Abstract
Community-level physiological profiling based upon fluorometric detection of oxygen consumption was performed on hydroponic rhizosphere and salt marsh litter samples by using substrate levels as low as 50 ppm with incubation times between 5 and 24 h. The rate and extent of response were increased in samples acclimated to specific substrates and were reduced by limiting nitrogen availability in the wells.
Characterization and classification of heterotrophic microbial communities based on rapid assessment of multiple sole-carbon-source use, termed community-level physiological profiling (CLPP) by Lehman et al. (9), were developed over 10 years ago (6). Although CLPP is a valuable method for assessing relative change in microbial communities when properly applied (3, 12), limitations due to selective enrichment resulting from the high substrate concentrations (i.e., >100 mM) and long incubation periods (typically 1 to 4 days) restrict its present use for assessing actual differences in the physiological capabilities of microbial communities (2, 8, 14, 16). Development of more functionally relevant CLPP will depend on the application of new detection methods that allow for rapid and direct (e.g., O2 consumption or CO2 production) rather than indirect (e.g., tetrazolium dye reduction) assessment of respiration, with decreased requisite incubation times and substrate concentrations. In addition, the use of undefined, proprietary reagents should be minimized so that incubation conditions can be defined and manipulated by the investigator. 14C respirometric approaches limit enrichment effects by allowing for detection of physiological response with addition of very low substrate concentrations (i.e., nanomolar levels), but even simplified assay systems (13) still have the expense associated with disposal of radioactive wastes. CLPP approaches based on CO2 monitoring that limit incubation time have been developed (1) but require high substrate concentrations (i.e., 10 to 100 mM).
Tracking the rate of O2 consumption in the test medium may improve sensitivity, given the relatively low concentration and solubility of O2 in water. A recently developed, fluorescence-based microplate platform for assessing dissolved oxygen (BD Oxygen Biosensor System; BD Biosciences, Bedford, Mass.) (17) could enable rapid testing of multiple substrates; the effectiveness of this system to detect known shifts in substrate utilization by mixed microbial communities is the subject of this work. The BD Oxygen Biosensor System is based on an O2-sensitive fluorophore, 4,7-diphenyl-1,10-phenathroline ruthenium (II) chloride, absorbed into a silicone matrix that is permeable to O2 (17). The fluorescence of the ruthenium dye is quenched by the presence of O2, so the signal from the fluorophore-gel complex loaded on the bottom of the microplate wells increases in response to respiration in the overlying sample. In this work, samples of environmental systems suspended in sterile phosphate-buffered mineral salts (PBMS) (7 g of K2HPO4 liter−1, 3 g of KH2PO4 liter−1, 0.1 g of MgSO4 liter−1, 0.5 g of [NH4]2SO4 liter−1, 0.01 g of CaCl2 liter−1, 0.005 g of FeSO4 liter−1, 0.0025 g of MnSO4 liter−1, and 0.0025 g of Na2MoO4 liter−1) were inoculated into the wells.
In the first part of the work, the fluorescence responses of rhizosphere communities to two different types of surfactants were related to their previous exposure to these compounds and to independently collected data on surfactant degradation. These experiments were part of an ongoing project evaluating direct addition of human-hygiene water into hydroponic plant systems as an approach for water recycling on extended space missions, and specific description of the plant growth system is presented elsewhere (4). Briefly, wheat (Triticum aestivum L. cv. USU-Apogee) was grown by using nutrient film technique hydroponic culture inside a controlled environmental growth chamber. One hydroponic system served as a control with no continuous addition of surfactants, while the other systems received either sodium laureth sulfate (SLES) as Rhodapex ES-2 (25% SLES; Rhodia Inc., Cranbury, N.J.) or cocamidopropyl betaine (CAPB) as Mirataine BET C-30 (30% CAPB; Rhodia, Inc.), beginning 4 days after planting. Surfactant stock solutions (200 ml of 2,000 ppm) were added to the tanks in a continuous mode via a peristaltic pump over a 23-h period. Twenty days after planting, suspensions of rhizosphere communities were obtained by excising sections (∼1 by 1 cm) of root mat from the front of the trays and by hand shaking in 25 ml of sterile PBMS containing 2-mm glass beads for 2 min. A single rhizosphere sample was obtained from each treatment, and undiluted and diluted (1/5 and 1/25) rhizosphere suspensions were inoculated into the BD microplates and were read on a Dynex MFX Microplate Fluorometer at 485-nm excitation and 604-nm emission wavelengths with the top-reading mode. Plate contents were incubated at 30°C, and readings were obtained every 15 min for 120 h. Plates were not shaken during the incubation period. On day 21, a single pulse of 400 ml of a 1,000-ppm SLES and 1,000-ppm CAPB solution was added to all treatments to determine surfactant decay rates based on analysis of nutrient solution for SLES by using ion pair chromatography with suppressed conductivity detection (11) or for CAPB by using high-pressure liquid chromatography linked to electrospray ionization-mass spectrometry (10).
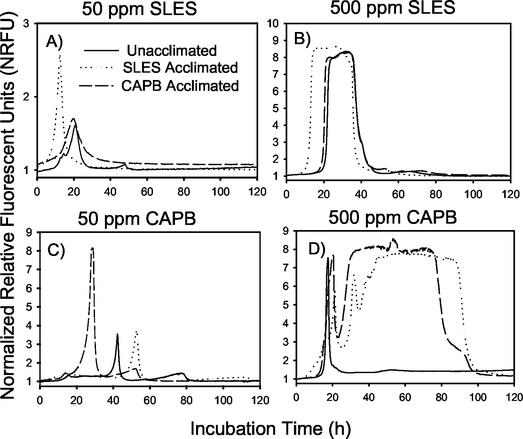
Samples from acclimated systems produced a more pronounced peak in fluorescence more rapidly when incubated in the BD systems at the 50-ppm level of surfactant (Fig. 1A and C; Table 1), corresponding to the higher rates of degradation in the same hydroponic systems as independently assessed by chemical analysis (Table 2). Both the extent and rate of fluorescence were dependent on inoculum density (Table 1), so comparisons among treatments were made with samples of approximately equivalent density (5.38 to 5.58 log cells ml−1 based on acridine orange direct counts [7] [Fig. 1]). For example, the extent of response (i.e., peak response) to 50 parts of SLES per million was greater in rhizosphere samples from SLES-acclimated systems (2.58 normalized relative fluorescent units [NRFU]) than in samples from the unacclimated (1.61 NRFU) or CAPB-acclimated (1.71 NRFU) systems and the rate of response (i.e., time to peak) was faster in the SLES-acclimated samples (12.5 h) than in the unacclimated (20.75 h) or CAPB-acclimated (19.75 h) samples. The greater SLES utilization in the BD microplate assay corresponds to a higher rate of SLES disappearance within the SLES-acclimated system (3.61 ppm h−1) than in the unacclimated or CAPB-acclimated systems (0.11 to 0.12 ppm h−1). Similarly, samples from the CAPB-acclimated system showed a greater response toward 50 parts of CAPB per million in the BD microplate assay than did samples from either the unacclimated or SLES-acclimated systems (Fig. 1B and D), corresponding to a higher rate of CAPB disappearance from the CAPB-acclimated systems. Biofilm samples removed from polyvinyl chloride coupons incubated in the nutrient delivery tanks showed a fluorescent response to acclimation similar to that seen with the rhizosphere samples (data not shown).
FIG. 1.
Fluorescence response of rhizosphere samples to 50 parts of SLES per million, 500 parts of SLES per million, 50 parts of CAPB per million, and 500 parts of CAPB per million. Data are presented for samples from hydroponic systems previously exposed to no surfactant (control), SLES, and CAPB. Inoculum density is approximately equivalent (5.38 to 5.58 log cells ml−1) for all samples. Origin of y axis equals 0.75 NRFU.
TABLE 1.
Response of samples from the rhizosphere of hydroponically grown wheat to SLES in the BD Oxygen Biosensor System
| Treatmenta | [C]b in ppm | Inoculum density (log cells ml−1)c | Minimum response time (h)d | Peake (NRFU) | Time to peak (h) | Areaf | Log-linear modelg results
|
Logistic modelh results
|
||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lag (h) | μmax | R2 | Lag (h) | μmax | R2 | |||||||
| Unacclimated | 50 | 5.58 | 11.75 | 1.61 | 20.75 | 2.45 | 9.1 | 1.04 | 0.930 | |||
| 4.88 | 14.25 | 1.69 | 23.50 | 6.31 | 18.0 | 1.10 | 0.986 | |||||
| 4.17 | 19.00 | 1.73 | 28.25 | 6.68 | 21.6 | 1.09 | 0.981 | |||||
| SLES acclimated | 50 | 6.78 | 0.50 | 6.10 | 5.75 | 133.8 | 2.0 | 1.66 | 0.995 | |||
| 6.08 | 2.25 | 2.96 | 8.75 | 130.6 | 4.0 | 1.25 | 0.991 | |||||
| 5.38 | 3.75 | 2.58 | 12.50 | 134.5 | 7.0 | 1.18 | 0.991 | |||||
| CAPB acclimated | 50 | 6.17 | 1.50 | 1.90 | 15.25 | 134.3 | 0.9 | 1.04 | 0.995 | |||
| 5.48 | 2.50 | 1.71 | 19.75 | 137.2 | 8.8 | 1.05 | 0.991 | |||||
| 4.78 | 14.25 | 1.65 | 24.00 | 128.3 | 15.4 | 1.06 | 0.997 | |||||
| Unacclimated | 500 | 5.58 | 10.75 | 7.11 | 19.00 | 239.5 | 16.6 | 2.27 | 0.993 | 18.2 | 2.45 | 0.998 |
| 4.88 | 8.75 | 7.33 | 22.75 | 252.6 | 20.0 | 2.21 | 0.991 | 21.8 | 1.92 | 0.998 | ||
| 4.17 | 18.00 | 7.33 | 27.00 | 238.9 | 24.0 | 2.10 | 0.986 | 26.0 | 2.06 | 0.999 | ||
| SLES acclimated | 500 | 6.78 | 0.75 | 8.31 | 5.00 | 278.2 | 2.0 | 2.55 | 0.977 | 3.67 | 3.27 | 0.996 |
| 6.08 | 2.25 | 8.29 | 8.25 | 292.7 | 4.7 | 2.10 | 0.989 | 9.77 | 2.29 | 0.998 | ||
| 5.38 | 4.50 | 8.21 | 12.25 | 294.6 | 7.4 | 1.88 | 0.982 | 12.87 | 1.79 | 0.999 | ||
| CAPB acclimated | 500 | 6.17 | 2.50 | 8.32 | 14.0 | 304.1 | 10.8 | 1.99 | 0.993 | 12.93 | 2.08 | 0.996 |
| 5.48 | 1.75 | 8.25 | 17.25 | 298.1 | 14.9 | 2.45 | 0.978 | 16.58 | 3.16 | 0.998 | ||
| 4.78 | 8.50 | 8.00 | 22.75 | 269.0 | 18.2 | 2.17 | 0.979 | 24.14 | 2.28 | 0.998 | ||
Samples obtained from systems receiving no surfactant (unacclimated) or either SLES or CAPB.
Concentration of SLES added to the wells within the BD Oxygen Biosensor plates.
Single samples removed from each system were inoculated into microplates after no, 1/5, or 1/25 dilution, resulting in three different inoculum densities.
Time for normalized relative fluorescence to increase by 10% (i.e., to 1.1).
Maximum normalized relative fluorescence.
Peak area calculated with Sigma Plot function.
Results from linear regression analysis of log-transformed normalized relative fluorescence versus time; lag is the x intercept; μmax is the slope (retransformed into linear space). The linear portion of the data was selected for analysis based on visual inspection.
Results from curve fitting to four-parameter logistic curve Y = Y0 + [a/1 + (x/x0)b]. Lag estimate equals x0; μmax estimate equals 1/b; a is a fitting parameter.
TABLE 2.
Results of linear regression analysis of surfactant decay data from pulse addition studiesa
| Surfactant | Treatment | Slope (ppm h−1) | R2 |
|---|---|---|---|
| SLES | Unacclimated | −0.12 | 0.90 |
| SLES acclimated | −3.67 | 0.95 | |
| CAPB acclimated | −0.11 | 0.76 | |
| CAPB | Unacclimated | −0.011 | 0.66 |
| SLES acclimated | −0.010 | 0.49 | |
| CAPB acclimated | −0.184 | 0.98 |
Analysis was performed on nonzero surfactant concentration (log transformed) versus time after addition. Treatment refers to the previous exposure of the system.
At the 500-ppm level, interpretation of the fluorescence response was less clear. The lag in SLES response remained shorter in acclimated samples as estimated by various means, but little difference existed in the peak value due to the asymptotic nature of the response, apparently due to the maintenance of low dissolved O2 as a result of sustained respiration (Fig. 1B). The asymptotic level of fluorescence (∼8 NRFU) was the maximal fluorescence response as determined from sodium sulfite (100 mM) controls. Differences among samples were ambiguous at the 500-ppm level of CAPB due to pronounced secondary peaks and overall complexity in the signal (Fig. 1D). While the degradation pathway of the CAPB molecule is not well defined, multiple steps (with different reaction rates) are likely required for complete oxidation of the entire molecule and probably cause the multiple peaks (15).
Various response parameters describing either the delay (i.e., minimum response time, time to peak, or lag estimated from the log-linear model) or peak in fluorescence appear suitable for comparison of samples (details in Table 1). Certain models (i.e., logistic model) are only applicable to the higher substrate concentration (500 ppm) in which an asymptotic level of fluorescence is observed, while definition of both the extent and rate of peak response is more readily applied to the lower substrate concentrations (50 ppm) due to the clear definition of a peak (visualization of response curves in Fig. 1). Further studies are needed to assess the most suitable variables for consistently discriminating among communities; the various parameters reported in Table 1 reflect a number of the different analytical approaches to evaluating the microplate data.
Lower substrate concentrations are preferred in physiological assays, given the lower potential for selective enrichment, and in this study the 50-ppm level provided more easily interpretable data. Substrate concentrations below 50 ppm (i.e., 5 and 10 ppm) were tested but did not produce a detectable response (data not shown).
A stimulatory effect of acclimation was not observed when rhizosphere communities were incubated in Biolog plates containing SLES or CAPB as a sole carbon source. Rhizosphere samples were obtained during three replicate studies and were prepared as described above, with the exception that sterile deionized water rather than PBMS was used as a diluent, given the concentration of nutrients already present in Biolog plates, and was inoculated into Biolog MT plates. The absorbance was negligible after 72 h of incubation for both surfactants at 5,000 and 50 ppm for all treatments (Table 3). The response was greater at 500 ppm for both surfactants, but no consistent differences were apparent between the acclimated and unacclimated systems (Table 3). The lack of correlation between the response in the Biolog CLPP approach and known differences in specific carbon source utilization has been previously observed (5).
TABLE 3.
Response of rhizosphere community samples from hydroponic systems either unacclimated (no surfactant added) or acclimated to SLES or CAPB in Biolog NT plates
| Community | Surfactant | Substrate concn (ppm) | A590a |
|---|---|---|---|
| Unacclimated | SLESb | 5,000 | 0.18 (0.14) |
| 500 | 0.40 (0.14) | ||
| 50 | 0.16 (0.73) | ||
| SLES acclimated | 5,000 | 0.14 (0.15) | |
| 500 | 0.36 (0.10) | ||
| 50 | 0.09 (0.08) | ||
| CAPB acclimated | 5,000 | 0.08 (0.10) | |
| 500 | 0.22 (0.09) | ||
| 50 | 0.04 (0.06) | ||
| Unacclimated | CAPBc | 5,000 | 0.30 (0.10) |
| 500 | 0.31 (0.12) | ||
| 50 | 0.16 (0.09) | ||
| SLES acclimated | 5,000 | 0.21 (0.20) | |
| 500 | 0.44 (0.28) | ||
| 50 | 0.07 (0.08) | ||
| CAPB acclimated | 5,000 | 0.22 (0.10) | |
| 500 | 0.46 (0.12) | ||
| 50 | 0.05 (0.08) |
A590 with subtraction of blank, after 72 h of incubation. Data are presented as mean and standard deviation of three replicates.
SLES was added to wells at the designated concentration.
CAPB was added to wells at the designated concentration.
Follow-up tests were performed with salt marsh plant litter decomposition communities to determine (i) if detectable patterns of response could be produced for a variety of readily assimilated substrates (i.e., amino acids, monosaccharides, and organic acids) and (ii) if the extent of response was sensitive to relevant biochemical factors that affect substrate utilization (i.e., N availability). Litter bags (20 by 20 cm) filled with ∼20 g of locally collected, air-dried black needle rush, Juncus roemarianus, were placed in a restored marsh within the Merritt Island National Wildlife Refuge in July 2001 and were sampled after 42 days of incubation. Suspensions produced by blending in 0.85% NaCl were diluted 1:10 into PBMS (to reduce the particulate content, yielding an inoculum density of ∼1.5 × 106 cells ml−1) either with or without (NH4)2SO4 to evaluate N effects on substrate response.
The fluorescence response to all of the substrates with the addition of nitrogen followed the same logarithmic increase to a peak succeeded by a rapid decrease to baseline levels, as observed in the previous rhizosphere testing at the 50-ppm concentrations of surfactants. The time to peak ranged from 8 to 18 h for all of the substrates tested, except phenylalanine, which showed a lag in maximum response of 45 h, and peak response ranged from 1.4 to 8.2 NRFU (Table 4). The removal of nitrogen from the test media eliminated the response to all the non-N-containing substrates but caused no consistent change in response to the amino acids, as would be predicted under nitrogen-limiting conditions.
TABLE 4.
Fluorescent response from marsh litter samplesa
| Type of carbon source | Carbon source (concn in mM) | Results
|
|||
|---|---|---|---|---|---|
| With Nitrogen
|
Without Nitrogen
|
||||
| Peak (NRFU) | Time to peak (h) | Peak (NRFU) | Time to peak (h) | ||
| Carbohydrate | Fructose (1) | 3.64 | 14.0 | 1.11 | 63.5 |
| 5.74 | 9.25 | 1.06 | 68.0 | ||
| Glucose (1) | 7.38 | 11.0 | 1.04 | 68.0 | |
| 4.01 | 7.45 | 1.06 | 68.0 | ||
| Cellobiose (0.5) | 8.14 | 13.25 | 1.04 | 68.0 | |
| 8.23 | 11.25 | 1.01 | 68.0 | ||
| Rhamnose (1) | 3.36 | 17.75 | 1.08 | 68.0 | |
| 3.32 | 16.15 | 1.04 | 68.0 | ||
| Amino acid | Glutamic acid (1) | 1.75 | 13.00 | 4.26 | 11.75 |
| 1.58 | 10.25 | 2.15 | 10.75 | ||
| Asparagine (1) | 1.67 | 12.0 | 1.76 | 11.0 | |
| 1.77 | 9.25 | 1.58 | 9.50 | ||
| Phenylalanine (1) | 4.38 | 45.75 | 5.71 | 46.00 | |
| 8.01 | 44.75 | 2.62 | 45.00 | ||
| Alanine (1) | 2.39 | 18.00 | 2.11 | 17.50 | |
| 3.53 | 13.00 | 2.04 | 12.50 | ||
| Organic acid | Glycolic acid (1) | 1.53 | 13.75 | 1.06 | 68.00 |
| 1.64 | 12.25 | 1.02 | 68.00 | ||
| Acetate (1) | 1.48 | 11.25 | 1.09 | 68.00 | |
| 1.42 | 8.25 | 1.06 | 68.00 | ||
Peak values are reported in NRFU. Two replicates were exposed to each carbon source both with and without the addition of 2.5 mg of [N]NH4 per liter in the media.
These initial data indicate that the BD microplates offer an approach for CLPP with lower potential for selective enrichment than presently employed technology possesses. Detectable responses were observed at substrate addition levels 10 to 100 times less than those found in CLPP approaches based on either CO2 monitoring (1) or redox dyes (6). The time-to-peak response was consistently less than 20 h and as short as 5 to 6 h, indicating that a standard incubation time of 24 h could be employed. This incubation time is shorter than the 2- to 3-day assay period typically used for CLPP based on Biolog plates but is longer than the several-hour incubation used in the CO2-monitoring approach. The BD plates allow for complete definition and concomitant manipulation of chemical factors such as N levels that may affect substrate utilization. Further testing with bottom-reading fluorometry may allow for testing of intact communities (e.g., biofilm coupons and small litter bags) compared to suspensions of organisms as in the present study. While such extensions of the assay will not allow for estimation of in situ rates of substrate utilization feasible with radiotracer approaches, they will provide a CLPP approach with increasing physiological relevance. Depending on the research question, microbial ecologists can then choose CLPP approaches targeted at phenotypic potential and/or phenotypic expression.
Acknowledgments
Funding for this project was provided by the National Aeronautics and Space Administration Office of Physical and Biological Research as part of its Advanced Life Support project.
REFERENCES
- 1.Degens, B. P., L. A Schipper, G. P. Sparling, and L. C. Duncan. 2001. Is the microbial community in a soil with reduced catabolic diversity less resistant to stress or disturbance? Soil Biol. Biochem. 33:1143-1153. [Google Scholar]
- 2.Di Giovanni, G. D., L. S. Watrud, R. J. Seidler, and F. Widmer. 1999. Fingerprinting of mixed bacterial strains and BIOLOG gram-negative (GN) substrate communities by enterobacterial repetitive intergenic consensus sequence-PCR (ERIC-PCR). Curr. Microbiol. 38:217-223. [DOI] [PubMed] [Google Scholar]
- 3.Garland, J. L., and R. M. Lehman. 1999. Dilution/extinction of community phenotypic characters to estimate relative structural diversity in mixed communities. FEMS Microbiol. Ecol. 30:333-343. [DOI] [PubMed] [Google Scholar]
- 4.Garland, J. L., L. H. Levine, N. C. Yorio, J. L. Adams, and K. L. Cook. 2000. Graywater processing in recirculating hydroponic systems: phytotoxicity, surfactant degradation, and bacterial dynamics. Wat. Res. 12:3075-3086. [Google Scholar]
- 5.Garland, J. L. K. L. Cook, C. A. Loader, and B. A. Hungate. 1997. The influence of microbial community structure and function and community-level physiological profiles, p. 171-183. In H. Insam and A. Rangger (ed.), Microbial communities: functional versus structural approaches. Springer-Verlag KG Berlin, Berlin, Germany.
- 6.Garland, J. L., and A. L. Mills. 1991. Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level sole-carbon-source utilization. Appl. Environ. Microbiol. 57:2351-2359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hobbie, J. E., R. J. Daley, and S. Jasper. 1977. Use of Nuclepore filters for counting bacteria by fluorescence microscopy. Appl. Environ. Microbiol. 33:1225-1228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Konopka, A., L. Oliver, and R. F. Turco. 1998. The use of carbon substrate utilization patterns in environmental and ecological microbiology. Microb. Ecol. 35:103-115. [DOI] [PubMed] [Google Scholar]
- 9.Lehman, R. M., F. S. Colwell, D. B. Ringelberg, and D. C. White. 1995. Combined microbial community-level analyses for quality assurance of terrestrial subsurface cores. J. Microbiol. Methods 22:263-281. [Google Scholar]
- 10.Levine, L. H., J. L. Garland, and J. V. Johnson. 2002. HPLC/ESI-quadrupole ion trap mass spectrometry for characterization and direct quantification of amphoteric and nonionic surfactants in aqueous samples. Anal. Chem. 74:2064-2071. [DOI] [PubMed] [Google Scholar]
- 11.Levine, L. H., J. E. Judkins, and J. L. Garland. 2000. Determination of anionic surfactants during wastewater recycling process by ion pair chromatography with suppressed conductivity detection. J. Chromatogr. 874:207-215. [DOI] [PubMed] [Google Scholar]
- 12.Mills, A. L., and J. L. Garland. 2002. Application of physiological profiles to assess community properties, p. 135-146. In C. J. Hurst, R. L. Crawford, G. R. Knudsen, M. J. McInerney, and L. D. Stetzenbach (ed.), Manual of environmental microbiology. ASM Press, Washington, D.C.
- 13.Reid, B. J., C. J. A. MacLeod, P. H. Lee, A. W. J. Morriss, J. D. Stokes, and K. T. Semple. 2001. A simple 14C-respirometric method for assessing microbial catabolic potential and contaminant bioavailability. FEMS Microbiol. Lett. 191:141-146. [DOI] [PubMed] [Google Scholar]
- 14.Smalla, K., U. Wachtendorf, H. Heuer, W.-T. Liu, and L. Forney. 1998. Analysis of BIOLOG GN substrate utilization patterns by microbial communities. Appl. Environ. Microbiol. 64:1220-1225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Swisher, R. D. 1987. Surfactant biodegradation, 2nd ed. Marcel Dekker, New York, N.Y.
- 16.Winding, A., and N. B. Hendriksen. 1997. Biolog substrate utilization assay for metabolic fingerprints of soil bacteria: incubation effects, p. 195-205. In H. Insam and A. Rangger (ed.), Microbial communities: functional versus structural approaches. Springer-Verlag KG Berlin, Berlin, Germany.
- 17.Wodnicka, M., R. D. Guarino, J. J. Hemperly, M. R. Thomas, D. Stitt, and J. B. Pitner. 2000. Novel fluorescent technology platform for high throughput cytotoxicity and proliferation assays. J. Biomol. Screen. 5:141-152. [DOI] [PubMed] [Google Scholar]