Abstract
Analysis of the human Rab6A gene structure reveals the presence of a duplicated exon, and incorporation of either of the two exons by alternative splicing is shown to generate two Rab6 isoforms named Rab6A and Rab6A′, which differ in only three amino acid residues located in regions flanking the PM3 GTP-binding domain of the proteins. These isoforms are ubiquitously expressed at similar levels, exhibit the same GTP-binding properties, and are localized to the Golgi apparatus. Overexpression of the GTP-bound mutants of Rab6A (Rab6A Q72L) or Rab6A′ (Rab6A′ Q72L) inhibits secretion in HeLa cells, but overexpression of Rab6A′ Q72L does not induce the redistribution of Golgi proteins into the endoplasmic reticulum. This suggests that Rab6A′ is not able to stimulate Golgi-to-endoplasmic reticulum retrograde transport, as described previously for Rab6A. In addition, Rab6A′ interacts with two Rab6A partners, GAPCenA and “clone 1,” but not with the kinesin-like protein Rabkinesin-6, a Golgi-associated Rab6A effector. Interestingly, we found that the functional differences between Rab6A and Rab6A′ are contingent on one amino acid (T or A at position 87). Therefore, limited amino acid substitutions within a Rab protein introduced by alternative splicing could represent a mechanism to generate functionally different isoforms that interact with distinct sets of effectors.
INTRODUCTION
In eukaryotic cells, membrane traffic along biosynthetic and endocytic pathways requires the coordinated activity of a large number of protein families. One of these families, the Rab subfamily of small GTPases, plays an important role in targeting and docking of transport vesicles or transport intermediates to their correct destinations (Novick and Zerial, 1997; for reviews, see Chavrier and Goud, 1999; Gonzalez and Scheller, 1999). It is now established that Rab proteins perform this task in at least three ways, i.e., by 1) linkage of transport vesicles to the actin or microtubule cytoskeleton via an interaction with molecular motors; 2) recruitment of docking complexes that anchor transport vesicles to their target membranes; and 3) regulation of the assembly of SNARE complexes.
In mammalian cells, more than 30 Rab family members have been identified, and probably almost 50 members exist (M. Seabra, personal communication). Most of the Rabs are ubiquitous, and their abundance likely reflects the multiplicity of transport steps that need to be regulated within a single cell. In addition, some Rabs are involved in specialized cell functions, such as regulated secretion (Lledo et al., 1994 [Rab3A]; Geppert and Sudhof, 1998) in neuronal and neuroendocrine cells and transcytosis (Rab17 [Hunziker and Peters, 1998; Zacchi et al., 1998]) in epithelial cells. The estimated number of 30–50 Rab GTPases includes proteins that are closely related to each other and are called Rab isoforms (e.g., Rab1A and -B and Rab3A, -B, -C, and -D). The amino acid identity between each isoform is in the range of 80–95%, the differences being mainly in domains located at the N or C terminus of the proteins. The precise functions of the different isoforms remain largely to be established. Some Rab isoforms have distinct subcellular localization or are not expressed at the same levels within a given cell type. For instance, Rab3A and -C are primarily expressed in neuronal and neuroendocrine cells (Darchen et al., 1990; Fischer von Mollard et al., 1990), whereas Rab3D appears to be the most abundant isoform in adipocytes (Baldini et al., 1995). Distinct functional properties of Rab3A and -B have been reported in PC12 cells and in insulin-secreting cells (Weber et al., 1994; Iezzi et al., 1999). Alternatively, Rab isoforms may be functionally redundant. It is documented, for instance, that Rab1A and -B (92% identity) fulfill the same function in the regulation of endoplasmic reticulum (ER)-to-Golgi transport (Nuoffer et al., 1994). Yeast YPT31 and YPT32 are phenotypically redundant (Benli et al., 1996), and the gene products of YPT51/YPT52/YPT53 have at least overlapping functions (Singer Kruger et al., 1994).
Only limited information is available on the genomic organization of mammalian Rab genes (Wichmann et al., 1989; Baumert et al., 1993; Lai et al., 1994; Barbosa et al., 1995; Zheng et al., 1997). Based on their nucleotide sequences, however, it is likely that Rab isoforms are encoded by distinct genes. This holds true for yeast Ypt proteins, because YPT31 and YPT32, as well as YPT51, YPT52, and YPT53, have been mapped to different chromosomes (Lazar et al., 1997). Here, we show that Rab isoforms may also be generated by alternative splicing of two homologous exons within the same gene. Such a mechanism is documented for the Rab6A gene, which encodes a ubiquitous Rab that regulates transport at the level of the Golgi apparatus (Martinez et al., 1994, 1997; Girod et al., 1999; White et al., 1999). In contrast to other Rab isoforms described so far, the two Rab6 isoforms (termed Rab6A and Rab6A′) differ only in three amino acids flanking the conserved PM3 domain involved in GTP binding (Valencia et al., 1991). Both proteins appear to be localized to the Golgi complex and display similar biochemical activities. Interestingly, the overexpression of the GTP-bound mutant Rab6A′ Q72L does not redistribute Golgi membranes into the ER, as was shown for Rab6A Q72L (Martinez et al., 1997). In addition, Rab6A′ does not interact with Rabkinesin-6, a molecular motor previously shown to be an effector of Rab6A (Echard et al., 1998). We discuss the functional implications of the presence of two Rab6A isoforms in mammalian cells.
MATERIALS AND METHODS
Library Screening and DNA Sequencing
Partial sequences encoding Rab6A′ were obtained by a reverse transcription (RT)-PCR–based cloning approach with degenerate primers matching conserved domains PM3 and G2 of Rab proteins in human umbilical vein endothelial cells (HUVECs) (de Leeuw et al., 1998) or with degenerate primers [forward, 5′-GGCGGCGGCTC-GAGGGI(A0.2/G0.8)(G0.2/A0.8) II (A0.2/G0.2/C0.6)IIII(A0.2/T0.2/G0.6)(G0.2/C0.2/T0.6)(A0.2/T0.8)GGIAA(A0.5/G0.5)(A0.5/T0.5)C-3′; reverse, 5′-GGCG-GCGGATCCTTC(C0.5/ T0.5)TGICC(A0.5/ T0.5)GCIGT(A0.5/G0.5)TCCCA-3′] corresponding to the conserved GTP-binding regions PM1 and PM3 in Caco-2 cells. The partial Rab6A′ amplified product from HUVECs was radiolabeled by random oligonucleotide priming (Feinberg and Vogelstein, 1983) and used as a probe to screen a HUVEC cDNA library (de Leeuw et al., 1998) consisting of ∼8 × 104 independent clones. Sequences from positive clones were determined according to the Sanger method (T7 sequencing kit, Pharmacia, Piscataway, NJ). Human Rab6A′ sequence data have been submitted to the DDBJ/EMBL/GenBank database under accession number AF198616.
PCR-based Genomic Analysis
To analyze the genomic organization of the Rab6A gene, specific primers (Eurogentec, Oligold, Seraing, Belgium) were constructed for Rab6A′ sequence (5′→3′, forward, AATCAGGCTTCAGCTG; reverse, TCGTAAACTACTACAGCTG) and Rab6A sequence (5′→3′, forward, ACAGTACGATTGCAATTA; reverse, CCACAGTGGAGTCACGA) based on their cDNA sequence differences. Furthermore, two primers (5′→3′, forward, CAGGCAACAATTGGC; reverse, ATCCACTTTGTAGTTTGC) flanking the specific sequences of Rab6A and Rab6A′ were generated. Combinations of forward and reverse primers (400 pmol) were used to perform PCR on human genomic DNA (100 ng) in a 50-μl volume in the presence of PCR buffer (50 mM KCl, 10 mM Tris-HCl, pH 8.3), 1.5 mM MgCl2, 200 μM dNTPs, and 5 U of Taq polymerase. The reactions were overlaid with 20 μl of mineral oil, transferred to a thermal cycler, and incubated for 35 cycles of 94°C for 1 min, 50°C for 1 min, and 72°C for 6 min with a final extension at 72°C for 10 min. PCR samples (10 μl) were loaded onto a 2% agarose gel, and amplified products were subsequently extracted from the gel with a gel extraction kit (Qiagen, Chatsworth, CA), cloned in pGEM-T vector (Promega, Madison, WI), transformed to Escherichia coli Ag1 cells, isolated, and sequenced.
RT-PCR Analysis
Total RNAs from various frozen human tissues and cultured cell lines were prepared according to the guanidinium isothiocyanate-phenol-chloroform extraction method (Chirgwin et al., 1979). RNA (1.5 μg) was dissolved in 34 μl of distilled water, and random hexamers (2 μg; Pharmacia) were added. The reaction mixtures were subsequently incubated at 70°C for 5 min and on ice for 2 min, and 24 μl of RT mix (1 μl of RNAsin [10 U], 6 μl of 0.1 M DTT, 4 μl of 10 mM dNTP mix, 12 μl of 5× RT buffer) was added. Then the reaction mixtures were separated into two equal volumes and incubated for 1 h at 42°C with (RT+) or without (RT−) Superscript reverse transcriptase (100 U; Life Technologies/BRL, Grand Island, NY) to ensure cDNA-specific amplification products. For expression analysis, primers were constructed matching sequences in the 5′ untranslated region (UTR) (5′→3′, forward, ATGTCCACGGG-CGGA) and 3′ UTR (5′→3′, reverse, CTGAAGAAGGTTGAAGA-TG) of both Rab6A and Rab6A′ and used to perform PCR on one-sixth of the generated single-stranded DNA in the presence of PCR reagents for 35 cycles of 94°C for 1 min, 50°C for 30 s, and 72°C for 1 min. Undigested or PstI-digested PCR products were analyzed by electrophoresis on a 2% agarose gel.
Cells, Media, and Cell Culture
Caco-2 TC7 cells were cultured and harvested 1 d after growth (log phase), after 70% confluence (undifferentiated cells), or 5 d after confluence (differentiated cells) with DMEM (Life Technologies/BRL) supplemented with 20% FCS and 1% nonessential amino acids (Life Technologies/BRL). To allow the differentiation of HT-29 cells, cells were grown without d-glucose (Darmoul et al., 1992). HeLa cells were cultured as described previously (Martinez et al., 1994). All other cell lines used for expression analysis were grown in DMEM supplemented with 10% FCS. Fibroblasts were freshly isolated from human uterus.
GST Fusion Protein Expression and [α-32P]GTP Blot Overlay Assay
The coding regions of Rab6A and Rab6A′ were amplified with N-terminal and C-terminal encoding primers (forward, 5′-CGGGATCCATGTCCACGGGCGGAGA-3′; reverse, 5′-CCGCTCGAG-CGGTTAGCAGGA-3′, containing a BamHI and a XhoI restriction site, respectively) and subsequently cloned into the BamHI–XhoI sites of the vector pGEX (Pharmacia). Transformed E. coli DH5α cells were grown to an OD600 of 0.3–0.5 at 37°C and induced with 0.1 mM isopropyl-1-thio-β-d-galactopyranoside for 18 h at 30°C. Cells were isolated by centrifugation, frozen, thawed, and lysed by sonication for 15 s on ice. After treatment with 1% Triton X-100, lysates were spun at 10,000 rpm for 10 min, and supernatant was collected. GST fusion proteins were adsorbed on a glutathione–Sepharose column, washed, and eluted with 10 mM glutathione in 10 mM Tris-HCl (pH 7.4). Purified protein samples (1 μg/sample) were layered in duplicate experiments on 12.5% polyacrylamide gels containing SDS and subjected to electrophoresis. One gel was stained with Coomassie brilliant blue, whereas the second was electrophoretically transferred onto nitrocellulose membranes. The blot was incubated with 1 nM [α-32P]GTP (1 μCi of [α-32P]GTP per ml), as described previously (Celis, 1998).
Assay of GTPγS Binding
Purified GST fusion protein samples (500 ng/assay) were diluted to 30 μl with 20 mM Tris buffer (pH 8.0), 1 mM EDTA, 1 mM DTT, and 0.1% Triton X-100. To each sample, 30 μl of GTPγS-binding mix (20 mM Tris buffer, pH 8.0, 1 mM EDTA, 2 mM DTT, 2 μM GTPγS, and ∼1.5 × 107 cpm of [35S]GTPγS) was added. Nonspecific binding was assayed with samples containing 0.1 mM unlabeled GTPγS. The samples were incubated at 30°C for 0, 1, 5, 15, 30, 60, or 120 min and terminated by the addition of 2 ml of ice-cold washing buffer (20 mM Tris-HCl, pH 8.0, 25 mM MgCl2,100 mM NaCl). Samples were filtered through nitrocellulose membranes (NC45, Schleicher & Schuell, Keene, NH), subsequently washed four times in ice-cold washing buffer, air dried, and counted in a water-compatible scintillation mixture (Opti-Fluor, Packard, Meriden, CT). As a control for the calculation of the amount of GTPγS bound to the proteins, 15 μl of the GTPγS-binding mix was counted in duplicate. All samples were assayed in duplicate.
Transient Expression, Pulse-Chase Experiments, and Immunoblotting
Human Rab6A and Rab6A′ proteins were expressed from the eukaryotic expression vector pCDNA3 (Invitrogen, Carlsbad, CA), which was modified at the 5′ end of the multiple cloning site with a synthetic DNA fragment that entails an initiator AUG codon followed by an N-terminal cMyc epitope tag and an EcoRI site. Full-length Rab6A and Rab6A′ were subcloned in-frame as a EcoRI–XhoI restriction fragment downstream from the cMyc tag sequence. For colocalization studies, the Rab6A′ coding region was cloned in-frame and downstream of green fluorescent protein (GFP) encoding cDNA in the mammalian expression vector pEGFP-N1 (Clontech, Palo Alto, CA) according to the strategy described previously (White et al., 1999). A modified GFP cDNA was placed downstream of sequences encoding the cytoplasmic, transmembrane, and stalk regions of human N-acetylglucosaminyltransferase I in the mammalian expression vector pRcCMV (Shima et al., 1997).
HeLa cells were grown on glass coverslips or plated on a 10-cm dish and cultured to 70% confluence. Cells were transiently transfected with 0.5 μg of cMyc-tagged Rab6A or Rab6A′ containing plasmid DNA or cotransfected with 0.5 μg of N-acetylglucosaminyltransferase I with the use of Lipofectamine Plus reagent according to the manufacturer's instructions (Life Technologies, Rockville, MD).
For immunoblotting, cells from the 10-cm dish were harvested by scraping in ice-cold PBS and lysed directly in an equal volume of 2× sample buffer (100 mM Tris-HCl, pH 6.8, 200 mM DTT, 4% SDS, 0.2% bromphenol blue, 20% glycerol). Protein lysates were layered on a 12.5% SDS-polyacrylamide gel, subjected to electrophoresis, and transferred to nitrocellulose membranes by Western blotting. After blocking with 5% nonfat dry milk in 10 mM Tris-HCl (pH 8.0), 150 mM NaCl, and 0.05% Tween 20 (TBST), the blot was incubated with anti-cMyc mAb 9E10 (hybridoma culture supernatant; Kari et al., 1986) for 2 h at room temperature. Incubations with primary and secondary antibodies (0.06 μg/ml HRP-conjugated AffiniPure (Jackson Immunoresearch, Westgrove, PA) goat anti-mouse immunoglobulin G; 1:10,000 dilution) and subsequent washes were done in TBST. Labeled bands were visualized with the use of CPD Star chemiluminescence according to the manufacturer (Tropix, Bedford, MA).
For transient expression in HeLa cells with the vaccinia system and pulse-chase experiments, we exactly followed the conditions described previously (Martinez et al., 1994, 1997). pGEM plasmids encoding Rab6A Q72L, Rab6A′ Q72L, and Rab6A′ Q72L A87T proteins were constructed with the use of PCR and verified by sequencing. PCR products were cloned into the pGEM-1 vector (Promega) with identical restriction sites and 5′ (Kozak's sequence) and 3′ noncoding regions to allow comparable expression levels.
Immunofluorescence Assay
Transfected HeLa cells, cultured on 24-wells plates on glass coverslips, were washed with PBS, fixed in 1% paraformaldehyde for 1 h at room temperature, and washed twice in PBS, 0.05% Tween 20 (PBST). After 15 min of methanol permeabilization, cells were incubated with anti-cMyc mAb 9E10 (1:25 dilution) for 1 h at room temperature and washed four times for 5 min in PBST. Specific labeling was detected by subsequent incubation with Texas Red–conjugated goat anti-mouse immunoglobulin G (1:75 dilution, 10 μg/ml; Jackson ImmunoResearch Laboratories, West Grove, PA) in PBST for 1 h and washing with PBST. Cells were mounted in Mowiol (Sigma Chemical, St. Louis, MO) and observed with the use of a confocal laser scanning microscope (MRC 1000, Bio-Rad, Richmond, CA).
HeLa cells were transfected for 5 h with pGEM control, pGEM Rab6A Q72L, pGEM Rab6A′ Q72L, or Rab6A′ Q72L A87T and processed for immunofluorescence as described previously (Martinez et al., 1997).
Two-Hybrid Experiments
The yeast reporter strain L40, which contains the reporter genes HIS3 and LacZ, was cotransformed with pLexA-Rab6A Q72L, pLexA-Rab6A′ Q72L, or Rab6A′ Q72L A87T and pGADGH-GAPCenA/Rab6 interacting domain (Cuif et al., 1999), pGADGH-Rabkinesin-6 (Echard et al., 1998), or pGADGH-clone 1. After 3 d at 30°C on selective medium, cotransformants were patched on drop-out medium lacking tryptophan and leucine (DO W-L-) and replicated on drop-out medium lacking tryptophan and leucine (DO W-L-) and drop-out medium lacking tryptophan, leucine, and histidine (DO W-L-H-). Transformation, analysis, and media were as described by Janoueix-Lerosey et al. (1995).
Overlay Experiments
Equal amounts (0.2 nmol) of purified histidine-tagged 174 domain (amino acids 529–665 of Rabkinesin-6; Echard et al., 1998) were run separately on SDS-PAGE and transferred onto nitrocellulose membranes in a carbonate buffer (3 mM Na2CO3, 10 mM NaHCO3, pH 9.8). Proteins were then renatured for 1.5 h at room temperature in 50 mM NaOH-HEPES, pH 7.2, 5 mM Mg acetate, 100 mM K acetate, 3 mM DTT, 10 mg/ml BSA, 0.1% Triton X-100, and 0.3% Tween 20. Radiolabeled [α-32P]GTP (50 μCi) was exchanged for 1 h at 30°C on bacterially purified GST-tagged Rab6A or Rab6A′ (2 nmol) by 4 mM EDTA, and the reaction was stopped with 10 mM MgCl2. After the renaturation step, each membrane was incubated for 1 h at room temperature with either radiolabeled Rab6A or Rab6A′ in 10 ml of binding buffer (12.5 mM NaOH-HEPES, pH 7.2, 1.5 mM Mg acetate, 75 mM K acetate, 1 mM DTT, 2 mg/ml BSA, 0.05% Triton X-100, 0.05% 3-([3-chloramidopropyl]dimethylammonio)-2-hydroxy-1-propanesulfonate). Membranes were washed separately two times (10 min each) in 20 mM Tris-HCl, pH 7.4, 100 mM NaCl, 20 mM MgCl2, and 0.005% Triton X-100 and autoradiographed.
RESULTS
Identification and Cloning of Rab6A′
We previously used a RT-PCR–based approach to characterize Rab proteins expressed in human epithelial colon carcinoma (Caco-2) cells and in HUVECs. Primer sets corresponding to the conserved GTP-binding domains PM3 and G2 (according to the nomenclature of Valencia et al. [1991]) revealed the presence of 28 different partial cDNAs encoding small GTPases in HUVECs (de Leeuw et al., 1998). A variety of partial sequences encoding Rab proteins were also identified in Caco-2 cells with the use of primers corresponding to the PM1 and PM3 domains (Opdam et al., 2000). These two independent analyses led to the identification of a partial sequence (Figure 1A, product 1 from Caco-2 cells, product 2 from HUVECs) showing high identity with human Rab6 cDNA. However, mismatches in a region close to the PM3 domain were found between the two nonoverlapping sequences and the Rab6 sequence. By using product 2 as a probe, we then isolated a full-length cDNA from a human endothelial cell library. The cDNA sequence contained the partial sequences of the previously amplified products 1 and 2 and revealed a stretch of 96 base pairs (bp) that was not identical to the Rab6 sequence (referred as Rab6A′, Rab6 being Rab6A; Figure 1A). Nevertheless, the 5′ and 3′ UTRs and the coding region flanking the stretch of mismatches were identical to that of the Rab6A cDNA.
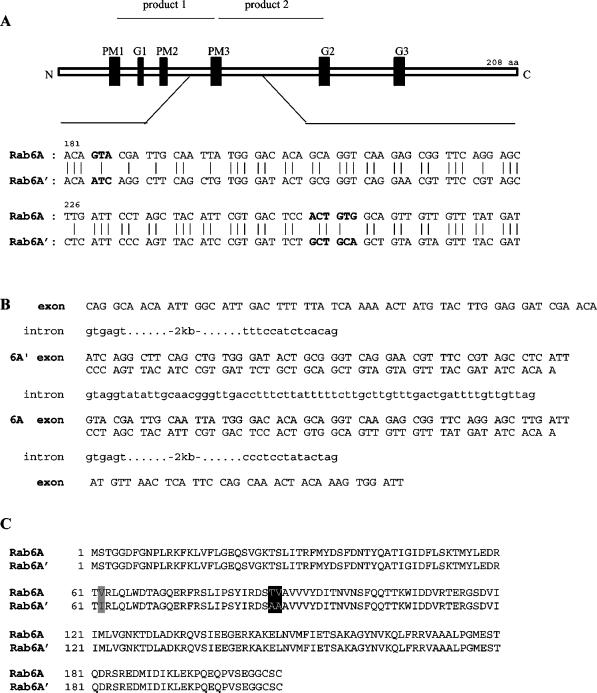
Figure 1.
Rab6A gene organization reveals the presence of a duplicated exon. (A) Alignment of the divergent sequence stretch between human Rab6A and Rab6A′ cDNA sequences. Product 1 and product 2 represent partial sequences that were amplified by RT-PCR with degenerate Rab-specific primers on RNA from Caco-2 cells and HUVECs, respectively. Product 2 was used as a probe to isolate the corresponding Rab6A′ cDNA sequence from a human endothelial cell cDNA library. The 5′ and 3′ UTRs of both sequences are identical. Consensus domains for phosphate/magnesium binding (PM1–PM3) and guanine nucleotide binding (G1–G3) are depicted as black boxes. Nucleotide triplets that lead to amino acid divergence are in bold. (B) Partial genomic organization of the Rab6A gene reveals a duplicated exon separated by an intron of 66 bp. The exon for Rab6A′ sequence precedes the exon for Rab6A. (C) Amino acid sequence alignment of the Rab6A isoforms. Nonconservative amino acid changes are boxed in black, and conservative substitutions are boxed in gray.
Rab6A Gene Organization
The facts that identical UTRs were found in Rab6A and Rab6A′ cDNAs and that the nucleotide changes were restricted to only one relatively long region make it unlikely that the Rab6A′ sequence resulted from a duplication of the Rab6A gene or was due to the existence of a Rab6A pseudogene. Rather, Rab6A′ sequence could be the result of either allelic polymorphism or mutually exclusive incorporation of distinct exons by alternative splicing of the Rab6A gene. To address this issue, we set up a genomic PCR-based strategy to elucidate the organization of the Rab6A gene. Based on the Rab6A and Rab6A′ cDNA sequences, forward and reverse primers specific for either of the two sequences were constructed (see MATERIALS AND METHODS). Two nonspecific primers flanking the stretch of divergence were also designed. We then performed PCR on human genomic DNA with the use of different combinations of primer sets (our unpublished results). The partial genomic organization of the Rab6A gene shown in Figure 1B revealed two homologous but distinct exons separated by an intron of 66 bp. In addition, the exon encoding Rab6A′ precedes the exon encoding Rab6A, and both are flanked by an intron of ∼2 kilobases. The intron boundary sequences conform to the GT-AG rule (Krawczak et al., 1992). We confirmed the presence of the two exons by performing additional PCRs on different human genomic DNA sources (our unpublished results). Therefore, Rab6A′ is generated by alternative use of a homologous but distinct exon within the Rab6A gene.
Comparison of the predicted protein sequence of Rab6A′ with the Rab6A sequence revealed only three amino acid differences in two regions flanking the PM3 domain: one conservative substitution (I instead of V at position 62) and two nonconservative changes (the “TV motif”: A for T and A for V at positions 87 and 88, respectively) (Figure 1C).
During the course of this study, a new isoform of human Rab6, termed Rab6C, was characterized in human platelets (Fitzgerald and Reed, 1999). The amino acid sequence of Rab6A′ is identical to that of Rab6C. However, because we confirm in our study that Rab6A′ (Rab6C) and Rab6A are products of the same gene, we believe that it is more appropriate to retain the Rab6A′ nomenclature.
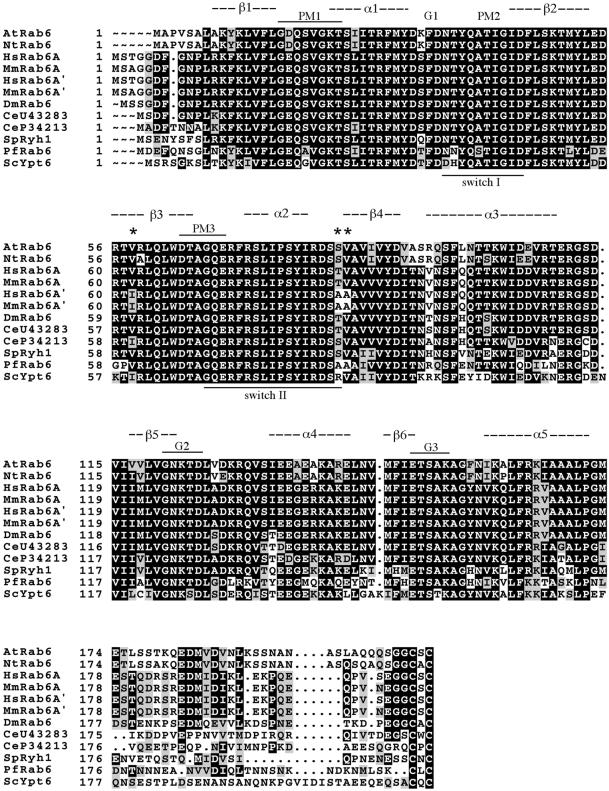
Figure 2 shows an alignment of Rab6 proteins identified in various species. A search in the expressed sequence tag databases revealed two partial mouse Rab6 clones that exhibit similar sequence divergence as the human clones. Their deduced amino acid sequences correspond exactly to their human counterparts, one bearing the TV motif, the other one the AA motif (Figure 2). It is of note that only one Rab6 gene is present in the yeast Saccharomyces cerevisiae, encoding the Ypt6 protein (Strom et al., 1993), as well as in the parasite Plasmodium falciparum (de Castro et al., 1996; Templeton and Kaslow, 1998). Interestingly, P. falciparum Rab6 contains the AA motif, and in Ypt6, a nonconservative amino acid (R) replaces the first A. Two putative Rab6 proteins exist in Caenorhabditis elegans, both with a TV or TV-like motif (Figure 2). It also should be pointed out that in all organisms, Rab6 proteins have either a valine or an isoleucine at position 62.
Figure 2.
Sequence alignment of Rab6 proteins from different species. Identical amino acids are boxed in black, and conservative substitutions are boxed in gray. The black stars indicate residue divergences between human (or mouse) Rab6A and Rab6A′. Consensus domains for phosphate/magnesium binding (PM1–PM3) and guanine nucleotide binding (G1–G3) and putative secondary structural elements (α helices, β strands) are depicted above the sequence alignment. At, Arabidopsis thaliana (accession number CAB38902); Nt, Nicotiana tabacum (accession number L29273); Hs, Homo sapiens (accession number M28212); Mm, Mus musculus (accession number AB041575); Dm, Drosophila melanogaster (accession number D84314); Ce, Caenorhabditis elegans (accession numbers U43283 and P34213); Sp, Schizosaccharomyces pombe (accession number X52475); Pf, Plasmodium falciparum (accession number X92977); Sc, Saccharomyces cerevisiae (accession number Q99260). MmRab6A′ sequence is derived from overlapping expressed sequence tags (accession numbers BE457736.1 and AA170363.1).
Rab6A and Rab6A′ Are Ubiquitously Expressed
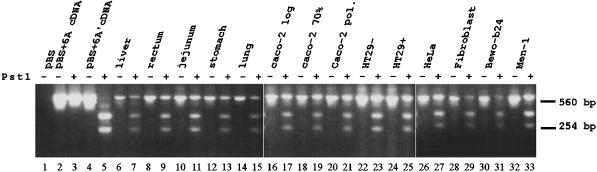
The existence of two Rab6A isoforms prompted us to analyze their expression patterns. A RT-PCR on total RNAs from a wide variety of adult human tissues and cell lines was performed with the use of primers corresponding to specific sequences present in the 5′ and 3′ UTRs (Figure 3). We took advantage of the presence of a unique PstI restriction site in the Rab6A′ sequence to distinguish between the expression of both forms. As shown in Figure 3 (lanes 1–5), only Rab6A′ cDNA was sensitive to PstI digestion, and the addition of an excess of restriction enzyme ensures complete digestion. Amplification with the specific primers resulted in an expected product of 675 bp, present in all tissues and cell lines examined (lanes PstI −). Subsequent digestion of the product resulted in three bands (lanes PstI +): the undigested Rab6A product and two lower fragments (405 and 270 bp) corresponding to the digested Rab6A′ cDNA sequence. The results shown in Figure 3 indicate that Rab6A and Rab6A′ RNAs are coexpressed in all tissues and cell lines examined. Interestingly, no striking differences in expression ratios between the forms were observed. In addition, no differences in the expression of Rab6A and Rab6A′ RNAs were found in polarized versus nonpolarized Caco-2 and HT29 cells (lanes 17, 19, and 21 and lanes 23 and 25, respectively). However, it should be pointed out that Rab6A′ (Rab6C) appears to be the prominent Rab6 isoform in human platelets (Fitzgerald and Reed, 1999).
Figure 3.
Mutually exclusive alternative splicing of the duplicated exon within the Rab6A gene results in the ubiquitous expression of two Rab6A isoforms. The coding regions of Rab6A isoforms were amplified by RT-PCR on various human tissues and cell lines with primers corresponding to sequences in 5′ and 3′ UTRs (Pst −, product of 675 bp). Products were digested with PstI (Pst +) to distinguish between Rab6A (675 bp) and Rab6A′ (400 and 275 bp) expression. Complete and specific digestion was controlled with excess amounts of Rab6A and Rab6A′ cDNA (lanes 1–5).
Rab6A and Rab6A′ Show Similar GTP-binding Activities
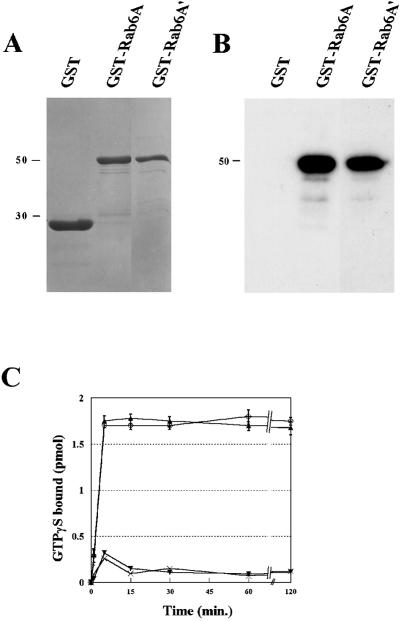
We next compared the biochemical properties of human Rab6A and Rab6A′. GST fusion proteins were purified from E. coli (Figure 4A) and analyzed by GTP overlay. Both proteins were found to bind [α-32P]GTP to a similar extent (Figure 4B). To measure the kinetics of guanine nucleotide binding, fusion proteins were incubated for various times with [35S]GTPγS. As shown in Figure 4C, the two forms bind the radiolabeled nucleotide in a time-dependent, saturable manner at 30°C (∼0.2 pmol of [35S]GTPγS per pmol of GST–Rab6A and GST–Rab6A′). No significant difference was observed between Rab6A and Rab6A′.
Figure 4.
Rab6A and Rab6A′ show similar GTP-binding properties. Samples of GST–Rab6A and GST–Rab6A′ fusion proteins or GST alone (1 μg/lane) were subjected to SDS-PAGE and stained with Coomassie blue (A) or transferred to nitrocellulose membranes (B), followed by incubation with 1 nM [α-32P]GTP. (C) For time-course studies of [35S]GTPγS binding, 500 ng of GST–Rab6A (▴) or GST–Rab6A′ (○) were incubated with tracer [35S]GTPγS at 30°C. As a control, GTPγS binding was measured to GST (×) or to GST-Rab6A′ in the presence of an excess of unlabeled GTPγS (▾). At the indicated times, binding activities were measured. The results show means of two independent experiments.
Rab6A displays a very low, almost undetectable GTPase activity (Yang et al., 1993; Cuif et al., 1999). We found that this was also the case for Rab6A′ (our unpublished results).
Both Rab6A Isoforms Are Localized to the Golgi Apparatus
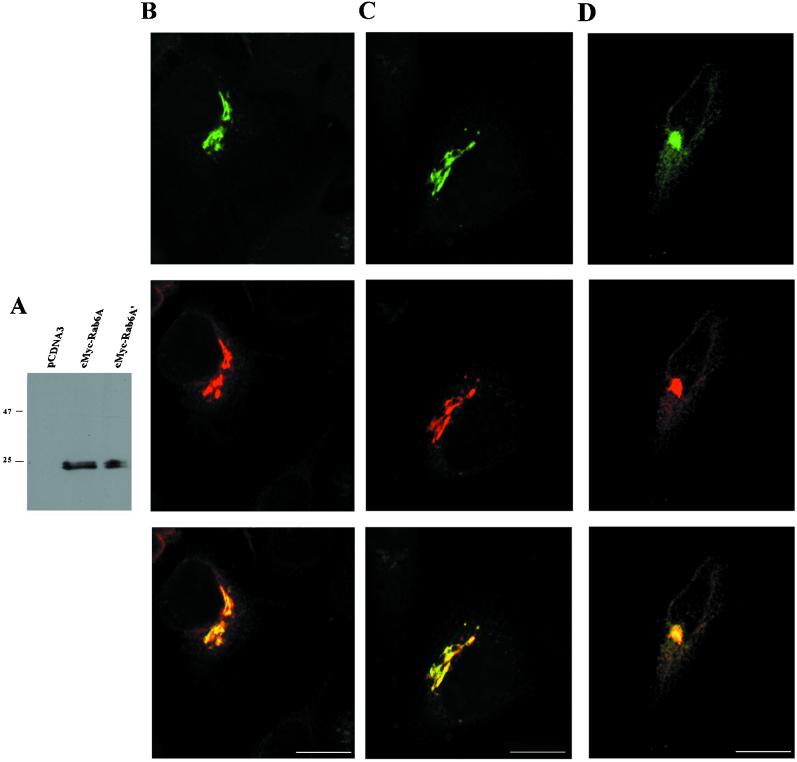
Endogenous Rab6 has been localized to membranes of the Golgi apparatus and the trans-Golgi network (Goud et al., 1990; Antony et al., 1992). Such a localization has been obtained with a specific polyclonal antibody raised against bacterially expressed Rab6A. However, this antibody cannot distinguish between the two Rab6A isoforms (our unpublished results). We expressed cMyc-tagged Rab6A and Rab6A′ in COS-1 (our unpublished results) and HeLa cells. As shown in Figure 5A, Rab6A and Rab6A′ migrated at the same apparent molecular mass on SDS-PAGE (25 kDa, containing 2 kDa of cMyc tag), a value that is consistent with the apparent molecular mass of endogenous Rab6 detected with the anti-Rab6A antibody (Goud et al., 1990). In the Western blot shown in Figure 5A, Rab6A and Rab6A′ appeared as a doublet, which likely corresponds to unprenylated and prenylated forms of the proteins (Yang et al., 1993).
Figure 5.
Both Rab6A and Rab6A′ colocalize at the Golgi apparatus. (A) HeLa cells were transfected with empty, cMyc–Rab6A-, or cMyc–Rab6A′-encoding plasmids, and cell lysates were subjected to immunoblotting with anti-cMyc antibody. (B–D) HeLa cells were cotransfected with plasmids encoding GFP-coupled N-acetylglucosaminyltransferase I and cMyc–Rab6A (column B), GFP-coupled N-acetylglucosaminyltransferase I and cMyc–Rab6A′ (column C), or GFP-coupled Rab6A′ and cMyc–Rab6A (column D). Cells were fixed with 1% paraformaldehyde, immunostained with anti-cMyc antibodies, and viewed with a confocal microscope. Upper panels, localization of GFP fusions (green); middle panels, c-Myc staining (red); lower panels, merged images. Scale bar, 10 μm.
The expression of tagged proteins also enabled us to follow the localization of individual proteins by immunofluorescence. For both isoforms, the staining with anti-cMyc antibody in HeLa cells (Figure 5, B and C) overlaps with the signal obtained with the cotransfected, GFP-coupled medial Golgi marker N-acetylglucosaminyltransferase I (Shima et al., 1997). This indicates that both endogenous Rab6A′ and Rab6A associate with Golgi membranes. To examine their mutual localization pattern, differentially tagged forms of Rab6A′ (GFP-coupled) and Rab6A (cMyc-tagged) were coexpressed in HeLa cells. As seen in Figure 5D, GFP–Rab6A′ colocalized with cMyc–Rab6A in the Golgi area of transfected cells. We also observed a weak reticular staining outside of the Golgi apparatus that likely corresponds to the ER. It should be noted that in this region, Rab6A and Rab6A′ are closely juxtaposed but not completely colocalized. A similar pattern was observed when tags of the isoforms were reversed (our unpublished results).
Both Rab6A Q72L and Rab6A′ Q72L Inhibit the Secretory Pathway
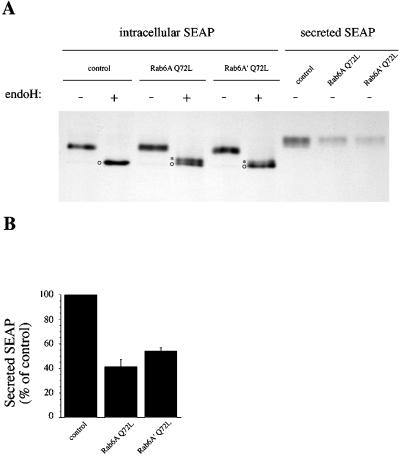
The overexpression of the GTP-bound form of Rab6A (Rab6A Q72L) leads to a strong inhibition of the secretion of both luminal (e.g., secreted alkaline phosphatase [SEAP]) and transmembrane (e.g., hemagglutinin of influenza virus) secretory markers (Martinez et al., 1994). To investigate Rab6A′ function, HeLa cells were transfected with the GTP-bound mutant of Rab6A′ (Rab6A′ Q72L) and then metabolically pulse-labeled and chased for 2 h, and the secretion of SEAP into the extracellular medium was measured. As shown in Figure 6, A and B, Rab6A′ Q72L inhibits secretion to a similar extent as Rab6A Q72L (∼50% inhibition compared with control conditions). As documented previously, the block in secretion at the level of the Golgi apparatus induced by Rab6A Q72L results in unusual (partially) endoglycosidase H–resistant forms of intracellular SEAP that are not detected in control cells (Figure 6A, asterisks). A similar effect was obtained by overexpressing Rab6A′ Q72L, although the partially endoglycosidase H–resistant form of SEAP was reproducibly less prominent in Rab6A′ Q72L than in Rab6A Q72L conditions.
Figure 6.
Both Rab6A Q72L and Rab6A′ Q72L inhibit SEAP secretion and affect N-glycosylation. (A) HeLa cells were cotransfected for 4 h with SEAP plasmids (encoding the SEAP marker) and pGEM-1 (control), Rab6A Q72L–, or Rab6A′ Q72L–encoding plasmids. Cells were then metabolically labeled for 10 min with [35S]methionine and [35S]cysteine and chased for 2 h, and SEAP was immunoprecipitated either from extracellular medium (secreted SEAP) or from the cells (intracellular SEAP). Intracellular SEAP was finally digested (+) or not (−) with endoglycosidase H (endoH) to investigate SEAP processing by Golgi glycosylation enzymes. Asterisks, partially endoH-resistant form; circles, fully endoH-sensitive form. (B) Quantification of secreted SEAP after a 2-h chase (three independent experiments, mean ± SD). Results are expressed as percent of secreted SEAP in control conditions.
Rab6A′ Q72L Does Not Redistribute Golgi Resident Proteins into the ER
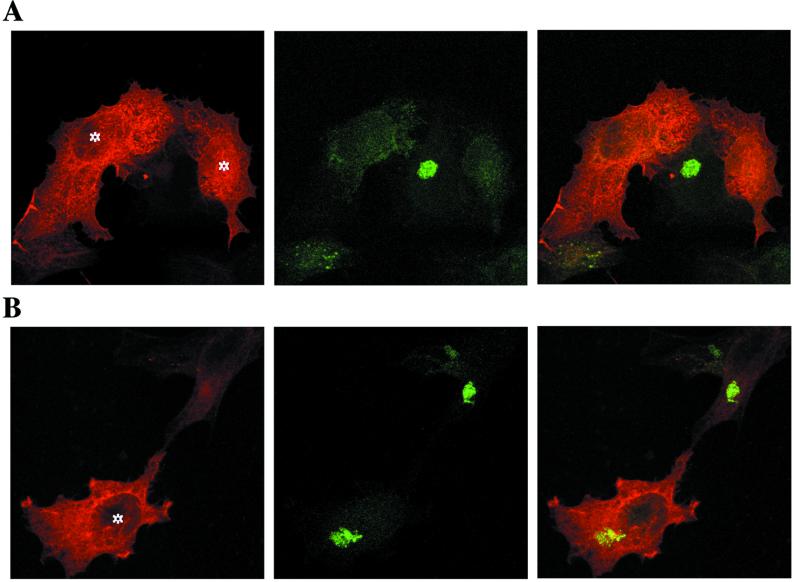
Another striking effect of the overexpression of Rab6A Q72L is a microtubule-dependent redistribution of Golgi proteins into the ER (Martinez et al., 1997). This effect is likely due to a stimulation by Rab6A Q72L of retrograde transport between the Golgi and ER, progressively relocating Golgi resident proteins into the ER. Indeed, it was shown recently that Rab6A controls a COPI-independent retrograde pathway between the Golgi and ER and that dominant negative forms of Rab6A (GDP-bound) block the continuous recycling of Golgi resident proteins through the ER (Girod et al., 1999; White et al., 1999). It was of interest to investigate whether the overexpression of Rab6A′ Q72L would induce the same phenotypic effect as Rab6A Q72L. Interestingly, the overexpression of Rab6A′ Q72L led to no detectable change in the morphology of Golgi membranes (Figure 7B), whereas under the same conditions, Rab6A Q72L–overexpressing cells induced a complete loss of Golgi structures (Figure 7A), as documented previously (Martinez et al., 1994, 1997).
Figure 7.
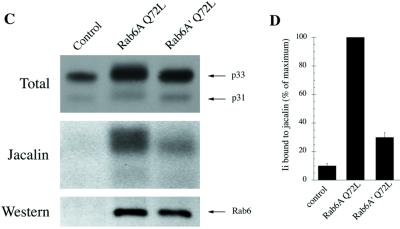
Rab6A′ Q72L does not redistribute Golgi proteins into the ER. HeLa cells transfected with GTP-bound Rab6A Q72L– (A) or Rab6A′ Q72L– (B) encoding plasmids were fixed 5 h after transfection with 4% paraformaldehyde. Cells were double labeled with a mAb to the medial Golgi antigen CTR433 (middle panel, green staining) and anti-Rab6 polyclonal antibody (left panel, red staining). The right panel shows the superimposition of the two labeling patterns. Asterisks indicate the cells that highly overexpressed Rab6A or Rab6A′ mutants (both recognized by the anti-Rab6 antibody). In Rab6A′ Q72L–overexpressing cells, the Golgi apparatus remains intact, whereas Rab6A Q72L overexpression leads to the redistribution of Golgi proteins into the ER (the compact Golgi staining is lost). It should be pointed out that Rab6 staining (red) appears diffuse because the bulk of overexpressed Rab6 remains in the cytosol (Martinez et al., 1994). (C) HeLa cells cotransfected for 4 h with Ii and pGEM-1 (control), Rab6A Q72L–, or Rab6A′ Q72L–encoding plasmids were then metabolically labeled for 10 min with [35S]methionine and [35S]cysteine and chased for 3 h. Ii (p31/p33) was immunoprecipitated: one-tenth of immune precipitates were analyzed by SDS-PAGE (Total), and nine-tenths was subjected to jacalin chromatography (Jacalin). Glycosylated immunoprecipitated Ii forms from Rab6A Q72L transfected cells migrate with a slower mobility than in control or Rab6A′ Q72L transfected cells (Total). Rab6A and Rab6A′ are overexpressed at similar levels, as shown by Western blotting before the immunoprecipitation step (Western). Note that endogenous Rab6 is not detected in the control lane because of the small amount of total proteins loaded on the gel. (D) Quantification of Ii bound to jacalin columns (three independent experiments, mean ± SD). For each experiment, results are expressed as percent of Ii bound to jacalin in Rab6A Q72L conditions (percent of maximum).
The Rab6A Q72L–induced relocalization of Golgi enzymes, including O-glycosyltransferases, into the ER allows the glycosylation of ER resident proteins, such as the invariant chain (Ii) of major histocompatibility complex class II molecules expressed in HeLa cells. Such an effect can be monitored by SDS-PAGE, because O-glycosylated Ii molecules (both the p33 and p31 forms) migrate with a slower mobility than non-O-glycosylated Ii forms (Figure 7C, Total) (Martinez et al., 1997). In contrast, no shift in mobility of Ii molecules was observed in Rab6A′ Q72L–overexpressing cells compared with control cells (Figure 7C, Total). The different effects induced by Rab6A Q72L and Rab6A′ Q72L on Ii glycosylation were measured quantitatively by chromatography on jacalin columns (Figure 7, C, Jacalin, and D), a lectin that binds specifically to the central motif of O-linked glycans. As documented previously (Martinez et al., 1997), ∼10-fold more Ii taken from Rab6A Q72L–expressing cells was retained on jacalin columns compared with control (Figure 7D). A much lower proportion of Ii was bound to jacalin in Rab6A′ Q72L–expressing cells. Nevertheless, the amount of Ii was significantly higher in these cells than in control cells (approximately twofold). The observed effects are not due to differences in the overexpression of Rab6 isoforms, because the expression levels of both proteins were equivalent on Western blots (Figure 7C, Western).
Together, these results (Figures 6 and 7) indicate that although it is localized to the Golgi, Rab6A′ does not function in the same way as Rab6A (see DISCUSSION).
Rab6A′ Q72L Does Not Interact with Rabkinesin-6
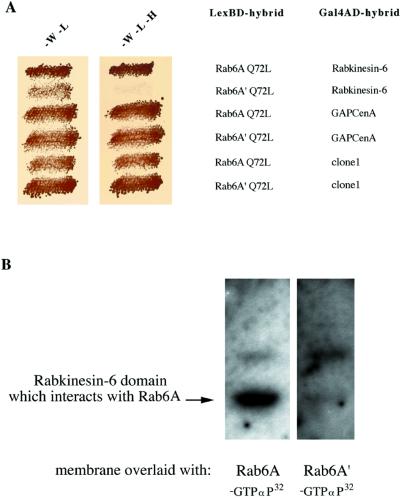
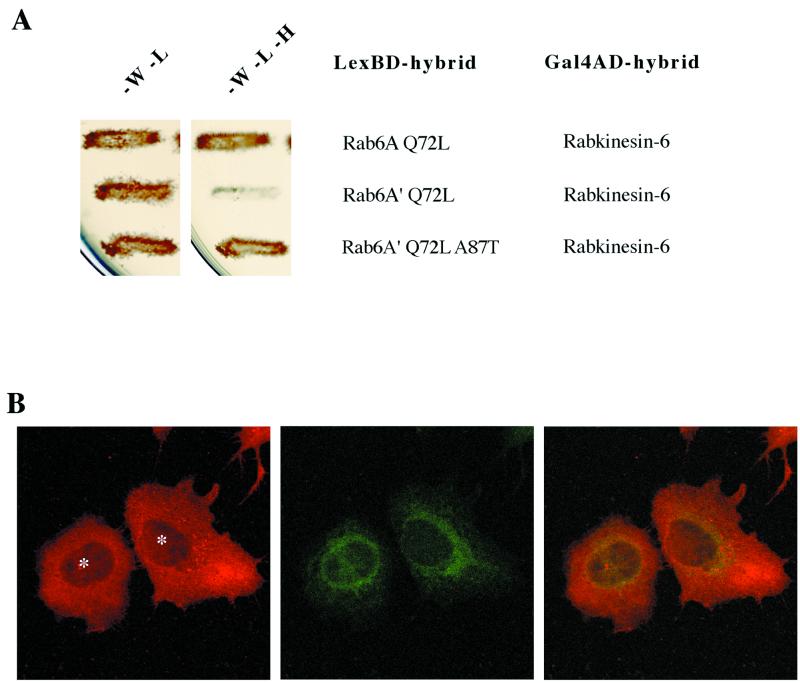
To further address Rab6A′ function, we investigated the interaction of Rab6A′ with three known Rab6A effectors identified in a yeast two-hybrid screen with Rab6A Q72L as bait: Rabkinesin-6 (Echard et al., 1998), a kinesin-like protein associated with the Golgi apparatus; GAPCenA (Cuif et al., 1999), a Rab6 GTPase activating protein; and clone 1, a 150-kDa cytosolic protein with an extensive coiled-coil domain whose exact function is still unknown (F. Jollivet, I. Janoueix-Lerosey, and B. Goud, unpublished results). The three proteins preferentially interact with the GTP-bound conformation of Rab6A. The interaction pattern of Rab6A′ Q72L with these three proteins was monitored with the use of the yeast two-hybrid assay. Although Rab6A′ was found to interact with both GAPCenA and clone 1 with a strength similar to that found for Rab6A, no interaction was detected with Rabkinesin-6 (Figure 8A). The absence of interaction between Rabkinesin-6 and Rab6A′ was further confirmed by overlay experiments. The 136 amino acids of the Rabkinesin-6 domain involved in the interaction with Rab6A (Echard et al., 1998) were incubated with either radiolabeled Rab6A–[α-32P]GTP (Figure 8B, left lane) or Rab6A′–[α-32P]GTP (Figure 8B, right lane). No interaction between Rab6A′ and the Rab6A-interacting domain of Rabkinesin-6 was observed, whereas a strong signal was detected with Rab6A. The lack of interaction between Rab6A′ and Rabkinesin-6 was also confirmed by coimmunoprecipitation experiments in HeLa cells overexpressing Rab6A′ and Rabkinesin-6 (our unpublished results).
Figure 8.
Rab6A′ Q72L does not interact with Rabkinesin-6. (A) The S. cerevisiae reporter strain L40 was cotransformed with either pLexA–Rab6A Q72L or pLexA–Rab6A′ Q72L and pGADGH–GAPCenA/Rab6-interacting domain, pGADGH–Rabkinesin-6, or pGADGH–clone 1. Transformants were patched for 3 d at 30°C on selective medium lacking tryptophan and leucine (left) or lacking tryptophan, leucine, and histidine (right). Growth in the right panel indicates an interaction between the encoded proteins. Western blot analysis on yeast lysates showed comparable expression levels of Rab6A and Rab6A′ (our unpublished results). (B) The same amounts of bacterially purified Rabkinesin-6 domain (amino acids 529–665), which directly binds to Rab6A, were independently run on SDS-PAGE and transferred onto nitrocellulose membranes. After a renaturation step, each membrane was incubated with either Rab6A (left panel) or Rab6A′ (right panel) previously exchanged with radiolabeled GTP. Membranes were washed separately and autoradiographed in parallel.
The A87T Mutation in Rab6A′ Q72L Restores the Ability of the Protein to Interact with Rabkinesin-6 and to Redistribute Golgi Proteins into the ER
In an attempt to identify those residues that are critical to the interaction with Rabkinesin-6, we replaced by site-directed mutagenesis the amino acids specific to Rab6A′ with the corresponding residues of Rab6A. For this purpose, we made the following constructs: Rab6A′ Q72L A87T, Rab6A′ Q72L A87T A88V, and Rab6A′ I62V Q72L A87T A88V (which is identical to Rab6A Q72L). Interestingly, the single substitution of alanine 87 to threonine in Rab6A′ was already sufficient to restore the interaction of the protein with Rabkinesin-6 (Figure 9A). In addition, this construct, when overexpressed in HeLa cells, induced the loss of Golgi structures, as Rab6A Q72L does (Figures 8A and 9B). The same results were obtained by overexpressing Rab6A′ Q72L A87T A88V or Rab6A′ I62V Q72L A87T A88V (our unpublished results). Therefore, these experiments indicate that the functional differences between Rab6A and Rab6A′ are dependent on residue A or T at position 87.
Figure 9.
Rab6A′ Q72L A87T interacts with Rabkinesin-6 and redistributes Golgi proteins into the ER. (A) The S. cerevisiae reporter strain L40 was cotransformed with pLexA–Rab6A Q72L, pLexA–Rab6A′ Q72L, or pLexA–Rab6A′ Q72L A87T and pGADGH–Rabkinesin-6. Transformants were analyzed as described in the legend to Figure 8. (B) HeLa cells transfected with Rab6A′ Q72L A87T–encoding plasmid were fixed 5 h after transfection with 4% paraformaldehyde. Cells were double labeled with a mAb to the medial Golgi antigen CTR433 (middle panel, green staining) and anti-Rab6 polyclonal antibody (left panel, red staining). The right panel shows the superimposition of the two labeling patterns. Asterisks indicate the cells that highly overexpressed Rab6A′ Q72L A87T mutant.
DISCUSSION
Generation of Rab6A Isoforms by Alternative Splicing
We have shown that two homologous but distinct exons exist within the Rab6A gene. Mutually exclusive alternative splicing of these exons results in the constitutive expression of two Rab6A isoforms. To our knowledge, this is the first example of such a mechanism within the Rab gene family. Alternative mRNA splicing has been suggested for the Rab28 gene based on an insertion of 95 bp found in a homologous Rab28 cDNA sequence (Brauers et al., 1996). However, this insertion is believed to be generated by an additional exon rather than by mutually exclusive incorporation of a duplicated exon. The genomic organization of only four Rab genes (Rab1, Rab3A, Rab11B, and S10) has been reported so far (Wichmann et al., 1989; Baumert et al., 1993; Lai et al., 1994; Zheng et al., 1997). It remains to be established whether alternative splicing of duplicated exons occurs for other members of the Rab family.
Splicing isoforms generated from duplicated exons and/or spliced variants generated from a limited set of genes have now been reported for a variety of genes and/or gene families, including some involved in intracellular transport. For instance, exon duplication was documented for chicken and human SNAP-25 genes (Bark, 1993; Bark and Wilson, 1994). The switch between the exons appears to result in a modification of membrane-binding properties of the SNAP-25 protein. Another example is the dynamin gene family, whose products play an essential role in clathrin- and non-clathrin-coated vesicle formation (McNiven, 1998). The three known genes give rise to at least 25 different spliced variants, which are differentially distributed in cells and tissues. Alternative splicing may then represent a mechanism to generate functional diversity in proteins involved in the complex events underlying intracellular transport. As shown here in the case of Rab proteins, alternative splicing may allow the generation of Rab isoforms interacting with distinct sets of effectors.
The TV Motif
Although Rab6A and Rab6A′ differ by three amino acids (an isoleucine instead of a valine at position 62 and two alanines instead of threonine and valine at positions 87 and 88, respectively), we found that the residue at position 87 is critical for the interaction with Rabkinesin-6. Therefore, it is likely that threonine 87 of Rab6A is involved directly in the interaction with Rabkinesin-6. This is consistent with the recent work of Ostermeier and Brunger (1999) on the crystal structure of the Rab3A/Rabphilin-3A complex, showing that a methionine residue at position 96 of Rab3A (corresponding to the threonine 87 in Rab6A) interacts directly with the SGAWFF structural element of Rabphilin-3A. In addition, the P. falciparum Rab6 structure shows that alanine 85 of this protein (corresponding to threonine 87 in the human Rab6A) is located at the surface of the molecule (Chattopadhyay et al., 2000). It is also notable that each member of the Rab family identified thus far contains either a valine or an isoleucine residue at the corresponding position 62 of Rab6. However, the amino acids corresponding to the TV/AA motifs of Rab6A/Rab6A′ are highly variable among members of the Rab family. The TV/AA residues are precisely positioned between the putative α2 helix and the β4 strand of Rab6A/Rab6A′ (Figure 2). The α2 helix corresponds to the switch II region and is predicted to change dramatically in conformation depending on whether Ras-like GTPases are bound to GDP or GTP (Krengel et al., 1990; Pai et al., 1990; Tong et al., 1991). Amino acid substitutions in this region, therefore, may modulate the conformational change induced by the binding of nucleotides and subsequently affect the interaction between Rab6A isoforms and their effectors.
It is noteworthy that the TV motif of Rab6A is part of a putative serine/threonine phosphorylation motif [RxxT(V)] (Hardie, 1993). In addition, both Rab6A and Rab6A′ contain PKC phosphorylation consensus sites, and indeed, Rab6C (Rab6A′) has recently been shown to be phosphorylated in platelets after thrombin activation (Fitzgerald and Reed, 1999). This raises the possibility that the interactions of Rab6A and Rab6A′ with their respective effectors could also be regulated by protein kinases. For instance, phosphorylation by a PKC-independent mechanism on the threonine of the Rab6A TV motif could specifically regulate interaction with Rab6A, but not with Rab6A′, effectors.
The Function of Rab6 Isoforms
A striking difference between Rab6A and Rab6A′ is the inability of the latter to induce in its GTP-bound conformation a redistribution of Golgi membranes into the ER, suggesting that both proteins fulfill distinct functions. The phenotypic effect of Rab6A Q72L is consistent with a role for this protein in Golgi-to-ER retrograde transport. Nevertheless, Rab6A′ was shown to impair secretion of an anterograde cargo, as Rab6A Q72L does. In addition, although it does not affect Golgi morphology at the immunofluorescence level, Rab6A′ Q72L induces significant O-glycosylation of invariant chain expressed in HeLa cells. As documented previously, the bulk of Ii is associated with ER compartments, but it likely cycles between the ER and early Golgi compartments. The precise function of Rab6A′ remains to be established, but a tentative hypothesis is that Rab6A′ may in fact regulate a retrograde transport pathway between late and early Golgi compartments. Activation of this pathway by Rab6A′ Q72L could induce a relocalization of medial/late Golgi enzymes in the early Golgi (increasing the amount of glycosylated Ii) and indirectly affect anterograde transport of a secretory marker.
Does Rab6A′ Represent the “Ancestral” Form of Rab6?
Another important difference between Rab6A and Rab6A′ is the fact that Rab6A′ does not interact with Rabkinesin-6. Although not yet proven directly, it is likely that Rabkinesin-6 participates in the long-range movement of tubular transport carriers between the Golgi and the ER, which carry Rab6A on their membranes. The finding that the replacement of alanine 87 to threonine in Rab6A′ restores the ability of the protein to both interact with Rabkinesin-6 and redistribute Golgi proteins into the ER strongly supports this hypothesis. If Rab6A′ plays a role in short-range retrograde transport within the Golgi complex, interaction with a motor protein may not be critical for its function. In addition, it is noteworthy that only one copy of Rab6 is present in lower eukaryotes such as S. cerevisiae (Ypt6) and P. falciparum and that no Rabkinesin-6 homologues appear to be present in these organisms. Moreover, yeast Ypt6 and P. falciparum Rab6 do not contain the TV motif present in Rab6A that was found in this study to be critical for interaction with Rabkinesin-6. These observations raise the interesting possibility that the duplication of a Rab exon has occurred during evolution to generate an isoform (Rab6A) able to interact with Rabkinesin-6 to regulate a transport route involving Rabkinesin-6 function. In support of this hypothesis, we recently found that Rab6A is involved in an alternative Golgi-to-ER retrograde pathway (Girod et al., 1999; White et al., 1999). This pathway is distinct from the COPI-dependent retrograde pathway documented so far in all eukaryotic cells, including yeast, and used by ER resident proteins carrying the KDEL/HDEL or KKXX motifs to be retrieved from the Golgi (Pelham, 1995). It is conceivable that such a route, used in mammalian cells by Shiga toxin to reach the ER, and likely by Golgi enzymes to recycle through this compartment, does not exist in lower eukaryotes.
In conclusion, it remains to be established whether alternative splicing of duplicated exons arises in other genes encoding for Rab GTPases. If so, this genetic mechanism could represent a more general system used by eukaryotic cells to generate functional diversity at the interface between closely related Rab proteins and their effectors, allowing the cells to handle the increased complexity of membrane traffic and intracellular transport during evolution.
ACKNOWLEDGMENTS
We thank François Darchen, Gordon Langsley, and Bé Wieringa for critical reading of the manuscript. This work was supported by grants from the Human Frontier Science Program and the Ligue Nationale Contre le Cancer (to B.G.). H.J.P.C. de Leeuw was supported by the Netherlands Heart Foundation (Grant 93.086).
REFERENCES
- Antony C, Cibert C, Geraud G, Santa Maria A, Maro B, Mayau V, Goud B. The small GTP-binding protein rab6p is distributed from medial Golgi to the trans-Golgi network as determined by a confocal microscopic approach. J Cell Sci. 1992;103:785–796. doi: 10.1242/jcs.103.3.785. [DOI] [PubMed] [Google Scholar]
- Baldini G, Scherer PE, Lodish HF. Nonneuronal expression of Rab3A: induction during adipogenesis and association with different intracellular membranes than Rab3D. Proc Natl Acad Sci USA. 1995;92:4284–4288. doi: 10.1073/pnas.92.10.4284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barbosa MD, Johnson SA, Achey K, Gutierrez MJ, Wakeland EK, Zerial M, Kingsmore SF. The Rab protein family: genetic mapping of six Rab genes in the mouse. Genomics. 1995;30:439–444. doi: 10.1006/geno.1995.1262. [DOI] [PubMed] [Google Scholar]
- Bark IC. Structure of the chicken gene for SNAP-25 reveals duplicated exon encoding distinct isoforms of the protein. J Mol Biol. 1993;233:67–76. doi: 10.1006/jmbi.1993.1485. [DOI] [PubMed] [Google Scholar]
- Bark IC, Wilson MC. Human cDNA clones encoding two different isoforms of the nerve terminal protein SNAP-25. Gene. 1994;139:291–292. doi: 10.1016/0378-1119(94)90773-0. [DOI] [PubMed] [Google Scholar]
- Baumert M, Fischer von Mollard G, Jahn R, Sudhof TC. Structure of the murine rab3A gene: correlation of genomic organization with antibody epitopes. Biochem J. 1993;293:157–163. doi: 10.1042/bj2930157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benli M, Doring F, Robinson DG, Yang X, Gallwitz D. Two GTPase isoforms, Ypt31p and Ypt32p, are essential for Golgi function in yeast. EMBO J. 1996;15:6460–6475. [PMC free article] [PubMed] [Google Scholar]
- Brauers A, Schurmann A, Massmann S, Muhl Zurbes P, Becker W, Kainulainen H, Lie C, Joost HG. Alternative mRNA splicing of the novel GTPase Rab28 generates isoforms with different C-termini. Eur J Biochem. 1996;237:833–840. doi: 10.1111/j.1432-1033.1996.0833p.x. [DOI] [PubMed] [Google Scholar]
- Celis JE. Cell Biology: A Laboratory Handbook. San Diego: Academic Press; 1998. [Google Scholar]
- Chattopadhyay D, Langsley G, Carson M, Recacha R, DeLucas L, Smith C. Structure of the nucleotide-binding domain of Plasmodium falciparumRab6 in the GDP-bound form. Acta Crystallogr D Biol Crystallogr. 2000;56:937–944. doi: 10.1107/s0907444900007575. [DOI] [PubMed] [Google Scholar]
- Chavrier P, Goud B. The role of ARF and rab GTPases in membrane transport. Curr Opin Cell Biol. 1999;11:466–475. doi: 10.1016/S0955-0674(99)80067-2. [DOI] [PubMed] [Google Scholar]
- Chirgwin JM, Przybyla AE, MacDonald RJ, Rutter WJ. Isolation of biologically active ribonucleic acid from sources enriched in ribonuclease. Biochemistry. 1979;18:5294–5299. doi: 10.1021/bi00591a005. [DOI] [PubMed] [Google Scholar]
- Cuif MH, Possmayer F, Zander H, Bordes N, Jollivet F, Couedel Courteille A, Janoueix-Lerosey I, Langsley G, Bornens M, Goud B. Characterization of GAPCenA, a GTPase activating protein for Rab6, part of which associates with the centrosome. EMBO J. 1999;18:1772–1782. doi: 10.1093/emboj/18.7.1772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Darchen F, Zahraoui A, Hammel F, Monteils MP, Tavitian A, Scherman D. Association of the GTP-binding protein Rab3A with bovine adrenal chromaffin granules. Proc Natl Acad Sci USA. 1990;87:5692–5696. doi: 10.1073/pnas.87.15.5692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Darmoul D, Lacasa M, Baricault L, Marguet D, Sapin C, Trotot P, Barbat A, Trugnan G. Dipeptidyl peptidase IV (CD 26) gene expression in enterocyte-like colon cancer cell lines HT-29 and Caco-2: cloning of the complete human coding sequence and changes of dipeptidyl peptidase IV mRNA levels during cell differentiation. J Biol Chem. 1992;267:4824–4833. [PubMed] [Google Scholar]
- de Castro FA, Ward GE, Jambou R, Attal G, Mayau V, Jaureguiberry G, Braun Breton C, Chakrabarti D, Langsley G. Identification of a family of Rab G-proteins in Plasmodium falciparumand a detailed characterization of pfrab6. Mol Biochem Parasitol. 1996;80:77–88. doi: 10.1016/0166-6851(96)02670-9. [DOI] [PubMed] [Google Scholar]
- de Leeuw HP, Koster PM, Calafat J, Janssen H, van Zonneveld AJ, van Mourik JA, Voorberg J. Small GTP-binding proteins in human endothelial cells. Br J Haematol. 1998;103:15–19. doi: 10.1046/j.1365-2141.1998.00965.x. [DOI] [PubMed] [Google Scholar]
- Echard A, Jollivet F, Martinez O, Lacapere JJ, Rousselet A, Janoueix-Lerosey I, Goud B. Interaction of a Golgi-associated kinesin-like protein with Rab6. Science. 1998;279:580–585. doi: 10.1126/science.279.5350.580. [DOI] [PubMed] [Google Scholar]
- Feinberg AP, Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983;132:6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Fischer von Mollard G, Mignery GA, Baumert M, Perin MS, Hanson TJ, Burger PM, Jahn R, Sudhof TC. rab3 is a small GTP-binding protein exclusively localized to synaptic vesicles. Proc Natl Acad Sci USA. 1990;87:1988–1992. doi: 10.1073/pnas.87.5.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fitzgerald ML, Reed GL. Rab6 is phosphorylated in thrombin-activated platelets by a protein kinase C-dependent mechanism: effects on GTP/GDP binding and cellular distribution. Biochem J. 1999;342:353–360. [PMC free article] [PubMed] [Google Scholar]
- Geppert M, Sudhof TC. RAB3 and synaptotagmin: the yin and yang of synaptic membrane fusion. Annu Rev Neurosci. 1998;21:75–95. doi: 10.1146/annurev.neuro.21.1.75. [DOI] [PubMed] [Google Scholar]
- Girod A, Storrie B, Simpson JC, Johannes L, Goud B, Roberts LM, Lord JM, Nilsson T, Pepperkok R. Evidence for copI-independent transport routes from the Golgi complex to the endoplasmic reticulum. Nat Cell Biol. 1999;1:423–430. doi: 10.1038/15658. [DOI] [PubMed] [Google Scholar]
- Gonzalez L, Jr, Scheller RH. Regulation of membrane trafficking: structural insights from a Rab/effector complex. Cell. 1999;96:755–758. doi: 10.1016/s0092-8674(00)80585-1. [DOI] [PubMed] [Google Scholar]
- Goud B, Zahraoui A, Tavitian A, Saraste J. Small GTP-binding. protein associated with Golgi cisternae. Nature. 1990;345:553–556. doi: 10.1038/345553a0. [DOI] [PubMed] [Google Scholar]
- Hardie DG. Protein Phosphorylation. Oxford, UK: IRL Press at Oxford University Press; 1993. [Google Scholar]
- Hunziker W, Peters PJ. Rab17 localizes to recycling endosomes and regulates receptor-mediated transcytosis in epithelial cells. J Biol Chem. 1998;273:15734–15741. doi: 10.1074/jbc.273.25.15734. [DOI] [PubMed] [Google Scholar]
- Iezzi M, Escher G, Meda P, Charollais A, Baldini G, Darchen F, Wollheim CB, Regazzi R. Subcellular distribution and function of Rab3A, B, C, and D isoforms in insulin-secreting cells. Mol Endocrinol. 1999;13:202–212. doi: 10.1210/mend.13.2.0228. [DOI] [PubMed] [Google Scholar]
- Janoueix-Lerosey I, Jollivet F, Camonis J, Marche PN, Goud B. Two-hybrid system screen with the small GTP-binding protein Rab6: identification of a novel mouse GDP dissociation inhibitor isoform and two other potential partners of Rab6. J Biol Chem. 1995;270:14801–14808. doi: 10.1074/jbc.270.24.14801. [DOI] [PubMed] [Google Scholar]
- Kari B, Lussenhop N, Goertz R, Wabuke Bunoti M, Radeke R, Gehrz R. Characterization of monoclonal antibodies reactive to several biochemically distinct human cytomegalovirus glycoprotein complexes. J Virol. 1986;60:345–352. doi: 10.1128/jvi.60.2.345-352.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krawczak M, Reiss J, Cooper DN. The mutational spectrum of single base-pair substitutions in mRNA splice junctions of human genes: causes and consequences. Hum Genet. 1992;90:41–54. doi: 10.1007/BF00210743. [DOI] [PubMed] [Google Scholar]
- Krengel U, Schlichting L, Scherer A, Schumann R, Frech M, John J, Kabsch W, Pai EF, Wittinghofer A. Three-dimensional structures of H-ras p21 mutants: molecular basis for their inability to function as signal switch molecules. Cell. 1990;62:539–548. doi: 10.1016/0092-8674(90)90018-a. [DOI] [PubMed] [Google Scholar]
- Lai F, Stubbs L, Artzt K. Molecular analysis of mouse Rab11b: a new type of mammalian YPT/Rab protein. Genomics. 1994;22:610–616. doi: 10.1006/geno.1994.1434. [DOI] [PubMed] [Google Scholar]
- Lazar T, Gotte M, Gallwitz D. Vesicular transport: how many Ypt/Rab-GTPases make a eukaryotic cell? Trends Biochem Sci. 1997;22:468–472. doi: 10.1016/s0968-0004(97)01150-x. [DOI] [PubMed] [Google Scholar]
- Lledo PM, Johannes L, Vernier P, Zorec R, Darchen F, Vincent JD, Henry JP, Mason WT. Rab3 proteins: key players in the control of exocytosis. Trends Neurosci. 1994;17:426–432. doi: 10.1016/0166-2236(94)90017-5. [DOI] [PubMed] [Google Scholar]
- Martinez O, Antony C, Pehau Arnaudet G, Berger EG, Salamero J, Goud B. GTP-bound forms of rab6 induce the redistribution of Golgi proteins into the endoplasmic reticulum. Proc Natl Acad Sci USA. 1997;94:1828–1833. doi: 10.1073/pnas.94.5.1828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martinez O, Schmidt A, Salamero J, Hoflack B, Roa M, Goud B. The small GTP-binding protein rab6 functions in intra-Golgi transport. J Cell Biol. 1994;127:1575–1588. doi: 10.1083/jcb.127.6.1575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McNiven MA. Dynamin: a molecular motor with pinchase action. Cell. 1998;94:151–154. doi: 10.1016/s0092-8674(00)81414-2. [DOI] [PubMed] [Google Scholar]
- Novick P, Zerial M. The diversity of Rab proteins in vesicle transport. Curr Opin Cell Biol. 1997;9:496–504. doi: 10.1016/s0955-0674(97)80025-7. [DOI] [PubMed] [Google Scholar]
- Nuoffer C, Davidson HW, Matteson J, Meinkoth J, Balch WE. A GDP-bound of rab1 inhibits protein export from the endoplasmic reticulum and transport between Golgi compartments. J Cell Biol. 1994;125:225–237. doi: 10.1083/jcb.125.2.225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Opdam FJM, Kamps G, Croes H, van Bokhoven H, Ginsel LA, Fransen JAM. Expression of Rab small GTPases in epithelial Caco-2 cells: Rab21 is an apically located GTP-binding protein in polarised intestinal epithelial cells. Eur J Cell Biol. 2000;79:308–316. doi: 10.1078/S0171-9335(04)70034-5. [DOI] [PubMed] [Google Scholar]
- Ostermeier C, Brunger AT. Structural basis of Rab effector specificity: crystal structure of the small G protein Rab3A complexed with the effector domain of rabphilin-3A. Cell. 1999;96:363–374. doi: 10.1016/s0092-8674(00)80549-8. [DOI] [PubMed] [Google Scholar]
- Pai EF, Krengel U, Petsko GA, Goody RS, Kabsch W, Wittinghofer A. Refined crystal structure of the triphosphate conformation of H-ras p21 at 1.35 Å resolution: implications for the mechanism of GTP hydrolysis. EMBO J. 1990;9:2351–2359. doi: 10.1002/j.1460-2075.1990.tb07409.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pelham HR. Sorting and retrieval between the endoplasmic reticulum and Golgi apparatus. Curr Opin Cell Biol. 1995;7:530–535. doi: 10.1016/0955-0674(95)80010-7. [DOI] [PubMed] [Google Scholar]
- Shima DT, Haldar K, Pepperkok R, Watson R, Warren G. Partitioning of the Golgi apparatus during mitosis in living HeLa cells. J Cell Biol. 1997;137:1211–1228. doi: 10.1083/jcb.137.6.1211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singer Kruger B, Stenmark H, Dusterhoft A, Philippsen P, Yoo JS, Gallwitz D, Zerial M. Role of three rab5-like GTPases, Ypt51p, Ypt52p, and Ypt53p, in the endocytic and vacuolar protein sorting pathways of yeast. J Cell Biol. 1994;125:283–298. doi: 10.1083/jcb.125.2.283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strom M, Vollmer P, Tan TJ, Gallwitz D. A yeast GTPase-activating protein that interacts specifically with a member of the Ypt/Rab family. Nature. 1993;361:736–739. doi: 10.1038/361736a0. [DOI] [PubMed] [Google Scholar]
- Templeton TJ, Kaslow DC. An amino terminal intron indicates that transcripts for the Plasmodium falciparumGolgi network marker Rab6 do not encode alternative amino termini. Mol Biochem Parasitol. 1998;94:149–153. doi: 10.1016/s0166-6851(98)00038-3. [DOI] [PubMed] [Google Scholar]
- Tong LA, de Vos AM, Milburn MV, Kim SH. Crystal structures at 2.2 Å resolution of the catalytic domains of normal ras protein and an oncogenic mutant complexed with GDP. J Mol Biol. 1991;217:503–516. doi: 10.1016/0022-2836(91)90753-s. [DOI] [PubMed] [Google Scholar]
- Valencia A, Chardin P, Wittinghofer A, Sander C. The ras protein family: evolutionary tree and role of conserved amino acids. Biochemistry. 1991;30:4637–4648. doi: 10.1021/bi00233a001. [DOI] [PubMed] [Google Scholar]
- Weber E, et al. Expression and polarized targeting of a rab3 isoform in epithelial cells. J Cell Biol. 1994;125:583–594. doi: 10.1083/jcb.125.3.583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- White J, Johannes J, Mallard F, Girod A, Grill S, Reinsch S, Keller P, Echard A, Goud B, Stelzer E. Rab6 coordinates a novel Golgi to ER retrograde transport pathway in live cells. J Cell Biol. 1999;147:743–759. doi: 10.1083/jcb.147.4.743. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wichmann H, Disela C, Haubruck H, Gallwitz D. Nucleotide sequence of the mouse ypt1 gene encoding a ras-related GTP-binding protein. Nucleic Acids Res. 1989;17:6737–6738. doi: 10.1093/nar/17.16.6737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang C, Mollat P, Chaffotte A, McCaffrey M, Cabanie L, Goud B. Comparison of the biochemical properties of unprocessed and processed forms of the small GTP-binding protein, rab6p. Eur J Biochem. 1993;217:1027–1037. doi: 10.1111/j.1432-1033.1993.tb18334.x. [DOI] [PubMed] [Google Scholar]
- Zacchi P, Stenmark H, Parton RG, Orioli D, Lim F, Giner A, Mellman I, Zerial M, Murphy C. Rab17 regulates membrane trafficking through apical recycling endosomes in polarized epithelial cells. J Cell Biol. 1998;140:1039–1053. doi: 10.1083/jcb.140.5.1039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng JY, Koda T, Arimura Y, Kishi M, Kakinuma M. Structure and expression of the mouse S10 gene. Biochim Biophys Acta. 1997;1351:47–50. doi: 10.1016/s0167-4781(97)00022-5. [DOI] [PubMed] [Google Scholar]