Abstract
Using model strains in which we ectopically express the cyanobacterial clock protein KaiC in cells from which the clock genes kaiA, kaiB and/or kaiC are deleted, we found that some features of circadian clocks in eukaryotic organisms are conserved in the clocks of prokaryotic cyanobacteria, but others are not. One unexpected difference is that the circadian autoregulatory feedback loop in cyanobacteria does not require specific clock gene promoters as it does in eukaryotes, because a heterologous promoter can functionally replace the kaiBC promoter. On the other hand, a similarity between eukaryotic clock proteins and the cyanobacterial KaiC protein is that KaiC is phosphorylated in vivo. The other essential clock proteins KaiA and KaiB modulate the status of KaiC phosphorylation; KaiA inhibits KaiC dephosphorylation and KaiB antagonizes this action of KaiA. Based upon an analysis of clock mutants, we conclude that the circadian period in cyanobacteria is determined by the phosphorylation status of KaiC and also by the degradation rate of KaiC. These observations are integrated into a model proposing rhythmic changes in chromosomal status.
Keywords: circadian/clock/cyanobacteria/degradation/phosphorylation
Introduction
Circadian (daily) biological clocks are found throughout the eukaryotes, where they help to keep track of the daily cycle. At least one group of prokaryotes, the eubacterial cyanobacteria, are also circadian timekeepers (Johnson et al., 1996). The fundamental properties of circadian clocks in eukaryotes and in cyanobacteria are the same: surprisingly precise self-sustained oscillations with an approximately 24 h period that are temperature compensated and entrainable by environmental cycles. Are the mechanisms that accomplish this timing similar in diverse organisms, or have many different solutions to the challenge of circadian timekeeping evolved? The comparison of the molecular bases of circadian clocks between cyanobacteria and eukaryotes can address this question.
In eukaryotes, a web of evidence supports a model that proposes autoregulatory feedback loops of central clock gene expression (Hardin et al., 1990; Dunlap, 1999; Young and Kay, 2001). In Drosophila, mammals and Neurospora, where the model is most highly developed, there is an ensemble of different molecular components that are believed to participate in the feedback loop. For example, in Drosophila and mammals, the clock proteins PERIOD, TIMELESS and CRYTOCHROME negatively feed back upon the per, tim and cry promoters, while CLOCK functions as a positive element (Darlington et al., 1998; Gekakis et al., 1998; Kume et al., 1999; Young and Kay, 2001; Reppert and Weaver, 2002). In Neurospora, the clock protein FRQ represses its own transcription (Dunlap, 1999). There are rhythms in the abundance of eukaryotic clock proteins, and the turnover/stability of these proteins is likely to be an important determinant of circadian period (Ruoff et al., 1996). Eukaryotic clock proteins such as PER and FRQ are post-translationally modified by phosphorylation as a prelude to degradation, and modulating the phosphorylation of FRQ and PER concomitantly alters the circadian period (Liu et al., 2000; Görl et al., 2001; Kloss et al., 2001; Young and Kay, 2001; Reppert and Weaver, 2002).
In eukaryotes, ∼5–10% of genes appear to be regulated by the circadian clockwork; in contrast, essentially all promoters are rhythmically regulated by the circadian clock in the prokaryotic cyanobacterium Synechococcus elongatus PCC 7942 (Liu et al., 1995). A cluster of three genes in S.elongatus, named kaiA, kaiB and kaiC, encodes essential circadian clock components (Ishiura et al., 1998). A promoter for the kaiA gene (kaiAp) gives rise to a monocistronic kaiA mRNA, whereas the kaiBC promoter (kaiBCp) produces a dicistronic kaiBC mRNA (Ishiura et al., 1998). Even though there are no sequence similarities between the kai genes and any known eukaryotic clock genes, some features of kai gene regulation are reminiscent of PER and FRQ regulation. The kaiA and kaiBC transcripts (Ishiura et al., 1998) and the KaiB and KaiC proteins are rhythmically abundant (Xu et al., 2000; Iwasaki et al., 2002). The KaiA, KaiB and KaiC proteins interact with each other, and KaiC can be phosphorylated (Iwasaki et al., 1999, 2002; Nishiwaki et al., 2000; Williams et al., 2002). Continuous overexpression of KaiC repressed kaiBCp, suggesting negative feedback of KaiC on its promoter, while pulses of KaiC expression reset the phase of the rhythms (Ishiura et al., 1998; Xu et al., 2000).
Taken together with analogous data from eukaryotic clocks, these results have been used to support a hypothesis wherein there is negative feedback control of kaiBC expression by the KaiC protein acting upon kaiBCp to generate a circadian oscillation in which the stability of KaiC is influenced by its phosphorylation status and thereby modulates period length (Ishiura et al, 1998; Nishiwaki et al., 2000). This hypothesis implies that the biochemistry of the clockwork in cyanobacteria is similar to that in eukaryotes and predicts that (i) rhythmicity is dependent upon kaiBCp function, (ii) the status of KaiC phosphorylation in vivo alters clock properties and (iii) the rate of KaiC degradation will influence period. While testing those predictions, we unexpectedly found that the kaiBC promoter is not specifically required for the circadian oscillator to operate in cyanobacteria. KaiA inhibits the dephosphorylation of KaiC and KaiB antagonizes KaiA’s effect. Clock speed is influenced by both the extent of KaiC phosphorylation and the rate of KaiC degradation. These results are integrated into a model proposing rhythmic changes in chromosomal status that are regulated by a KaiC-containing protein complex.
Results
kaiBCp is not required for circadian rhythmicity
The rationale of the experimental strategy described in this paper was to express KaiC in a pulsatile fashion under the control of the derepressible trc promoter (trcp) and measure the degradation of KaiC after removal of the inducer (IPTG). We expressed wild-type KaiC (KaiCWT) in strains in which other genes have been deleted (e.g. kaiA, kaiB, cikA, sasA). Alternatively, we expressed versions of KaiC that confer different circadian periods (KaiCWT or the mutant KaiCC22a and KaiCC55a) in the same genetic background that lacks the endogenous kaiC gene, namely ΔkaiC (Figure 1A). The original goal was to ascertain parameters that might affect the degradation rate of KaiC and thereby affect the circadian period. This strategy was successful towards that goal, but in addition we made two unexpected observations: kaiBCp is not specifically required for clock function and the status of KaiC phosphorylation in vivo is dependent upon KaiA and KaiB.
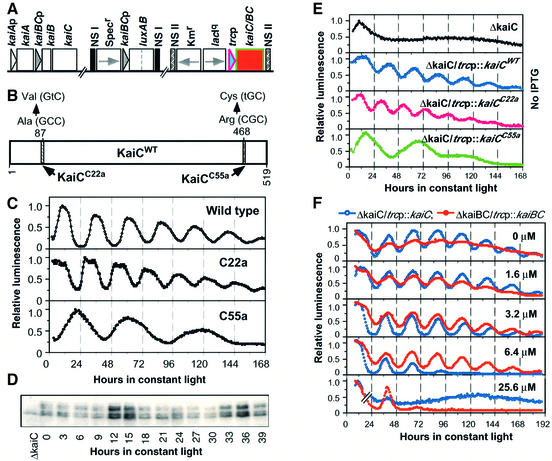
Fig. 1. kaiBCp can be functionally replaced by a heterologous promoter. (A) A generalized diagram for the strains used in this paper. Three sites in the chromosome are shown: (i) the wild-type kaiABC cluster (kaiABCWT); (ii) the kaiBCp::luxAB or psbAIp::luxAB reporters in neutral site I (NS I); and (iii) an IPTG-derepressible promoter driving expression of KaiC or KaiB and KaiC together (BC) in NS II, shown as the red box. Specr, spectinomycin resistance gene; Kmr, kanamycin resistance gene; lacIq, a lactose repressor gene; trcp, an IPTG-derepressible heterologous promoter (Amann et al., 1988). Many of the strains used in this paper had in-frame deletions of kaiA, kaiB and/or kaiC, as described in Materials and methods. For example, the ΔkaiC/trcp::kaiC strain has an in-frame deletion of the endogenous kaiC gene with the trcp::kaiC construct in the ectopic NS II. Boxes denote the coding regions and triangles represent the promoter sequences. Arrows indicate the transcriptional directions. (B) The KaiC protein (KaiCWT) sequence, indicating the locations of the mutations conferring long-period (KaiCC55a) and short-period (KaiCC22a) phenotypes. (C) Bioluminescence rhythms in wild-type, C22a and C55a strains. (D) Immunoblot showing rhythm of KaiC abundance in the ΔkaiC/trcp::kaiCWT strain in constant light in the presence of 0.5 µM IPTG. (E) Recovery of rhythmicity in the KaiC-null strains (ΔkaiC). Top trace: loss of rhythmicity in ΔkaiC. Traces 2–4: recovery of rhythmicity in ΔkaiC cells transformed with trcp::kaiC in the absence of the inducer, IPTG. Trace 1 is ΔkaiC/trcp::kaiCWT resulting in a wild-type period, trace 2 is ΔkaiC/ trcp::kaiCC22a resulting in a short period, and Trace 3 is ΔkaiC/trcp::kaiCC55a resulting in a long period. (F) Titration of rhythmicity in the ΔkaiC/trcp::kaiCWT (blue) and ΔkaiBC/trcp::kaiBCWT (red) strains with IPTG dose. The concentration of IPTG (µM) is shown in the upper right corner of each panel. For (C–F), luminescence expression was monitored in constant light (LL) and the maximum levels of luminescence were normalized to 1. Reporter constructs were psbAIp::luxAB for (C) and kaiBCp::luxAB for (E) and (F).
As described in the Introduction, previous results supported a model of autoregulatory feedback upon clock gene promoters in cyanobacteria, as shown by the fact that the essential clock protein KaiC represses its own transcription when KaiC is overexpressed (Ishiura et al., 1998). But is that phenomenon due to negative feedback specifically upon the kaiBC promoter, as would be expected if the cyanobacterial clockwork were organized like those in eukaryotes? While testing the temporal gene expression patterns of strains with in-frame deletions in the kai genes, we found that the promoter which normally drives kaiC expression (kaiBCp) is not specifically required for rhythmic clock gene expression.
The promoters for the kaiBC and psbAI genes are both rhythmically active, as monitored with the kaiBCp::luxAB and/or psbAIp::luxAB reporters (Ishiura et al., 1998). The patterns for psbAIp::luxAB reporter expression in wild type (period ≈25 h), the short period mutant C22a (period ≈22 h) and the long period mutant C55a (period ≈55 h) are shown in Figure 1C (mutation sites depicted in Figure 1B). In-frame deletion of the kaiC gene (ΔkaiC) abolishes rhythmic activity of the kaiBCp::luxAB reporter (top panel of Figure 1E). Unexpectedly, rhythmicity of the ΔkaiC strain is restored when the kaiC gene is replaced under the control of the heterologous, derepressible trc promoter that is derived from Escherichia coli (Amann et al., 1988; right part of Figure 1A). As shown in the bottom three panels of Figure 1E, the kaiBCp::luxAB reporter is rhythmically active when KaiC is expressed under the control of trcp (as is the psbAIp::luxAB reporter; data not shown). This effect is observed for expression of KaiCWT and of the mutants KaiCC22a and KaiCC55a in the ΔkaiC strain, where the period-specific effects of the mutant KaiCs are preserved when driven from trcp.
The data shown in Figure 1E were monitored from cells in the absence of the inducer, IPTG. We have found that trcp is slightly ‘leaky’, so that there is a measurable amount of KaiC expressed in the absence of IPTG (not shown). A small induction of KaiCWT in the ΔkaiC/trcp::kaiCWT strain above this level with low concentrations of IPTG (0–5 µM) also allows sustained rhythmicity, but higher concentrations of IPTG dampen (e.g. 6.4 µM) and then abolish (≥25.6 µM) rhythmicity (Figure 1F). The observation that KaiC overexpression by higher concentrations of IPTG abolishes rhythmicity is equivalent to that previously published for overexpression of KaiC in cells with an intact kaiABC operon (Ishiura et al., 1998), and indicates that KaiC expressed from the trc promoter is active even though it is transcribed away from its normal site. Even more importantly, the loss of rhythmicity at higher IPTG concentrations demonstrates that the autoregulatory repression of kaiC transcription does not require that the endogenous kaiC promoter drive KaiC expression. Overexpression of KaiCC22a and KaiCC55a also abolishes rhythmicity (not shown). Immunoblot analysis confirmed that KaiCWT protein is rhythmically abundant in the ΔkaiC/trcp::kaiCWT strain (Figure 1D), as had been found previously in wild-type cells (Xu et al., 2000). Therefore, not only is the kaiBC promoter rhythmically active when KaiC is expressed from a heterologous promoter, but KaiC turns over cyclically.
The results with the ΔkaiC/trcp::kaiCWT strain suggested that the kaiBC promoter was not necessary for rhythmic expression. However, in this strain, KaiB is still expressed under the control of kaiBCp. To eliminate the possibility that kaiBCp was needed only to drive KaiB expression, we created a ΔkaiBC/trcp::kaiBC strain in which kaiB and kaiC were both deleted from their original site and expressed ectopically from the heterologous trcp. Similar results were obtained with the ΔkaiBC/trcp::kaiBC strain as with the ΔkaiC/trcp::kaiC strain, except that slightly higher concentrations of IPTG were required to elicit robust rhythmicity (Figure 1F). At IPTG concentrations of 50 µM and higher, rhythmicity was abolished in the ΔkaiBC/trcp::kaiBC strain (not shown). Similar results were obtained by Y.Nakahira and T.Kondo (personal communication) with a slightly different construct. These data show that kaiBCp is not specifically required for rhythmic expression in cyanobacteria.
In vivo degradation of different KaiC alleles
The observation that KaiC is rhythmically abundant and that pulses of KaiC expression elicit phase resetting led to the hypothesis that KaiC turnover might be an important determinant of clock period and phase (Xu et al., 2000). To test this hypothesis, we investigated KaiC stability. The rationale of our approach was to express KaiC as pulses in ΔkaiC strains and to monitor the degradation of the induced KaiC. We chose this approach over the more common one of monitoring degradation in the presence of transcriptional or translational inhibitors (e.g. chloramphenicol) because in the long time-course experiments we intended to use, those inhibitors might also affect the turnover of proteases (or kinases/phosphatases) and thereby alter the normal degradation rate. Figure 2 illustrates that this method was successful; after a 3 h pulse of the inducer IPTG followed by washout, KaiC was degraded in vivo (Figure 2A). The concentration of IPTG that we used (100 µM) induced KaiC levels that were ∼10-fold higher than normal endogenous levels (Xu et al., 2000).
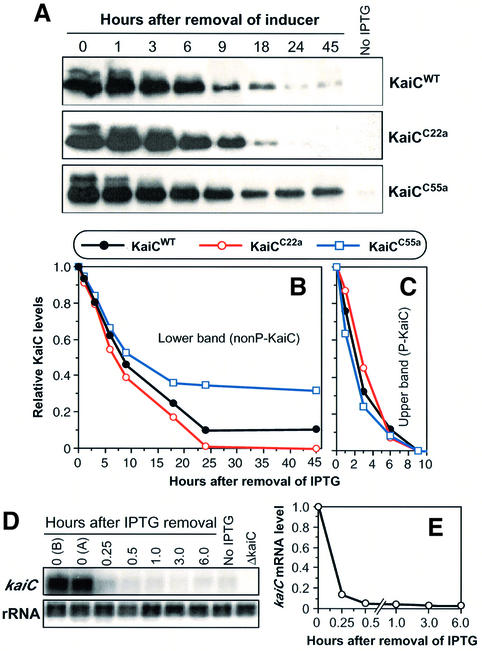
Fig. 2. KaiC pulse/degradation assay for wild-type and mutant KaiCs. (A) Degradation of wild-type and mutant KaiCs following a 3 h induction of KaiC expression. After a 3 h pulse of the inducer IPTG in light, the levels of KaiC proteins were determined at different times by immunoblotting. In each case, the strain was ΔkaiC into which the trcp::kaiC construct was inserted, creating the strains ΔkaiC/trcp::kaiCWT, ΔkaiC/ trcp::kaiCC55a or ΔkaiC/trcp::kaiCC22a. Top panel, wild-type KaiCWT; middle panel, short-period KaiCC22a; bottom panel, long-period KaiCC55a. (B and C) Densitometry of the KaiC bands shown in (A). (B) Lower bands, nonP-KaiC. (C) Upper bands, P-KaiC. The maximum levels were normalized to 1. (D) kaiC mRNA levels before and after IPTG washout in the trcp strains (data from ΔkaiC/trcp::kaiCWT is depicted, but all strains were equivalent). Northern blot analysis shows the kaiC mRNA and rRNA (to confirm equal loading) signals at various times (in hours) after IPTG washout. Time-0 controls are ‘0 (B)’, which is after 3 h in IPTG but before IPTG washout, and ‘0 (A)’, which is immediately after IPTG washout (kept at 0°C). (E) Densitometry of the averages of kaiC mRNA degradation from seven different strains in the pulsed-KaiC assay. At this scale, the standard deviation bars are smaller than the diameter of the symbols. The maximum levels of kaiC mRNA were normalized to 1.
Two clear KaiC bands were visualized by immunoblotting. Figure 2A shows that the two bands disappear at different rates after the removal of IPTG in cells of the same background (ΔkaiC) expressing KaiCWT, KaiCC22a or KaiCC55a. As we will show by the data in Figure 3, the upper band is phosphorylated KaiC (P-KaiC), a conclu sion that was also reached by Iwasaki et al. (2002). Densitometric quantification of the immunoblots shown in Figure 2A are presented in Figure 2B and C. The upper band degrades at about the same rate for KaiCWT, KaiCC22a and KaiCC55a (Figure 2C), with the following half-lives: KaiCWT 2.0 h, KaiCC22a 2.3 h and KaiCC55a 1.5 h. On the other hand, there was a significant difference in the degradation rates of the lower band of wild-type versus mutant KaiCs (Figure 2B). The rate of degradation of the lower band in the first 18 h was log-linear, with the following half-lives: KaiCWT 8.8 h, KaiCC22a 6.7 h and KaiCC55a 13.9 h. After 18 h, the rate of degradation was no longer log-linear, but there is a large difference in the persistence of the ‘tail’ of the decline in KaiC levels among KaiCWT, KaiCC22a and KaiCC55a that correlates with the period of the clock in vivo (C22a < wild type < C55a; Figure 2B). Therefore, not only do pulses of KaiC degrade in vivo, but also the relative degradation rates of non-phosphorylated KaiC (nonP-KaiC) are correlated with the rates of the clock in the corresponding strains.
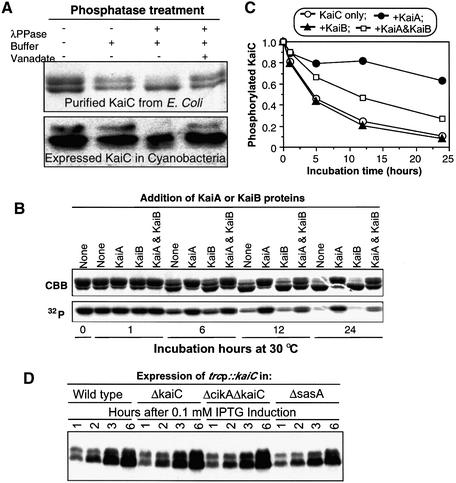
Fig. 3. KaiA and KaiB modulate KaiC dephosphorylation. (A) Treatment with lambda phosphatase. Upper panel, purified KaiC::His6 was incubated in vitro with or without lambda protein phosphatase (λPPase) in the presence or absence of the phosphatase inhibitor vanadate, followed by SDS–PAGE. Lower panel, the same treatment as in the upper panel, but the samples were extracts from cyanobacterial cells (strain ΔkaiC/trcp::kaiCWT expressing KaiC as in Figure 2A), and the signals were immunoblots using anti-KaiC antibody. (B) Dephosphorylation of purified KaiC::His6 in vitro (30°C). KaiC was autophosphorylated in vitro with [γ-32P]ATP, then allowed to dephosphorylate in the presence or absence of KaiA and/or KaiB. Following SDS–PAGE, KaiC was detected by Coomassie Brilliant Blue staining (CBB, upper panel) and autoradiography (32P, lower panel). Bands labeled with 32P correspond to the upper bands in CBB. (C) Quantification of the radioactive signals depicted in (B). The maximum levels of phosphorylated KaiC were normalized to 1. (D) In vivo phosphorylation of KaiC is independent of SasA and CikA. KaiCWT expression was induced by IPTG for 1, 2, 3 or 6 h in the following strains harboring the trcp::kaiCWT construct: wild type, ΔkaiC, ΔcikAΔkaiC or ΔsasA (assay performed in the light).
This estimation of the relative degradation rates of KaiC depends upon knowing the degradation rate of kaiC mRNA. We therefore estimated the rate of kaiC mRNA degradation by northern blot analyses under the same IPTG pulse/washout protocol in each of the trcp-driven KaiC strains shown in Figures 2, 4 and 5. The results for all the strains were almost exactly the same, with a kaiC mRNA half-life of 5–6 min. The data for the ΔkaiC/trcp::kaiCWT strain are shown in Figure 2D and E, where the half-life was 5.5 min. These mRNA half-lives are so short that they can be ignored in the estimation of KaiC protein half-life. Because these KaiCs have been overexpressed ∼10-fold relative to their usual intracellular concentrations, we cannot be sure that the absolute degradation rates of the mutant versus wild-type KaiCs calculated by this method are truly reflective of their normal in vivo half-lives. In this paper, the method used will be to estimate relative degradation rates, not absolute degradation rates. The fact that the degradation of pulses of KaiCWT versus KaiCC22a versus KaiCC55a is in the predicted order relative to period length (Figure 2) validates this method as a relative in vivo gauge of KaiC degradation rate.
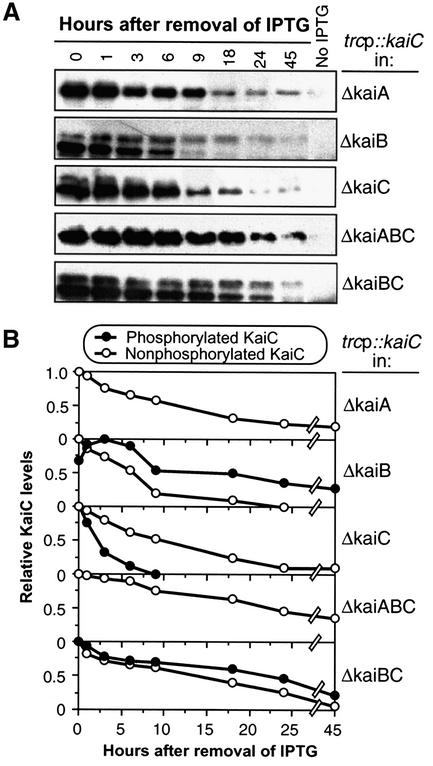
Fig. 4. KaiC phosphorylation/degradation is influenced by KaiA and KaiB in vivo. (A) KaiC degradation and phosphorylation assay was performed as in Figure 2A except that the KaiC was expressed in different kai-null backgrounds (ΔkaiA, ΔkaiB, ΔkaiC, ΔkaiABC or ΔkaiBC) harboring the trcp::kaiCWT construct. For comparison, the upper panel in Figure 2A (KaiCWT) is shown again, labeled here as ‘ΔkaiC’. (B) Degradation of phosphorylated and unphosphorylated KaiC in different genetic backgrounds quantified by densitometry. The maximum levels were normalized to 1.
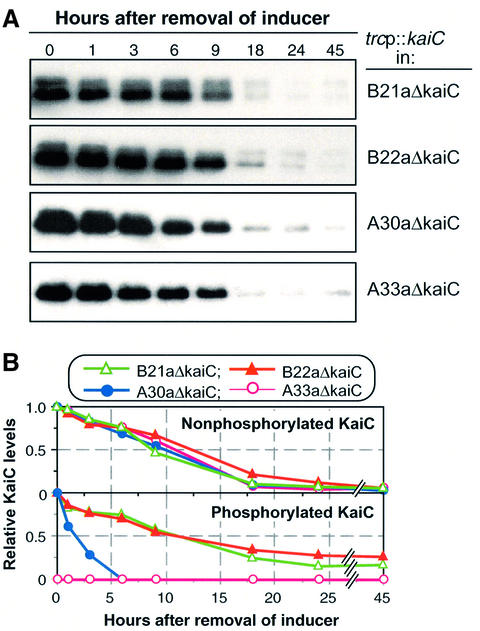
Fig. 5. KaiCWT phosphorylation/degradation in kaiA and kaiB period mutants. (A) KaiC degradation and phosphorylation assay was performed as in Figure 2A except that KaiCWT was expressed from trcp::kaiCWT in kaiA or kaiB period mutants (B21a, B22a, A30a or A33a) in which the endogenous kaiC gene was inactivated. (B) Quantification by densitometry of the nonP-KaiCWT (upper traces) and P-KaiCWT (lower traces) levels depicted in (A). The maximum levels were normalized to 1.
KaiA and KaiB influence KaiC dephosphorylation in vitro
An obvious feature of the KaiC immunoblots in Figure 2A is the double bands seen at the initial time points. Nishiwaki et al. (2000) reported that purified recombinant KaiC can autophosphorylate in vitro, and Iwasaki and coworkers have reported KaiC that is phosphorylated in vitro or in vivo has a lower mobility on SDS–PAGE than nonP-KaiC (Iwasaki et al., 2002). We have confirmed those conclusions by digestion with lambda phosphatase. As shown in Figure 3A, recombinant KaiC purified from E.coli and allowed to autophosphorylate in vitro exhibits two bands in SDS–PAGE. The upper band is converted to the lower band by treatment with lambda phosphatase in vitro, and this conversion is inhibited by vanadate (Figure 3A). This phosphorylation also occurs in cyanobacterial cells in vivo. When extracts of cyanobacterial cells were treated with lambda phosphatase in the absence of vanadate, the upper band is dephosphorylated and disappears (Figure 3A). Therefore, the upper KaiC bands are phosphorylated KaiC that have lower mobility in our SDS–PAGE system.
Iwasaki et al. (2002) and Williams et al. (2002) have reported that KaiA appears to stimulate KaiC autophosphorylation in vitro and that KaiB antagonizes this stimulation. We decided to test whether KaiC that has been phosphorylated in vitro can spontaneously dephosphorylate in vitro. We found that this prediction was correct: KaiC that has been autophosphorylated with [γ-32P]ATP undergoes dephosphorylation in vitro, as shown in Figure 3B where the 32P label and the upper band disappears progressively during the incubation. The dephosphorylation estimated by the loss of the upper band occurs in the presence of unlabeled Mg2+-ATP, suggesting that the rate of dephosphorylation exceeds that of new autophosphorylation. Significantly, KaiA inhibits KaiC dephosphorylation in vitro and KaiB antagonizes the KaiA inhibition (Figure 3B and C). KaiB alone has no effect on KaiC dephosphorylation.
KaiA and KaiB influence KaiC phosphorylation and degradation in vivo
Although KaiC is able to autophosphorylate in vitro, it is likely that other proteins might influence its phosphorylation status in vivo. Two obvious candidates that might be expected to influence KaiC’s phosphorylation in vivo are (i) the histidine kinase SasA that interacts with KaiC (Iwasaki et al., 2000) and (ii) the bacteriophytochrome CikA that has conserved histidine kinase domains and has been reported to be involved in the light input pathway (Schmitz et al., 2000). Using deletion strains of cikA and sasA, however, we found that neither gene was required for in vivo phosphorylation of KaiC (Figure 3D).
On the other hand, we found a very significant effect of KaiA and KaiB on the in vivo phosphorylation of KaiCWT in our pulsed-KaiC assay. As shown in Figure 4, when the kaiA gene was deleted (in ΔkaiA and ΔkaiABC), only the nonP-KaiC band was observed in the pulsed-KaiC assay. The rate of nonP-KaiCWT degradation was similar in the ΔkaiA and ΔkaiABC strains, and was nearly the same as in the ΔkaiC strain. The opposite pattern was observed in the absence of KaiB (in ΔkaiB and ΔkaiBC); there, the lifetime of the P-KaiCWT band was enhanced. In the ΔkaiB strain, the rate of nonP-KaiCWT degradation is somewhat faster than in the ΔkaiC strain. But the dramatic result is that in the absence of KaiB, P-KaiCWT persists 10-fold longer (∼2 h half-life in the ΔkaiC strain, ∼20 h half-life in the ΔkaiB and ΔkaiBC strains). The results shown in Figure 4 suggest that KaiA and KaiB have antagonistic effects on the phosphorylation status of KaiC and the degradation of P-KaiC in vivo.
We also monitored the patterns of KaiC phosphorylation in strains in which the kaiA and kaiB genes were mutated to produce circadian period phenotypes. The endogenous kaiC gene was deleted from these mutant strains and the trcp::kaiC construct was inserted into a neutral site. The period mutants were: (i) for the kaiA gene, A30a (30 h period) and A33a (33 h period); and (ii) for the kaiB gene, B21a (21 h period) and B22a (22 h period) (Ishiura et al., 1998). As shown in Figure 5A, the period mutants have a phenotype of KaiC phosphorylation patterns similar to their corresponding deletion strains (i.e. ΔkaiA and ΔkaiB): B21a/B22a exhibit enhanced P-KaiCWT lifetime, and A30a/A33a show a reduced level of P-KaiCWT. On the original blot, there is a clear upper band in A30a at the intial time points, but at a reduced level compared with the wild-type trace in Figure 2. There is no detectable upper band in A33a, suggesting a correlation between the P-KaiC:nonP-KaiC ratio and the severity of the period phenotype in the kaiA mutants. Therefore, it appears that the B21a, B22a, A30a and A33a mutations all reduce the activity of the corresponding proteins, since the KaiC phosphorylation phenotypes of these mutant strains tend towards those of the matching deletion strains (ΔkaiA and ΔkaiB). Despite the striking differences in the phosphorylation patterns, however, there are no significant differences in the degradation rate of total KaiCWT in these mutant backgrounds (Figure 5B).
Discussion
Degradation of KaiC
We previously reported that KaiC is rhythmically abundant and that KaiC pulses reset the phase of the clock in cyanobacteria (Xu et al., 2000). In the present study, we confirmed the observation of Iwasaki et al. (2002) that KaiC is phosphorylated in vivo. Iwasaki et al. (2002) additionally reported that KaiC is rhythmically phosphorylated. In retrospect, we realize that the multiple KaiC bands we observed in our previous time-course study of wild-type cells (Xu et al., 2000) are indicative of rhythms of KaiC phosphorylation in which P-KaiC predominates in the subjective night (upper bands). The abundance of nonP-KaiC is also rhythmic, although the nonP-KaiC band was obscured in our previous investigation by a cross-reacting antigen (Xu et al., 2000). Because KaiC (P-KaiC and nonP-KaiC) is rhythmically abundant and KaiC pulses reset the phase of the clock in cyanobacteria, KaiC turnover was proposed to be an important determinant of clock period, and therefore differences in the degradation rate of KaiC might be expected to affect the period (Xu et al., 2000).
The results shown in Figure 2 partially support that hypothesis, and also support the validity of the pulsed-KaiC protocol as a relative gauge of degradation rate in vivo. In the cases of the KaiC mutants KaiCC22a and KaiCC55a, the differences in degradation rate of nonP-KaiC are consistent with a significant contribution to the observed differences in period. However, the observed differences in degradation rates are probably not due to differences in the phosphorylation of KaiCWT versus KaiCC22a versus KaiCC55a, because there are no correlative trends in the phosphorylation patterns of KaiCWT, KaiCC22a versus KaiCC55a (Figure 2B versus C). The conclusion that differences in KaiC phosphorylation do not directly lead to differences in degradation is supported by the data illustrated in Figures 4 and 5. In Figure 4, there is not a large difference in the degradation rate of total KaiCWT in ΔkaiA versus ΔkaiBC despite dramatic differences in phosphorylation status. Moreover, in Figure 5 the lifetime of P-KaiC is clearly longer in the B21a and B22a mutants (short period mutants), and shorter in the A30a and A33a mutants (long period mutants). These differences do not appear to be reflected in major changes in the degradative rate of total KaiC that correlate with period. On the other hand, the dramatic differences in phosphorylation status of KaiC in the short period backgrounds (kaiB mutants, B21a and B22a) compared with the long period backgrounds (kaiA mutants, A30a and A33a) led to the hypothesis that the period differences among these strains could be due to the effects of KaiA and KaiB on the status of KaiC phosphorylation (see below).
Phosphorylation and dephosphorylation of KaiC
KaiC has been found to autophosphorylate in vitro (Nishiwaki et al, 2000), and phosphorylated form(s) of KaiC are present in cyanobacterial cells in vivo (Figures 3–5; Xu et al., 2000; Iwasaki et al., 2002). The putative kinases CikA and SasA that may be involved in the circadian system of cyanobacteria do not appear to influence KaiC phosphorylation in our pulsed-KaiC assay, but the phosphorylation status of KaiC is influenced by KaiA and KaiB. Recent studies have reported that KaiA stimulates KaiC autophosphorylation in vitro 3- to 4-fold (Iwasaki et al., 2002; Williams et al., 2002). Moreover, the inclusion of KaiB inhibits this KaiA stimulation by 50% in vitro (Williams et al., 2002). It is possible that our data on P-KaiC dephosphorylation in vitro (Figure 3) can account for the results of the other studies; for example, perhaps the apparent KaiA stimulation of KaiC autophosphorylation is merely due to an inhibition of dephosphorylation. A quantitative analysis of the relative actions of KaiA in stimulating KaiC phosphorylation as opposed to inhibiting KaiC dephosphorylation is required. For the time being, we will adopt the interpretation that KaiA both stimulates KaiC autophosphorylation and inhibits its dephosphorylation. Additionally, KaiB antagonizes both of these effects of KaiA (Figure 6).
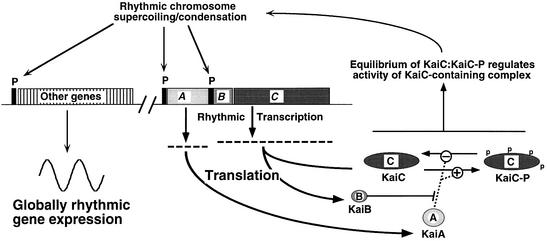
Fig. 6. Model of the circadian system in cyanobacteria. KaiA, KaiB and KaiC are transcribed and translated from the kaiABC cluster using two promoters: kaiAp and kaiBCp. KaiA promotes the phosphorylation of KaiC and inhibits its dephosphorylation, while KaiB antagonizes the actions of KaiA. KaiC phosphorylation is coincident with the formation of a KaiC-containing complex that mediates rhythmic and global changes in the status of the chromosome. These changes in chromosomal status influence the transcriptional activity of all promoters in the chromosome so that there are global circadian changes in gene expression. ‘Other genes’ includes heterologous genes (and heterologous promoters such as trcp).
In vivo, there is no detectable upper band of KaiC when the kaiA gene is deleted (Figure 4; Iwasaki et al., 2002). In the kaiA mutants, there is either no detectable P-KaiC (A33a) or the P-KaiC is much less stable (A30a; Figure 5). Together with the in vitro studies, this observation suggests that in the absence of KaiA the rate of KaiC autophosphorylation is low, and that P-KaiC is very unstable. The presence of KaiB significantly reduces the lifetime of P-KaiC in vivo, since the upper KaiC band persisted in the ΔkaiB, ΔkaiBC, B21a and B22a strains. Therefore, in the absence of robustly active KaiB, the action of KaiA is not antagonized so there are relatively high and stable levels of P-KaiC. Therefore, the level of KaiC phosphorylation is clearly a function of antagonistic interplay between KaiA and KaiB activity levels.
In eukaryotes, there is considerable evidence for regulation of clock protein turnover by phosphorylation acting as a prelude to degradation. However, these processes do not appear to be linked in cyanobacteria (see above; Figures 2, 4 and 5). The rate constants that drive KaiC phosphorylation and dephosphorylation are different from those driving KaiC degradation, since the P-KaiC:nonP-KaiC ratio is not constant throughout the KaiC decay curve in wild-type backgrounds (Figure 2). This difference in rate constants appears to be mediated by KaiB, since the P-KaiC:nonP-KaiC ratio is constant during KaiC degradation in ΔkaiBC (Figure 4), B21a and B22a (Figure 5). There appears to be an inverse correlation between the half-life of P-KaiCWT and period length in the kaiA and kaiB mutant backgrounds (Figure 5), and no correlation between the half-life of nonP-KaiCWT and period length. Therefore, while degradation of total KaiC is positively correlated with period (Figure 2), the rate of P-KaiC dephosphorylation is not correlated with the rate of degradation of total KaiC (Figure 4). The overall conclusion is that KaiC influences period length by both its phosphorylation status (Figure 5) and by the rate of nonP-KaiC degradation (Figure 2), but that these two processes are not directly linked (Figures 4 and 5).
The kaiBC promoter is not specifically required for circadian rhythmicity
One of the most significant observations of this study is the fact that the circadian clockwork operates even when KaiB and KaiC are expressed from a heterologous promoter (trcp). Therefore, specific cis elements in kaiBCp are not necessary for this clockwork, in striking contrast to the situation with eukaryotic clock genes. Elevating KaiC expression above a critical level by the application of higher concentrations of IPTG abolishes rhythmicity, indicating that the negative feedback observed in wild-type remains active in the ΔkaiC/trcp::kaiC and ΔkaiBC/trcp::kaiBC strains. Therefore, negative feedback upon KaiB and KaiC production does not specifically require the endogenous promoter (kaiBCp); apparently all that is required to enable rhythmicity is the expression of KaiB and KaiC within a critical window of intracellular concentrations.
Unlike the situation in eukaryotes, circadian gene expression patterns in this cyanobacterium are global; essentially all promoters in this organism are regulated rhythmically (Liu et al., 1995), including that of the heterologous promoter, conIIp (Katayama et al., 1999). The rhythmic orchestration of gene expression is apparently not mediated directly by regulation of sigma factors, since inactivation (singly or pairwise) of any of the four known group 2 sigma factor genes from this cyanobacterium does not eliminate rhythmically global gene expression (Nair et al., 2002). We previously proposed that this global regulation is based on rhythmic topological changes in the chromosome that influence the activity of all promoters, including those that drive expression of KaiC from either endogenous or heterologous promoters (Mori and Johnson, 2001). Our revised model (Figure 6) proposes that KaiC might mediate both its own negative feedback regulation and global regulation of the cyanobacterial genome by rhythmically controlling the condensation and/or supercoiling status of the entire cyanobacterial chromosome. The chromosome of bacteria has a highly organized architecture based on condensation and coiling of DNA, and it is well known that changes in the local supercoiling status of DNA can affect the transcription rate of prokaryotic genes (Trun and Marko, 1998).
In cyanobacteria, we postulate that oscillations in the torsional stress and/or condensation of the chromosome promote rhythmic modulation of the transcription rates of all genes (Figure 6), accounting for global regulation of gene expression (Liu et al., 1995). By this interpretation, gene-specific cis elements that mediate rhythmic gene expression might be (at least partially) responsive to chromosomal status rather than exclusively to trans factors. In addition, heterologous promoters such as trcp that are integrated into the chromosome are driven rhythmically, since they are also subjected to the oscillating chromosomal status. Therefore, the results shown in Figure 1 provide supporting evidence for our working hypothesis that global expression is not strictly controlled by promoter-specific sequences, but might be largely mediated by an oscillating chromosome.
Role of KaiC in the cyanobacterial clockworks
We suggest that KaiC (or most likely, a KaiC-containing complex) is a key player in regulating these changes of chromosomal status. We hypothesize that the ratio of P-KaiC:nonP-KaiC reflects the formation and activity of clock-active KaiC. There are several lines of evidence to support this hypothesis. First, there is a clear rhythm of P-KaiC over the circadian time course (see above; Xu et al., 2000; Iwasaki et al., 2002). Secondly, the ratio of P-KaiC:nonP-KaiC is very significantly different between the B21a/B22a (short period) and the A30a/A33a (long period) strains that have differing period phenotypes. Therefore, the ‘speed’ of the clock is correlated with the P-KaiC:nonP-KaiC ratio in these kaiA and kaiB mutants. Thirdly, in studies with purified KaiC, we have found that KaiC forms hexameric complexes in vitro that bind DNA (Mori et al., 2002). The formation of these hexamers (i) is ATP dependent and (ii) increases in concert with an increased P-KaiC:nonP-KaiC ratio (Mori et al., 2002). KaiC is a member of a gene family in which other members such as RecA and DnaB form hexamers in the active state that act upon DNA, implying that KaiC might also act directly on DNA (Leipe et al., 2000).
Therefore, the model illustrated in Figure 6 suggests that KaiC forms hexameric complexes concomitantly with phosphorylation of KaiC subunits; KaiA promotes this process and KaiB partially inhibits this process. We propose that this hexameric complex is the active form of KaiC, and mediates rhythmic changes in chromosomal torsion (probably in conjunction with other proteins in a supramolecular assembly). The acceleration of the clock in the B21a/B22a strains is proposed to be due to the increased proportion of P-KaiC, indicative of more rapid complex formation, whereas the opposite slowing of the clock in the A30a/A33a strains is due to the decreased proportion of P-KaiC and a concomitant reduction in the rate of KaiC complex formation. The rate at which KaiC degrades can also influence period length. In wild-type cells, there are rhythms of KaiB and KaiC abundance, but KaiA is constitutively present (Xu et al., 2000). Therefore, KaiA is present to promote KaiC phosphorylation as soon as KaiC is synthesized. However, KaiB is co-translated with KaiC and therefore may promote a dynamic equilibrium in the P-KaiC:nonP-KaiC ratio. The result is a rhythm of the ratio between P-KaiC and nonP-KaiC (Iwasaki et al., 2002), which we postulate to modulate the assembly of a clock-active KaiC-containing complex (Figure 6).
Materials and methods
Growth conditions and luminescence assays
Cyanobacterial strains were grown at 30°C in continuous white light (LL) on modified BG-11 medium (Kondo et al., 1993) containing appropriate antibiotics (20 µg/ml spectinomycin, 5 µg/ml kanamycin, 7.5 µg/ml chloramphenicol and/or 2 µg/ml gentamicin). For solid plates, 1.5% agar was added to the medium. For liquid cultures, cell suspensions were bubbled with sterile air. Cell densities of liquid cultures were estimated from the absorbance at 750 nm. Luminescence rhythms were measured as described previously for liquid cultures (Kondo et al., 1993) or for single colonies on agar (Kondo et al., 1994) in LL after a 12 h dark pulse to synchronize the clocks in the population.
kaiA, kaiB and kaiC mutants
The wild-type reporter strain, AMC149, was as described by Kondo et al. (1993). Six different period mutants were used: two mutations in KaiC (C55a with period ≈55 h and C22a with period ≈22 h), two KaiA mutants (A30a and A33a with 30 h and 33 h periods, respectively) and two KaiB mutants (B21a and B22a with 21 h and 22 h periods, respectively). A30a, A33a, B21a, B22a and C22a were previously reported by Ishiura et al. (1998). C55a is a new long-period mutant that was isolated in our laboratory using the method described by Kondo et al. (1994). C55a is rescued by wild-type kaiC DNA and results from a single point mutation in kaiC, as revealed by DNA sequence analysis (Figure 1B).
Generation of gene knockout strains
Kai-null strains were made in the laboratory of Dr S.S.Golden by in-frame deletions as follows: ΔkaiA (with a deletion of amino acids 22–255), ΔkaiB (amino acids 11–79 deleted), ΔkaiC (amino acids 76–410 deleted) and ΔkaiBC (amino acids 11–79 of kaiB deleted and amino acids 76–410 of kaiC deleted). The kaiABC-null strain (ΔkaiABC) was created by replacement of the kaiABC DNA region with a kanamycin resistance gene (Ishiura et al., 1998). To produce the strains A30aΔkaiC, A33aΔkaiC, B21aΔkaiC and B22aΔkaiC, the kaiC gene was inactivated by insertion of a 1.4 kb of DNA fragment carrying a gentamycin resistance gene into the EcoRV site of the kaiC coding region in the kaiA or kaiB mutants A30a, A33a, B21a or B22a, respectively. The ΔcikAΔkaiC strain was made by replacement of a 1.3 kb of BglII–BamHI fragment within the cikA coding region with a gentamicin resistance gene in the ΔkaiC strain. The ΔsasA strain was obtained by replacement of a DraIII–SmaI fragment harboring its major coding region with a 1.34 kb DNA fragment encoding a kanamycin resistance gene. All deletion strains were confirmed by PCR analysis of the corresponding genomic DNA. Deletion strains were further characterized by immunoblot assays for the presence of the relevant proteins and/or luminescence measurement for reporter rhythmicity.
Construction of the kaiBCp::luxAB reporter strains
For monitoring the promoter activity of the kaiBC gene by luminescence, a transcriptional fusion construct (kaiBCp::luxAB) was made. The kaiBC promoter (kaiBCp) that includes the kaiB sequence from positions –479 to +23 was fused to the luxAB gene cassette. For the wild-type strain, the gene fusion psbAIp::luxAB (spectinomycin resistance) in the neutral site I (NS I) of AMC149 was replaced with the kaiBCp::luxAB construct (kanamycin resistance) to give rise to a new wild-type reporter strain, AMC149/kaiBCp. For the gene knockout strains used in this study (i.e. kai-null, ΔcikA and ΔsasA strains), the kaiBCp::luxAB construct was put into NS I with a selection marker of spectinomycin resistance.
Cloning and expression of wild-type KaiC, KaiBC or mutant KaiCs
The region encoding KaiCWT, KaiCC22a or KaiCC55a was amplified with NdeI ends using Pfu DNA polymerase (Stratagene, La Jolla, CA) from the genomic DNA of the strains AMC149, C22a and C55a, respectively. A genomic DNA fragment harboring the start codon of the KaiB coding region through the stop codon of KaiC coding region was amplified with NdeI ends from wild-type kai template. The DNA fragments were then cloned into a NS I vector containing spectinomycin resistance or a NS II vector containing kanamycin resistance. In both cases, the expression of wild-type KaiC, KaiBC (KaiB and KaiC) or mutant KaiC was under the control of an IPTG-inducible trcp regulated by the repressor lacIq (Amann et al., 1988). The linker sequence between the trc promoter and the kaiB or kaiC coding region was 5′-AGGAAACAGACCATGGAATTCGAG GTCAGGGAGGAATAACAT-3′ (functional ribosome-binding site is underlined). The corresponding phenotypes were confirmed by expressing the cloned KaiCs in the ΔkaiC strain. Then, trcp::kaiCWT, trcp::kaiCC22a, trcp::kaiCC55a or trcp::kaiBC were integrated into NS I of the strains ΔkaiABC and ΔsasA, or into NS II of the strains AMC149, ΔkaiA, ΔkaiB, ΔkaiC, ΔkaiBC, B21aΔkaiC, B22aΔkaiC, A30aΔkaiC, A33aΔkaiC and ΔcikAΔkaiC, to generate the strains described in Results.
KaiC and/or kaiC mRNA assays: degradation, immunoblots and northern blots
A single of colony of a gene-null and/or kai-mutant strain expressing trcp::kaiCWT, trcp::kaiCC22a or trcp::kaiCC55a was grown in 30 ml of BG-11 medium with appropriate antibiotics until the OD750 reached 0.4. The cultures were transferred into 300 ml fresh medium and further grown in constant light (LL) to OD750 = 0.2. After a 12 h dark pulse, the cultures were treated with or without 0.1 mM IPTG for 3 h at 30°C in the light with air bubbling. The inducer IPTG was removed by washing cultures four times with medium at 4°C. The washed pellets were resuspended in fresh medium (30°C) to an OD750 of 0.2, and the cultures were released to normal growth conditions with addition of fresh medium to maintain the same cell density. At various time points, ∼40 ml of these cultures were collected, and the samples snap-frozen in liquid nitrogen and stored at –80°C to be used for later immunoblot or RNA blot analyses of the kaiC gene.
For immunoblot analysis, total proteins were extracted from cyanobacteria and immunoblot analysis for the KaiC protein was performed as described previously (Xu et al., 2000) with a further purified antiserum.
For northern blot assays, total RNA was prepared using a RiboPure™-Bacteria RNA isolation kit (Ambion, Inc., Austin, TX). Normalization of the RNA loading (1 µg per lane) was confirmed by ribosome RNA assays. An [α-32P]dCTP-labeled 940 bp fragment corresponding to amino acids 76–410 of KaiC, which were deleted in the ΔkaiC strain, was used as probe for northern assays. Quantitation of the blot densities was performed with an IS-1000 digital imaging system using Spot Density Tool software (Alpha Innotech, San Leandro, CA) after extensive standardization using films of known densities or scanned with a PhosphorImager and analyzed with ImageQuant™ software (Molecular Dynamics, Sunnyvale, CA).
Phosphatase treatment
For phosphatase treatment of cyanobacterial extracts, the concentrated extracts (10 µg/µl protein) from the IPTG-induced ΔkaiC strain expressing trcp::kaiC were prepared in the following buffer: 50 mM HEPES pH 7.4, 130 mM KCl, 5% glycerol, 2.5 mM EDTA, 10 µg/ml aprotinin, 10 µg/ml leupeptin, 2 µg/ml pepstatin and 5 mM phenylmethylsulfonyl fluoride. For phosphatase treatment of purified recombinant KaiC, His6-tagged KaiC protein was purified as described elsewhere (Mori et al., 2002). About 1 µg of purified KaiC::His6 or 30 µg of total protein from extracts was diluted in 1× phosphatase buffer (New England Biolabs, Beverly, MA). For the samples that included phosphatase, 1000 U of lambda protein phosphatase (λPPase; New England Biolabs) were added, and the reaction was incubated at 30°C for 1 h in the presence or absence of 40 mM sodium vanadate. The reaction was stopped by addition of 2× SDS loading buffer and boiled for 5 min, and an aliquot of the sample subjected to SDS–PAGE followed by Coomassie Brilliant Blue staining (for purified KaiC) or immunoblot analysis (for cyanobacterial extracts).
KaiC dephosphorylation in vitro
Purified KaiC-His6 (186 µg) was incubated with 100 µCi [γ-32P]ATP (6000 Ci/mmol) in 100 µl of 20 mM Tris–HCl pH 8, 1 mM ATP, 1 mM dithiothreitol, 5% (v/v) glycerol, 100 mM NaCl, at 4°C for 24 h and 30°C for 15 min. Non-incorporated [γ-32P]ATP was then removed by centrifugation through a Bio-Gel P-30 spin column (Bio-Rad) and ultrafiltration with a Microcon-50 centrifugal filter (Millipore, Billerica, MA). For the dephosphorylation reaction, phosphorylated KaiC-His6 (13.3 µg) was incubated in 100 µl of 22 mM Tris–HCl pH 8, 5.5% (v/v) glycerol, 5 mM 2-mercaptoethanol, 170 mM NaCl, 5.5 mM ATP, 5 mM MgCl2 in the presence or absence of purified KaiA (2.75 µg) and/or KaiB (7.87 µg) at 30°C. Aliquots (20 µl) were withdrawn at 0, 1, 6, 12 and 24 h of incubation, and mixed with 4 µl of 6× SDS–PAGE sample loading buffer. SDS–PAGE (10% T/2.6% C) was run at room temperature. Gels were fixed and stained with Coomassie Brilliant Blue R-250, dried and then exposed to a storage phosphor screen (Eastman Kodak, Rochester, NY) for 22.75 h. The screen was scanned with a PhosphorImager, and relative amounts of radioactivity in each band were quantified with ImageQuant.
Acknowledgments
Acknowledgements
We dedicate this paper to the memory of Dr Gisela Mosig in recognition of the inspiration she provided to this study. We thank Drs Jayna L.Ditty, Stanly B.Williams and Susan S.Golden for the gift of strains that carry in-frame deletions of the kai genes, and for the construct to make the ΔcikA strain. We also thank Drs Takao Kondo and Masahiro Ishiura for the C22a, B21a, B22a, A30a and A33a strains. Finally, we thank Drs Stanly Williams, Hideo Iwasaki, Susan Golden and Takao Kondo for sharing the results of their experiments prior to publication. We are grateful for support from the National Science Foundation (grant MCB-9874371) and the National Institutes of Health (grants MH01179, MH43836 and GM67152).
References
- Amann E., Ochs,B. and Abel,K.J. (1988) Tightly regulated tac promoter vectors useful for the expression of unfused and fused proteins in Escherichia coli. Gene, 69, 301–315. [DOI] [PubMed] [Google Scholar]
- Darlington T.K., Wager-Smith,K., Ceriani,M.F., Staknis,D., Gekakis,N., Steeves,T.D.L., Weitz,C.J., Takahashi,J.S. and Kay S.A. (1998) Closing the circadian loop: CLOCK-induced transcription of its own inhibitors per and tim. Science, 280, 1599–1603. [DOI] [PubMed] [Google Scholar]
- Dunlap J.C. (1999) Molecular bases for circadian clocks. Cell, 96, 271–290. [DOI] [PubMed] [Google Scholar]
- Gekakis N., Stankis,D., Nguyen,H.B., Davis,F.C., Wilsbacher,L.D., King,D.P., Takahashi,J.S. and Weitz,C.J. (1998) Role of the CLOCK protein in the mammalian circadian mechanism. Science, 280, 1564–1569. [DOI] [PubMed] [Google Scholar]
- Görl M., Merrow,M., Huttner,B., Johnson,J., Roenneberg,T. and Brunner,M. (2001) A PEST-like element in FREQUENCY determines the length of the circadian period in Neurospora crassa. EMBO J., 20, 7074–7084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hardin P.E., Hall,J.C. and Rosbash,M. (1990) Feedback of the Drosophila period gene product on circadian cycling of its messenger RNA levels. Nature, 343, 536–540. [DOI] [PubMed] [Google Scholar]
- Ishiura M., Kutsuna,S., Aoki,S., Iwasaki,H., Andersson,C.R., Tanabe,A., Golden,S.S., Johnson,C.H. and Kondo,T. (1998) Expression of a gene cluster kaiABC as a circadian feedback process in cyanobacteria. Science, 281, 1519–1523. [DOI] [PubMed] [Google Scholar]
- Iwasaki H., Taniguchi,Y., Ishiura,M. and Kondo,T. (1999) Physical interactions among circadian clock proteins KaiA, KaiB and KaiC in cyanobacteria. EMBO J., 18, 1137–1145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwasaki H., Williams,S.B., Kitayama,Y., Ishiura,M., Golden,S.S. and Kondo,T. (2000) A KaiC-interacting sensory histidine kinase, SasA, necessary to sustain robust circadian oscillation in cyanobacteria. Cell, 101, 223–233. [DOI] [PubMed] [Google Scholar]
- Iwasaki H., Nishiwaki,T., Kitayama,Y., Nakajima,M. and Kondo,T. (2002) KaiA-stimulated KaiC phosphorylation in circadian timing loops in cyanobacteria. Proc. Natl Acad. Sci. USA, 99, 15788–15793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson C.H., Golden,S.S., Ishiura,M. and Kondo,T. (1996) Circadian clocks in prokaryotes. Mol. Microbiol., 21, 5–11. [DOI] [PubMed] [Google Scholar]
- Katayama M., Tsinoremas,N.F., Kondo,T. and Golden,S.S. (1999) cpmA, a gene involved in an output pathway of the cyanobacterial circadian system. J. Bacteriol., 181, 3516–3524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kloss B., Price,J.L., Saez,L., Blau,J., Rothenfluh,A. and Young,M.W. (1998) The Drosophila clock gene double-time encodes a protein closely related to human casein kinase Iε. Cell, 94, 97–107. [DOI] [PubMed] [Google Scholar]
- Kloss B., Rothenfluh,A., Young,M.W. and Saez,L. (2001) Phosphorylation of PERIOD is influenced by cycling physical associations of DOUBLE-TIME, PERIOD and TIMELESS in the Drosophila clock. Neuron, 30, 699–706. [DOI] [PubMed] [Google Scholar]
- Kondo T., Strayer,C.A., Kulkarni,R.D., Taylor,W.R., Ishiura,M., Golden,S.S. and Johnson,C.H. (1993) Circadian rhythms in prokaryotes: luciferase as a reporter of circadian gene expression in cyanobacteria. Proc. Natl Acad. Sci. USA, 90, 5672–5676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kondo T., Tsinoremas,N.F., Golden,S.S., Johnson,C.H., Kutsuna,S. and Ishiura,M. (1994) Circadian clock mutants of cyanobacteria. Science, 266, 1233–1236. [DOI] [PubMed] [Google Scholar]
- Kume K., Zylka,M., Sriram,S., Shearman,L.P., Weaver,D.R., Jin,X., Maywood,E.S., Hastings,M.H. and Reppert,S.M. (1999) mCRY1 and mCRY2 are essential components of the negative limb of the circadian feedback loop. Cell, 98, 193–205. [DOI] [PubMed] [Google Scholar]
- Leipe D.D., Aravind,L., Grishin,N.V. and Koonin,E.V. (2000) The bacterial replicative helicase DnaB evolved from a RecA duplication. Genome Res., 10, 5–16. [PubMed] [Google Scholar]
- Liu Y., Loros,J. and Dunlap,J.C. (2000) Phosphorylation of the Neurospora clock protein FREQUENCY determines its degradation rate and strongly influences the period length of the circadian clock. Proc. Natl Acad. Sci. USA, 97, 234–239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y., Tsinoremas,N.F., Johnson,C.H., Lebedeva,N.V., Golden,S.S., Ishiura,M. and Kondo,T. (1995) Circadian orchestration of gene expression in cyanobacteria. Genes Dev., 9, 1469–1478. [DOI] [PubMed] [Google Scholar]
- Mori T. and Johnson,C.H. (2001) Circadian programming in cyanobacteria. Semin. Cell Dev. Biol., 12, 271–278. [DOI] [PubMed] [Google Scholar]
- Mori T., Saveliev,S.V., Xu,Y., Stafford,W.F., Cox,M.M., Inman,R.B. and Johnson,C.H. (2002) Circadian clock protein KaiC forms ATP-dependent hexameric rings and binds DNA. Proc. Natl Acad. Sci. USA, 99, 17203–17208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nair U., Ditty,J.L., Min,H. and Golden,S.S. (2002) Roles for sigma factors in global circadian regulation of the cyanobacterial genome. J. Bacteriol., 184, 3530–3538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishiwaki T., Iwasaki,H., Ishiura,M. and Kondo,T. (2000) Nucleotide binding and autophosphorylation of the clock protein KaiC as a circadian timing process of cyanobacteria. Proc. Natl Acad. Sci. USA, 97, 495–499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reppert S.M. and Weaver,D. (2002) Coordination of circadian timing in mammals. Nature, 418, 935–941. [DOI] [PubMed] [Google Scholar]
- Ruoff P., Mohsenzadeh,S. and Rensing,L. (1996) Circadian rhythms and protein turnover: the effect of temperature on the period lengths of clock mutants simulated by the Goodwin oscillator. Naturwissenschaften, 83, 514–517. [DOI] [PubMed] [Google Scholar]
- Schmitz O., Katayama,M., Williams,S.B., Kondo,T. and Golden,S.S. (2000) CikA, a bacteriophytochrome that resets the cyanobacterial circadian clock. Science, 289, 765–768. [DOI] [PubMed] [Google Scholar]
- Trun N.J. and Marko,J.F. (1998) Architecture of a bacterial chromosome. ASM News, 64, 276–283. [Google Scholar]
- Williams S.B., Vakonakis,I., Golden,S.S. and LiWang,A.C. (2002) Structure and function from the circadian clock protein KaiA of Synechococcus elongatus: a potential clock input mechanism. Proc. Natl Acad. Sci. USA, 99, 15357–15362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu Y., Mori,T. and Johnson,C.H. (2000) Circadian clock-protein expression in cyanobacteria: rhythms and phase-setting. EMBO J., 19, 3349–3357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young M.W. and Kay,S.A. (2001) Time zones: a comparative genetics of circadian clocks. Nat. Rev. Genet., 2, 702–715. [DOI] [PubMed] [Google Scholar]