Abstract
Phosphatidylinositol (4,5)-bisphosphate [PI(4,5)P2] is an important factor for a variety of cellular functions ranging from cell signaling to actin cytoskeletal dynamics and endocytic membrane traffic. Here, we have identified the clathrin adaptor complex AP-2 as a regulator of phosphatidylinositol 4-phosphate 5-kinase (PIPK)-mediated PI(4,5)P2 synthesis. AP-2 directly interacts with the kinase core domain of type I PIPK isozymes via its μ2-subunit in vitro and in native protein extracts. Endocytic cargo protein binding to μ2 leads to a potent stimulation of PIPK activity. These data thus identify a positive feedback loop consisting of endocytic cargo proteins, AP-2μ, and PIPK type I which may provide a specific pool of PI(4,5)P2 dedicated to clathrin/AP-2-dependent receptor internalization.
Keywords: phosphoinositides, sorting motifs, clathrin, endocytosis
Phosphoinositides have pleiotropic functions in cell physiology including the regulation of signal transduction, membrane traffic (1), and the organization of the actin cytoskeleton (2). Phosphatidylinositol (4,5)-bisphosphate [PI(4,5)P2], a phosphoinositide concentrated at the plasma membrane, is required for the generation of diacylglycerol (DAG) and inositol 1,4,5-trisphosphate (IP3), remodelling of the actin cytoskeleton (2, 3), clathrin-dependent (4) and -independent pathways of cell entry (5), the exo–endocytic cycling of synaptic and neurosecretory vesicles (6–8), and serves as a substrate for the synthesis of phosphatidylinositol 3,4,5-trisphosphate [PI(3,4,5)P3]. Biochemical, genetic, and cell biological data suggest that PI(4,5)P2 plays an essential role in clathrin-dependent endocytosis of plasma membrane proteins including nutrient (4) and growth factor receptors, postsynaptic ion channels, as well as synaptic vesicle proteins (9–11). Endocytic adaptor proteins like the heterotetrameric AP-2 complex (via its α and μ2 subunits) (12, 13), AP180/CALM, epsin (14), Dab2, and HIP1/1R as well as the large GTPase dynamin (15) all bind directly to PI(4,5)P2. In addition to serving as a membrane attachment site for endocytic proteins, PI(4,5)P2 also plays an active role by allosterically eliciting a conformational change within AP-2 that is required for its interaction with sorting signals of transmembrane proteins (16) and to stably associate with the plasmalemma (17). Impairment of PI(4,5)P2 hydrolysis by deletion of the phosphoinositide–phosphatase synaptojanin (18–20) results in the accumulation of clathrin-coated pits and vesicles, suggesting that the membrane concentration of PI(4,5)P2 controls the stability of endocytic clathrin coats.
Three isoforms of type I phosphatidylinositol 4-phosphate 5-kinase (PIPK) have been identified in mammals (α, β, γ), which generate PI(4,5)P2 by phosporylating PI(4)P at the D-5 position of the inositol ring (21, 22). Although isoform-specific functions of type I PIPKs have been shown to contribute to distinct cellular processes such as formation of focal adhesions (23, 24), at present it is largely unclear how the specific pool of PI(4,5)P2 needed for clathrin adaptor recruitment and receptor internalization is provided. On the basis of overexpression and rescue experiments, it has been suggested that human PIPK type Iβ is the major contributor to PI(4,5)P2 levels in constitutive pathways of clathrin/AP-2-dependent endocytosis (25), whereas type Iα (26) and γ isoforms (27) may regulate stimulation-induced endocytic pathways in nonneuronal and neuronal cell types, respectively. Additional regulatory mechanisms such as ARF6-mediated stimulation of PIPK activities (28, 29) may also contribute to the synthesis of a pool of PI(4,5)P2 dedicated to endocytosis. However, given the pleiotropic nature of ARF6-dependent effector pathways (5), this effect may be insufficient to confer specificity.
In the present study, we have set out to identify regulators of PIPK-mediated PI(4,5)P2 formation. We find that the endocytic clathrin adaptor complex AP-2, via its μ2-subunit, directly interacts with the kinase core domain of type I PIPK isozymes in vitro and in native protein extracts. Endocytic cargo protein binding to μ2 leads to potent stimulation of PIPK activity. These data thus identify a positive feedback loop consisting of endocytic cargo proteins, AP-2μ, and PIPK type I, which may provide a specific pool of PI(4,5)P2 dedicated to clathrin/AP-2-dependent receptor internalization.
Results
PIPK Type Iγ Is Functionally Associated with Clathrin/AP-2-Mediated Receptor Internalization.
To better understand the relationship between PIPK-mediated PI(4,5)P2 synthesis and clathrin/AP-2-mediated receptor internalization we generated HEK293 flip-in cells allowing for the regulated inducible expression of PIPK type Iγ. Consistent with the essential role of PI(4,5)P2 in clathrin-dependent endocytosis (4, 18), we observed an increase in the initial rates of transferrin uptake after doxycycline-induced overexpression of PIPK Iγ, suggesting that PI(4,5)P2 is rate-limiting for internalization under these conditions (Fig. 6, which is published as supporting information on the PNAS web site). This finding is similar to what has been reported for other type I PIPK isozymes (25). Conversely, inhibition of PI(4,5)P2-synthesis by knockdown of all three mammalian type I (α, β, γ) PIPK isoforms resulted in a partial depletion of endocytic AP-2 adaptors from plasmalemmal coated pits (see Fig. 2A; ref. 29). We then analyzed the subcellular localization of PIPK Iα or γ, AP-2, and endocytic cargo receptors (i.e., EGFR) accumulated in coated pits at low temperature by deconvolution fluorescence microscopy. EGF-containing AP-2-coated pits appeared to form along but not exactly coinciding with plasmalemmal sites enriched in PIPK Iα or γ. PIPK Iα or γ (Fig. 1; see ref. 27) colocalized only partially with AP-2 and internalizing EGF, suggesting that regulatory mechanisms may be involved in providing the pool of PI(4,5)P2 needed for clathrin/AP-2-dependent endocytosis. Therefore, we set out to identify such putative regulators of PIPK type Iγ.
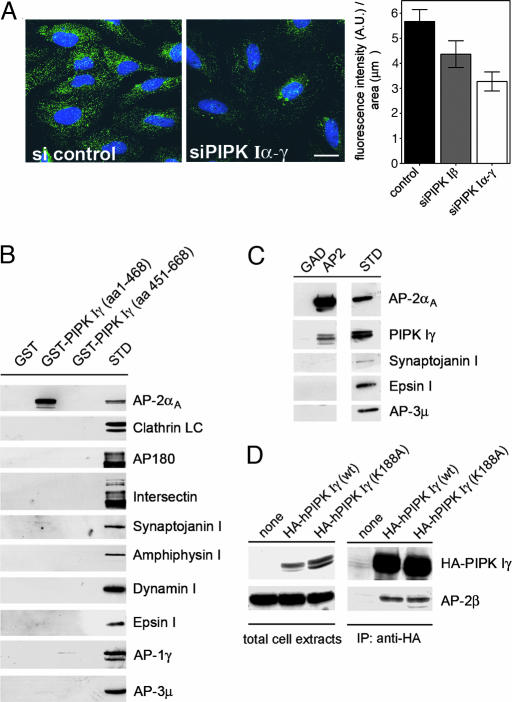
Fig. 2.
PIPK Iγ directly interacts with AP-2μ. (A) RNAi-mediated knockdown of PIPK I attenuates AP-2-coated pit staining in HeLa cells. (Left and Center) Fixed cells were analyzed for AP-2α-staining. Total fluorescence intensity was quantified and normalized to the area covered by the cells. (Right) Data are depicted as mean (±SD). (Scale bar, 20 μm.) (B) The kinase core domain of PIPK Iγ interacts with AP-2 but not AP-1 or AP-3. Immunoblot analysis of material affinity-purified using GST, GST-tagged PIPK Iγ kinase (1–468), or tail domains (451–668) is shown. γ- and μ3-Adaptins are subunits of AP-1 or AP-3 complexes, respectively. STD, 2.5% of the total amount of Triton X-100-extracted rat brain lysates added to the assay. (C) PIPK Iγ coimmunoprecipitates with AP-2 from rat brain lysates. Detergent-lysed rat brain extracts were subjected to immunoprecipation using monoclonal antibodies against AP-2α. Samples were washed and analyzed by immunoblotting for the proteins indicated. STD, 5% of the total amount of Triton X-100-extracted rat brain lysates added to the assay. (D) AP-2 coimmunoprecipitates with HA-PIPK Iγ or a kinase-dead mutant (K188A) from transfected fibroblasts. Detergent extracts of transfected HEK293 cells expressing wild-type PIPK Iγ or a kinase-inactive mutant (K188A) were subjected to immunoprecipitation using anti-HA antibodies. Aliquots of the total cell extracts and the immunoprecipitated material were analyzed by immunoblotting with antibodies against AP-2β or HA-PIPK Iγ. Cells not expressing HA-PIPK Iγ (none) were taken as a control.
Fig. 1.
Cargo-containing clathrin/AP-2-coated pits form along PIPK-positive plasma membrane areas. (A) COS7 fibroblasts expressing HA-PIPK type Iα (blue) were incubated with Texas red-labeled EGF at 8°C for 60 min, washed, fixed, and immunostained for AP-2α (green). (Scale bar, 10 μm.) (B) Boxed area in A. The majority of EGF-positive AP-2-coated pits emanates from PIPK Iγ-containing membrane patches. Arrows denote spots of colocalization of AP-2 with both EGF (Lower Left), and PIPK Iγ (Lower Right). (Scale bar, 10 μm.) (C and D) Same as in A and B upon expression of HA-PIPK Iγ (K188A).
PIPK Iγ Directly and Specifically Interacts with AP-2 via Its μ2-Adaptin Subunit.
To this aim, we performed affinity chromatography using GST fusion proteins comprising either the kinase core or tail domains of PIPK Iγ using detergent-extracted rat brain lysates. Proteins that were specifically retained by the kinase core or tail domain chimeras but not by GST were then analyzed by SDS/PAGE, staining with Coomassie blue, followed by tryptic digest and MALDI mass spectrometry (MS) analysis. Using this approach, we identified the α and β subunits of the clathrin adaptor complex AP-2 as binding partner of the kinase core domain of PIPK Iγ (Fig. 2B and Fig. 7, which is published as supporting information on the PNAS web site). Immunoblot analysis indicated that the PIPK Iγ catalytic domain specifically retained AP-2, but not AP-1 or AP-3, complexes localized to TGN/endosomal membranes in most cell types. Other endocytic proteins such as clathrin, AP180, intersectin, synaptojanin 1, amphiphysin 1, dynamin 1, or epsin were not retained. None of these factors associated with GST or GST-PIPK Iγ-tail (Fig. 2B), which by contrast did interact with talin (Fig. 7) as expected (23, 24). Occassionally, very small amounts of AP-2 were found in GST-PIPK Iγ-tail pulldowns, which may be owed to the presence of a Yxxø-based motif within PIPK Iγ–p90, as suggested by a recent report (ref. 30; published while this study was under review). PIPK Iγ coimmunoprecipitated with native endogenous AP-2 complexes isolated from brain synaptosomal lysates (Fig. 2C). Together with the observed colocalization of both proteins at synapses (Fig. 8, which is published as supporting information on the PNAS web site), this finding suggests that both proteins interact in situ. The interaction between AP-2 and PIPK Iγ was further corraborated in fibroblasts transfected with wild-type or kinase-inactive variants of PIPK Iγ (Fig. 2D). The latter data also show that binding of PIPK Iγ to AP-2 is independent of its lipid kinase activity. Cells transiently overexpressing HA-PIPK Iα or γ were unable to undergo clathrin-mediated internalization of fluorescently labeled transferrin or EGF (Fig. 9 A and B, which is published as supporting information on the PNAS web site). This phenotype was accompanied by sequestration of AP-2 into intracellular aggregates that also contained HA-PIPK Iγ (Fig. 9C). Overexpression of kinase-inactive mutants of PIPK Iα or Iγ also inhibited transferrin endocytosis, albeit less potently (Fig. 9B). Thus, PIPK I-mediated AP-2 sequestration and excess PI(4,5)P2 synthesis (18–20) both contribute to these effects.
We then went on to better define this interaction at the molecular level. First, we synthesized the individual subunits of AP-2 by coupled transcription/translation in the presence of [35S]methionine in vitro and offered them to GST or to the GST-PIPK Iγ core domain fusion protein. Only the μ2 subunit of AP-2 was capable of specifically binding to PIPK Iγ core domain (Fig. 3A), suggesting a direct interaction. The C-terminal signal-binding domain of μ2 (C-μ2; residues 157–435) (13, 16) fused to GST (GST-μ2) was sufficient for the interaction with all three (α, β, γ) type I PIPK isozymes in pulldown assays using either in vitro translated (Fig. 10B, which is published as supporting information on the PNAS web site) or native proteins expressed in fibroblasts (Fig. 3B). GST-μ2 did not bind to PI kinase type IIα, a PI(4)P-synthesizing enzyme localized to the Golgi complex and to synaptic vesicles (31) (Fig. 10B). Moreover, PIPK type I enzymes specifically coimmunoprecipitated with C-μ2-EGFP [a soluble fusion protein between C-μ2 (residues 164–435) and EGFP] from transfected cells (Fig. 3C). C-μ2–EGFP was absent from HA-antibody immunoprecipitates derived from cells devoid of HA-PIPK (not shown). The association was also independent of the small GTPase ARF6 known to bind to PIPK type I and perhaps AP-2 (32) (Fig. 10A). Indeed, recombinant C-μ2 could directly bind to PIPK type Iγ purified from stably transfected HEK293 cells (Fig. 10C) with a KD of ≈515 nM as determined by surface plasmon resonance measurements (Fig. 3D). The binding site for PIPK type I within C-μ2 is distinct from that for tyrosine-based endocytic internalization motifs or for PI(4,5)P2 as mutants defective in association with either of these molecules retained their ability to interact with PIPK Iγ (see Fig. 5B and Fig. 11B, which is published as supporting information on the PNAS web site). The activation loop of PIPK Iγ was not sufficient for AP-2μ binding (Fig. 11A). Collectively, our data indicate that AP-2μ directly and specifically interacts with the kinase core domain of PIPK type I enzymes.
Fig. 3.
Interaction of type I PIP kinases with AP-2. (A) PIPK Iγ binds to AP-2 via its μ2-subunit. GST or GST-PIPK Iγ (1–468) were incubated with [35S]-radiolabeled AP-2 subunits synthesized individually by coupled transcription/translation in vitro. After extensive washes, bound proteins were eluted and analyzed by SDS/PAGE and autoradiography. STD, 20% of the total amount of radiolabeled protein added to the assay. (B) AP-2μ associates with all three isoforms of PIPK type I from a detergent extract of transfected fibroblasts. COS7 cells were transfected with HA-tagged PIPK I enzymes; a detergent extract was affinity-purified by using GST or GST-C-μ2, and the affinity-purified material was analyzed by immunoblotting using antibodies recognizing clathrin heavy chain (HC) or HA. (C) Soluble C-μ2-EGFP coimmunoprecipitates with HA-tagged PIPK type I isoforms from transfected fibroblasts. Detergent extracts of transfected Cos7 cells coexpressing soluble C-μ2 (residues 164–435)-EGFP together with PIPK I (α, β, or γ) were subjected to immunoprecipitation using anti-HA-antibodies. Aliquots of the immunoprecipitated material or the supernatants were analyzed with antibodies against EGFP, HA, or AP-1γ. (D) Binding of His6-C-μ2 to the PIPK Iγ kinase core domain determined by surface plasmon resonance. GST-PIPK Iγ (amino acid 1–468) or GST were immobilized on a sensor chip, and His6-C-μ2 was injected at concentrations between 62 nM and 1 μM (lower to upper curves). KD was calculated from the kinetic rate constants derived from the sensograms.
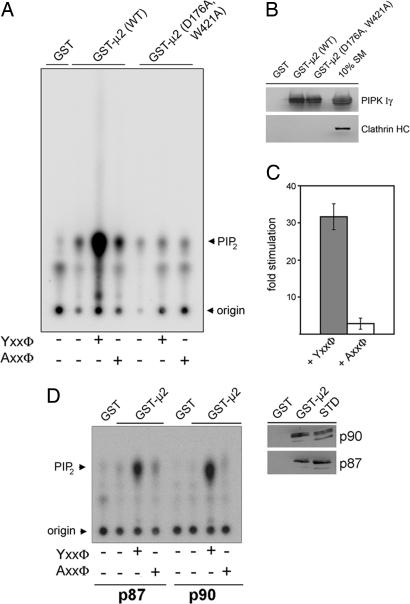
Fig. 5.
AP-2μ-tyrosine motif sorting signal complexes activate the PI(4,5)P2-synthesizing activity of PIPK Iγ. (A–C) AP-2μ bound to tyrosine-based endocytic sorting signals stimulates PIPK Iγ-mediated PI(4,5)P2 synthesis. PIPK Iγ was affinity-purified from detergent lysates of HEK293 flip-in cells using GST-μ2 or a Yxxø-motif binding-defective mutant [GST-μ2 (D176A,W421A)]. An aliquot of this material was subjected to SDS/PAGE and immunoblotting. (B) Radiolabeled PI(4,5)P2 synthesized in the presence or absence of peptides together with [32P]ATP and PI(4)P was analyzed by TLC and quantitative phosphorimage analysis. Representative data are shown in A. Quantifications are depicted in C. Data (mean stimulation ± SD) derived from three independent experiments were normalized to the amount of PI(4,5)P2 synthesized by GST-μ2-associated PIPK Iγ in the absence of peptide. (D) PI(4,5)P2 synthesis by the p87 and p90 isoforms of PIPK Iγ is stimulated equally well by AP-2μ–Yxxø complexes (Left). The experiment was essentially done as described in A. Similar amounts of p87 or p90 were found to be associated with GST-μ2 (Right). STD, 5% of the total amount of Triton X-100-extracted HEK293 cell lysates added to the assay.
PIPK Iγ Preferentially Interacts with AP-2μ Bound to Endocytic Cargo Proteins.
Because AP-2 binding to endocytic sorting signals within the cytoplasmic domains of transmembrane receptors is modulated by the PI(4,5)P2 content of the membrane (17), we were interested to see whether PIPK Iγ preferentially associated with soluble or membrane-bound AP-2 complexes. To this aim, the association between PIPK Iγ and AP-2 was studied in HEK293 flip-in cells induced with doxycycline. AP-2 could be coimmunoprecipitated with PIPK Iγ efficiently, if detergent-lysed membrane fractions were used as a starting material. By contrast, little AP-2 was present in PIPK Iγ-precipitates obtained from a cytosolic pool (Fig. 4A). The ear domain of AP-2α binds to soluble accessory proteins whereas AP-2μ associates with the cytoplasmic domains of cargo membrane proteins (16). Therefore, we asked whether PIPK Iγ might preferentially associate with AP-2 bound to accessory proteins or cytoplasmic receptor domains. To this aim, we compared the amounts of AP-2 or PIPK Iγ retained on GST fusion proteins comprising the α-adaptin ear domain binding site of stonin 1 (residues 1–33) (33) or the cytoplasmic tail (CT) of the EGF receptor (GST-EGFR). Although AP-2 bound equally well to both fusion proteins, PIPK Iγ was efficiently retained only by the GST-EGFR chimera (Fig. 4B) but not by GST-stonin 1. Consistent with the formation of a heteromeric complex between AP-2, PIPK, and endocytic cargo membrane proteins, EGFR and PIPK Iγ could be coimmunoprecipitated with AP-2 from lysates of EGFR- and PIPK Iγ-overexpressing fibroblasts (Fig. 4C). Similar results were seen for other endocytic cargo proteins such as the cytoplasmic tail of the AMPA-type glutamate receptor subunit GluR2 (data not shown). PIPK Iγ did not bind to cargo protein cytoplasmic domains in the absence of AP-2μ (not shown). We conclude that PIPK Iγ preferentially associates with AP-2μ bound to endocytic cargo proteins.
Fig. 4.
PIPK Iγ preferentially associates with AP-2 bound to membranes or endocytic receptor sorting signals. (A) AP-2-PIPK Iγ complexes can be isolated preferentially from membrane fractions. Detergent extracts of HEK293 cells inducibly expressing PIPK Iγ were subjected to immunoprecipitation using anti-HA affinity tag antibodies. Aliquots of the material immunoprecipitated from total cell lysates or cytosolic or membrane fractions were analyzed by immunoblotting with antibodies against AP-2α, talin, or HA-PIPK Iγ. STD, detergent extracted rat brain lysates (15 μg of total protein). (B) PIPK Iγ associates with AP-2 bound to the cytoplasmic tail of the EGFR. GST-stonin 1(1–33), the GST-tagged cytoplasmic tail of the EGF receptor (EGFR), or GST were used for affinity purification from detergent-extracted rat brain lysates. Samples were analyzed by SDS/PAGE and immunoblotting for AP-2μ or PIPK Iγ. STD, 5% of the total amount of Triton X-100-extracted rat brain lysates added to the assay. (C) AP-2 coimmunoprecipitates with both EGFR and PIPK Iγ. Detergent extracts of Cos7 cells that had been mock-treated or transfected with plasmids encoding HA-PIPK Iγ and the EGFRs were subjected to immunoprecipitation using antibodies against AP-2α. Aliquots of the supernatant or pellet fractions were analyzed by immunoblotting for AP-2α, EGFR, or HA-PIPK Iγ.
AP-2μ-Endocytic Cargo Complexes Potently Activate the PI(4,5)P2-Synthesizing Activity of PIPK Iγ.
To analyze the effect of AP-2μ in the presence or absence of endocytic cargo, protein-derived peptides on PIPK Iγ-mediated PI(4,5)P2-synthesis lipid kinase activity assays from HEK293 flip-in cell lysates were performed. Addition of His6-C-μ2 (157–435) in combination with a functional AP-2μ-binding tyrosine motif peptide (Yxxø) to lysates of doxycycline-induced HEK293 cells potently stimulated formation of radiolabeled PI(4,5)P2. This was strictly dependent on the presence of AP-2μ. His6-C-μ2 (157–435) alone or together with a mutated nonfunctional peptide (Axxø) produced little or no stimulation of PI(4,5)P2 synthesis (Fig. 12A, which is published as supporting information on the PNAS web site). Moreover, if a functional tyrosine motif peptide was added to PIPK Iγ affinity-purified by using wild-type GST-μ2, potent, >25-fold activation of lipid kinase activity was observed (Fig. 5A and C), whereas the mutant Axxø peptide did not elicit such an effect. This finding could be attributed to PIPK Iγ activation because addition of the peptide had not effect on binding of PIPK Iγ to AP-2 assayed by immunoprecipitation (Fig. 12C). If lysates prepared from cells not treated with doxycycline were taken instead, no PI(4,5)P2 synthesis was observed (Fig. 12B), indicating that PI(4,5)P2 formation is indeed mediated by PIPK Iγ. A GST-μ2 mutant defective in binding to tyrosine-based sorting motifs retained its ability to associate with PIPK Iγ (Fig. 5B) but was unable to stimulate PI(4,5)P2 synthesis upon addition of the Yxxø peptide (Fig. 5A), suggesting that endocytic sorting motif recognition was a prerequisite for enzyme activation. The p87 and p90 variants of PIPK Iγ were both stimulated to similar extents by AP-2μ–cargo peptide complexes, reemphasizing the fact that the interaction is mainly mediated by the kinase core but not the alternatively spliced tail domains (Fig. 5D) that may, however, contribute to stabilizing an AP-2–PIPK Iγ–p90 complex in the absence of cargo (30). Collectively, these data suggest that PIPK Iγ activity is potently stimulated by AP-2μ–cargo complexes.
Discussion
Our combined data identify AP-2μ–cargo sorting signal complexes as potent regulators of PIPK type I-mediated PI(4,5)P2-synthesis. By directly associating with the kinase core domain of type I but not type II enzymes and with endocytic sorting signals present in membrane cargo proteins, such as the EGFR, AP-2μ potently stimulates PIPK activity in vitro. Our observation that all type I PIPK isozymes are subject to AP-2μ–cargo complex activation suggests that we have unraveled a general mechanism by which endocytic AP adaptor–cargo complexes may regulate the synthesis of PI(4,5)P2, which in turn is needed for stable association of AP-2 and other adaptors including AP180/CALM, and epsin with the plasma membrane and for endocytic coated pit formation. AP-2-μ–cargo complexes may synergize with other regulators of PIPK-mediated PI(4,5)P2-synthesis, including phospholipase D (34), ARF6 (28, 29), Rho family GTPases (3), and in the case of PIPK Iγ–p90, talin (23, 24). The cooperative action of these proteins together with AP-2μ–cargo complexes could thus provide an important positive feedback loop resulting in the generation of a pool of PI(4,5)P2 dedicated to clathrin/AP-2-dependent receptor internalization. Consistent with this scenario, live cell imaging studies have shown that clathrin/AP-2-coated pits are stabilized by the concomitant presence of membrane cargo (35). Accordingly, all type I PIPK isozymes have been implicated in different pathways of clathrin/AP-2-dependent endocytosis within nonneuronal cells (25, 26) and at synapses (8).
At the moment we can only speculate about the exact mechanism by which PIPK activation occurs. One possibility might be that binding of sorting signals within cargo membrane proteins to μ2-adaptin triggers AP-2 dimerization, which in turn could aid the formation of hyperactive PIPK dimers at the plasmalemma (Fig. 13, which is published as supporting information on the PNAS web site). In agreement with this hypothesis, we have observed that (i) crosslinking by specific divalent antibodies potently stimulates PIPK Iγ activity (Fig. 14A, which is published as supporting information on the PNAS web site) and that (ii) cargo sorting signal binding to AP-2μ facilitates dimerization of intact native AP-2 (36) as well as that of C-μ2 (Fig. 14B) in vitro, in agreement with the observed crystallographic C-μ2-Yxxø dimer (37). Similar results have been reported for the oligomeric assembly of the homologous clathrin adaptor complex AP-1 bound to liposomal membranes (38). Furthermore, crystallographic studies have revealed that phosphatidylinositol 5-phosphate 4-kinase, another member of the phosphoinositide kinase superfamily, is a disk-shaped homodimer (39), suggesting that the active form of phosphoinositide kinases might be dimers. Thus, it will be interesting to see how exactly AP-2μ–cargo complexes trigger activation of PIPK type I and whether similar regulatory mechanisms operate for adaptor-mediated protein transport at TGN/endosomal membranes.
Materials and Methods
Antibodies.
A list of antibodies used in this study is provided in Supporting Text, which is published as supporting information on the PNAS web site.
Recombinant Proteins.
GST- or His6-tagged fusion were purified according to published procedures (13).
Plasmid DNA and Site-Directed Mutagenesis.
See Supporting Text for further information.
Affinity Purification and Immunoprecipitation.
Affinity purification (28, 33) and immunoprecipitations (17) were done as described.
PIPK Activity Assays.
A detailed description can be found in Supporting Text. Briefly, cell extracts or cytosol were either incubated with GST-fusion proteins or assayed directly. Samples were incubated in kinase buffer (50 μl) containing 10 μg of PI4P, 200 μM ATP, and 10 μCi of [γ-32P]ATP for 20 min at 37°C. Lipids were extracted and analyzed as described (28).
Fluorescence Microscopy.
Images were acquired by using a motorized Carl Zeiss Axiovert 200M inverted microscope equipped with the Stallion System (Intelligent Imaging). Images of neurons were processed by deconvolution.
Cell Culture, Transfection, Endocytosis Assays, and SPR Experiments.
See Supporting Text for further information.
Supplementary Material
Acknowledgments
We are grateful to Drs. P. De Camilli (Yale University School of Medicine, New Haven, CT), P. Mc Pherson (Michigan State University, East Lansing, MI), and R. Jahn (Max Planck Institute, Göttingen, Germany) for providing antibodies; Dr. H. Ishihara (Tohoku University, Sendai, Japan), S. Cotecchia (Lausanne, Switzerland), and Dr. A. Ullrich (Max Planck Institute, Martinsreid, Germany) for plasmids; and Dr. B. Schmid (Göttingen, Germany) for performing MALDI-MS analysis. This study was supported by Deutsche Forschungsgemeinschaft Grants HA 2686/2-1, SFB 449/A11, and GRK 1123 and the European Molecular Biology Organization (EMBO YIP Award) (to V.H.).
Abbreviations
- DAG
diacylglycerol
- IP3
inositol 1,4,5-trisphosphate
- PI(4,5)P2
phosphatidylinositol (4,5)-bisphosphate
- PIPK
phosphatidylinositol 4-phosphate 5-kinase
- AP-2
adaptor protein 2.
Footnotes
Conflict of interest statement: No conflicts declared.
This paper was submitted directly (Track II) to the PNAS office.
References
- 1.Martin T. F. Curr. Opin. Cell Biol. 2001;13:493–499. doi: 10.1016/s0955-0674(00)00241-6. [DOI] [PubMed] [Google Scholar]
- 2.Yin H. L., Janmey P. A. Annu. Rev. Physiol. 2003;65:761–789. doi: 10.1146/annurev.physiol.65.092101.142517. [DOI] [PubMed] [Google Scholar]
- 3.Weernink P. A., Meletiadis K., Hommeltenberg S., Hinz M., Ishihara H., Schmidt M., Jakobs K. H. J. Biol. Chem. 2004;279:7840–7849. doi: 10.1074/jbc.M312737200. [DOI] [PubMed] [Google Scholar]
- 4.Jost M., Simpson F., Kavran J. M., Lemmon M. A., Schmid S. L. Curr. Biol. 1998;8:1399–1402. doi: 10.1016/s0960-9822(98)00022-0. [DOI] [PubMed] [Google Scholar]
- 5.Donaldson J. G. J. Biol. Chem. 2003;278:41573–41576. doi: 10.1074/jbc.R300026200. [DOI] [PubMed] [Google Scholar]
- 6.Gong L. W., Di Paolo G., Diaz E., Cestra G., Diaz M. E., Lindau M., De Camilli P., Toomre D. Proc. Natl. Acad. Sci. USA. 2005;102:5204–5209. doi: 10.1073/pnas.0501412102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Milosevic I., Sorensen J. B., Lang T., Krauss M., Nagy G., Haucke V., Jahn R., Neher E. J. Neurosci. 2005;25:2557–2565. doi: 10.1523/JNEUROSCI.3761-04.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Di Paolo G., Moskowitz H. S., Gipson K., Wenk M. R., Voronov S., Obayashi M., Flavell R., Fitzsimonds R. M., Ryan T. A., De Camilli P. Nature. 2004;431:415–422. doi: 10.1038/nature02896. [DOI] [PubMed] [Google Scholar]
- 9.Galli T., Haucke V. Sci. STKE. 2004 doi: 10.1126/stke.2642004re19. 2004, re19. [DOI] [PubMed] [Google Scholar]
- 10.Wenk M. R., De Camilli P. Proc. Natl. Acad. Sci. USA. 2004;101:8262–8269. doi: 10.1073/pnas.0401874101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Haucke V. Biochem. Soc. Trans. 2005;33:1285–1289. doi: 10.1042/BST0331285. [DOI] [PubMed] [Google Scholar]
- 12.Gaidarov I., Keen J. H. J. Cell Biol. 1999;146:755–764. doi: 10.1083/jcb.146.4.755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Rohde G., Wenzel D., Haucke V. J. Cell Biol. 2002;158:209–214. doi: 10.1083/jcb.200203103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ford M. G., Mills I. G., Peter B. J., Vallis Y., Praefcke G. J., Evans P. R., McMahon H. T. Nature. 2002;419:361–366. doi: 10.1038/nature01020. [DOI] [PubMed] [Google Scholar]
- 15.Vallis Y., Wigge P., Marks B., Evans P. R., McMahon H. T. Curr. Biol. 1999;9:257–260. doi: 10.1016/s0960-9822(99)80114-6. [DOI] [PubMed] [Google Scholar]
- 16.Bonifacino J. S., Traub L. M. Annu. Rev. Biochem. 2003;72:395–447. doi: 10.1146/annurev.biochem.72.121801.161800. [DOI] [PubMed] [Google Scholar]
- 17.Honing S., Ricotta D., Krauss M., Spate K., Spolaore B., Motley A., Robinson M., Robinson C., Haucke V., Owen D. J. Mol. Cell. 2005;18:519–531. doi: 10.1016/j.molcel.2005.04.019. [DOI] [PubMed] [Google Scholar]
- 18.Cremona O., Di Paolo G., Wenk M. R., Luthi A., Kim W. T., Takei K., Daniell L., Nemoto Y., Shears S. B., Flavell R. A., et al. Cell. 1999;99:179–188. doi: 10.1016/s0092-8674(00)81649-9. [DOI] [PubMed] [Google Scholar]
- 19.Harris T. W., Hartwieg E., Horvitz H. R., Jorgensen E. M. J. Cell Biol. 2000;150:589–600. doi: 10.1083/jcb.150.3.589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Verstreken P., Koh T. W., Schulze K. L., Zhai R. G., Hiesinger P. R., Zhou Y., Mehta S. Q., Cao Y., Roos J., Bellen H. J. Neuron. 2003;40:733–748. doi: 10.1016/s0896-6273(03)00644-5. [DOI] [PubMed] [Google Scholar]
- 21.Ishihara H., Shibasaki Y., Kizuki N., Wada T., Yazaki Y., Asano T., Oka Y. J. Biol. Chem. 1998;273:8741–8748. doi: 10.1074/jbc.273.15.8741. [DOI] [PubMed] [Google Scholar]
- 22.Fruman D. A., Meyers R. E., Cantley L. C. Annu. Rev. Biochem. 1998;67:481–507. doi: 10.1146/annurev.biochem.67.1.481. [DOI] [PubMed] [Google Scholar]
- 23.Di Paolo G., Pellegrini L., Letinic K., Cestra G., Zoncu R., Voronov S., Chang S., Guo J., Wenk M. R., De Camilli P. Nature. 2002;420:85–89. doi: 10.1038/nature01147. [DOI] [PubMed] [Google Scholar]
- 24.Ling K., Doughman R. L., Firestone A. J., Bunce M. W., Anderson R. A. Nature. 2002;420:89–93. doi: 10.1038/nature01082. [DOI] [PubMed] [Google Scholar]
- 25.Padron D., Wang Y. J., Yamamoto M., Yin H., Roth M. G. J. Cell Biol. 2003;162:693–701. doi: 10.1083/jcb.200302051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Barbieri M. A., Heath C. M., Peters E. M., Wells A., Davis J. N., Stahl P. D. J. Biol. Chem. 2001;276:47212–47216. doi: 10.1074/jbc.C100490200. [DOI] [PubMed] [Google Scholar]
- 27.Wenk M. R., Pellegrini L., Klenchin V. A., Di Paolo G., Chang S., Daniell L., Arioka M., Martin T. F., De Camilli P. Neuron. 2001;32:79–88. doi: 10.1016/s0896-6273(01)00456-1. [DOI] [PubMed] [Google Scholar]
- 28.Krauss M., Kinuta M., Wenk M. R., De Camilli P., Takei K., Haucke V. J. Cell Biol. 2003;162:113–124. doi: 10.1083/jcb.200301006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Aikawa Y., Martin T. F. J. Cell Biol. 2003;162:647–659. doi: 10.1083/jcb.200212142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Bairstow S. F., Ling K., Su X., Firestone A. J., Carbonara C., Anderson R. A. J. Biol. Chem. 2006;281:20632–20642. doi: 10.1074/jbc.M601465200. [DOI] [PubMed] [Google Scholar]
- 31.Guo J., Wenk M. R., Pellegrini L., Onofri F., Benfenati F., De Camilli P. Proc. Natl. Acad. Sci. USA. 2003;100:3995–4000. doi: 10.1073/pnas.0230488100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Paleotti O., Macia E., Luton F., Klein S., Partisani M., Chardin P., Kirchhausen T., Franco M. J. Biol. Chem. 2005;280:21661–21666. doi: 10.1074/jbc.M503099200. [DOI] [PubMed] [Google Scholar]
- 33.Walther K., Diril M. K., Jung N., Haucke V. Proc. Natl. Acad. Sci. USA. 2004;101:964–969. doi: 10.1073/pnas.0307862100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Arneson L. S., Kunz J., Anderson R. A., Traub L. M. J. Biol. Chem. 1999;274:17794–17805. doi: 10.1074/jbc.274.25.17794. [DOI] [PubMed] [Google Scholar]
- 35.Ehrlich M., Boll W., Van Oijen A., Hariharan R., Chandran K., Nibert M. L., Kirchhausen T. Cell. 2004;118:591–605. doi: 10.1016/j.cell.2004.08.017. [DOI] [PubMed] [Google Scholar]
- 36.Haucke V., Krauss M. Eur. J. Cell Biol. 2002;81:647–653. doi: 10.1078/0171-9335-00289. [DOI] [PubMed] [Google Scholar]
- 37.Owen D. J., Evans P. R. Science. 1998;282:1327–1332. doi: 10.1126/science.282.5392.1327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Meyer D. M., Crottet P., Maco B., Degtyar E., Cassel D., Spiess M. Mol. Biol. Cell. 2005;16:4745–4754. doi: 10.1091/mbc.E05-06-0568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Rao V. D., Misra S., Boronenkov I. V., Anderson R. A., Hurley J. H. Cell. 1998;94:829–839. doi: 10.1016/s0092-8674(00)81741-9. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.