Abstract
The human immunodeficiency virus type 1 (HIV-1) Gag precursor protein Pr55Gag drives the assembly and release of virus-like particles in the infected cell. The capsid (CA) domain of Gag plays an important role in these processes by promoting Gag-Gag interactions during assembly. The C-terminal domain (CTD) of CA contains two dileucine-like motifs (L189/L190 and I201/L202) implicated in regulating the localization of Gag to multivesicular bodies (MVBs). These dileucine-like motifs are located in the vicinity of the CTD dimer interface, a region of CA critical for Gag-Gag interactions during virus assembly and CA-CA interactions during core formation. To study the importance of the CA dileucine-like motifs in various aspects of HIV-1 replication, we introduced a series of mutations into these motifs in the context of a full-length, infectious HIV-1 molecular clone. CA mutants LL189,190AA and IL201,202AA were both severely impaired in virus particle production because of a variety of defects in the binding of Gag to membrane, Gag multimerization, and CA folding. In contrast to the model suggesting that the CA dileucine-like motifs regulate MVB targeting, the IL201,202AA mutation did not alter Gag localization to the MVB in either HeLa cells or macrophages. Revertants of single-amino-acid substitution mutants were obtained that no longer contained dileucine-like motifs but were nevertheless fully replication competent. The varied phenotypes of the mutants reported here provide novel insights into the interplay among Gag multimerization, membrane binding, virus assembly, CA dimerization, particle maturation, and virion infectivity.
Human immunodeficiency virus type 1 (HIV-1) assembly and release are driven by the Gag polyprotein precursor Pr55Gag, whose expression is sufficient for the formation of virus-like particles (VLPs) (13, 45). Pr55Gag is composed of four major domains—matrix (MA), capsid (CA), nucleocapsid (NC), and p6—and two small spacer peptides (SP1 and SP2) located C terminal to CA and NC, respectively. Each of these domains plays an important role in VLP production. MA contains the determinants in Gag that interact directly with the lipid bilayer and thus regulate the binding of Gag to membrane. The CA domain is composed of two structural units, i.e., the N-terminal domain (NTD), which plays a crucial role in virion maturation (see below), and the C-terminal domain (CTD), which bears residues necessary for Gag-Gag oligomerization (17, 18). Specifically, the CA CTD induces Gag dimerization and mutations in residues that form the CA CTD dimer interface severely disrupt Gag assembly and virus particle production (19, 49). NC also promotes Gag oligomerization during assembly, largely because of the ability of basic residues within NC to interact with viral RNA enabling the nucleic acid to serve as a scaffold along which Gag molecules can align and multimerize (2, 3, 5, 7, 30). CA alone can assemble into tubular structures in vitro without NC, and mutations in the CTD dimer interface disrupt this assembly reaction (4, 12, 18, 19, 21, 22, 48). These findings suggest that the CA CTD is critical for mediating the initial Gag-Gag contacts during assembly. Finally, p6 bears a “late” domain that promotes particle budding and release by interacting with cellular endosomal sorting machinery, most notably the endosomal sorting complex required for transport 1 (ESCRT-I) component Tsg101 (9, 29).
In most commonly studied cell types (e.g., HeLa, 293T, COS, and T-cell lines), HIV-1 assembly occurs predominantly on the plasma membrane. However, in other cell types, particularly the physiologically relevant primary monocyte-derived macrophage (MDM), assembly and budding take place in an intracellular compartment identified as the multivesicular body (MVB) or late endosome (31, 36, 40, 43). The itinerary that Gag follows to reach its destination in the cell remains ill defined; it has been suggested that even in cell types in which assembly and budding occur predominantly at the plasma membrane, Gag traffics though an endosomal or endocytic compartment before reaching the cell surface (10, 20, 32). A highly basic domain in MA and a region of MA referred to as the “80's domain” play an important role in Gag targeting to the plasma membrane. Mutations in these sequences lead to retargeting of virus assembly from the plasma membrane to the MVB in HeLa cells and in T-cell lines, thereby recapitulating the phenotype observed for wild-type (WT) Gag in MDMs (16, 36, 38).
During or immediately after virus release from the infected cell, the viral protease (PR) cleaves Pr55Gag at sites between the major Gag domains to generate the mature Gag proteins MA, CA, NC, and p6. Cleavage of Gag by PR leads to a morphological transition in the virus particle, a process known as maturation, which is essential for virus infectivity (13). During maturation, the CA protein undergoes a conformational transformation and reassembles into helical arrays in which the CA NTD forms hexameric rings that are linked into a continuous lattice by the CTD (25, 51). Mutations have been described throughout CA that interfere with the formation of conical cores and hence block virus infectivity (49).
Lindwasser and Resh (27) observed that fusing a large region of HIV-1 Gag encompassing the CA CTD, SP1, and NC to a cytoplasmic tail deletion-containing derivative of CD4 resulted in rapid internalization of the CD4-Gag chimera. These results suggested that the CA CTD-SP1-NC region of Gag might contain a motif(s) responsible for the association of Gag with cellular endocytic machinery. Inspection of this region of Gag revealed two di-Leu-like motifs (L189/L190 and I201/L202) within the CA CTD. Although the role of these di-Leu-like motifs in the internalization of CD4-Gag chimeras was not tested, mutation of these motifs severely disrupted VLP production and reduced MVB localization of Gag in COS cells treated with the drug U18666A, which induces the redistribution of cholesterol from the plasma membrane to MVBs. The authors proposed that these motifs, particularly I201/L202, might be responsible for the association of Gag with MVBs (27). However, these residues also lie in the vicinity of the CA CTD dimer interface, raising the possibility that mutations at these positions might disrupt virus particle production by impeding CA-mediated Gag-Gag interactions.
To investigate the role of the CA di-Leu-like motifs in multiple aspects of HIV-1 replication, we introduced both single- and double-amino-acid substitutions into these motifs in the context of a full-length, infectious HIV-1 molecular clone. We examined the effects of the mutations on virus assembly, Gag targeting, membrane binding and multimerization, virus maturation, replication, and infectivity. Our results indicate that mutation of the CA di-Leu-like motifs severely inhibits virus particle production for a number of widely varied reasons, including impaired Gag-membrane binding, disrupted Gag oligomerization, and altered CA folding. While mutations in the di-Leu-like motifs markedly inhibited virus assembly and release, they did not affect targeting of Gag to the MVB in either HeLa cells or primary human MDMs.
MATERIALS AND METHODS
Cells, transfections, and infections.
HeLa and TZM-bl cells (the latter were obtained from J. Kappes through the NIH AIDS Research and Reference Reagent Program) (42) were cultured in Dulbecco modified Eagle medium as described previously (24). Jurkat and MT-4 T-cell lines were maintained as previously described (24). Monocyte-derived macrophages were prepared and cultured as previously described (14). Plasmid DNA was purified with the plasmid purification maxi prep kit (QIAGEN, Valencia, CA), and transfections were performed with the Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA) by following the manufacturer's protocol. Jurkat and MT-4 T cells were transfected by the DEAE-dextran transfection protocol as previously reported (24). For infection assays, virus stocks were prepared by HeLa cell transfection as described above. Transfected-cell supernatants were passed through a 0.45-μm-pore-size filter, normalized for reverse transcriptase (RT) activity, and used in infections as indicated. TZM-bl cells were infected in the presence of DEAE-dextran at a final concentration of 20 μg/ml.
Mutagenesis and DNA cloning.
CA mutants were constructed in the pNL4-3 backbone with the Quick Change site-directed mutagenesis kit (Stratagene, La Jolla, CA). Primer sequences will be made available on request. The PR− versions of the above mutants were constructed by introducing the ApaI (pNL4-3 nucleotide [nt] 2006)-to-EcoRI (pNL4-3 nt 5743) fragment from pNL4-3/PR− (23) into the CA mutant pNL4-3 derivatives. The 1GA/FLAG-tagged derivatives were constructed by inserting the Sph1 (pNL4-3 nt 1443)-to-Apa1 (pNL4-3 nt 2006) fragment of the CA single and double mutants into pNL4-3/1GA/FLAG (35, 38).
Radioimmunoprecipitation and Western blot analysis.
Methods used for metabolic labeling of HeLa cells, preparation of cell and virion lysates, and immunoprecipitation of viral proteins have been described in detail previously (15, 50). Briefly, transfected HeLa cells were metabolically labeled with [35S]Met/Cys and virions were pelleted by ultracentrifugation. Cell and virus lysates were immunoprecipitated with HIV immunoglobulin (HIV-Ig) obtained from NABI Biopharmaceuticals and the National Heart Blood and Lung Institute through the NIH AIDS Research and Reference Reagent Program and subjected to sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), followed by fluorography. For Western blotting, proteins were separated by SDS-PAGE (10% gel) and transferred to polyvinylidene difluoride membranes (Millipore). To detect hemagglutinin (HA)- and FLAG-tagged proteins, membranes were incubated with anti-HA7 (Sigma, St. Louis, MO) or anti-FLAGM2 (Sigma, St. Louis, MO) antibodies (Abs), followed by horseradish peroxidase (HRP)-conjugated anti-mouse Ig (Amersham Biosciences Uppsala, Sweden). The CA-related proteins in cells were detected with HIV-1 anti-p25/p24 SF2 antiserum (obtained from the Division of AIDS through the NIH AIDS Research and Reference Reagent Program), followed by HRP-conjugated anti-rabbit Ig (Amersham Biosciences, Uppsala, Sweden). To detect HIV-1 proteins in cells, membranes were incubated with HIV-Ig, followed by HRP-conjugated anti-human Ig (Amersham Biosciences, Uppsala, Sweden). Antigens recognized by Abs were detected with enhanced chemiluminescence reagents (Perkin-Elmer, Boston, MA). Quantification of bands was performed with the Alpha Innotech digital imager.
Gag membrane binding and multimerization assays.
Assays for Gag membrane binding and higher-order Gag multimerization were conducted as previously described (35, 39). Briefly, HeLa cells were transfected with the pNL4-3/PR− clones expressing either WT or mutant CA. Transfected cells were pulsed with [35S]Met/Cys for 5 min and chased for 15 min. Cells were then washed with cold phosphate-buffered saline, scraped, and sonicated on ice to achieve disruption of 90% of the cells. Postnuclear supernatants of cell homogenates were subjected to equilibrium flotation centrifugation at 35,000 rpm for 16 h with 85.5%, 65%, and 10% (wt/vol) sucrose gradients. Membrane-bound and non-membrane-bound fractions were then collected and lysed with 2× radioimmunoprecipitation assay buffer (280 mM NaCl, 16 mM Na2HPO4, 4 mM NaH2PO4, 2% NP-40, 1% sodium deoxycholate, 0.1% SDS, 20 mM iodoacetamide, protease inhibitors). Both membrane and nonmembrane fractions were then either denatured by boiling in the presence of 0.057 volume of 2× sample buffer (125 mM Tris-HCl [pH 6.8] containing 6% SDS, 10% 2-mercaptoethanol, and 20% glycerol) or left nondenatured. Immunoprecipitations were then performed on both denatured and nondenatured fractions with HIV-Ig as already described.
Fluorescence microscopy and electron microscopy (EM).
Fluorescence microscopy of transfected cells cultured in chamber slides (Nunc) was conducted as previously described (38). Briefly, cells were fixed 24 to 48 h posttransfection with 3.7% formaldehyde in 100 mM sodium phosphate buffer (pH 7.2) for 20 min, permeabilized with phosphate-buffered saline containing 0.1% Triton X-100, and incubated with anti-p17 monoclonal Ab (Advanced Biotechnologies Inc., Columbia, MD), followed by Texas Red-conjugated anti-mouse IgG. Cells were mounted with Aqua Poly Mount (Polysciences Inc., Warrington, PA) and examined with a Delta Vision RT microscope. When double staining for Gag and CD63 with two different mouse monoclonal Abs, Abs were fluorescently labeled with the Zenon One Alexa 488 or 594 IgG1 labeling kit (Molecular Probes) according to the manufacturer's protocol. To avoid dissociation and rebinding of Zenon One reagents, cells were incubated in the fixation buffer after incubation with each Ab. Fluorescence microscopy in primary MDMs was performed as previously described (33). Briefly, virus stocks prepared by transfecting HeLa cells with WT or IL-201,202AA mutant pNL4-3, pCMVNLGagPolRRE (37), and pHCMV-G (52) were used to infect MDMs. Cells were fixed postinfection, costained with anti-p17 and anti-CD63 Abs, and examined with a DeltaVision RT microscope. Ten Gag-positive cells were analyzed, and costaining between Gag and CD63 in both HeLa cells and MDMs was determined with the softWoRx colocalization module. This tool generates a product image of two channels after subtracting a threshold value for each. A scatter plot is then created on the basis of the two intensities on a pixel-by-pixel basis. The Pearson coefficient of correlation indicates how closely the two intensities are colocalized (full colocalization is 1.0) and is defined as the sum of products of the standard scores of two measures divided by the degrees of freedom. Fixation of cells, preparation of samples, and EM were performed as described previously (16).
RESULTS
Mutation of the HIV-1 CA di-Leu-like motifs leads to severe defects in virus assembly and release.
To define the role of the CA di-Leu-like motifs in HIV-1 assembly and release, we introduced mutations into these motifs in the context of the full-length molecular clone pNL4-3 (Fig. 1). Initially, we substituted Leu189/Leu190 and Ile201/Leu202 for alanines to generate mutants LL189,190AA and IL201,202AA, respectively. As a control, we also included the assembly-deficient WM184,185AA mutant (39), which contains substitutions in two key residues of the CA CTD dimer interface (49). HeLa cells were transfected in parallel with WT or mutant molecular clones and were metabolically labeled with [35S]Met/Cys. Levels of cell-associated and virion-associated HIV-1 proteins were determined by radioimmunoprecipitation of cell and virion lysates with HIV-Ig (see Materials and Methods). The LL189,190AA and IL201,202AA mutations reduced virus particle production approximately 10-fold and 5-fold, respectively (Fig. 2A). As previously reported (39), the WM184,185AA mutant also displayed severely impaired virus particle production. In addition to defective particle production, the CA mutants showed abnormal processing of the Pr55Gag precursor to the Gag processing intermediate p41Gag and the mature CA protein p24 (Fig. 2A). Specifically, the mutants showed elevated levels of p41Gag relative to Pr55Gag, and the LL189,190AA mutant displayed an increased ratio of Pr55Gag to p24 (CA). Interestingly, the IL201,202AA mutations also resulted in the appearance of an additional protein product smaller than 24 kDa (Fig. 2A).
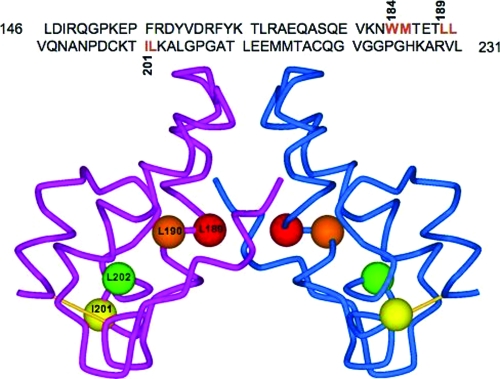
FIG. 1.
Sequence and structure of the HIV-1 CA CTD. The amino acid sequence of the CA CTD is shown at the top. The CTD dimer interface residues W184/M185 and the putative di-Leu-like motifs L189/L190 and I201/L202 are indicated in red. At the bottom is shown a structural model of a dimer of the HIV-1 CA CTD (provided by S. R. Durell) with the locations of L189, L190, I201, and L202 indicated by the red, orange, yellow, and green balls, respectively. Residue L189 lies at the CA dimer interface; residues L190 and L202 are buried, while residue I201 is exposed. The structure was constructed with data from 1A43.pdb (51).
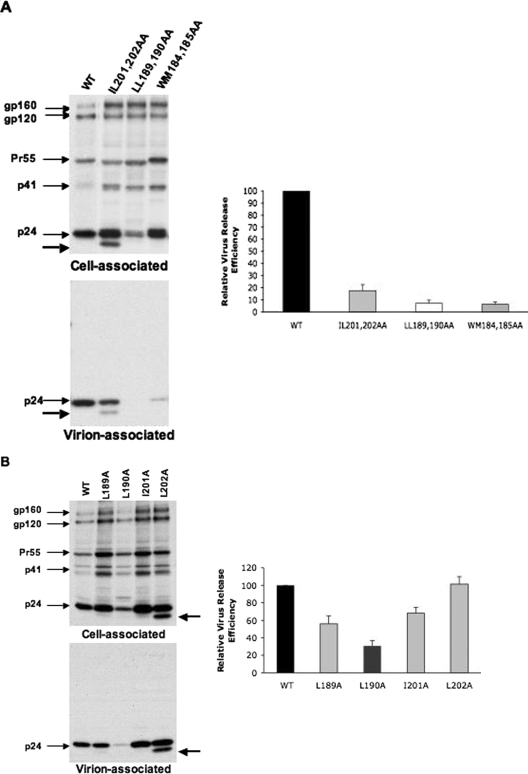
FIG. 2.
The CA mutants LL189,190AA and IL201,202AA are severely defective in virus assembly and release. HeLa cells were transiently transfected with WT pNL4-3 (WT) or derivatives containing the indicated (A) CA double or (B) CA single mutations. Transfected HeLa cells were metabolically labeled with [35S]Met/Cys, and virions were pelleted by ultracentrifugation. Cell and virion lysates were immunoprecipitated with HIV-Ig and analyzed by SDS-PAGE, followed by phosphorimager analysis. The positions of the Env precursor gp160, the mature surface Env glycoprotein gp120, the Gag precursor Pr55Gag, the Gag processing intermediate p41Gag, and p24 (CA) are shown. The ∼20-kDa protein product detected with the IL201,202AA and L202A mutants is indicated by an unmarked arrow. Virus release efficiency was calculated from at least three independent experiments as the amount of radiolabeled, virion-associated Gag as a fraction of the total (cell plus virion) radiolabeled Gag protein detected. Error bars show standard deviations.
To evaluate further the roles of Leu189, Leu190, Ile201, and Leu202 in virus assembly and release, we individually mutated each of these residues to Ala. As depicted in Fig. 2B, the CA single mutants also showed various degrees of defects in virus assembly and release, with the L190A mutant being the most compromised. Interestingly, the CA single mutants also exhibited defects in Gag processing. In addition, the smaller protein product associated with the CA double mutant IL201,202AA (Fig. 2A) could also be seen with the L202A mutant (Fig. 2B). These data thus indicate that both the CA double and single mutants exhibit various degrees of defects in HIV assembly and release, with the double mutants being affected more severely than the single mutants.
The IL201,202AA and L202A mutants display aberrant CA processing.
As shown in Fig. 2A and B, a protein product smaller than 24 kDa was associated with the CA double mutant IL201,202AA and the single mutant L202A. A similar phenotype has been observed previously upon mutation of buried hydrophobic residues in the CA NTD (e.g., F40A) (47). It is possible that mutation of CA202 induces protein misfolding, thereby exposing normally buried sites within CA to cleavage by PR. To determine whether the <24-kDa protein product is CA related, lysates from cells transfected with the LL189,190AA, IL201,202AA, and WM184,185AA double mutants or the L202A single mutant were subjected to SDS-PAGE and probed with either anti-p24 Ab or HIV-Ig. In both instances, a protein product of approximately 20 kDa was visible only for the double mutant IL201,202AA and the single mutant L202A (data not shown), indicating that this species is indeed CA related. To further address the issue of abnormal processing of CA by PR, we generated PR− versions of the CA double and single mutants and performed immunoprecipitations following transfection of HeLa cells with these PR− clones. The PR− versions of the IL201,202AA and L202A mutants no longer showed the presence of the ∼20-kDa CA-related product (data not shown). These data support the hypothesis that mutation of CA202 induces CA misfolding and aberrant processing by PR. VLP release in the context of the PR− clones was also severely inhibited by the LL189,190AA, IL201,202AA, and WM184,185AA double mutants (data not shown).
Since the IL201,202AA and L202A mutants appeared to show defects in CA protein folding, we sought to examine whether these mutants would release non-particle-associated CA from virus-expressing cells. To examine this possibility, HeLa cells were transfected with the LL189,190AA, IL201,202AA, and WM184,185AA double mutants or the L202A single mutant. Following metabolic radiolabeling, released virus was pelleted by ultracentrifugation and both the virus pellet lysate and the postvirus supernatant were immunoprecipitated with HIV-Ig (Fig. 3). Cells transfected with WT pNL4-3 or the LL189,190AA or WM184,185AA double mutant released barely detectable levels of non-particle-associated CA. In contrast and interestingly, the IL201,202AA mutant released large amounts of non-particle-associated p24 (CA). The L202A mutant released lower, but detectable, levels of free CA. Overall, these findings suggest that the severe assembly defects observed for the IL201,202AA mutant could be a consequence of CA misfolding and/or instability.
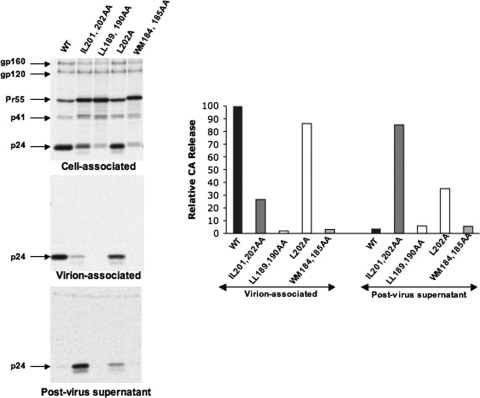
FIG. 3.
Mutation of L202 leads to the release of free CA and CA-derived product. HeLa cells transfected with WT pNL4-3 or the indicated CA mutants were metabolically labeled with [35S]Met/Cys, and virions were pelleted by ultracentrifugation. Cell- and virion-associated material and the supernatant obtained from virus pelleting (postvirus supernatant) were immunoprecipitated with HIV-Ig and analyzed by SDS-PAGE, followed by phosphorimager quantification. Results from one representative experiment are shown; similar results were obtained in an independently repeated experiment.
Replication kinetics of the CA mutants in T-cell lines; isolation of viral revertants.
To characterize further the phenotypes of the CA mutants, virus replication kinetics were evaluated in T-cell lines. This analysis revealed that neither of the CA double mutants showed detectable replication, even when transfected cultures were maintained for 3 months (data not shown). We also observed that none of the single mutants was able to replicate in Jurkat T cells (data not shown). In contrast, in the more permissive MT-4 T-cell line, the L189A, L190A, and I201A single mutants all replicated, albeit with markedly delayed kinetics. The L202A mutant did not replicate in MT-4 cells (Fig. 4A).
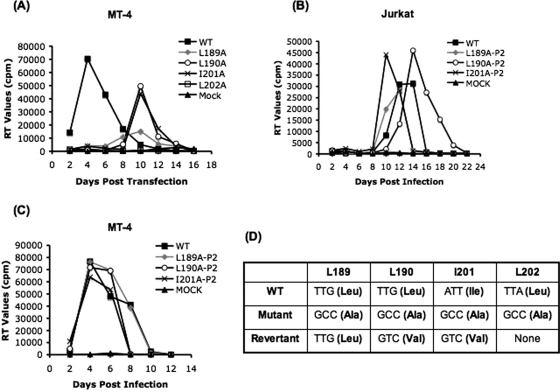
FIG. 4.
Replication kinetics of CA mutants and isolation of viral revertants. (A) MT-4 T cells were transfected with either WT pNL4-3 or the indicated CA single mutants. Cells were split every 2 days; supernatant was reserved at each time point for RT analysis. Repassage of CA single mutants and isolation of viral revertants. (B) Jurkat and (C) MT-4 T-cell lines were infected with WT or CA mutant virus (normalized for RT activity) obtained from the MT-4 replication experiment shown in panel A. Culture supernatant was reserved every 2 days for RT analysis. Cell pellets were also harvested for DNA isolation, PCR amplification, and sequencing of the gag gene. P2 after the mutant name denotes passage 2. (D) Sequence profile of revertant viruses obtained from repassage of CA single mutants in Jurkat and MT-4 cells. The codons found in WT pNL4-3, the mutant derivatives, and the PCR-amplified viral DNA are indicated; the encoded amino acid is indicated in parentheses. For L202, no revertants were obtained (none).
To determine whether the delayed replication observed for the L189A, L190A, and I201A single mutants represented the emergence of viral revertants, virus stocks were collected during the peak of RT activity and used to reinfect both Jurkat and MT-4 cells. The CA single mutants that were previously unable to grow in Jurkat cells now replicated with kinetics comparable to those of the WT (Fig. 4B). In MT-4 cells, the repassaged viruses replicated with WT kinetics (Fig. 4C). These results suggested that the CA single mutants reverted during their passage in MT-4 cells. To confirm this prediction and to characterize potential revertants, viral DNA was isolated from infected Jurkat and MT-4 cells. The Gag coding region was then amplified by PCR and sequenced. The sequencing data indicated that the L189A mutant had reverted back to the WT codon (Leu), whereas the L190A and I201A mutants acquired a Val codon in place of the original mutant (Ala) codon (Fig. 4D). The same changes were identified upon sequencing of viral DNAs derived from both Jurkat and MT-4 cells (data not shown). Viruses carrying the L190V or I201V mutation were assembled and released as efficiently as the WT and also showed WT replication kinetics in Jurkat T cells (data not shown). Single-cycle infectivity data indicated that the CA double mutants and the L189A, L190A, I201A, and L202A mutants were severely defective while the L190V and I201V mutants displayed WT infectivity (data not shown). These findings indicate that the L189L190 and I201L202 motifs are not required for full Gag function in that L190 and I201 can be replaced with Val with no adverse effects on assembly, release, or virus replication.
Immunofluorescence microscopy and EM reveal distinct defects for the IL201,202AA and LL189,190AA mutants.
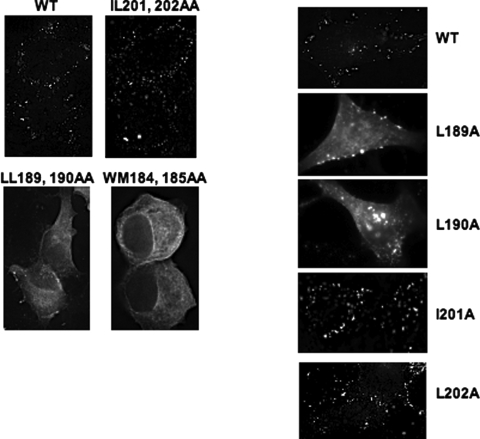
To more rigorously elucidate the mechanism by which mutation of CA residues 189, 190, 201, and 202 induced defects in virus production and replication, HeLa cells were transfected in parallel with WT or mutant molecular clones and stained with anti-p17 (MA) Ab. Immunofluorescence analysis revealed that cells transfected with WT pNL4-3 or the IL201,202AA mutant showed cell surface punctate staining typical of WT Gag (Fig. 5). In contrast, the LL189,190AA mutant showed an accumulation of cytosolic, non-plasma membrane-associated Gag with a lack of cell surface puncta. This pattern of Gag staining was suggestive of a defect in Gag membrane binding and/or multimerization. For comparison, we analyzed the CA CTD dimer interface mutant WM184,185AA. Mutation of CA residue 184 or 185 has previously been shown to induce a severe defect in CA-CA and Gag-Gag interactions (18, 49) and also to impair membrane binding (39). The WM184,185AA mutant showed a pattern of Gag staining essentially indistinguishable from that of LL189,190AA (Fig. 5). We also determined the Gag localization pattern of the single mutants. The L189A and L190A mutants showed diffuse cytosolic staining similar to that of the LL189,190AA double mutant and the presence of some cell surface punctate staining. The I201A and L202A mutants exhibited a WT localization pattern, with a considerable amount of cell surface punctate Gag staining (Fig. 5). Overall, the immunofluorescence data suggested that mutation of CA189 and CA190 induced defects in Gag multimerization and/or membrane binding.
FIG. 5.
Subcellular localization of CA-mutant Gag. HeLa cells were transfected with WT pNL4-3 or the indicated CA-mutant derivatives. Cells were fixed 24 h posttransfection and stained with anti-HIV-1 p17 (MA) Ab followed by fluorescence microscopy with a DeltaVision RT microscope.
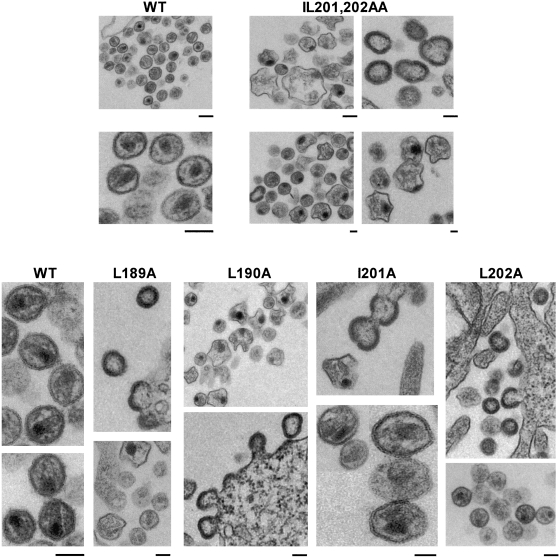
To examine the effects of the CA mutations on virus particle morphology, we transfected HeLa cells with WT pNL4-3 or the CA mutant derivatives and performed thin-section transmission EM. pNL4-3-transfected cultures showed large numbers of released virus particles, many of which contained conical, condensed cores and which were ∼100 to 120 nm in diameter (Fig. 6). In contrast, virus particles produced by the IL201,202AA mutant were highly distorted, were morphologically immature or possessed acentric cores, and were abnormally large (approximately 300 to 400 nm in diameter). No budding or released virus particles were observed for the LL189,190AA mutant. EM analysis of the CA single mutants also showed gross defects in particle morphology and a failure to form conical cores (Fig. 6).
FIG. 6.
EM analysis of CA-mutant virions. HeLa cells were transfected with WT pNL4-3 or the indicated CA-mutant derivatives. Cells were fixed at 2 days posttransfection and observed by transmission EM. The bar under each panel represents a 100-nm scale.
Assembly and release of the IL201,202AA but not the LL189,190AA mutant can be efficiently rescued by coexpression with WT Gag.
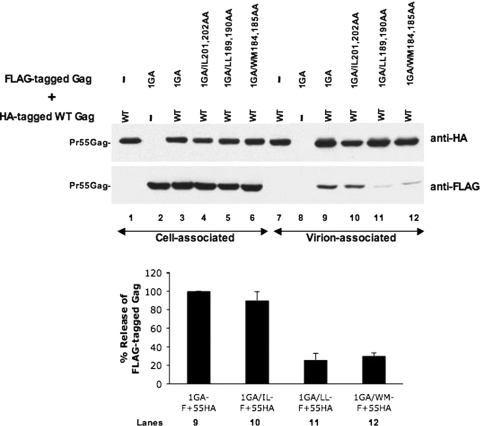
The immunofluorescence analysis presented above suggested that the LL189,190AA mutant may be defective for Gag multimerization. To investigate this possibility directly, we used a previously described assay (2, 34) to evaluate the ability of CA mutant Gag to be rescued upon coexpression with WT Gag. In this assay, Gag mutants are analyzed in the context of a C-terminally epitope-tagged, nonmyristylated (1GA) (16) pNL4-3 derivative. Because of the elimination of the N-terminal myristylation signal, these Gags are unable to bind membrane and assemble when expressed alone. However, upon coexpression with WT Gag bearing a different C-terminal epitope tag, the nonmyristylated Gag protein is incorporated into virions via coassembly with WT Gag. The extent to which the nonmyristylated mutant Gag protein is recovered in VLP fractions provides a measurement of the efficiency with which it can engage in Gag-Gag interactions. To perform this analysis, FLAG-tagged versions of pNL4-3/1GA expressing either WT or mutant CA were cotransfected with a pNL4-3 derivative encoding HA-tagged WT Gag. Rescue of the mutant (FLAG-tagged) Gag protein by the WT (HA-tagged) Gag protein was then monitored by quantitative immunoblotting with anti-HA and anti-FLAG Abs. The results indicated that no particle-associated, FLAG-tagged Pr55Gag was released upon single expression of 1GA Gag (Fig. 7, lane 8). However, coexpression with HA-tagged WT Gag led to efficient rescue of FLAG-tagged 1GA Gag in the virion-associated fraction (Fig. 7, lane 9). Similarly, the 1GA/IL201,202AA mutant could be rescued almost as efficiently as the 1GA Gag protein containing WT CA (Fig. 7, lane 10). In contrast, the 1GA/LL189,190AA and 1GA/WM184,185AA mutants displayed three- to fivefold reductions in the ability to coassemble with WT Gag and be recovered in virion-associated fractions (Fig. 7, lanes 11 and 12). These data reinforce our immunofluorescence findings and confirm that the LL189,190AA mutant is defective in Gag-Gag interactions to an extent similar to that of the WM184,185AA CTD dimer interface mutant. Interestingly, all of the CA single mutants could be efficiently rescued (data not shown), consistent with their less severe phenotype relative to the double mutants.
FIG. 7.
Analysis of the rescue of CA mutant Gag into VLPs upon coexpression with WT Gag. HeLa cells were transfected with FLAG-tagged versions of either 1GA/pNL4-3 or 1GA/pNL4-3 derivatives containing the indicated CA double mutations. Rescue of the nonmyristylated (1GA) Gag protein into VLPs was measured by cotransfection with WT HA-tagged Gag. Cell and virus lysates were subjected to SDS-PAGE, followed by immunoblotting with anti-HA or anti-FLAG Ab. Quantification of Pr55Gag bands was performed with the Alpha Innotech digital imaging system. Virus release efficiency was calculated as the percent release of FLAG-tagged 1GA Gag, with the amount of FLAG-tagged 1GA Gag containing WT CA rescued into VLPs by HA-tagged Gag being given a relative value of 100%. Lane numbers under the graph correspond to the respective lanes in the anti-FLAG Western blot. In the graph, 1GA-F, 1GA/IL-F, 1GA/LL-F, and 1GA/WM-F refer to FLAG-tagged 1GA Gag proteins containing WT, IL201,202AA, LL189,190AA, and WM184,185AA CA, respectively. 55HA denotes HA-tagged WT Gag. Data represent averages from at least three independent experiments ± the standard deviations.
The LL189,190AA mutant is defective in membrane binding.
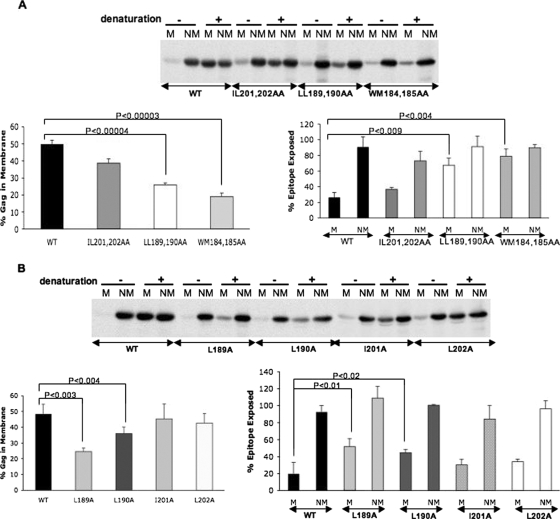
We previously demonstrated that the WM184,185AA dimer interface mutant displays a defect in membrane binding that is most likely linked to its inability to multimerize efficiently (39). The similarities observed here between the WM184,185AA and LL189,190AA mutants in Gag localization and Gag multimerization suggested that the LL189,190AA mutant was also defective in membrane binding, as reported previously (27). To evaluate the ability of the CA double mutants to bind membrane, we performed membrane flotation assays with sucrose gradients (Fig. 8A). This analysis indicated that while the IL201,202AA mutant Gag protein showed membrane binding ability similar to that of the WT, the amount of LL189,190AA Gag recovered in membrane fractions was approximately half that of WT Gag. The membrane binding ability of the WM184,185AA mutant was likewise reduced approximately twofold (Fig. 8A).
FIG. 8.
Effects of CA mutations on membrane binding and Gag multimerization. HeLa cells were transfected with pNL4-3/PR− versions of the (A) CA double or (B) CA single mutants. Transfected cells were pulse-chase labeled with [35S]Met/Cys and sonicated, and postnuclear supernatants were subjected to membrane flotation centrifugation on sucrose gradients (see Materials and Methods). The membrane (M) and nonmembrane (NM) fractions were isolated and immunoprecipitated with HIV-Ig either with (+) or without (−) prior denaturation. Pr55Gag bands were quantified by phosphorimager analysis. The graphs in the lower left of panels A and B indicate membrane binding efficiency. Percent epitope exposure (lower right), which is a measurement of higher-order Gag multimerization (39), was determined by calculating the percentage of Gag recovered by immunoprecipitation from nondenatured relative to denatured fractions. Note that epitope masking is observed predominantly with membrane-associated Gag. P values represent significant differences between the WT and mutant Gag proteins as determined by the Student t test.
We previously described an assay that measures higher-order, in vivo Gag multimerization (39). This assay is based on the observation that membrane-bound HIV-1 Gag is poorly detected by immunoprecipitation prior to denaturation. Thus, comparison of the amount of Gag detected with versus without denaturation (i.e., the amount of Gag that is “epitope exposed”) provides a measure of higher-order Gag multimerization. As part of our membrane flotation analysis presented above, we also determined the degree of epitope exposure for the CA double mutants. Consistent with the VLP rescue assay shown in Fig. 7, we observed that the IL201,202AA mutant displayed a level of epitope masking (i.e., higher-order Gag multimerization) similar to that of WT in the membrane fraction (Fig. 8A). In contrast, the LL189,190AA double mutant showed significantly increased epitope exposure in the membrane fraction, confirming its defect in Gag multimerization. Consistent with our earlier data (39) and similar to the LL189,190AA double mutant, a high percentage of WM184,185AA mutant Gag could be immunoprecipitated from the membrane-bound fractions without denaturation (Fig. 8A).
We also applied the membrane flotation and epitope exposure assays to the CA single mutants. While the I201A and L202A mutants did not show statistically significant defects, the L189A mutant displayed a significant reduction in membrane binding (Fig. 8B) and a small, but significant, increase in epitope exposure. The L190A mutant exhibited an intermediate phenotype, with a small reduction in membrane binding (Fig. 8B). Altogether, the membrane flotation and epitope masking data presented in Fig. 8 reinforce the results of our immunofluorescence microscopy and Gag rescue assays and indicate that the CA mutant LL189,190AA but not IL201,202AA is defective in Gag multimerization and membrane binding.
The IL-201,202AA mutant does not disrupt MVB targeting.
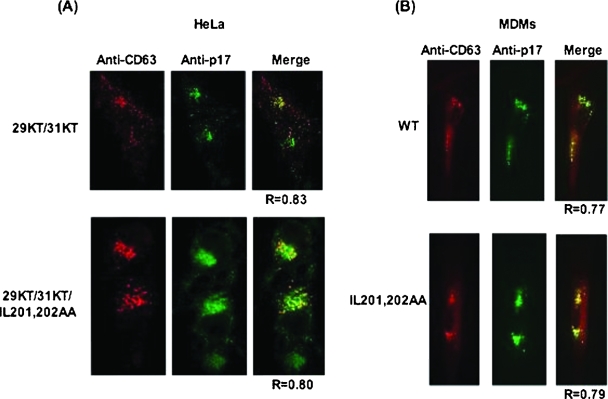
The immunofluorescence data presented in Fig. 6 indicate that the pattern of Gag localization for IL201,202AA is indistinguishable from that of the WT, suggesting that this mutant is not defective in Gag trafficking. However, since it was suggested that the I201/L202 di-Leu-like motif may play an important role in the association of Gag with endocytic machinery (27), we sought to determine whether the IL201,202AA mutation would affect the MVB localization of Gag. As mentioned in the introduction, we have previously demonstrated that mutations in the highly basic domain of MA (e.g., 29KT/31KT) induce the relocalization of Gag to a CD63+ late endosomal or MVB compartment (36, 38). If the I201/L202 di-Leu-like motif were important for the association of Gag with MVB, we would predict that the IL201,202AA mutation would disrupt MVB localization in the context of 29KT/31KT-mutant MA. To test this hypothesis, we constructed a pNL4-3 derivative containing both the MA (29KT/31KT) and CA (IL201,202AA) mutations. HeLa cells were transfected with this molecular clone or with the 29KT/31KT MA mutant clone. Transfected cells were fixed and costained with anti-p17 (MA) and anti-CD63 Abs (Fig. 9A). As we reported previously (36), the 29KT/31KT MA-mutant Gag protein largely localized to the MVB (R = 0.82 ± 0.05) (Fig. 9A, top). Interestingly, the CA IL201,202AA mutation did not significantly alter Gag localization to the MVB (R = 0.80 ± 0.04) (Fig. 9A, bottom). We also introduced the CA LL189,190AA mutation into the 29KT/31KT MA mutant clone. This mutant showed a cytosolic distribution of Gag (data not shown) similar to that seen with the LL189,190AA mutant (Fig. 5), consistent with the defect in membrane binding imposed by disruption of the L189/L190 motif.
FIG. 9.
MVB localization of the IL201,202AA CA mutant. (A) HeLa cells were transfected with pNL4-3/29KT/31KT (top) or with the derivative containing the IL201,202AA CA mutation (bottom). (B) WT NL4-3 or the IL201,202AA CA-mutant derivative was expressed in primary human MDMs (Materials and Methods). Cells were fixed 24 h posttransfection, permeabilized, and stained with anti-HIV-1 p17 (MA) (green) and anti-CD63 (red) Abs, followed by microscopy with a DeltaVision RT microscope. The softWoRx colocalization module was used to determine costaining between Gag and CD63 on an average of 8 to 10 cells. One representative field for each is depicted. When data were compiled and averaged for all of the cells analyzed, the Gag/CD63 colocalization in the context of WT CA was R = 0.82 ± 0.05 in HeLa cells and R = 0.7028 ± 0.17 in MDMs; for the IL201,202AA CA mutant, it was R = 0.80 ± 0.04 in HeLa cells and 0.6826 ± 0.102 in MDMs. R = Pearson coefficient of correlation.
To extend these findings to a cell type in which WT Gag is predominantly MVB targeted, we examined the localization of WT Gag or the IL201,202AA mutant in primary human MDMs (Fig. 9B). Again, we found no difference in the CD63 colocalization of the IL201,202AA mutant relative to that of the WT (R = 0.7028 ± 0.17 for WT and 0.6826 ± 0.102 for IL201,202AA). Together, these findings indicate that disruption of the I201/L202 di-Leu-like motif does not prevent the targeting of Gag to the late endosome or MVB.
DISCUSSION
In this study, we demonstrate that mutations in the putative di-Leu-like motifs in the HIV-1 CA CTD lead to severe defects in virus assembly and release, virion maturation, and infectivity. Several of the single-amino-acid mutants reverted in culture; analysis of these revertants indicated that Leu-to-Val substitutions at both residues 190 and 201 were well tolerated, despite the fact that these mutations disrupt the integrity of the di-Leu-like motifs. Finally, the IL201,202AA mutant efficiently localized to the MVB in both HeLa cells and MDMs, thus demonstrating that the I201/L202 di-Leu-like motif is not required for MVB targeting.
The defects induced by the mutations described here emphasize the role of the CA CTD dimer interface in both virus particle production and virion maturation. Structural modeling revealed that while residue I201 is surface exposed, L202 is buried (Fig. 1). We therefore speculate that mutation of the I201/L202 motif disrupted normal CA folding, making a previously buried site(s) accessible for cleavage by PR. Interestingly, the L202A mutation did not significantly affect virus particle production but impaired virion maturation, virus replication, and infectivity, suggesting that mutation of L202 predominantly affected CA-CA interactions following particle release, rather than Gag-Gag interactions during assembly. We note that aberrant cleavage of CA by PR has also been observed upon mutation of buried residues in the CA NTD (47). The release of free CA observed with mutants containing substitutions in CA residue 202 (Fig. 3) may result from a loss of virion stability following aberrant cleavage of the mutant CA by PR.
The double mutant LL189,190AA showed clear defects in Gag membrane binding and multimerization, consistent with previous results (27). Residue L189 lies in the CA dimer interface (Fig. 1), a domain whose function in promoting CA-CA interactions both in vivo and in vitro is well established (8, 11, 18, 19, 28, 49). The dimer interface forms a hydrophobic core that includes residues Val181, Trp184, Met185, and Leu189 (18, 19, 49). Consistent with these observations, the L189A single mutant showed more-severe defects in membrane binding than did the L190A mutant (Fig. 9B). These defects can be attributed to the fact that the CTD dimer interface links the hexameric rings formed by the CA NTD into a continuous lattice and that disruption of residues in this domain impair CA-CA interactions (25). Our data obtained with both the LL189,190AA and WM184,185AA mutants support the proposal of von Schwedler and coworkers (49) that the CA CTD dimer interface is important not only for CA-CA interactions during core maturation but also for Gag-Gag interactions during particle assembly.
The role of Gag multimerization in Gag-membrane binding remains to be fully elucidated. Several studies comparing full-length and C-terminally truncated Gag constructs suggested that Gag multimerization mediated by NC may enhance membrane binding (41, 44). By using membrane flotation centrifugation techniques that separate membrane-associated Gag from non-membrane-bound Gag oligomers, we observed that a Gag truncation mutant containing only MA and the CA NTD displayed efficient membrane binding at steady state (34) but reduced membrane binding kinetics (37). Mutations in SP1 that inhibit Gag multimerization have also been reported to reduce membrane binding (26), and dimerization of the Rous sarcoma virus MA increased its capacity to associate with membrane in vitro (6). Structural studies with HIV-1 MA have demonstrated that low-order multimerization leads to greater exposure of the myristate moiety, thereby presumably increasing membrane binding ability (46). We recently reported that a severely assembly-deficient mutant lacking all of the basic residues in NC displayed WT levels of membrane binding whereas the WM184,195AA CA dimer interface mutant was membrane binding defective (39). The data presented here that were obtained with the LL189,190AA mutant support and extend our proposal that Gag-Gag contacts mediated by the CA CTD, but not higher-order Gag assembly mediated by the basic residues in NC, are required for efficient and stable binding of Gag to membrane.
As mentioned in the introduction, Lindwasser and Resh (27) observed that the IL201,202AA mutation severely disrupted VLP production and reduced MVB localization of Gag in COS cells treated with the drug U18666A. They proposed that the defect in particle production exhibited by this mutant Gag resulted from its failure to traffic to the MVB. Alternatively, they suggested that the effect of the IL201,202AA mutation on virus assembly and release might be distinct from a role in Gag trafficking. Our data support the latter hypothesis. The results presented here suggest that the impaired virus particle production induced by mutations in the L189/L190 and I201/L202 motifs is due to a diversity of defects in Gag-membrane binding, multimerization, and folding. We also find that MVB localization of IL201,202AA Gag is not affected in HeLa cells in the context of a MA mutation that disrupts plasma membrane targeting (Fig. 9A) or in MDMs in the context of otherwise WT Gag (Fig. 9B). Finally, the fact that Leu-to-Val substitutions at both residues L190 and I201 are well tolerated argues against a requirement in HIV-1 replication for di-Leu-like motifs at these positions (1).
Our results do not exclude the possibility that the CA CTD di-Leu-like motifs could promote the internalization of CD4-Gag chimeras, nor do they contradict the observation that these motifs affect Gag endocytosis in cells treated with U18666A (27). However, our data suggest that these motifs do not play a major role in Gag trafficking under physiological conditions. Ongoing studies will attempt to define further the viral determinants involved in Gag targeting and elucidate the cellular machinery that regulates the localization of HIV-1 assembly.
Acknowledgments
We thank S. Ablan for expert technical assistance, Ferri Soheilian for EM support, and Vineet KewalRamani, Judith Levin, Akira Ono, and members of the Freed lab for critical reviews of the manuscript. We also thank S. R. Durell for providing the CA CTD model shown in Fig. 1.
The HIV-Ig and anti-p24 Abs were obtained through the AIDS Research and Reference Reagent Program, Division of AIDS, NIAID, NIH. This research was supported by the Intramural Research Program of the Center for Cancer Research, National Cancer Institute, NIH, and by the Intramural AIDS Targeted Antiviral Program. This project was funded in part with federal funds from the National Cancer Institute, NIH, under contract N01-CO-12400.
The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the U.S. Government.
REFERENCES
- 1.Bonifacino, J. S., and L. M. Traub. 2003. Signals for sorting of transmembrane proteins to endosomes and lysosomes. Annu. Rev. Biochem. 72:395-447. [DOI] [PubMed] [Google Scholar]
- 2.Burniston, M. T., A. Cimarelli, J. Colgan, S. P. Curtis, and J. Luban. 1999. Human immunodeficiency virus type 1 Gag polyprotein multimerization requires the nucleocapsid domain and RNA and is promoted by the capsid-dimer interface and the basic region of matrix protein. J. Virol. 73:8527-8540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Campbell, S., and A. Rein. 1999. In vitro assembly properties of human immunodeficiency virus type 1 Gag protein lacking the p6 domain. J. Virol. 73:2270-2279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Campbell, S., and V. M. Vogt. 1995. Self-assembly in vitro of purified CA-NC proteins from Rous sarcoma virus and human immunodeficiency virus type 1. J. Virol. 69:6487-6497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Cimarelli, A., S. Sandin, S. Hoglund, and J. Luban. 2000. Basic residues in human immunodeficiency virus type 1 nucleocapsid promote virion assembly via interaction with RNA. J. Virol. 74:3046-3057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Dalton, A. K., P. S. Murray, D. Murray, and V. M. Vogt. 2005. Biochemical characterization of Rous sarcoma virus MA protein interaction with membranes. J. Virol. 79:6227-6238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Dawson, L., and X. F. Yu. 1998. The role of nucleocapsid of HIV-1 in virus assembly. Virology 251:141-157. [DOI] [PubMed] [Google Scholar]
- 8.del Alamo, M., J. L. Neira, and M. G. Mateu. 2003. Thermodynamic dissection of a low affinity protein-protein interface involved in human immunodeficiency virus assembly. J. Biol. Chem. 278:27923-27929. [DOI] [PubMed] [Google Scholar]
- 9.Demirov, D. G., and E. O. Freed. 2004. Retrovirus budding. Virus Res. 106:87-102. [DOI] [PubMed] [Google Scholar]
- 10.Dong, X., H. Li, A. Derdowski, L. Ding, A. Burnett, X. Chen, T. R. Peters, T. S. Dermody, E. Woodruff, J. J. Wang, and P. Spearman. 2005. AP-3 directs the intracellular trafficking of HIV-1 Gag and plays a key role in particle assembly. Cell 120:663-674. [DOI] [PubMed] [Google Scholar]
- 11.Dorfman, T., A. Bukovsky, A. Ohagen, S. Hoglund, and H. G. Gottlinger. 1994. Functional domains of the capsid protein of human immunodeficiency virus type 1. J. Virol. 68:8180-8187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ehrlich, L. S., B. E. Agresta, and C. A. Carter. 1992. Assembly of recombinant human immunodeficiency virus type 1 capsid protein in vitro. J. Virol. 66:4874-4883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Freed, E. O. 1998. HIV-1 Gag proteins: diverse functions in the virus life cycle. Virology 251:1-15. [DOI] [PubMed] [Google Scholar]
- 14.Freed, E. O., G. Englund, and M. A. Martin. 1995. Role of the basic domain of human immunodeficiency virus type 1 matrix in macrophage infection. J. Virol. 69:3949-3954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Freed, E. O., and M. A. Martin. 1994. Evidence for a functional interaction between the V1/V2 and C4 domains of human immunodeficiency virus type 1 envelope glycoprotein gp120. J. Virol. 68:2503-2512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Freed, E. O., J. M. Orenstein, A. J. Buckler-White, and M. A. Martin. 1994. Single amino acid changes in the human immunodeficiency virus type 1 matrix protein block virus particle production. J. Virol. 68:5311-5320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Gamble, T. R., F. F. Vajdos, S. Yoo, D. K. Worthylake, M. Houseweart, W. I. Sundquist, and C. P. Hill. 1996. Crystal structure of human cyclophilin A bound to the amino-terminal domain of HIV-1 capsid. Cell 87:1285-1294. [DOI] [PubMed] [Google Scholar]
- 18.Gamble, T. R., S. Yoo, F. F. Vajdos, U. K. von Schwedler, D. K. Worthylake, H. Wang, J. P. McCutcheon, W. I. Sundquist, and C. P. Hill. 1997. Structure of the carboxyl-terminal dimerization domain of the HIV-1 capsid protein. Science 278:849-853. [DOI] [PubMed] [Google Scholar]
- 19.Ganser-Pornillos, B. K., U. K. von Schwedler, K. M. Stray, C. Aiken, and W. I. Sundquist. 2004. Assembly properties of the human immunodeficiency virus type 1 CA protein. J. Virol. 78:2545-2552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Gould, S. J., A. M. Booth, and J. E. Hildreth. 2003. The Trojan exosome hypothesis. Proc. Natl. Acad. Sci. USA 100:10592-10597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gross, I., H. Hohenberg, C. Huckhagel, and H. G. Kräusslich. 1998. N-terminal extension of human immunodeficiency virus capsid protein converts the in vitro assembly phenotype from tubular to spherical particles. J. Virol. 72:4798-4810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Gross, I., H. Hohenberg, and H. G. Kräusslich. 1997. In vitro assembly properties of purified bacterially expressed capsid proteins of human immunodeficiency virus. Eur. J. Biochem. 249:592-600. [DOI] [PubMed] [Google Scholar]
- 23.Huang, M., J. M. Orenstein, M. A. Martin, and E. O. Freed. 1995. p6Gag is required for particle production from full-length human immunodeficiency virus type 1 molecular clones expressing protease. J. Virol. 69:6810-6818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kiernan, R. E., A. Ono, G. Englund, and E. O. Freed. 1998. Role of matrix in an early postentry step in the human immunodeficiency virus type 1 life cycle. J. Virol. 72:4116-4126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Li, S., C. P. Hill, W. I. Sundquist, and J. T. Finch. 2000. Image reconstructions of helical assemblies of the HIV-1 CA protein. Nature 407:409-413. [DOI] [PubMed] [Google Scholar]
- 26.Liang, C., J. Hu, J. B. Whitney, L. Kleiman, and M. A. Wainberg. 2003. A structurally disordered region at the C terminus of capsid plays essential roles in multimerization and membrane binding of the Gag protein of human immunodeficiency virus type 1. J. Virol. 77:1772-1783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lindwasser, O. W., and M. D. Resh. 2004. Human immunodeficiency virus type 1 Gag contains a dileucine-like motif that regulates association with multivesicular bodies. J. Virol. 78:6013-6023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Melamed, D., M. Mark-Danieli, M. Kenan-Eichler, O. Kraus, A. Castiel, N. Laham, T. Pupko, F. Glaser, N. Ben-Tal, and E. Bacharach. 2004. The conserved carboxy terminus of the capsid domain of human immunodeficiency virus type 1 Gag protein is important for virion assembly and release. J. Virol. 78:9675-9688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Morita, E., and W. I. Sundquist. 2004. Retrovirus budding. Annu. Rev. Cell Dev. Biol. 20:395-425. [DOI] [PubMed] [Google Scholar]
- 30.Muriaux, D., J. Mirro, D. Harvin, and A. Rein. 2001. RNA is a structural element in retrovirus particles. Proc. Natl. Acad. Sci. USA 98:5246-5251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Nguyen, D. G., A. Booth, S. J. Gould, and J. E. Hildreth. 2003. Evidence that HIV budding in primary macrophages occurs through the exosome release pathway. J. Biol. Chem. 278:52347-52354. [DOI] [PubMed] [Google Scholar]
- 32.Nydegger, S., M. Foti, A. Derdowski, P. Spearman, and M. Thali. 2003. HIV-1 egress is gated through late endosomal membranes. Traffic 4:902-910. [DOI] [PubMed] [Google Scholar]
- 33.Ono, A., S. D. Ablan, S. J. Lockett, K. Nagashima, and E. O. Freed. 2004. Phosphatidylinositol (4,5) bisphosphate regulates HIV-1 Gag targeting to the plasma membrane. Proc. Natl. Acad. Sci. USA 101:14889-14894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ono, A., D. Demirov, and E. O. Freed. 2000. Relationship between human immunodeficiency virus type 1 Gag multimerization and membrane binding. J. Virol. 74:5142-5150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ono, A., and E. O. Freed. 1999. Binding of human immunodeficiency virus type 1 Gag to membrane: role of the matrix amino terminus. J. Virol. 73:4136-4144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ono, A., and E. O. Freed. 2004. Cell-type-dependent targeting of human immunodeficiency virus type 1 assembly to the plasma membrane and the multivesicular body. J. Virol. 78:1552-1563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Ono, A., and E. O. Freed. 2001. Plasma membrane rafts play a critical role in HIV-1 assembly and release. Proc. Natl. Acad. Sci. USA 98:13925-13930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ono, A., J. M. Orenstein, and E. O. Freed. 2000. Role of the Gag matrix domain in targeting human immunodeficiency virus type 1 assembly. J. Virol. 74:2855-2866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ono, A., A. A. Waheed, A. Joshi, and E. O. Freed. 2005. Association of human immunodeficiency virus type 1 Gag with membrane does not require highly basic sequences in the nucleocapsid: use of a novel Gag multimerization assay. J. Virol. 79:14131-14140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Pelchen-Matthews, A., B. Kramer, and M. Marsh. 2003. Infectious HIV-1 assembles in late endosomes in primary macrophages. J. Cell Biol. 162:443-455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Platt, E. J., and O. K. Haffar. 1994. Characterization of human immunodeficiency virus type 1 Pr55gag membrane association in a cell-free system: requirement for a C-terminal domain. Proc. Natl. Acad. Sci. USA 91:4594-4598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Platt, E. J., K. Wehrly, S. E. Kuhmann, B. Chesebro, and D. Kabat. 1998. Effects of CCR5 and CD4 cell surface concentrations on infections by macrophagetropic isolates of human immunodeficiency virus type 1. J. Virol. 72:2855-2864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Raposo, G., M. Moore, D. Innes, R. Leijendekker, A. Leigh-Brown, P. Benaroch, and H. Geuze. 2002. Human macrophages accumulate HIV-1 particles in MHC II compartments. Traffic 3:718-729. [DOI] [PubMed] [Google Scholar]
- 44.Sandefur, S., V. Varthakavi, and P. Spearman. 1998. The I domain is required for efficient plasma membrane binding of human immunodeficiency virus type 1 Pr55Gag. J. Virol. 72:2723-2732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Swanstrom, R., and J. W. Wills. 1997. Synthesis, assembly, and processing of viral proteins, p. 263-334. In J. M. Coffin, S. H. Hughes, and H. E. Varmus (ed.), Retroviruses. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. [PubMed]
- 46.Tang, C., E. Loeliger, P. Luncsford, I. Kinde, D. Beckett, and M. F. Summers. 2004. Entropic switch regulates myristate exposure in the HIV-1 matrix protein. Proc. Natl. Acad. Sci. USA 101:517-522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Tang, S., T. Murakami, B. E. Agresta, S. Campbell, E. O. Freed, and J. G. Levin. 2001. Human immunodeficiency virus type 1 N-terminal capsid mutants that exhibit aberrant core morphology and are blocked in initiation of reverse transcription in infected cells. J. Virol. 75:9357-9366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.von Schwedler, U. K., T. L. Stemmler, V. Y. Klishko, S. Li, K. H. Albertine, D. R. Davis, and W. I. Sundquist. 1998. Proteolytic refolding of the HIV-1 capsid protein amino-terminus facilitates viral core assembly. EMBO J. 17:1555-1568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.von Schwedler, U. K., K. M. Stray, J. E. Garrus, and W. I. Sundquist. 2003. Functional surfaces of the human immunodeficiency virus type 1 capsid protein. J. Virol. 77:5439-5450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Willey, R. L., J. S. Bonifacino, B. J. Potts, M. A. Martin, and R. D. Klausner. 1988. Biosynthesis, cleavage, and degradation of the human immunodeficiency virus 1 envelope glycoprotein gp160. Proc. Natl. Acad. Sci. USA 85:9580-9584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Worthylake, D. K., H. Wang, S. Yoo, W. I. Sundquist, and C. P. Hill. 1999. Structures of the HIV-1 capsid protein dimerization domain at 2.6 Å resolution. Acta Crystallogr. D Biol. Crystallogr. 55(Pt. 1):85-92. [DOI] [PubMed] [Google Scholar]
- 52.Yee, J. K., T. Friedmann, and J. C. Burns. 1994. Generation of high-titer pseudotyped retroviral vectors with very broad host range. Methods Cell Biol. 43(Pt. A):99-112. [DOI] [PubMed] [Google Scholar]