proc. Natl. Acad. Sci. USA. 2006;103:10098–10103. doi: 10.1073/pnas.0603727103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Hoffman J. A., Kafatos F. C., Janeway C. A., Jr., Ezekowitz R. A. B. Science. 1999;284:1313–1318. doi: 10.1126/science.284.5418.1313. [DOI] [PubMed] [Google Scholar]
- 3.Menezes H., Jared C. Comp. Biochem. Physiol. C Toxicol. Pharmocol. 2002;132:1–7. doi: 10.1016/s1532-0456(02)00043-1. [DOI] [PubMed] [Google Scholar]
- 4.Nürnberger T., Brunner F., Kemmerling B., Piater L. Immunol. Rev. 2004;198:249–266. doi: 10.1111/j.0105-2896.2004.0119.x. [DOI] [PubMed] [Google Scholar]
- 5.Parker J. Trends Plant. Sci. 2003;8:245–247. doi: 10.1016/S1360-1385(03)00105-5. [DOI] [PubMed] [Google Scholar]
- 6.Zipfel C., Robatzek S., Navarro L., Oakeley E. J., Jones J. D. G., Felix G., Boller T. Nature. 2004;428:764–767. doi: 10.1038/nature02485. [DOI] [PubMed] [Google Scholar]
- 7.Pearce G., Strydom D., Johnson S., Ryan C. A. Science. 1991;253:895–897. doi: 10.1126/science.253.5022.895. [DOI] [PubMed] [Google Scholar]
- 8.Pearce G., Moura D. S., Stratmann J., Ryan C. A. Nature. 2001;411:817–820. doi: 10.1038/35081107. [DOI] [PubMed] [Google Scholar]
- 9.Pearce G., Ryan C. A. J. Biol. Chem. 2003;278:30044–30050. doi: 10.1074/jbc.M304159200. [DOI] [PubMed] [Google Scholar]
- 10.Hahlbrock K., Scheel D., Logemann E., Nurnberger T., Parniske M., Reinnold S., Sacks W. R., Schmelzer E. Proc. Natl. Acad. Sci. USA. 1995;92:4150–4157. doi: 10.1073/pnas.92.10.4150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.van den Askerveken G. G. J. M., Vassen J. A. M. J., De Wit P. J. G. M. Plant Physiol. 1993;103:91–96. doi: 10.1104/pp.103.1.91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kammpren S. Curr. Opin. Plant Biol. 2001;4:295–300. doi: 10.1016/s1369-5266(00)00176-x. [DOI] [PubMed] [Google Scholar]
- 13.Kunze G., Zipfel C., Robatzek S., Niehaus K., Boller T., Felix G. Plant Cell. 2004;16:3496–3507. doi: 10.1105/tpc.104.026765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Navarro L., Zipfel C., Rowland R., Keller I., Robatzek S., Boller T., Jones J. D. G. Plant Physiol. 2004;135:1113–1128. doi: 10.1104/pp.103.036749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Stratmann J., Scheer J., Ryan C. A. Proc. Natl. Acad. Sci. USA. 2000;97:8862–8867. doi: 10.1073/pnas.97.16.8862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Scheer J. M., Ryan C. A. Proc. Natl. Acad. Sci. USA. 2002;99:9585–9590. doi: 10.1073/pnas.132266499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Shiu S. H., Bleecker A. B. Proc. Natl. Acad. Sci. USA. 2001;98:10763–10768. doi: 10.1073/pnas.181141598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jinn T.-L., Stone J. M., Walker J. C. Genes Dev. 2000;14:108–117. [PMC free article] [PubMed] [Google Scholar]
- 19.Clark S. E., Williams R. W., Meyerowitz E. M. Cell. 1997;89:575–585. doi: 10.1016/s0092-8674(00)80239-1. [DOI] [PubMed] [Google Scholar]
- 20.DeYoung B. J., Bickle K. L., Schrage K. J., Muskett P., Patel K., Clark S. E. Plant J. 2006;45:1–6. doi: 10.1111/j.1365-313X.2005.02592.x. [DOI] [PubMed] [Google Scholar]
- 21.Scheer J. M., Pearce G., Ryan C. A. Proc. Natl. Acad. Sci. USA. 2003;100:10114–10117. doi: 10.1073/pnas.1432910100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ron M., Avini A. Plant Cell. 2004;16:1604–1615. doi: 10.1105/tpc.022475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chinchilla D., Bauer Z., Ragenass M., Boller T., Felix G. Plant Cell. 2006;18:465–476. doi: 10.1105/tpc.105.036574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Zipfel C., Kunze G., Chinchilla C., Caniard A., Boller T., Felix G. Cell. 2006;125:749–760. doi: 10.1016/j.cell.2006.03.037. [DOI] [PubMed] [Google Scholar]
- 25.Scheer J. M., Ryan C. A. Plant Cell. 1999;11:1525–1535. doi: 10.1105/tpc.11.8.1525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Scheer J. M., Ryan C. A. Anal. Biochem. 2001;298:130–132. doi: 10.1006/abio.2001.5384. [DOI] [PubMed] [Google Scholar]
- 27.Shevchenko A., Wilm M., Vorn O., Mann M. Anal. Chem. 1996;68:850–858. doi: 10.1021/ac950914h. [DOI] [PubMed] [Google Scholar]
- 28.Alonso J. M., Stepanova A. N., Leisse T. J., Kim C. J., Chen H., Shinn P., Stevenson D. R., Zimmerman K., Barajas P., Cheuk R., et al. Science. 2003;301:653–657. doi: 10.1126/science.1086391. [DOI] [PubMed] [Google Scholar]
- 29.Gallie D. R., Sleat D. E., Watts J. W., Turner P. C., Wilson T. M. Nucleic Acids Res. 1987;15:3257–3273. doi: 10.1093/nar/15.8.3257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Nakayama H., Yoshida K., Ono H., Murooka Y., Shinmyo A. Plant Physiol. 2000;122:1239–1248. doi: 10.1104/pp.122.4.1239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Thompson J. D., Higgins D. G., Gibson T. J. Nucleic Acids Res. 1994;22:4673–4680. doi: 10.1093/nar/22.22.4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Page R. D. Comput. Appl. Biosci. 1996;12:357–358. doi: 10.1093/bioinformatics/12.4.357. [DOI] [PubMed] [Google Scholar]