Abstract
Background
Aminopeptidase N (APN) type proteins isolated from several species of lepidopteran insects have been implicated as Bacillus thuringiensis (Bt) toxin-binding proteins (receptors) for Cry toxins. We examined brush border membrane vesicle (BBMV) proteins from the mosquito Anopheles quadrimaculatus to determine if APNs from this organism would bind mosquitocidal Cry toxins that are active to it.
Results
A 100-kDa protein with APN activity (APNAnq 100) was isolated from the brush border membrane of Anopheles quadrimaculatus. Native state binding analysis by surface plasmon resonance shows that APNAnq 100 forms tight binding to a mosquitocidal Bt toxin, Cry11Ba, but not to Cry2Aa, Cry4Ba or Cry11Aa.
Conclusion
An aminopeptidase from Anopheles quadrimaculatus mosquitoes is a specific binding protein for Bacillus thuringiensis Cry11Ba.
Background
The main African vectors of malaria are in the Anopheles gambiae complex mosquitoes [1]. In general, all species of Anopheles have been found to be susceptible to a certain extent to infection by some strain of human plasmodia [2]. Studies on lepidopteran insects revealed several types of Bt toxin-binding proteins (receptors): aminopeptidase N (APN) -like proteins [3,4]; cadherin-like proteins [5,6]; a glycoconjugate [7] and glycolipids [8]. In mosquitoes, two types of receptors were discovered: a protein with maltase activity from Culex. pipiens that binds the Bin toxin of Bacillus sphaericus [9], and a 65 kDa protein of unknown function (lacking aminopeptidase activity) from Aedes aegypti that binds Cry4Ba and Cry11Aa [10]. Two APNs have been identified in Ae. aegypti but not associated with binding Cry proteins [11]
APNs (EC 3.4.11.2) are exopeptidases that cleave single amino acids from the N-terminus of a polypeptide. APNs are expressed in many tissues including the brain, the lung, blood vessels, primary cultures of fibroblasts [12], and have the highest levels in intestinal and kidney brushborder membranes [13]. APNs belong to the M1 family of zinc metallopeptidases [14], which includes related enzymes like aminopeptidase A [15], aminopeptidase B [16,17], and leukotriene A4 hydrolase [18]. APNs have also been implicated as cellular receptors for human, canine, and feline coronaviruses [19].
In this study, intestinal APN from An. quadrimaculatus larvae was isolated and tested for binding ability to different mosquitocidal Cry toxins (Cry2Aa, Cry4Ba, Cry11Aa, and Cry11Ba). Membrane proteins were extracted from An. quadrimaculatus brush border membrane vesicles (BBMV) and separated by anion-exchange chromatography. Fractions containing APN activity were pooled and purified by size-exclusion chromatography. A 100-kDa protein with APN activity was isolated from the BBMV and its N-terminal sequence was determined to be AQLEDYRLNDDVRPTAYRIE. This protein was used to screen different mosquitocidal Cry toxins binding via Biacore analysis. From the screening, it was discovered that only Cry11Ba was able to bind the APN. A protein BLAST search limited to the arthropod database revealed three highly homologous An. gambiae APNs based on the N-terminal sequence.
Results
Purification of An. quadrimaculatus aminopeptidase N
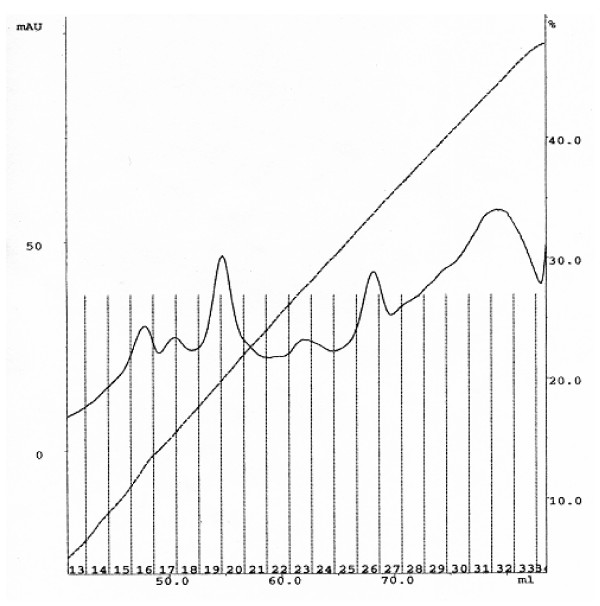
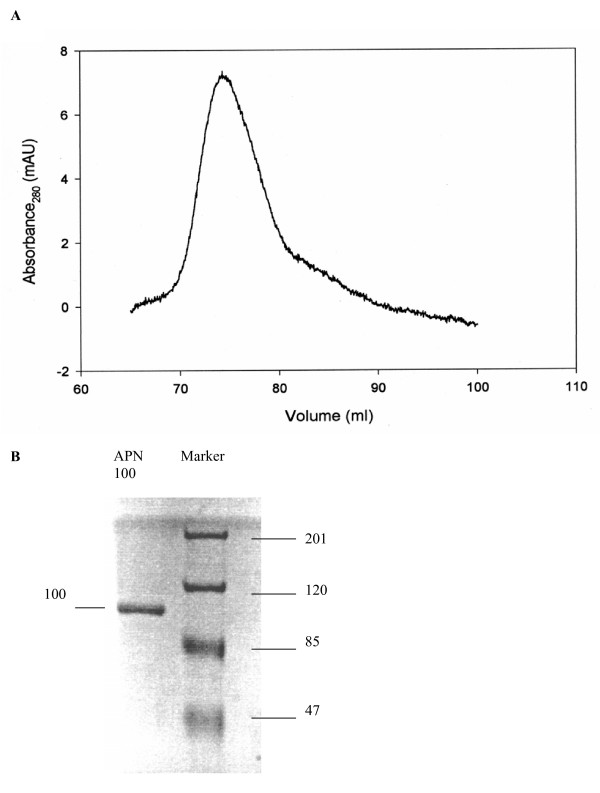
SPR analysis requires purified ligands and analytes to be used. Solubilized An. quadrimaculatus BBMV proteins were separated by anion-exchange chromatography and all elution fractions were tested for APN activity. Fractions 19–21 and 24–34 showed APN activity. Fractions 19–21 were made up of a single peak, and fractions 24–34 were made up of at least two peaks (Fig. 1). Fractions 19–21 were pooled, concentrated, and purified further by size-exclusion chromatography. A single peak was eluted at around 75 ml of run volume that correspond to a protein size of about 100 kDa (Fig. 2A). This peak was collected and was determined to hold APN activity. SDS-PAGE analysis of the protein also indicated a size of 100 kDa (Fig. 2B) and the 100 kDa protein was highly purified. The 100 kDa protein was named APNAnq 100.
Figure 1.
Separation of An. quadrimaculatus aminopeptidase N from solubilized BBMV proteins by anion-exchange chromatography. The UV absorbance at 280 nm (mAU) is indicated at the top left corner, and the percent conductivity of buffer B (%) is indicated at the top right corner. Collected fractions are shown at the bottom in 2-ml intervals. Run volume is indicated at the bottom (ml). Fractions 19–21 and 24–34 contain APN activity.
Figure 2.
(A) Further purification of APN fractions (fractions 19–21) from anion-exchange chromatography of An. quadrimaculatus BBMV by size-exclusion chromatography. A single peak was eluted at 75 ml elution volume, corresponding to 100 kDa. (B) SDS-PAGE of purified APN (APNAnq 100) obtained in (A) above. The estimated sizes of the protein bands are indicated on both sides of the gel in kDa.
Determination of binding affinity by SPR analysis
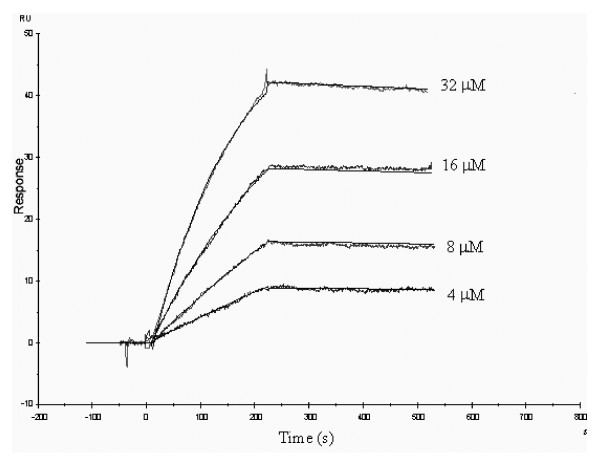
Initially, APNAnq 100 was evaluated for binding by SPR analysis to four Cry toxins (Cry2Aa, Cry4Ba, Cry11Aa and Cry11Ba), which were previously determined in this laboratory to have mosquitocidal activity towards An. quadrimaculatus (data not shown). Only Cry11Ba bound significantly to APNAnq 100. Further analysis of real-time binding kinetic of Cry11Ba to APNAnq 100 was performed at different analyte concentrations (Fig. 3), followed by global fitting of all the response curves. A 1:1 binding stoichiometry, including a drifting-baseline correction, produced the following apparent rate constants of the bimolecular interaction: ka = 184.0 M-1s-1 (± 1.0) and kd = 1.03 × 10-7 s-1 (± 4.01 × 10-6), KD = 0.56 nM. More complex binding models, such as 2-site independent binding (A + B1 ↔ AB1; A + B2 ↔ AB2), and 2-site sequential binding (A + B ↔ AB ↔ AB*) also gave as good fitting as the simple 1:1 binding (A + B ↔ AB) with χ2 = 0.112 (data not shown).
Figure 3.
Real-time binding of Cry11Ba to An. quadrimaculatus APNAnq 100. Experimental curves (jagged line) are shown overlaid with fitted curves (smooth line) obtained with the 1:1 Langmuir binding with drifting baseline model. The overlaid BIAcore response curves are shown for Cry11Ba toxin injections at 4, 8, 16, 32 μM as indicated.
N-terminal sequence of APNAnq100
A twenty amino acid residue sequence (AQLEDYRLNDDVRPTAYRIE) was obtained from N-terminal sequencing of purified APNAnq 100. Data mining for similar sequences in the arthropod databases revealed high homology (80–85% identities) with 3 conceptual translated proteins from An. gambiae (Table 1). A BLAST search using the first protein's full amino acid sequence from An. gambiae (accession no. EAA08760.1) revealed homology with many aminopeptidases from organisms of other genera (data not shown). This would suggest that the three proteins from An. gambiae have aminopeptidase activity.
Table 1.
Amino acid sequence similarities of the N-terminal sequence of APNAnq 100 from An. quadrimaculatus with three protein sequences from An. gambiae obtained through a BLAST search.
| Source identity Acc. No.b | Amino acid sequencea | % identity |
| An. quadrimaculatus | 1-AQLEDYRLNDDVRPTAYRIE-20 | NAc |
| An. gambiae EAA08760.1 | 42-AQLEDYRLNDDVWPTHYDIE-61 | 85 |
| An. gambiae EAA08929.1 | 53-AQLEEYRLNDDVWPTHYDIE-72 | 85 |
| An. gambiae EAA08763.1 | 45-AQPEDYRLNDDVWPTHYDIE-64 | 80 |
a The numbers flanking the sequences represent residue position in the protein.
b The accession no. in protein database.
c NA- Not applicable.
Analysis of the N-terminal region with the program SignalP (http://www.cbs.dtu.dk/) predicted that the most probable cleavage site for the signal peptide sequence was between position 25 and 26 for EAA08760.1; between position 27 and 28 for EAA08763.1; and between position 28 and 29 for EAA08929.1. However, the sequences of the proteins shown in Table 2 start at positions further downstream from the predicted cleavage sites, which suggested that there might have been further processing of the N-terminal region of the An. quadrimaculatus APN. Analysis of the C-terminal region for possible glycosylphosphatidylinositol (GPI) anchor sites using the program Big-PI Predictor (http://mendel.imp.univie.ac.at/gpi/gpi_server.html) found no potential GPI-modification site for EAA08760.1. Potential GPI-modification sites were found at position 930 and 920 for EAA08763.1 and EAA08929.1, respectively. Analysis of the sequences using the program NetOGlyc 2.0 (http://www.cbs.dtu.dk/services/NetOGlyc/) [20] to reveal potential GalNAc O-glycosylation sites found 5 sites in EAA08760.1, 7 sites in EAA08763.1, and 6 sites in EAA08929.1. Analysis of the sequences using NetNGlyc 1.0 (http://www.cbs.dtu.dk/services/NetNGlyc/) [21] to reveal potential N-glycosylation sites found 2 sites in EAA08760.1, 8 sites in EAA08763.1, and 3 sites in EAA08929.1.
Table 2.
Putative aminopeptidases in An. gambiae that contain a conserved MAAVPDFSAGAMENWGLL sequence.
| No. | Accession no. | Protein length (residues) |
| 1 | EAA05382.1 | 649 |
| 2 | EAA01063.1 | 1800 |
| 3 | EAA13235.1 | 1691 |
| 4 | EAA09719.1 | 734 |
| 5 | EAA08912.1 | 811 |
| 6 | EAA02981.1 | 641 |
| 7 | EAA08915.1 | 870 |
| 8 | EAA08931.1 | 997 |
| 9 | EAA12046.1 | 955 |
| 10 | EAA10722.1* | 809 |
| 11 | EAA08434.1 | 990 |
| 12 | EAA08760.1 | 791 |
| 13 | EAA08910.1 | 614 |
| 14 | EAA08929.1 | 940 |
| 15 | EAA03210.1 | 639 |
| 16 | EAA08763.1 | 952 |
* HEXXH motif for the APN zinc-iron-binding site does not exist in this sequence, which would exclude this protein from the metallopeptidase family.
Another protein BLAST search was performed using the sequence of a known conserved region for aminopeptidases (MAAVPDFSAGAMENWGLL) [22], which yielded 16 homologous proteins from the An. gambiae genomic database (Table 2). This indicated that there are a large number of aminopeptidase isomers in these mosquitoes.
Discussion and conclusion
An aminopeptidase N (APN) type protein has been implicated as a Cry toxin-binding protein in several lepidopteran species: Manduca sexta [4], Bombyx mori [23,24], Lymantria dispar [25,26], Heliothis virescens [27], Plutella xylostella [28], Trichoplusia ni [29], Helicoverpa armigera [30] and Spodoptera litura [31]. Recently the binding epitopes of Cry1Aa to an APN from B. mori have been mapped by monoclonal antibody inhibition [32]. Thus, targeting APN for analysis as a possible toxin-binding protein is a reasonable approach.
The surface plasmon resonance (SPR) method allows analysis of bimolecular interaction in the native state, without a potentially interfering label [33]. Thus, since the Cry11Ba and APNAnq 100 interaction detected in this study represents tight (ca. 1 nM KD ) native-state binding, we propose that APNAnq 100 is a putative receptor for Cry11Ba. APNAnq 100 did not bind to Cry2Aa, Cry4Ba or Cry11Aa even though the toxins have insecticidal activity against An. quadrimaculatus. The specific binding of Cry11Ba to APNAnq 100 suggests that its mode of action would be different from Cry2Aa, Cry4Ba, or Cry11Aa.
The N-terminal sequence of APNAnq 100 showed high homology with three putative APNs from An. gambiae. One or more of these APNs could act as a binding protein for Cry11Ba.
Recently the binding epitopes of Cry1Aa to an APN from B. mori have been mapped by monoclonal antibody inhibition [32].
Methods
Preparation of mosquito brush border membrane vesicles (BBMV)
Fourth instars An. quadrimaculatus larvae were filtered with a nylon mesh, washed in distilled water, separated from large residual food particles, and dried briefly on a filter paper (Fisher) under vacuum suction. Harvested larvae were frozen at -70°C until needed. About 4–6 g of frozen larvae were homogenized in 8–12 ml of cold buffer A (300 mM mannitol, 5 mM EGTA, 17 mM Tris-HCl, pH 7.5). Larvae were homogenized by 40 strokes of Potter-Elvehjem PTFE pestle in glass tube at speed number 5 (~6000 rpm). BBMV were enriched through differential centrifugation by selective divalent-cation precipitations as described by Silva-Filha, et al [34]. The BBMV pellet was resuspended in 1 ml of ice-cold binding buffer (8 mM NaHPO4 , 2 mM KH2 PO4 , 150 mM NaCl, pH 7.4) supplemented with COMPLETE™ (Roche) protease inhibitor and homogenized by10 extrusions using a small Teflon pestle.The protein concentration of the BBMV was measured with the Coomassie protein assay reagent (Pierce), using BSA as the standard. The BBMV was kept at -70°C until needed.
Purification of An. quadrimaculatus aminopeptidase N (APN) from BBMV
Approximately 20 mg of BBMV was solubilized overnight at 4°C in the binding buffer supplemented with 10 mg/ml of CHAPS (Roche). Later, the solution was vortexed briefly and centrifuged at 15,000 rpm in a JA-17 rotor at 4°C for 10 min. The supernatant was treated with PIPLC for 1 hr at 37°C. The supernatant was separated by anion-exchange chromatography (HiTrap 5 ml column, Pharmacia) by continuous salt gradient using two buffers: A, 20 mM Tris-Cl, pH 7.4, 0.4 mg/ml CHAPS; B, buffer A with 1 M NaCl. Two milliliters elution fractions were collected at a flow rate of 1 ml/min. A small fraction of each elution fraction was tested for the presence of APN activity using L-leucine p-nitroanilide (Sigma) as substrate. Neighboring fractions containing APN activities were pooled and concentrated using centricon (YM30, Millipore) according to the manufacturer. The pooled fractions were further purified by size exclusion chromatography (Superdex 200, Pharmacia) in 20 mM Tris, pH 7.4, 0.4 mg/ml CHAPS and concentrated as before. The quality of the sample was checked by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) as described by Laemmli [35].
Purification of Cry toxins
An E. coli clone of Cry2Aa (a grateful gift from Takashi Yamamoto) was used as a source of this gene. The cry2Aa gene was extracted by PCR and cloned into plasmid pHT600 and transformed into B. thuringiensis 4Q7, a plasmidless Cry- derivitive. The genes cry4Aa, cry4Ba, cry11Aa and cry11Ba were received in the same plasmid vector and host B. thuringiensis strain (gratefully donated by Armelle Delécluse). Single Bt colonies were inoculated into a 5 ml LB medium supplemented with 10 μg/ml erythromycin and grown overnight at 30°C in an incubator-shaker at 250 rpm. These cultures were inoculated into a 500 ml SSM medium [36] also supplemented with erythromycin and incubated a further 4 days until sporulation and autolysis. Bt crystals in the autolysed-cells suspension were purified as described previously [37] for purification of Cry toxins expressed in E. coli, except that the sonication steps were omitted. The crystals were solubilized in carbonate buffer (30 mM Na2 CO3 , 20 mM NaHCO3 , pH 10.0) supplemented with 10 mM dithiothreitol (Roche) at 37°C for 3 hours. Next, the solubilized toxin was incubated with 1/20 (v/v) 10 mg/ml trypsin (Sigma) at 37°C for 3 hours. The activated toxin was purified by FPLC using a Superdex 200 (Pharmacia) column in the carbonate buffer. Protein concentration was measured using the Coomassie protein assay reagent (Pierce) with bovine serum albumin as standard.
Biosensor analysis of toxin-APN affinities
All surface plasmon resonance (SPR) experiments were performed on a BIAcore 3000 machine (Biacore AB). An. quadrimaculatus APN in 20 mM ammonium acetate, pH 4.2, was immobilized on a CM5 sensor chip by amine-coupling method (Biacore AB). The flow buffer HBS-EP (10 mM HEPES, 150 mM NaCl, 3 mM EDTA, 0.005% polysorbate 20 (v/v), pH 7.4) (Biacore AB) was used at a flow rate of 30 μl/min. Multiple concentrations (4, 8, 16, and 32 μM) of Cry11Ba was injected across the flow cell containing the APN and one blank flow cell containing ethanolamine as a blocking agent. Surfaces were regenerated with 2 pulses of 10 μl of 10 mM NaOH, pH 11, at 100 μl/min or until the signal return to baseline. Signal responses from the blank flow cells were subtracted from all response curves and data were globally fitted using BIAevaluation Ver. 3.1 (Biacore AB). The curves were fitted to a simple 1:1 Langmuir binding model (A+B ↔ AB) to obtain apparent rate constants.
N-terminal sequencing and sequence similarity search
For N-terminal sequencing, proteins separated in SDS-PAGE were transferred onto PVDF membrane (Roche) by electro-transfer (Mini-PROTEAN™ II, Bio Rad) according to the manufacturer. The membrane was stained briefly with Coomassie Blue R-250 and destained in 50% methanol. Bands representing 100-kDa proteins were excised and sequencing was performed on an automated sequencer (Model 477A, Applied Biosystems) at USDA Forest Service Laboratory, Delaware, OH. Data mining was performed on the N-terminal sequence using the basic local alignment search tool (BLAST), an on-line tool, at the National Center for Biotechnology Information (NCBI) website. The search parameter was limited to arthropods. CLUSTAL W (http://npsa-pbil.ibcp.fr/cgi-bin/npsa_automat.pl?page=npsa_clustalw.html) was used to align the amino acid sequences.
Authors' contributions
MAFA and DHD planned the study and wrote the initial draft of the manuscript. MAFA conducted all experiments, except the N-terminal amino acid analysis. APV conducted the N-terminal amino acid analysis. All authors were involved in revising the manuscript and giving final approval of the version to be published.
Acknowledgments
Acknowledgements
We thank A. Curtiss for technical assistance. This work was supported by NIH grant to D.H. Dean and M.J. Adang (Grant # R01 AI 29092).
Contributor Information
Mohd Amir F Abdullah, Email: mamir@uga.edu.
Algimantas P Valaitis, Email: avalaitis@fs.fed.us.
Donald H Dean, Email: dean.10@osu.edu.
References
- Ye-Ebiyo Y, Pollack RJ, Spielman A. Enhanced development in nature of larval Anopheles arabiensis mosquitoes feeding on maize pollen. Am J Trop Med Hyg. 2000;63:90–93. doi: 10.4269/ajtmh.2000.63.90. [DOI] [PubMed] [Google Scholar]
- Bates M. The natural history of mosquitoes. New York , The Macmillan Company; 1949. p. 379. [Google Scholar]
- Knight PJK, Crickmore N, Ellar DJ. The receptor for Bacillus thuringiensis CryIA(c) delta-endotoxin in the brush border membrane of the lepidopteran Manduca sexta is aminopeptidase N. Mol Microbiol. 1994;11:429–436. doi: 10.1111/j.1365-2958.1994.tb00324.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sangadala S, Walters FS, English LH, Adang MJ. A mixture of Manduca sexta aminopeptidase and phosphatase enhances Bacillus thuringiensis insecticidal CryIA(c) toxin binding and 86Rb+-K+ efflux in vitro. J Biol Chem. 1994;269:10088–10092. [PubMed] [Google Scholar]
- Vadlamudi RK, Ji TH, Bulla LAJ. A specific binding protein from Manduca sexta for the insecticidal toxin of Bacillus thuringiensis subsp. berliner. J Biol Chem. 1993;268:12334–12340. [PubMed] [Google Scholar]
- Nagamatsu Y, Toda S, Yamaguchi F, Ogo M, Kogure M, Nakamura M, Shibata Y, Katsumoto T. Identification of Bombyx mori midgut receptor for Bacillus thuringiensis insecticidal CryIA(a) toxin. Biosci Biotechnol Biochem. 1998;62:718–726. doi: 10.1271/bbb.62.718. [DOI] [PubMed] [Google Scholar]
- Valaitis AP, Jenkins JL, Lee MK, Dean DH, Garner KJ. Isolation and partial characterization of gypsy moth BTR-270, an anionic brush border membrane glycoconjugate that binds Bacillus thuringiensis Cry1A toxins with high affinity. Arch Insect Biochem Physiol. 2001;46:186–200. doi: 10.1002/arch.1028. [DOI] [PubMed] [Google Scholar]
- Griffitts JS, Haslam SM, Yang T, Garczynski SF, Mulloy B, Morris H, Cremer PS, Dell A, Adang MJ, Aroian RV. Glycolipids as receptors for Bacillus thuringiensis crystal toxin. Science. 2005;307:922–925. doi: 10.1126/science.1104444. [DOI] [PubMed] [Google Scholar]
- Darboux I, Nielsen-LeRoux C, Charles JF, Pauron D. The receptor of Bacillus sphaericus binary toxin in Culex pipiens (Diptera: Culicidae) midgut: molecular cloning and expression. Insect Biochem Mol Biol. 2001;31:981–990. doi: 10.1016/S0965-1748(01)00046-7. [DOI] [PubMed] [Google Scholar]
- Buzdin AA, Revina LP, Kostina LI, Zalunin IA, Chestukhina GG. Interaction of 65- and 62-kDa proteins from the apical membranes of the Aedes aegypti larvae midgut epithelium with Cry4B and Cry11A endotoxins of Bacillus thuringiensis. Biochemistry (Moscow) 2002;67:540–546. doi: 10.1023/A:1015594127636. [DOI] [PubMed] [Google Scholar]
- Pootanakit K, Angsuthanasombat C, Panyim S. Identification of two isoforms of aminopeptidase N in Aedes aegypti larval midgut. J Biochem Molec Biol. 2004;36:508–513. doi: 10.5483/bmbrep.2003.36.5.508. [DOI] [PubMed] [Google Scholar]
- Sanderink GJ, Artur Y, Siest G. Human aminopeptidases: a review of the literature. J Clin Chem Clin Biochem. 1988;26:795–807. doi: 10.1515/cclm.1988.26.12.795. [DOI] [PubMed] [Google Scholar]
- Maroux S, Louvard D, Baratti J. The aminopeptidase from hog intestinal brush border. Biochim Biophys Acta. 1973;321:282–295. doi: 10.1016/0005-2744(73)90083-1. [DOI] [PubMed] [Google Scholar]
- Rawlings ND, Barrett AJ. Evolutionary families of metallopeptidases. Methods Enzymol. 1995;248:183–228. doi: 10.1016/0076-6879(95)48015-3. [DOI] [PubMed] [Google Scholar]
- Nanus DM, Engelstein D, Gastl GA, Gluck L, Vidal MJ, Morrison M, Finstad CL, Bander NH, Albino AP. Molecular cloning of the human kidney differentiation antigen gp160: human aminopeptidase A. Proc Natl Acad Sci U S A. 1993;90:7069–7073. doi: 10.1073/pnas.90.15.7069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukasawa KM, Fukasawa K, Kanai M, Fujii S, Harada M. Molecular cloning and expression of rat liver aminopeptidase B. J Biol Chem. 1996;271:30731–30735. doi: 10.1074/jbc.271.48.30731. [DOI] [PubMed] [Google Scholar]
- Cadel S, Foulon T, Viron A, Balogh A, Midol-Monnet S, Noel N, Cohen P. Aminopeptidase B from the rat testis is a bifunctional enzyme structurally related to leukotriene-A4 hydrolase. Proc Natl Acad Sci U S A. 1997;94:2963–2968. doi: 10.1073/pnas.94.7.2963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Funk CD, Radmark O, Fu JY, Matsumoto T, Jornvall H, Shimizu T, Samuelsson B. Molecular cloning and amino acid sequence of leukotriene A4 hydrolase. Proc Natl Acad Sci U S A. 1987;84:6677–6681. doi: 10.1073/pnas.84.19.6677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tresnan DB, Holmes KV. Feline aminopeptidase N is a receptor for all group I coronaviruses. Adv Exp Med Biol. 1998;440:69–75. doi: 10.1007/978-1-4615-5331-1_9. [DOI] [PubMed] [Google Scholar]
- Hansen JE, Lund O, Tolstrup N, Gooley AA, Williams KL, Brunak S. NetOglyc: Prediction of mucin type O-glycosylation sites based on sequence context and surface accessibility. Glycocon J. 1998;15:115–130. doi: 10.1023/A:1006960004440. [DOI] [PubMed] [Google Scholar]
- Gupta R, Jung E, Brunak S. Prediction of N-glycosylation sites in human proteins. In preparation. 2002.
- Gill SS, Cowles EA, Francis V. Identification, isolation, and cloning of a Bacillus thuringiensis CryIAc toxin-binding protein from the midgut of the lepidopteran insect Heliothis virescens. J Biol Chem. 1995;270:27277–27282. doi: 10.1074/jbc.270.45.27277. [DOI] [PubMed] [Google Scholar]
- Hua G, Tsukamoto K, Rasilo ML, Ikezawa H. Molecular cloning of a GPI-anchored aminopeptidase N from Bombyx mori midgut: a putative receptor for Bacillus thuringiensis CryIA toxin. Gene. 1998;214:177–185. doi: 10.1016/S0378-1119(98)00199-1. [DOI] [PubMed] [Google Scholar]
- Yaoi K, Nakanishi K, Kadotani T, Imamura M, Koizumi N, Iwahana H, Sato R. cDNA cloning and expression of Bacillus thuringiensis Cry1Aa toxin binding 120 kDa aminopeptidase N from Bomyx mori. Biochim Biophys Acta. 1999;1444:131–137. doi: 10.1016/s0167-4781(98)00250-4. [DOI] [PubMed] [Google Scholar]
- Lee MK, You TH, Young BA, Valaitis AP, Dean DH. Aminopeptidase N purified from gypsy moth BBMV is a specific receptor for Bacillus thuringiensis CryIAc toxin. Appl Environ Microbiol. 1996;62:2845–2849. doi: 10.1128/aem.62.8.2845-2849.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garner KJ, Hiremath S, Lehtoma K, Valaitis AP. Cloning and complete sequence characterization of two gypsy moth aminopeptidase-N cDNAs, including the receptor for Bacillus thuringiensis Cry1Ac toxin. Insect Biochem Mol Biol. 1999;29:527–535. doi: 10.1016/S0965-1748(99)00027-2. [DOI] [PubMed] [Google Scholar]
- Luo K, Sangadala S, Masson L, Mazza A, Brousseau R, Adang MJ. The Heliothis virescens 170 kDa aminopeptidase functions as "receptor A" by mediating specific Bacillus thuringiensis Cry1A d-endotoxin binding and pore formation. Insect Biochem Mol Biol. 1997;27:735–743. doi: 10.1016/S0965-1748(97)00052-0. [DOI] [PubMed] [Google Scholar]
- Nakanishi K, Yaoi K, Shimada N, Kadotani T, Sato R. Bacillus thuringiensis insecticidal Cry1Aa toxin binds to a highly conserved region of aminopeptidase N in the host insect leading to its evolutionary success. Biochim Biophys Acta. 1999;1432:57–63. doi: 10.1016/s0167-4838(99)00086-2. [DOI] [PubMed] [Google Scholar]
- Lorence A, Darzon A, Bravo A. Aminopeptidase dependent pore formation of Bacillus thuringiensis Cry1Ac toxin on Trichoplusia ni membranes. FEBS Lett. 1997;414:303–307. doi: 10.1016/S0014-5793(97)01014-4. [DOI] [PubMed] [Google Scholar]
- Ingle SS, Trivedi N, Prasad R, Kuruvilla J, Rao KK, Chhatpar HS. Aminopeptidase-N from the Helicoverpa armigera (Hubner) brush border membrane vesicles as a receptor of Bacillus thuringiensis Cry1Ac delta-endotoxin. Curr Microbiol. 2001;43:255–259. doi: 10.1007/s002840010297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Agrawal N, Malhotra P, Bhatnagar RK. Interaction of Gene-Cloned and Insect Cell-Expressed Aminopeptidase N of Spodoptera litura with Insecticidal Crystal Protein Cry1C. Appl Environ Microbiol. 2002;68:4583–4592. doi: 10.1128/AEM.68.9.4583-4592.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Atsumi S, Mizono E, Hara H, Nakanishi K, Kitami M, Miura N, Tabunoki H, Watanabe A, Sato R. Loatation of the Bombyx mori aminopeptidase N type 1 binding site on Bacillus thuringiensis Cry1Aa toxin. Appl Environ Microbiol. 2005;71:3966–3977. doi: 10.1128/AEM.71.7.3966-3977.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fägerstam LG, Frostell A, Karlsson R, Kullman M, Larsson A, Malmqvist M, Butt H. Detection of antigen-antibody interactions by surface plasmon resonance. Application to epitope mapping. J Mol Recognit. 1990;3:208–214. doi: 10.1002/jmr.300030507. [DOI] [PubMed] [Google Scholar]
- Silva-Filha MH, Nielsen-Leroux C, Charles JF. Binding kinetics of Bacillus sphaericus binary toxin to midgut brush-border membranes of Anopheles and Culexsp. mosquito larvae. Eur J Biochem. 1997;247:754–761. doi: 10.1111/j.1432-1033.1997.00754.x. [DOI] [PubMed] [Google Scholar]
- Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Schaeffer P, Millet J, Aubert JP. Catabolic repression of bacterial sporulation. Proc Natl Acad Sci USA. 1965;54:704–711. doi: 10.1073/pnas.54.3.704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee MK, Milne RE, Ge AZ, Dean DH. Location of a Bombyx mori receptor binding region on a Bacillus thuringiensis d-endotoxin. J Biol Chem. 1992;267:3115–3121. [PubMed] [Google Scholar]