Abstract
Six strains of Salmonella enterica serovar Typhi which were resistant to ampicillin, chloramphenicol, trimethoprim-sulfamethoxazole, streptomycin, tetracycline, and gentamicin were isolated in Korea. This multidrug resistance was transferred by a conjugative plasmid of about 50 kb. The plasmid harbored a class 1 integron, which included six resistance genes, aacA4b, catB8, aadA1, dfrA1, aac(6′)-IIa, and the novel blaP2, in that order. All of the isolates showed the same-size plasmids and the same ribotyping patterns, which suggests a clonal spread of these multidrug-resistant isolates.
The multidrug-resistant (MDR) Salmonella enterica serovar Typhi, which is resistant to the first-line antimicrobials, has become an important public health problem in Southeast Asia (12). In 2001, MDR Salmonella serovar Typhi strain CT18's resistance-determining plasmid pHCM1 was completely sequenced (9). The plasmid is 218,150 bp in size and belongs to incompatibility group H1. The clinically relevant resistance genes identified in pHCM1 were dfrA14 (trimethoprim), sul-2 (sulfonamide), catA1 (chloramphenicol), blaTEM-1 (ampicillin), and strAB (streptomycin).
The integrons found in clinical isolates of gram-negative bacteria generally contain one or more integrated cassettes, each including an antibiotic resistance gene, and a large number of gene cassettes containing different resistance genes have been identified (11). Although there are three distinct classes of integrons, those belonging to class 1 often contain an antimicrobial resistance gene in clinical isolates (11). For Salmonella serovar Typhi, the dfrA14 gene in pHCM1 was an integron gene cassette (9, 10).
In this report, we describe the MDR Salmonella serovar Typhi in which the multidrug resistance is mediated by a complex class 1 integron, including six resistance gene cassette.
Salmonella serovar Typhi isolates 552, 607, 703, 768, 2301, and 2306 were isolated from the blood of six patients with typhoid fever in 1999. Among the six isolates, five were isolated in April 1999 and one was isolated in July 1999. The residences of the patients from whom the strains were isolated were in different provinces: three in Kyongki, one in Kyeongnam, one in Kangwon, and one in Chungnam. The strains were tested for susceptibility to antimicrobials by a controlled disk diffusion technique according to the guidelines provided by the National Committee for Clinical Laboratory Standards (7). Six isolates were resistant to ampicillin, chloramphenicol, trimethoprim-sulfamethoxazole, tetracycline, streptomycin, gentamicin, and tobramycin, but all isolates were sensitive to amikacin and ciprofloxacin. Five of the six isolates were sensitive to nalidixic acid; isolate 552 was the exception.
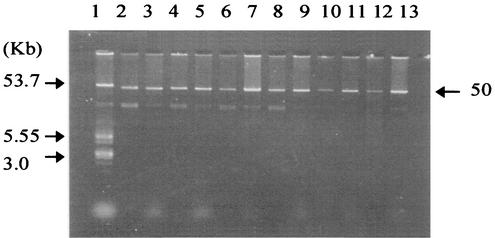
To test the transmissibility of the multidrug resistance of the isolates, a conjugation experiment was performed using Escherichia coli J53 Azir as a recipient and a selective medium containing 100 μg each of sodium azide and ampicillin/ml (8). With the transfer of ampicillin resistance, resistances againsttrimethoprim-sulfamethoxazole, chloramphenicol, tetracy-cline, streptomycin, gentamicin, and tobramycin were cotransferred in all six isolates, but nalidixic acid resistance was not achieved with the conjugation. Upon electrophoresis of plasmid DNAs, the transconjugants were found to acquire multidrug resistance via a plasmid with a size of about 50 kb (Fig. 1).
FIG. 1.
Agarose gel electrophoresis of the plasmid DNAs. Lane 1, plasmid size marker from E. coli V517; lane 2, Salmonella serovar Typhi isolate 552; lane 3, E. coli J53(pST552); lane 4, Salmonella serovar Typhi isolate 607; lane 5, E. coli J53(pST607); lane 6, Salmonella serovar Typhi isolate 703; lane 7, E. coli J53(pST703); lane 8, Salmonella serovar Typhi isolate 768; lane 9, E. coli J53(pST768); lane 10, Salmonella serovar Typhi isolate 2301; lane 11, E. coli J53(pST2301); lane 12, Salmonella serovar Typhi isolate 2306; lane 13, E. coli J53(pST2306).
To amplify the genes contained in the class 1 integron, the following oligonucleotide primers were synthesized and used for PCR: hep 58, 5′-TCA TGG CTT GTT ATG ACT GT-3′ (5′ upstream conserved sequences [5′-CS] of int-11), and hep 59, 5′-GTA GGG CTT ATT ATG CAC GC-3′ (3′ upstream CS [3′-CS] of qacEΔ1). The PCR amplification was carried out with the six isolates and their transconjugants as described elsewhere (17). Consequently, 4,686-bp PCR products were amplified from all isolates and their transconjugants. By using the PCR-Script+ Cam cloning kit (Stratagene, La Jolla, Calif.), the purified PCR product from strain 2301 was cloned into vector plasmid pBC SK, which encoded chloramphenicol resistance (pCST2301). This plasmid was then introduced into E. coli Epicurian Coli XL10-Gold ultracompetent cells (Stratagene) by chemical transformation (5). After selection with 30 μg of chloramphenicol/ml, the colonies were screened for insert size. With primers T3 and T7, which matched the nucleotides at the flanking regions of the multiple-cloning site in the vector, cycle sequencing (Perkin-Elmer Cetus, Norwalk, Conn.) of the DNA was initiated and continued by primer walking. The coding sequences [of aacA4b, dfrA1, aac(6′)-IIa, and blaP2] were confirmed by direct sequencing of PCR fragments amplified with the wild strain as a template by using the following primers: F211, 5′-ATG ACC TTG CGA TGC TCT-3′; B731, 5′-TCG ATG GAA GGG TTA GGC-3′; F2356, 5′-CTG AGG AAG AAT TGT GAA AC-3′; B2844, 5′-TGT TAA CCC TTT TGC CAG AT-3′; F2936, 5′-CAG AGC GAC CAT TTC ATG T-3′; B3563, 5′-AAT ATG GCT AAC ACC TGA AT-3′; F3602, 5′-CAA AAT TAT GGG GTT ACT TAC-3′; and B4561, 5′-TTG TTA GCC TTA TCA GCG CG-3′. The nucleotide sequence is available at GenBank (accession number AY123251).
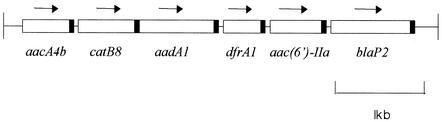
The 4,686-bp sequences contained cassettes with the following six resistance genes: aacA4b (4), catB8 (GenBank accession number AF227506), aadA1 (14), dfrA1 (2), aac(6′)-IIa (13), and the novel blaP2 (3), in that order. All six resistance genes had complete coding sequences and 59-base elements at the downstream end of each gene (Fig. 2). The β-lactamase encoded by blaP2 was a novel enzyme, and the deduced amino acid sequence was different in one amino acid from that of the N29 β-lactamase (pCS229-encoded carbenicillinase) identified in Proteus mirabilis, Val→Ile at amino acid 90 (3). Crude β-lactamase preparations from clinical isolates or their transconjugants were obtained as described previously (8), and isoelectric focusing was performed by the method of Matthew et al. (6). The pI value of the novel β-lactamase was 6.9, which was identical to that of N29 β-lactamase (3). In order to exclude the presence of TEM-related β-lactamase outside of the integron in the plasmid, a TEM-specific PCR was performed as described previously (8), and a negative result was obtained.
FIG. 2.
Arrangement of antibiotic resistance genes of Salmonella serovar Typhi isolate 2301 as deduced from the nucleic acid sequence described in the present work. The black boxes represent the 59-base elements, and the arrows indicate the orientation of the genes.
Because the primers used in our study were at the 5′ and 3′ CS of the class 1 integron between int-11 and qacEΔ1, our clone did not contain the int-11 and qacEΔ1 or sul-1 genes but did contain the amplified resistance genes between the 5′ CS and the 3′ CS. The result is striking in that this novel integron possessed six resistance gene cassettes. It is known that, when more than one cassette is present, the position of the cassette in the array influences the level of antibiotic resistance expressed by the cassette gene and that the resistance level is highest when the gene is in the first cassette and is reduced by the presence of individual upstream cassettes (1). Although the level of gene expression for this long operon was not studied, the resistance patterns of the isolates and their transconjugants and the results of the isoelectric focusing study showing the pI value of blaP2 (from the most distal gene cassette) suggested the expression of all of the resistance genes.
The wild strains and their transconjugants were resistant to tetracycline, but the tetracycline resistance gene was not included in our clone. To observe the structure of the resistance region of the plasmid, further analysis of the rest of the plasmid is needed.
In order to observe the clonal relationship of the six isolates, molecular typing was performed by using a ribotyping method, as described previously (8, 15). Six MDR isolates had eight bands on ribotyping and showed the same patterns. The same ribotyping and plasmid patterns of the six isolates suggested a clonal spread, although the residences of the patients from whom these strains were isolated were in different provinces throughout the Korean Peninsula. We speculate that at least five of six patients who developed the disease in April had eaten contaminated food that was distributed by the same company or that was served at the same restaurant, because the Republic of Korea is a small country, within a 24-h driving distance.
It is of great interest that this highly efficient assembly of gene cassettes was found in Salmonella serovar Typhi, a pathogen of community-acquired infection. Because human carriers are the source of Salmonella serovar Typhi infection, we suspect that the resistance plasmid might have been transferred from other bacteria to Salmonella serovar Typhi in the human carriers. The prevalence of Salmonella serovar Typhi and other Salmonella strains harboring this resistance plasmid in Korea has not yet been studied. However, we were unable to obtain an amplified PCR product with the same primers from several MDR Salmonella serovar Typhi strains which had different antibiograms and which had been isolated in different years.
Considering the spread of MDR Salmonella serovar Typhi in Southeast Asia and of Salmonella serovar Typhimurium DT104 in the United States and Europe (12, 16), this MDR isolate might be a serious threat to public health. Thus, further study of the prevalence of this MDR Salmonella isolate should be carried out and the occurrence of MDR isolates with similar antibiograms should be carefully monitored.
Acknowledgments
This study was supported by a grant (01-PJ1-PG3-20500-0064) from the Korea Health 21 R&D Project, Ministry of Health and Welfare, Seoul, Republic of Korea.
REFERENCES
- 1.Collis, C. M., and R. M. Hall. 1995. Expression of antibiotic resistance genes in the integrated cassettes of integrons. Antimicrob. Agents Chemother. 39:155-162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Fling, M. E., and C. Richards. 1983. The nucleotide sequence of the trimethoprim-resistant dihydrofolate reductase gene harbored by Tn7. Nucleic Acids Res. 11:5147-5158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ito, Y., and T. Hirano. 1997. Carbenicillin-hydrolysing penicillinase mediated by a plasmid of Proteus mirabilis and its relationship to the PSE-type enzymes of Pseudomonas aeruginosa. J. Appl. Microbiol. 83:175-180. [DOI] [PubMed] [Google Scholar]
- 4.Lambert, T., M. C. Ploy, and P. Courvalin. 1994. A spontaneous point mutation in the aac(6′)-Ib gene results in altered substrate specificity of aminoglycoside 6′-N-acetyltransferase of a Pseudomonas aeruginosa strain. FEMS Microbiol. Lett. 115:297-304. [DOI] [PubMed] [Google Scholar]
- 5.Maniatis, T., E. F. Fritsch, and J. Sambrook. 1982. Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
- 6.Matthew, M., M. Harris, M. J. Marshall, and G. W. Rose. 1975. The use of analytical isoelectric focusing for detection and identification of β-lactamases. J. Gen. Microbiol. 88:169-178. [DOI] [PubMed] [Google Scholar]
- 7.National Committee for Clinical Laboratory Standards. 1984. Performance standards for antimicrobial disk susceptibility test, vol. 4, p. 369-406. M2-M3, 3rd ed. National Committee for Clinical Laboratory Standards, Villanova, Pa.
- 8.Pai, H., S. Lyu, J. H. Lee, J. Kim, Y. Kwon, J.-W. Kim, and K. W. Choe. 1999. Survey of extended-spectrum β-lactamases in clinical isolates of Escherichia coli and Klebsiella pneumoniae: prevalence of TEM-52 in Korea. J. Clin. Microbiol. 37:1758-1763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Parkhill, J., et al. 2001. Complete genome sequence of a multidrug resistant Salmonella enterica serovar Typhi CT18. Nature 413:848-852. [DOI] [PubMed] [Google Scholar]
- 10.Partridge, S. R., G. D. Recchia, H. W. Stokes, and R. M. Hall. 2001. Family of class 1 integrons related to In4 from Tn1696. Antimicrob. Agents Chemother. 45:3014-3020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Recchia, G. D., and R. M. Hall. 1995. Gene cassettes: a new class of mobile element. Microbiology 141:3015-3027. [DOI] [PubMed] [Google Scholar]
- 12.Rowe, B., L. R. Ward, and E. J. Threlfall. 1997. Multidrug-resistant Salmonella Typhi: a worldwide epidemic. Clin. Infect. Dis. 24(Suppl. 1):S106-S109. [DOI] [PubMed] [Google Scholar]
- 13.Shaw, K. J., C. A. Cramer, M. Rizzo, R. Mierzwa, K. Gewain, G. H. Miller, and R. S. Hare. 1989. Isolation, characterization, and DNA sequence analysis of an AAC(6′)-II gene from Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 33:2052-2062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Sundstrom, L., P. Radstrom, G. Swedberg, and O. Skold. 1988. Site-specific recombination promotes linkage between trimethoprim- and sulfonamide resistance genes. Sequence characterization of dhfrV and sulI and a recombination active locus of Tn21. Mol. Gen. Genet. 213:191-201. [DOI] [PubMed] [Google Scholar]
- 15.Tarkka, E., H. Ahman, and A. Siitonen. 1994. Ribotyping as an epidemiologic tool for Escherichia coli. Epidemiol. Infect. 112:263-274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Threlfall, E. J. 2002. Antimicrobial drug resistance in Salmonella: problems and perspectives in food- and water-borne infections. FEMS Microbiol. Rev. 26:141-148. [DOI] [PubMed] [Google Scholar]
- 17.White, P. A., C. J. McIver, Y. M. Deng, and W. D. Rawlinson. 2000. Characterization of two new gene cassettes, aadA5 and dhrA17. FEMS Microbiol. Lett. 182:265-269. [DOI] [PubMed] [Google Scholar]