Abstract
To determine the prevalence and genotype of an extended-spectrum beta-lactamase and new chromosomal AmpC beta-lactamases among clinical isolates of Enterobacter species, we performed antibiotic susceptibility testing, pI determination, induction tests, transconjugation, enterobacterial repetitive consensus (ERIC) PCR, sequencing, and phylogenetic analysis. Among the 51 clinical isolates collected from a university hospital in Korea, 6 isolates have been shown to produce SHV-12 and inducible AmpC beta-lactamases. These also included three isolates producing TEM-1b and one strain carrying TEM-1b and CMY-type beta-lactamases with a pI of 8.0. The results from ERIC PCR revealed that six isolates were genetically unrelated, suggesting that dissemination of SHV-12 was responsible for the spread of resistance to extended-spectrum beta-lactams in Korea. Six genes of inducible AmpC beta-lactamases that are responsible for the resistance to cephamycins (cefoxitin and cefotetan), amoxicillin, cephalothin, and amoxicillin-clavulanic acid were cloned and characterized. A 1,165-bp DNA fragment containing the ampC genes was sequenced and found to have an open reading frame coding for a 381-amino-acid beta-lactamase. The nucleotide sequence of four ampC genes (blaEcloK992004.1, blaEcloK995120.1, blaEcloK99230, and blaEareK9911729) shared considerable homology with that of AmpC-type class C beta-lactamase genes of gram-negative bacteria, especially that of the chromosomal ampC gene (blaEcloMHN1) of Enterobacter cloacae MHN1 (99.9, 99.7, 99.6, and 99.6% identity, respectively). The sequences of two ampC genes (blaEcloK9973 and blaEcloK9914325) showed close similarity to the chromosomal ampC gene (blaEcloQ908R) of E. cloacae Q908R (99.7% identity). The results from phylogenetic analysis suggested that six ampC genes could originate from blaEcloMHN1 or blaEcloQ908R.
The beta-lactamases (EC 3.5.2.6) produced by the bacterium have been known to protect against the lethal effect of penicillins, cephalosporins, or monobactams on cell-wall synthesis. A variety of beta-lactamases have been classified into classes A, B, C, and D according to their amino acid homologies (4). AmpC beta-lactamases, mostly conferring resistance to many beta-lactam antibiotics (cephamycins and broad-spectrum cephalosporins), are included in class C beta-lactamases. Chromosomally encoded AmpC beta-lactamases are present in Enterobacter spp., Shigella spp., Serratia marcescens, Citrobacter freundii, Morganella morganii, Providencia spp., Pseudomonas aeruginosa, and Escherichia coli (16). Enterobacter species are becoming increasingly important as nosocomial pathogens (22). Risk factors for nosocomial Enterobacter infection include the prior use of antimicrobial agents, a prolonged hospital stay, a serious underlying illness, immunosuppression, and the presence of a foreign device (2). In view of the spreading risk of AmpC resistance determinants among enterobacterial isolates, it is necessary to elucidate the AmpC resistance mechanism. The present study was conducted to determine the prevalence and genotypes of an extended-spectrum beta-lactamase (ESBL) and new AmpC beta-lactamases among clinical isolates of Enterobacter spp. in South Korea. We constructed a phylogeny of ampC beta-lactamases, paying particular attention to the Enterobacter species-related ampC genes.
MATERIALS AND METHODS
Bacterial strains.
Five strains of Enterobacter cloacae and one strain of Enterobacter aerogenes, producing the new AmpC beta-lactamases, were isolated at Kosin Medical Center (Pusan, South Korea) during a 12-month period from June 1998 to May 1999, and their profiles were presented in Table 1. The isolates were identified by using the conventional techniques (6) and/or the Vitek GNI card (bioMérieux Vitek Inc., Hazelwood, Mo.). E. coli DH5α was used as the host strain for transformation, and E. coli J53 AzideR (11) was used as the recipient strain for transfer experiments by conjugation. E. coli ATCC 25922 was used as the MIC reference strain.
TABLE 1.
Profiles of six clinical isolates, their transformants, and host (E. coli DH5α) for transformationa
| Strain | Species | Type of specimen | Ward | Type of infection | MIC (mg/liter) of beta-lactam:
|
pI(s) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AMX | AUG | LOT | TAX | TAZ | ATM | FEP | IMP | MER | FOX | TTN | ||||||
| K9973 | E. cloacae | Urine | Nephrology | UTI | >256 | 128 | >256 | 32 | 256 | 256 | 4 | 0.25 | 0.25 | 256 | 256 | 6.0, 7.6, 8.9 |
| trf K9973 | E. coli | 128 | 128 | >256 | 8 | 16 | 4 | 0.125 | 0.25 | <0.06 | 8 | 4 | 8.9 | |||
| K99230 | E. cloacae | Sputum | Hepatobiliary | Pneumonia | >256 | 256 | >256 | 32 | 128 | 128 | 2 | 0.5 | 0.125 | 128 | 256 | 6.0, 7.6, 8.9 |
| trf K99230 | E. coli | 128 | 128 | >256 | 32 | 16 | 8 | 0.25 | 0.5 | <0.06 | 16 | 8 | 8.9 | |||
| K992004.1 | E. cloacae | Pus | GS | Wound infection | >256 | 256 | >256 | 16 | 256 | 256 | 4 | 0.25 | 0.125 | 256 | 256 | 7.6, 8.9 |
| trf K992004.1 | E. coli | 16 | 16 | 32 | 1 | 0.5 | 0.5 | 0.125 | 0.25 | <0.06 | 2 | 1 | 8.9 | |||
| K995120.1 | E. cloacae | Pus | NS | Wound infection | >256 | 128 | >256 | 32 | 128 | 128 | 2 | 1 | 0.125 | 256 | 256 | 7.6, 8.9 |
| trf K995120.1 | E. coli | 16 | 16 | 32 | 1 | 0.5 | 0.5 | 0.125 | 0.25 | <0.06 | 2 | 1 | 8.9 | |||
| K9911729 | E. aerogenes | Sputum | CS | Pneumonia | >256 | >256 | >256 | 32 | 64 | 128 | 1 | 0.5 | 0.25 | >256 | 128 | 6.0, 7.6, 8.0, 8.9 |
| trf K9911729 | E. coli | 128 | 128 | >256 | 32 | 32 | 16 | 0.25 | 0.5 | <0.06 | 16 | 16 | 8.9 | |||
| K9914325 | E. cloacae | Urine | O&G | UTI | >256 | 128 | >256 | 32 | 128 | 128 | 4 | 0.5 | 0.125 | 128 | 64 | 7.6, 8.9 |
| trf K9914325 | E. coli | 64 | 64 | >256 | 16 | 16 | 4 | 0.25 | 0.25 | <0.06 | 8 | 4 | 8.9 | |||
| DH5α | E. coli | 1 | 1 | 2 | <0.06 | 0.125 | <0.06 | 0.06 | 0.25 | <0.06 | <0.06 | <0.06 | ||||
AMX, amoxicillin; AUG, amoxicillin-clavulanic acid; LOT, cephalothin; TAX, cefotaxime; TAZ, ceftazidime; ATM, aztreonam; FEP, cefepime; IMP, imipenem; MER, meropenem; FOX, cefoxitin; TTN, cefotetan; GS, general surgery; NS, neonate surgery; CS, chest surgery; O&G, obstetrics and gynecology; UTI, urine tract infection; trf, transformant.
Susceptibility to beta-lactams.
Susceptibility was determined on Mueller-Hinton agar plates (Difco Laboratories, Detroit, Mich.) containing serially twofold-diluted beta-lactams as previously described (13). The results were interpreted by using the National Committee for Clinical Laboratory Standards criteria (18).
Conjugation experiments.
Conjugation experiments were performed as described previously (13) with sodium azide-resistant E. coli J53 AzideR as the recipient. Transconjugants were selected on Muller-Hinton agar supplemented with sodium azide (Sigma, St. Louis, Mo.) (150 mg/liter) to inhibit the growth of the donor strain and cefoxitin (20 mg/liter) to inhibit the growth of the recipient strain.
Cloning of ampC genes.
The genomic DNA of six clinical isolates was prepared with the Wizard genomic DNA purification kit (Promega, Madison, Wis.) and used as the template DNA in PCR analysis. Oligonucleotides AmpCF1 (5′-TCGGAATTCCGGAGGATTACTGATGATGA-3′) and AmpCR1 (5′-TTAGTCGACAATGTTTTACTGTAGCGCCTCG-3′) were used, respectively, as forward and backward primers. Both primers contained a tail with EcoRI (AmpCF1) and SalI (AmpCR1) recognition sequences (in boldface type). AmpCF1 and AmpCR1 are consensus sequences of the bla genes of MIR-1 (M37839), ACT (U58495), and three AmpCs of E. cloacae (GenBank accession no. X07274, X08081, and X08082). All restriction enzymes were purchased from Roche Applied Science (Mannheim, Germany). Since AmpCF1 and AmpCR1 primers were present, respectively, in the origin and in the end of the structural genes, all regulatory signals from the original genes were eliminated by the cloning strategy. In addition, primer AmpCF1 contains a mismatched base (underlined) to generate a consensus E. coli ribosome binding site. PCR amplifications were carried out on a DNA thermal cycler (model 2400; Perkin-Elmer Cetus, Norwalk, Conn.) as previously described (12). The expected PCR product of 1,165 bp was verified by agarose gel electrophoresis. The EcoRI/SalI-digested PCR products were ligated with the EcoRI/SalI-digested vector pHSG 398 (TaKaRa, Otsa, Shiga, Japan), which is the pUC type cloning vector plasmid carrying the chloramphenicol resistance gene. The ligation mixture was introduced by transformation in CaCl2 competent E. coli DH5α cells. Transformants were selected onto Luria-Bertani agar plates containing chloramphenicol (25 mg/liter) and ampicillin (50 mg/liter). The presence of the desired hybrid plasmid was purified by Wizard Plus Minipreps DNA purification system (Promega) and verified by restriction analysis and sequencing of the whole cloned ampC genes. DNA sequencing was performed by the direct sequencing method with an automatic sequencer (model 373A; Applied Biosystems, Weiterstadt, Germany), as previously described (14). Cloned ampC genes from a pHSG 398-based plasmid were expressed from the lac promoter. The expression of ampC genes was induced by isopropyl-β-d-thiogalactopyranoside (IPTG; 1 mM). To amplify and sequence TEM-, SHV-, and CMY-related genes from clinical isolates, the following primers were used: T1, T2, T3, and T4 (15) for blaTEM genes; S1, S2, S3, and S4 (15) for blaSHV; and C1, C2, C3, C4, CMYF1, and CMYR1 for blaCMY (13).
IEF analysis.
Isoelectric focusing (IEF) was performed in Ready Gel Precast IEF polyacrylamide gel (Bio-Rad, Hercules, Calif.) as previously described (13). Gels were developed with 0.5 mM nitrocefin (Oxoid, Basingstoke, United Kingdom). Inducibility of AmpC beta-lactamases was inferred from the intensity of IEF patterns for induced and uninduced beta-lactamase extracts by cefoxitin (50 mg/liter), as recommended by Pitout et al. (19).
ERIC PCR.
Enterobacterial repetitive consensus (ERIC) PCRs were performed in 50-μl volumes containing 10 ng of genomic DNA from six clinical isolates; 4 mM MgCl2; 50 pM concentrations of each primer (ERIC1R [5′-ATGTAAGCTCCTGGGGATTCAC-3′] and ERIC2 [5′-AAGTAAGTGACTGGGGTGAGCG-3′]) (24); 1.25 U of TaKaRa Ex Taq polymerase (TaKaRa); 0.2 mM (each) dATP, dCTP, dGTP, and dTTP; 25 mM TAPS [N-Tris(hydroxy)methyl-3-amino-propane sulfonic acid, pH 9.3]; 50 mM KCl; and 1 mM 2-mercaptoethanol. Amplification was carried out by using the following program: 95°C for 5 min followed by 35 cycles of 1 min at 52°C, 5 min at 70°C, and 1 min at 92°C. The final extension step was performed at 70°C for 10 min. The analysis of amplified products (10-μl aliquots) was performed in 2% Seakem LE agarose (BMA, Rockland, Maine).
DNA sequence analysis.
DNA sequence analysis was performed with DNASIS for Windows (Hitachi Software Engineering America Ltd., San Bruno, Calif.). Database similarity searches for both the nucleotide sequences and deduced protein sequences were carried out at the National Center for Biotechnology Information website (http://www.ncbi.nlm.nih.gov). Using PRODIST (protein distance matrix) and FITCH (Fitch-Margoliash and least-squares distance methods) from the PHYLIP package of Joseph Felsenstein (Department of Genetics at the University of Washington), a phylogenetic tree was constructed from CLUSTAL W multiple-sequence alignment (23) of 16 ampC genes which were chromosomal genes (EcloQ908R [X08081] of E. cloacae Q908R, EcloMNH1 [X08082] of E. cloacae MHN1, EcloCHE [AJ278994] of E. cloacae CHE, EcloGC1 [D44479] of E. cloacae GC1, EcloGN747 [AB016611] of E. cloacae GN747, EcloOUDhy [AJ278995] of E. cloacae OUDhy, EcloP99 [X07274] of E. cloacae P99, Eare1 [AF211348] of Enterobacter aerogenes, EcloK992004.1 [AF411144] of E. cloacae K992004.1, EcloK995120.1 [AF411145] of E. cloacae K995120.1, EcloK99230 [AF411146] of E. cloacae K99230, EareK9911729 [AF411147] of E. aerogenes K9911729, EcloK9973 [AF411148] of E. cloacae K9973, and EcloK9914325 [AF411149] of E. cloacae K9914325) and plasmid-borne genes (MIR-1 [M37839] of Klebsiella pneumoniae and ACT-1 [U58495] of K. pneumoniae MCQ-95 [GenBank accession numbers are in brackets]).
Nucleotide sequence accession number.
The ampC gene nucleotide sequence data appear in the GenBank nucleotide sequence database under the accession numbers AF411144 for blaEcloK992004.1, AF411145 for blaEcloK995120.1, AF411146 for blaEcloK99230, AF411147 for blaEareK9911729, AF411148 for blaEcloK9973, and AF411149 for blaEcloK9914325.
RESULTS AND DISCUSSION
Properties of six Enterobacter species.
Fifty-one clinical isolates were previously isolated from patients with various infections (12). Six of 51 clinical isolates were characterized by high levels of resistance to cefoxitin, cefotetan, cephalothin, amoxicillin, amoxicillin-clavulanic acid, aztreonam, and ceftazidime (Table 1). Resistance to broad-spectrum cephalosporins and broad-spectrum penicillins usually emerges in Enterobacter spp. that overproduce chromosomal beta-lactamases (19). This resistance pattern indicated that six clinical isolates produced an AmpC beta-lactamase. All the strains were focused at a pI of 8.9 by IEF (Table 1). According to PCR and sequencing results, the beta-lactamases of E. cloacae K992004.1 and E. cloacae K995120.1 were identical to the chromosomal AmpC beta-lactamase of E. cloacae MHN1 (4) and those of E. cloacae K99230, E. aerogenes K9911729, E. cloacae K9973, and E. cloacae K9914325, similar to that of E. cloacae MHN1 (98.5, 98.5, 98.2, and 98.2% identity of amino acid sequence) (Table 2). The AmpC beta-lactamases of all the strains were induced four- or fivefold by cefoxitin. Their blaAmpC genes (blaEcloK992004.1, blaEcloK995120.1, blaEcloK99230, blaEareK9911729, blaEcloK9973, and blaEcloK9914325) were not transferred to the E. coli J53 AzideR recipient in the transconjugation experiment. The genomic DNAs from six clinical isolates were hybridized with digoxigenin-labeled 1,165-bp PCR products between AmpCF1 and AmpCR1. These results indicated that they are chromosomal AmpC beta-lactamases. E. aerogenes K9911729 produced additional AmpC beta-lactamase with a pI of 8.0 (Table 1), which was closely related to CMY-1 (99.7% identity of amino acid sequence) (2) as a result of PCR and sequencing. Because of the presence of CMY-type beta-lactamases, MICs of amoxicillin-clavulanic acid and cefoxitin for E. aerogenes K9911729 were increased. The highest level of resistance to cefoxitin in E. cloacae K9911729 might be due to the additional presence of CMY-type beta-lactamase, the gene of which was transferred to the E. coli J53 AzideR recipient.
TABLE 2.
Nucleotide and amino acid differences between blaEcloMHN1, blaEcloK992004.1, blaEcloK995120.1, blaEcloK99230, blaEareK9911729, blaEcloGC1, blaEcloCHE, blaEcloQ908R, blaEcloK9973, blaEcloK9914325, and blaEcloP99
| Nucleotide no. | Amino acid no.a | Nucleotide (amino acid)b encoded by gene (accession no. or reference):
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| blaEcloMHN1 (X08082) | blaEcloK992004.1 (this study) | blaEcloK995120.1 (this study) | blaEcloK99230 (this study) | blaEareK9911729 (this study) | blaEcl oGC1 (D44479) | blaEcloCHE (AJ278994) | blaEcloQ908R (X08081) | blaEcloK9973 (this study) | blaEcloK9914325 (this study) | blaEcloP99 (X07274) | ||
| 21 | 3 | T (Ile) | T | T | T | T | A (Lys) | A (Lys) | ND | G (Arg) | G (Arg) | G (Arg) |
| 49 | 12 | G (Leu) | G | G | G | G | C | C | C | C | C | C |
| 53 | 14 | C (Leu) | C | C | C | C | A (Ile) | A (Ile) | A (Ile) | A (Ile) | A (Ile) | A (Ile) |
| 55 | 14 | C (Leu) | C | C | C | C | C (Ile) | T (Ile) | C (Ile) | C (Ile) | C (Ile) | C (Ile) |
| 74 | 21 | G (Ala) | G | G | G | G | A (Thr) | A (Thr) | A (Thr) | A (Thr) | A (Thr) | A (Thr) |
| 82 | 23 | A (Val) | A | G | A | A | G | G | G | G | G | G |
| 109 | 32 | C (Val) | C | C | C | C | A | C | C | C | C | C |
| 119 | 36 | G (Val) | G | G | G | G | G | G | G | G | G | A (Ile) |
| 185 | 58 | T (Ser) | T | T | C (Pro) | C (Pro) | C (Pro) | C (Pro) | C (Pro) | C (Pro) | C (Pro) | C (Pro) |
| 199 | 62 | G (Thr) | G | G | G | G | G | G | G | G | G | A |
| 262 | 83 | T (Gly) | T | C | T | T | T | T | T | T | T | T |
| 289 | 92 | T (Val) | T | T | T | T | G | T | T | T | T | T |
| 335 | 108 | C (Pro) | C | C | C | C | C | C | C | C | C | G (Ala) |
| 373 | 120 | G (Gln) | G | G | G | G | G | A | G | G | G | G |
| 466 | 151 | G (Leu) | G | G | G | G | G | G | C | G | G | G |
| 467 | 152 | C (Leu) | C | C | C | C | C | C | G (Val) | C | C | C |
| 571 | 186 | T (Ser) | T | T | C | C | T | T | T | T | T | T |
| 610 | 199 | T (Leu) | T | G | T | T | T | T | T | T | T | T |
| 646 | 211 | T (Val) | T | T | T | T | G | G | G | G | G | G |
| 697 | 228 | G (Ala) | G | G | G | G | I-9 | G | G | G | G | G |
| 715 | 234 | A (Gly) | A | A | A | A | T | T | T | T | T | T |
| 717 | 235 | T (Met) | T | T | C (Thr) | C (Thr) | T | T | T | T | T | T |
| 790 | 259 | G (Ala) | G | G | G | G | A | G | G | G | G | G |
| 796 | 261 | G (Glu) | G | G | A | A | G | G | G | G | G | G |
| 799 | 262 | G (Lys) | G | G | G | G | C (Asn) | C (Asn) | C (Asn) | C (Asn) | C (Asn) | C (Asn) |
| 805 | 264 | C (Ala) | C | C | C | C | T | T | T | T | T | T |
| 820 | 269 | G (Lys) | G | G | G | G | A | G | G | G | G | G |
| 928 | 305 | A (Glu) | A | A | A | A | G | G | G | G | G | G |
| 937 | 308 | C (Asp) | C | C | C | C | C | D-18 | C | C | C | C |
| 949 | 312 | G (Ala) | G | G | G | G | G | G | G | G | G | A |
| 961 | 316 | A (Leu) | A | A | A | A | G | G | G | G | G | G |
| 967 | 318 | G (Val) | A | G | G | G | G | G | G | G | G | G |
| 969 | 319 | C (Ala) | C | C | C | C | C | T (Val) | T (Val) | T (Val) | T (Val) | C |
| 1024 | 337 | T (Gly) | T | T | T | T | C | C | C | C | C | C |
| 1030 | 339 | C (Thr) | C | T | T | T | T | T | T | T | T | T |
| 1098 | 362 | A (Lys) | A | A | A | A | C (Thr) | C (Thr) | C (Thr) | C (Thr) | C (Thr) | C (Thr) |
Numbering is according to that of blaEcloMHN1 (X08082).
The encoded amino acid was indicated when there was a point mutation leading to an amino acid change relative to the EcloMHN1 sequence. Abbreviations: ND, not determined; I-9, 9 nucleotides (GGTGCGCGC) inserted between the 696th and 697th nucleotides; D-18, 18 nucleotides (AGTAAGGTAGCGCTGGCG) deleted between the 937th and 956th nucleotides. Abbreviations of gene names are given in Materials and Methods.
The double-disk synergy test (10) and the IEF results suggested that six clinical isolates produced an ESBL. All the strains were focused at a pI of 7.6 by IEF (Table 1), which was likely corresponding to that SHV-type beta-lactamase (4). According to PCR and sequencing results, the SHV-type beta-lactamase was SHV-12, an ESBL, and the beta-lactamase with a pI of 6.0 (Table 1) was TEM-1b (8).
Recently, ERIC PCR generated a characteristic genomic fingerprinting which can be used to reveal intra- and interspecies genotypic variations among members of the Enterobacteriaceae (5). Six clinical isolates generated distinct bands ranging in size from 0.6 to 5.1 kb (Fig. 1) and varied from 5 to 10 bands by ERIC PCR. There was a band (0.7 kb) common for all isolates. A different fingerprint pattern was observed for each clinical isolate. These results were confirmed by pulsed-field gel electrophoresis analysis with XbaI, showing that the differences among each clinical isolate in regard to the number of bands were more than 10. These results suggested that all isolates were genetically unrelated, indicating resistances to ESBLs spread out due not to an outbreak of a resistant strain but to the dissemination of SHV-12 beta-lactamase, the gene of which was carried on about 50-kb plasmids conjugally transferred to the E. coli J53 AzideR recipient. Six clinical isolates were from urines, pus, and sputa (Table 1), indicating that these were important sources of the spread. Stricter measures are required to control their further spread in Korea and elsewhere.
FIG. 1.
ERIC PCR patterns of genomic DNA from E. cloacae K9973 (lane 2), E. cloacae K9914325 (lane 3), E. cloacae K99230 (lane 4), E. aerogenes K9911729 (lane 5), E. cloacae K992004.1 (lane 6), and E. cloacae K995120.1 (lane 7). Lanes 1 (HindIII/EcoRI-digested phage λ) and 8 (100-bp stepwise ladder), DNA marker fragments (sizes in base pairs are indicated on the edge of the gel). ERIC PCR was performed with primers ERIC2 and ERIC1R.
All transformants containing ampC genes of six clinical isolates showed highly increased MICs of all beta-lactams except cefepime, imipenem, and meropenem by comparison with the host strain for transformation (Table 1). MICs of all tested beta-lactams for six clinical isolates were greater than those for all transformants. This interesting observation, which has also been reported by other investigators (2), may be due, in part, to the dissimilar genetic backgrounds of the cloning host strain and clinical isolates, resulting in decreased beta-lactamase expression and activity relative to that of the host strain. The MIC of amoxicillin was not reduced by the addition of clavulanic acid in all transformants, which likely corresponds to MIC patterns of strains producing Bush group 1 (class C) AmpC beta-lactamases (4). There were similar MIC patterns of all tested beta-lactams among transformants producing identical AmpC beta-lactamases (trf K9973 and trf K9914325, trf K99230 and trf K9911729, and trf K992004.1 and trf K995120.1) (Tables 1 and 2). MICs of all tested beta-lactams for trf K992004.1 and trf K995120.1 were lowest in six transformants. MICs of amoxicillin, amoxicillin-clavulanic acid, ceftazidime, cefotaxime, cefoxitin, imipenem, and meropenem for trf K992004.1 and trf K995120.1 were similar to those for the transformant harboring pBGMHN (ampC gene of E. cloacae MHN1) (17). MICs of cefoxitin and cefotetan for trf K99230 and trf K9911729 were 8- or 16-fold greater than the corresponding MICs for trf K992004.1 and trf K995120.1. MICs of cefoxitin and cefotetan for trf K9973 and trf K9914325 were fourfold greater than the corresponding MICs for trf K992004.1 and trf K995120.1. These results appear to represent the in vivo evolution of ampC genes (from blaEcloK992004.1 and blaEcloK995120.1 to blaEcloK99230 and blaEareK9911729 or from blaEcloK992004.1 and blaEcloK995120.1 to blaEcloK9973 and blaEcloK9914325) under the selective pressure of antimicrobial therapy (especially cefoxitin and cefotetan), as was the case with blaSHV-8 (21). Therefore, more prudent use of these antibiotics is necessary to reduce the spread of these resistant strains.
Sequence analysis.
The complete nucleotide sequences of the ampC gene (blaEcloK992004.1, blaEcloK995120.1, blaEcloK99230, blaEareK9911729, blaEcloK9973, and blaEcloK9914325) encoding the cephamycin-resistant beta-lactamase were obtained with three PCR products and primers. Only one large open reading frame was found, which corresponds to a putative protein of 381 amino acids. The ampC gene consisted of 1,165 nucleotides (Fig. 2). The deduced amino acid sequence carried the catalytic residues SXXK (here, serine-isoleucine-serine-lysine), with the initial serine at position 84, as is typical of class C beta-lactamases (7), the class C-type motif YXN (here, tyrosine-alanine-asparagine) at position 170, and the KTG (here, lysine-threonine-glycine) motif at position 335. GenBank, EMBL, and DDBJ database searches (BLASTN) revealed that the blaEcloK992004.1, blaEcloK995120.1, blaEcloK99230, and blaEareK9911729 genes had the greatest homology with the chromosomal ampC gene (blaEcloMHN1) of E. cloacae MHN1 (99.9, 99.7, 99.6, and 99.6% identity, respectively) (Table 2). The blaEcloK9973 and blaEcloK9914325 genes were similar to the chromosomal ampC gene (blaEcloQ908R) of E. cloacae Q908R (99.7% identity). This high degree of similarity indicates that blaEcloK992004.1, blaEcloK995120.1, blaEcloK99230, and blaEareK9911729 genes may originate from blaEcloMHN1. The blaEcloK9973 and blaEcloK9914325 genes may originate from blaEcloQ908R.
FIG. 2.
Sequence of the E. cloacae (EcloK992004.1, EcloK995120.1, EcloK99230, EcloK9973, and EcloK9914325) and E. aerogenes (EareK9911729) ampC genes. Different nucleotides are marked by underlining. Boxed nucleotides indicate start and stop codons. Abbreviations of gene names are given in Materials and Methods.
Multiple-sequence alignment of the deduced amino acid sequences of six ampC genes was performed with other class C beta-lactamases. It demonstrated that six AmpC beta-lactamases are most closely related to AmpC of E. cloacae MHN1 (98.1 to 100% identity of amino acid sequence), followed by MIR-1 (9) (86.6 to 87.1% identity), ACT-1 (3) (87.4 to 87.7% identity), and AmpC of E. aerogenes (20) (74.8 to 75.1% identity). At this time, the number of chromosomal class C beta-lactamases that have been described is 36 (1). The incidence of many cephamycinase producers increased the possibility that cephamycin-resistant microorganisms would spread out in the family Enterobacteriaceae.
Phylogenetic analysis.
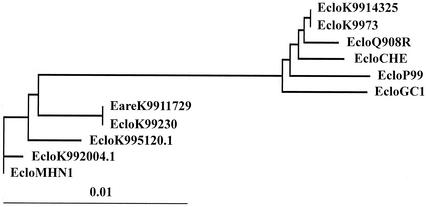
We selected 16 class C beta-lactamase genes that are closely related to the ampC genes of Enterobacter spp. The phylogenetic tree subclassified 16 class C beta-lactamase genes into subclasses 1a (EcloMNH1, EcloK992004.1, EcloK995120.1, EcloK99230, and EareK9911729), 1b (EcloGC1, EcloP99, EcloCHE, EcloQ908R, EcloK9973, and EcloK9914325), and 1c (EcloOUDhy, Eare1, EcloGN747, MIR-1, and ACT-1) (data not shown). This tree indicated that plasmid-mediated MIR-1 and ACT-1 are descended from an ampC gene such as EcloGN747, which was confirmed by a previous report (1). These results were confirmed by other methods: PRODIST-KITCH (Fitch-Margoliash and least-squares methods with evolutionary clock) and the neighbor-joining method from the PHYLIP package. We performed phylogenetic analysis with subclasses 1a and 1b (Fig. 3). MIC patterns of six transformants and phylogenetic analysis results indicated that there might be two in vivo evolution schemes: (i) from EcloMNH1 to EcloK992004.1 (EcloK995120.1) and from EcloK992004.1 (EcloK995120.1) to EcloK99230 (EareK9911729) and (ii) from EcloQ908R to EcloK9973 (EcloK9914325).
FIG. 3.
Phylogram of 11 ampC beta-lactamase genes. The branch length value represents relative phylogenetic distance. Abbreviations for taxon labels, accession numbers, gene locations (plasmid or chromosome), and organism names are given in Materials and Methods.
Acknowledgments
This work was supported by a research grant from the Korea Science and Engineering Foundation (1999-2-211-002-5) and in part by a research grant from Youngdong University (YU02-03) to S. H. Lee.
We thank J. Kim for the generous gift of E. coli J53 AzideR and B. H. Kim for technical assistance.
REFERENCES
- 1.Barlow, M., and B. G. Hall. 2002. Origin and evolution of the AmpC beta-lactamases of Citrobacter freundii. Antimicrob. Agents Chemother. 46:1190-1198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bauernfeind, A., I. Stemplinger, R. Jungwirth, R. Wilhelm, and Y. Chong. 1996. Comparative characterization of the cephamycinase blaCMY-1 gene and its relationship with other beta-lactamase genes. Antimicrob. Agents Chemother. 40:1926-1930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bradford, P. A., C. Urban, N. Mariano, S. J. Projan, J. J. Rahal, and K. Bush. 1997. Imipenem resistance in Klebsiella pneumoniae is associated with the combination of ACT-1, a plasmid-mediated AmpC beta-lactamase, and the loss of an outer membrane protein. Antimicrob. Agents Chemother. 41:563-569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bush, K., G. A. Jacoby, and A. A. Medeiros. 1995. A functional classification scheme for beta-lactamases and its correlation with molecular structure. Antimicrob. Agents Chemother. 39:1211-1233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Della-Costa, L. M., K. Irino, J. Rodrigues, I. N. G. Rivera, and L. R. Trabulsi. 1998. Characterization of diarrhoeagenic Escherichia coli clones by ribotyping and ERIC-PCR. J. Med. Microbiol. 47:227-234. [DOI] [PubMed] [Google Scholar]
- 6.Farmer, J. J., III. 1995. Enterobacteriaceae: introduction and identification, p. 438-449. In P. R. Murray, E. J. Baron, M. A. Pfaller, F. C. Tenover, and R. H. Yolken (ed.), Manual of clinical microbiology, 3rd ed. American Society for Microbiology, Washington, D.C.
- 7.Ghuysen, J. M. 1991. Serine beta-lactamases and penicillin-binding proteins. Annu. Rev. Microbiol. 45:37-67. [DOI] [PubMed] [Google Scholar]
- 8.Goussard, S., and P. Courvalin. 1991. Sequence of the genes blaT-1b and blaT-2. Gene 102:71-73. [DOI] [PubMed] [Google Scholar]
- 9.Jacoby, G., and J. Train. 1999. Sequence of the MIR-1 beta-lactamase gene. Antimicrob. Agents Chemother. 43:1759-1760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Jarlier, V., M. H. Nicolas, G. Fournier, and A. Philippon. 1988. Extended broad-spectrum beta-lactamase conferring transferable resistance to newer beta-lactam agents in Enterobacteriaceae: hospital prevalence and susceptibility patterns. Rev. Infect. Dis. 10:867-878. [DOI] [PubMed] [Google Scholar]
- 11.Kim, J., Y. Kwon, H. Pai, J. W. Kim, and D. T. Cho. 1998. Survey of Klebsiella pneumoniae strains producing extended-spectrum beta-lactamases: prevalence of SHV-12 and SHV-12a in Korea. J. Clin. Microbiol. 36:1446-1449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lee, S. H., and S. H. Jeong. 2002. Antibiotic susceptibility of bacterial strains isolated from patients with various infections. Lett. Appl. Microbiol. 34:215-221. [DOI] [PubMed] [Google Scholar]
- 13.Lee, S. H., J. Y. Kim, G. S. Lee, S. H. Cheon, Y. J. An, S. J. Jeong, and K. J. Lee. 2002. Characterization of blaCMY-11, an AmpC-type plasmid-mediated beta-lactamase gene in a Korean clinical isolate of Escherichia coli. J. Antimicrob. Chemother. 49:269-273. [DOI] [PubMed] [Google Scholar]
- 14.Lee, S. H., J. Y. Kim, S. H. Shin, S. K. Lee, M. M. Choi, I. Y. Lee, Y. B. Kim, J. Y. Cho, W. Jin, and K. J. Lee. 2001. Restriction fragment length dimorphism-PCR method for the detection of extended-spectrum beta-lactamases unrelated to TEM- and SHV-types. FEMS Microbiol. Lett. 200:157-161. [DOI] [PubMed] [Google Scholar]
- 15.Lee, S. H., S. H. Jeong, and K. J. Lee. 2001. Evolution of beta-lactamase genes identified by PCR with newly designed primers in Korean clinical isolates. Clin. Microbiol. Infect. 7:98-100. [DOI] [PubMed] [Google Scholar]
- 16.Livermore, D. M. 1995. Beta-lactamases in laboratory and clinical resistance. Clin. Microbiol. Rev. 8:557-584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Morisini, M. M., M. C. Negri, B. Shoichet, M. C. Baquero, F. Baquero, and J. Blazquez. 1998. An extended-spectrum AmpC-type beta-lactamase obtained by in vitro antibiotic selection. FEMS Microbiol. Lett. 165:85-90. [DOI] [PubMed] [Google Scholar]
- 18.National Committee for Clinical Laboratory Standards. 1999. Performance standards for antimicrobial susceptibility testing, 9th ed. Informational supplement. M100-S9. National Committee for Clinical Laboratory Standards, Wayne, Pa.
- 19.Pitout, J. D. D., K. S. Thomson, N. D. Hanson, A. F. Ehrhardt, P. Coudron, and C. C. Sanders. 1998. Plasmid-mediated resistance to expanded-spectrum cephalosporins among Enterobacter aerogenes strains. Antimicrob. Agents Chemother. 42:596-600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Preston, K. E., C. C. Radomski, and R. A. Venezia. 2000. Nucleotide sequence of the chromosomal ampC gene of Enterobacter aerogenes. Antimicrob. Agents Chemother. 44:3158-3162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Rasheed, J. K., C. Jay, B. Metchock,F. Berkowitz, L. Weigel, J. Crellin, C. Steward, B. Hill, A. A. Medeiros, and F. C. Tenover. 1997. Evolution of extended-spectrum beta-lactamase resistance (SHV-8) in a strain of Escherichia coli during multiple episodes of bacteremia. Antimicrob. Agents Chemother. 41:647-653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Sanders, W. E., Jr., and C. C. Sanders. 1997. Enterobacter spp.: pathogens poised to flourish at the turn of the century. Clin. Microbiol. Rev. 10:220-241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Thompson, J. D., D. G. Higgins, and T. J. Gibson. 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22:4673-4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Versalovic, J., T. Koeuth, and J. R. Lupski. 1991. Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res. 19:6823-6831. [DOI] [PMC free article] [PubMed] [Google Scholar]