Abstract
The retina is subject to degenerative conditions, leading to blindness. Although retinal regeneration is robust in lower vertebrates, regeneration does not occur in the adult mammalian retina. Thus, we have developed efficient methods for deriving retinal neurons from human embryonic stem (hES) cells. Under appropriate culture conditions, up to 80% of the H1 line can be directed to the retinal progenitor fate, and express a gene expression profile similar to progenitors derived from human fetal retina. The hES cell-derived progenitors differentiate primarily into inner retinal neurons (ganglion and amacrine cells), with functional glutamate receptors. Upon coculture with retinas derived from a mouse model of retinal degeneration, the hES cell derived retinal progenitors integrate with the degenerated mouse retina and increase in their expression of photoreceptor-specific markers. These results demonstrate that human ES cells can be selectively directed to a neural retinal cell fate and thus may be useful in the treatment of retinal degenerations.
Keywords: photoreceptors, eye development, neurogenesis
The neural retina is subject to a number of degenerative conditions, including retinitis pigmentosa, age-related macular degeneration, and glaucoma. Although there are a number of sources of progenitors for regeneration in nonmammalian vertebrates, these are greatly reduced or absent in the adult mammalian retina (1). By contrast, recent reports show that retinal progenitor cells can be derived from mouse ES cells (2, 3), and may provide an alternative to adult derived retinal stem cells. In other regions of the central nervous system, the transplantation of neurons derived from embryonic stem cells has led to some promising results. Dopaminergic neurons derived from mouse, monkey, and human embryonic stem cells have been shown to integrate into the brain after transplantation and partially restore function in animal models of Parkinson’s disease (4–8). Oligodendrocytes derived from ES cells can repair some of the damage caused by spinal cord trauma (9) as well as in mouse models of spinal demyelination (10–12).
Results
We have developed methods for deriving retinal neurons from human embryonic stem (hES) cells. The current molecular genetic model of vertebrate embryogenesis (13) suggests that there are several sequential induction steps. Forebrain development requires that both BMP and Wnt signaling are antagonized (14–19). Although the specific molecular signals required for eye field specification are not completely defined in any model system, insulin-like growth factor-1 (IGF-1) mRNA injections into Xenopus embryos specifically promote eye induction (20). Therefore, to direct the ES cells to an anterior neural fate, we treated embryoid bodies with a combination of noggin (a potent endogenous inhibitor of the BMP pathway) and Dickkopf-1 (dkk1; a secreted antagonist of the Wnt/β-catenin signaling pathway (14, 21)) and IGF-1. The embryoid bodies were cultured for 3 days in the three factors (Fig. 1A) and then transferred to six-well plates coated with either Matrigel or laminin where they were allowed to attach. The cells were then maintained in the same medium, with bFGF added, for an additional 3 weeks; we refer to this protocol as retinal determination (RD) conditions.
Fig. 1.
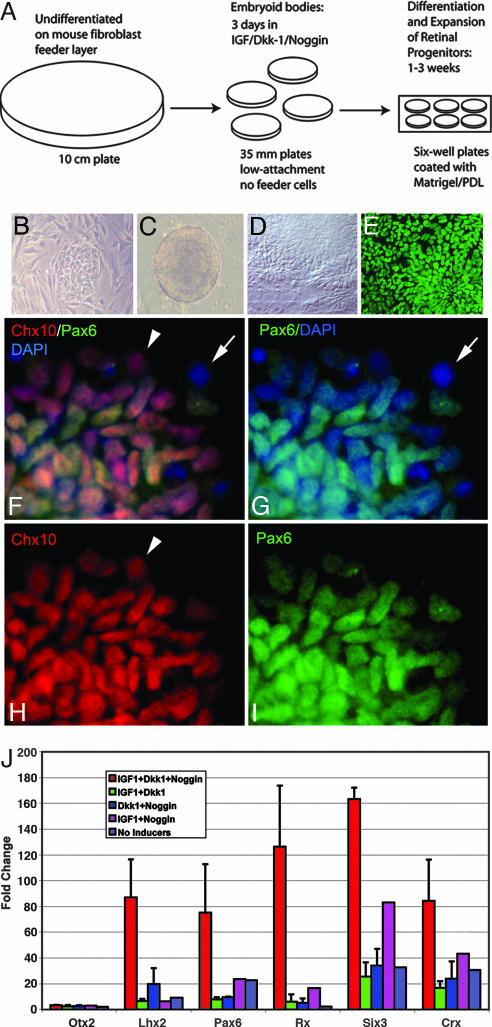
Efficient Retinal Induction of hES cells. (A) A schematic of the 3-week retinal determination protocol. (B–D) Bright-field phase images of the progression of undifferentiated hES cells (B) through embryoid body (C) to formation of neural rosettes (D). At the end of 3 weeks, ≈80% cells show immunoreactivity to retinal progenitor markers like Pax6 (E, G, and I) and Chx10 (F and H). (F–I) Coexpression of Pax6 (green) and Chx10 (red) by a group of cells labeled by DAPI in blue. Arrows in F and G point to a cell that expresses neither proteins, whereas arrowhead s in F and H point to a cell expressing Chx10 but not Pax6. For maximal induction of the eye-field transcription factors Pax6, Six3, and Rx, IGF1, Dkk1, and noggin are required. The experiment was performed by using the protocol and concentrations as described except that either IGF1, Dkk1, Noggin, or all three were omitted from the media. At the end of 1 week, comparison of the gene expression for the eye field transcription factors in each of the five cases (IGF1 + Dkk1+ Noggin, Dkk1+ noggin, IGF1 + noggin, IGF1 + Dkk1 and No inducers) was done by using quantitative RT-PCR (J). QPCR analysis of EFTFs at the end of 1 week under retinal determination conditions (n = 3; mean ± SEM) shows a 6–8 cycle (≈75- to 165-fold) increase in various retinal stem cell markers (Pax6, Lhx2, Rx, and Six3). None of the other conditions has a comparable inductive effect.
Previous work has shown that the presumptive eye field is defined by a group of transcription factors expressed in this region (eye field transcription factors; EFTFs), including ET, Rx, Pax6, Six3, Lhx2, tll and Optx2/Six6 (22). After the ES cells had been exposed to the RD conditions for one, two, or three weeks, we harvested the mRNA and analyzed the levels of expression for the EFTFs. Fig. 1J shows the relative expression levels for cells after 1 week in RD conditions for H-1 cell line (Hsf6 data in Fig. 5B, which is published as supporting information on the PNAS web site). We found a 6–8 cycle (75- to 165-fold) increase in the expression of the EFTFs, including Rx, Pax6, Lhx2, and Six3 (Fig. 1J) over the levels in the undifferentiated cells or cells differentiated without inducers present. By contrast, genes expressed in nonneural tissues or other regions of the CNS, like the cerebral cortex [Emx-1, Arx (not expressed)], or hindbrain (Engrailed-1) show no significant increase over the undifferentiated ES cells (Fig. 5A). Although previous reports indicated Noggin or Dkk-1 would promote anterior neural identity, the addition of IGF-1 specifically promotes retinal progenitor identity; leaving this factor out of the RD medium dramatically reduced the level of retinal progenitor gene expression (Fig. 1J). We also characterized the hES cells with immunofluorescence for retinal progenitor markers. Fig. 1 E–I shows the extensive labeling for Pax6 and Chx10, two transcription factors characteristic of retinal progenitors. Quantitative analysis of the cultures showed that 82% (SD ± 23%) of the cells were labeled with Pax6 antibodies at the end of 3 weeks in RD conditions. Of these, 86% coexpressed Chx10 (SD ± 14%). Most of the Pax6-labeled cells were also labeled with antibodies to Sox2 (data not shown).
These data demonstrate that a large fraction of the hES cells in cultures kept under RD conditions develop characteristics of retinal progenitors. To determine whether the hES cell-derived progenitors have the capacity for multilineage differentiation characteristic of retinal progenitors, we used immunofluorescence for specific types of retinal neurons, including HuC/D, Neurofilament-M, and Tuj-1 for ganglion and amacrine cells, Crx, Nrl, recoverin, S-opsin, and rhodopsin for photoreceptors, and Prox1 for amacrine and horizontal cells. All of these markers have been previously described in retinal neurons of various species, and their expression is shown in developing human retina in Figs. 6 and 7, which are published as supporting information on the PNAS web site. As noted above, most cells in the cultures express Pax6; however, some cells showed a distinct increase in the Pax6 level, characteristic of ganglion cells and amacrine cells (Fig. 2A). The cells that express a high level of Pax6 are also labeled with antibodies against HuC/D, another protein expressed by amacrine and ganglion cells (Fig. 2B). Primary cell cultures derived from a 78-day human fetal retina show a very similar pattern of labeling with Pax6 and HuC/D (Fig. 2 C and D), as do sections from the developing human retina (Fig. 6). Many cells in the hES cultures also label with other markers of ganglion and amacrine cells: Tuj-1 and Neurofilament-M (Fig. 2 E and F and Fig. 7 A and B), whereas other cells are labeled with photoreceptor-specific antibodies, including Crx, S-opsin, rhodopsin, and Nrl (Fig. 2 G–J and L), bipolar cell marker PKCα (Fig. 2K), and horizontal cell marker Prox-1 (Fig. 7 C–E). Twelve percent of all cells expressed Crx (SD ± 2.4), 12% of all cells expressed Hu C/D (SD ± 6.7), and 5.75% (SD ± 4.2) of the cells expressed Nrl (1,646 cells counted). S-opsin and rhodopsin were expressed in a very small percentage of cells (<0.01%). In addition to the antibody labeling, we analyzed the cultures for expression of genes associated with retinal photoreceptors by QPCR (Fig. 2N). There is a large and stable increase in the level of Crx (the earliest known photoreceptor marker) in the cells, as early as 1 week in RD conditions. Although the other photoreceptor differentiation markers, like the opsins, PDE-β, and recoverin, show only modest increases at 1 week, they steadily increase over the time in vitro (Fig. 2N). This finding is consistent with the developmental timing of expression of these genes in the retina: first Crx, then recoverin, and PDE-β, and lastly the opsin genes.
Fig. 2.
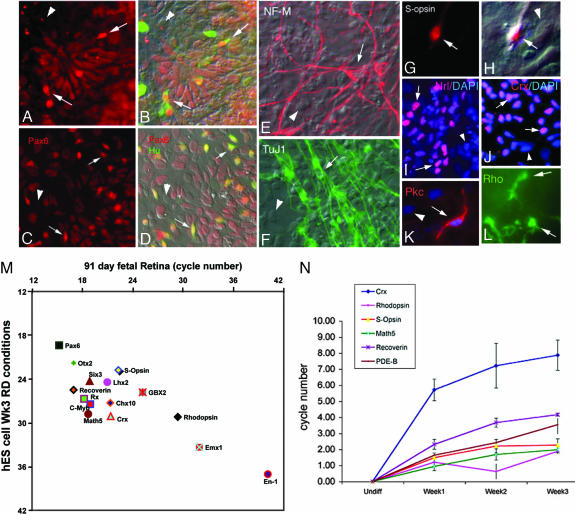
Multilineage differentiation of hES cell-derived retinal progenitors. (A) Two levels of Pax6 expression in these cells. Progenitors express lower levels of Pax6, whereas differentiating ganglion and amacrine cells express high levels of Pax6 and coexpress Hu C/D (green, B). (C and D) Similar pattern of Pax6 and Hu C/D expression in dissociated 78-day human fetal retinal cultures for comparison. (E and F) Human ES cells in RD conditions label for neurofilament-M (E) and Tuj-1 (F). (G and H) A cell expressing S-opsin in UV and Nomarski optics, respectively. (L) Cells labeled with the rod photoreceptor marker Rho-4D2 (rhodopsin). Many cells express the rod photoreceptor specific transcription factor, Nrl (red, I) or pan-photoreceptor marker, Crx (red, J); nuclei are also labeled with DAPI (blue). (K) Cell labeled with bipolar cell marker PKCα (red) with the nuclei labeled with DAPI (blue). In A–K, an arrow marks and labels cell in the field, whereas an arrowhead indicates a cell not expressing that protein. (M) Comparative QPCR analysis of expression of various genes between a 91 day human fetal retina and hES cells after 3 weeks under RD conditions, showing the correlation between hES cell derived retinal cells and retinal cells isolated from fetal human retina. (N) QPCR analysis of retinal differentiation genes (n = 3). The graph shows a steady increase in retinal neuronal markers Crx, Math5, S-opsin, rhodopsin, recoverin, and PDE-β over the 3 weeks of induction.
To more precisely determine the degree to which the hES cell-derived retinal progenitors resemble those from the developing retina, we collected mRNA from the retinas of a human fetus at 91 days after conception, an age when there are both progenitors and newly differentiating neurons. When we compare EFTF expression and the expression of genes specific to differentiating neurons, between the fetal retina and hES cells kept under RD conditions for 3 weeks, we find an excellent correlation (Fig. 2M). The degree of expression of the EFTFs is similar in the hES cell-derived retinal progenitors and those obtained from the fetal retina. However, although all of the photoreceptor markers are expressed in both groups of cells, their expression is somewhat reduced in the hES cell derived progenitors, as compared to the fetal-derived cells, suggesting that the RD conditions promote the expansion of the progenitor pool relative to the differentiating neurons and photoreceptors. These results show that hES cells, under RD conditions, develop a gene and protein expression profile highly reminiscent of normal human retinal progenitors, neurons, and photoreceptor cells.
To analyze the functional maturation of the retinal neurons produced in these cultures, we redissociated the cells and plated them at lower density. This method allowed us to analyze small clusters of cells with calcium imaging techniques. We found that some of the cells, particularly those with distinct neuron-like morphology, respond to glutamate and NMDA with substantial calcium fluxes (Fig. 3A–E). Because most inner retinal neurons have glutamate receptors, and retinal ganglion cells express NMDA receptors, these data lend further support to the immunofluorescent identification of ganglion cells and amacrine cells. In addition to functional glutamate receptors, cells with neuronal morphology displayed synaptophysin labeled puncta, consistent with synaptic development in vitro (Fig. 3 F and G).
Fig. 3.
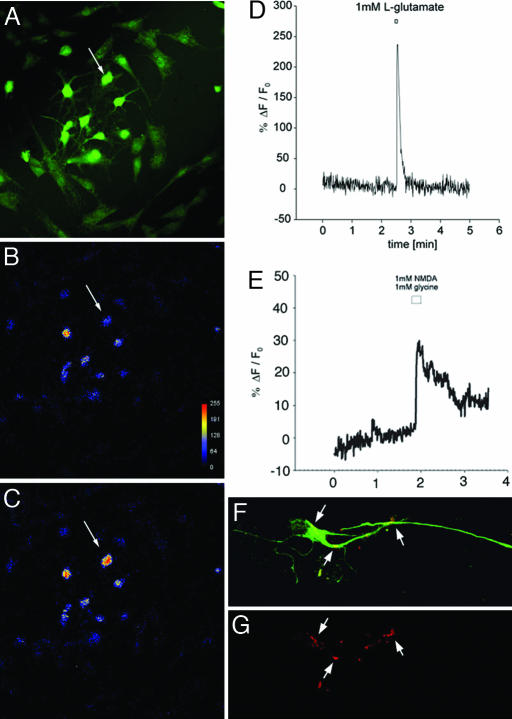
Glutamate and NMDA induced calcium changes in hES cell-derived retinal neurons. (A) Oregon Green BAPTA-1 a.m. loading of the hES derived neurons. (B) The same cells at baseline using a rainbow LUT palette. (C) The same field of cells is shown immediately following application of 1 mM glutamate. Arrows in A–C indicate a cell with large calcium transient after application. (D) The calcium change in the same cell expressed as a pseudoratio of fluorescence change expressed as a % ΔF/F0 over time. (E) A similar calcium change in a cell stimulated with 1 mM NMDA in the presence of 1 mM glycine, again expressed as a pseudoratio of fluorescence change. In each case at least four different preparations were analyzed (n = 4). (F and G) Cells with neuronal morphology and expressing internexin (green) show punctuate labeling with synaptophysin (red) antibody, a protein expressed in synapses.
Although the majority of well differentiated neurons in the cultures display characteristics of amacrine and ganglion cells, many cells also express markers for immature photoreceptors, including Crx and Nrl. However, only a very few cells express markers of more differentiated photoreceptors. Several groups have found that coculturing progenitors or stem cells with developing retina promotes photoreceptor differentiation (3). To stimulate photoreceptor differentiation from the hES cells, we cocultured the hES cell-derived progenitors with retinal explants from adult mice on 0.4-μm filters. The cocultures were allowed to survive up to 6 days, and analyzed for evidence of hES cell differentiation and integration into the mouse retina. The mice had GFP stably expressed in all cells (via the chicken β-actin promoter/CMV enhancer) and are easily distinguished from the human cells. In addition, we labeled the human cells with one of two different human-specific antibodies: a human nuclear antibody and a human mitochondrial antibody (Fig. 4G and Fig. 8 A and B, which is published as supporting information on the PNAS web site). The retinal progenitors derived from hES cells integrated with the mouse retina, and most express Pax6 (Fig. 4 A and B). We cocultured the hES cells with both wild-type mouse retinas and with retinas from mice with a mutation that causes photoreceptor degeneration (Aipl1−/−). In the cocultures with the Aipl1−/− retinas, recoverin immunoreactive cells, that resemble photoreceptors, are observed (Fig. 4 C–G); interestingly, recoverin-immunoreactive cells are rarely found when hES cells are cocultured with wild-type mouse retina (Fig. 4H). Coculturing the hES cell-derived progenitors also caused an increase in the number of cells labeled with the other photoreceptor markers: Rho-4D2 and Nrl (Fig. 4 I and J).
Fig. 4.
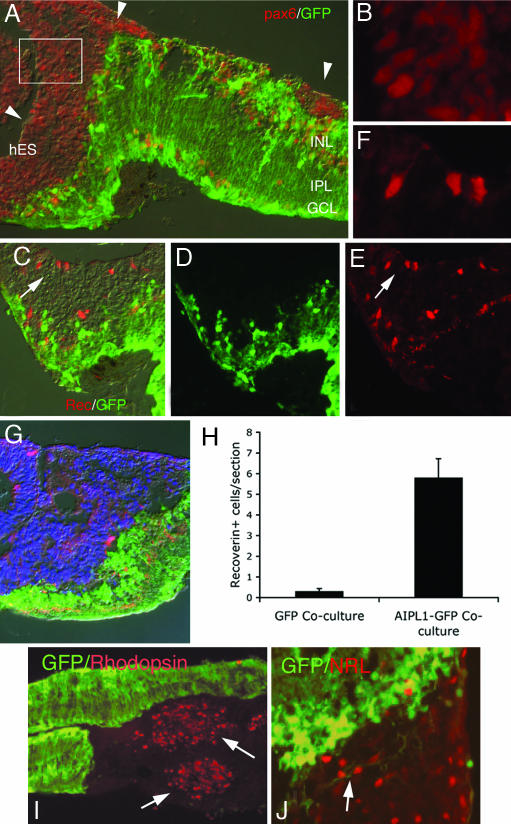
Explant coculture of hES-derived retinal progenitors with retinas of Aipl1−/− GFP mice. (A) The mouse retina in green (GFP) with the outer nuclear layer missing (degenerated). Many of the hES cells express Pax6. (B) An enlarged view of boxed area in a. (C–F) Recoverin expression, a marker of photoreceptors, in the cocultures. (C) A merged view of D and E, where D is showing mouse cells expressing GFP and E shows recoverin expression in a number of hES cell-derived neurons. Some mouse bipolars show recoverin and GFP co-expression. (F) A higher magnification of the region identified by the arrow in C–E showing recoverin expressing hES cell-derived retinal neurons. (G) Confirmation of the identity of the recoverin (red) expressing cells in hES region using colabeling with human specific nuclear marker (blue). (H) Graph showing number of recoverin expressing hES cell-derived neurons per section (n = 3; mean ± SEM) from cocultures with wild-type mice expressing GFP and Aipl1−/− GFP mice. (I) Transplanted human cells expressing rod photoreceptor marker, Rho-4D2. (J) Transplanted human cells expressing another rod photoreceptor specific marker, Nrl.
Discussion
Our data demonstrate that human ES cells can be directed to a retinal progenitor identity with high efficiency by using a combination of noggin, dkk1 and IGF-1. Ikeda et al. recently reported that mouse ES cells could be directed to an RX+/Pax6+ retinal progenitor identity at reasonable efficiency (26%) using a combination of dkk1, lefty (a nodal antagonist), FCS, and activin (3). This group has not found noggin to be effective in the mouse ES cells. Although we have not tested their specific conditions for the human ES cells, a key difference between human and mouse ES cells appears to be their different response to noggin. Other investigators have tested the Wnt or BMP antagonism in human ES to produce other anterior neural tissues, and so this combination is likely to generally promote production of various regions of anterior CNS, including cerebral cortex (23, 24). However, the addition of IGF-1 to the embryoid bodies specifically and efficiently directs the cells to a retinal progenitor identity, as evidenced by the fact that we found only very low levels of expression of genes associated with cerebral cortex or midbrain in our cultures. This result further strengthens the concept that information from model developmental systems can be applied in the design of conditions to direct hES cells to specific fates.
One of the most striking features of these cultures is that they are accelerated in “developmental time” over normal human embryological development. Less than 2 weeks after an undifferentiated state, the cells have acquired the characteristics of the eye field, and specifically the neural retina part of the optic cup. The optic vesicle does not become distinct in the human embryo until Streeter’s horizon 11 or 12 (25), or postovulatory day ≈24–26. The development of an inner “neuroblastic layer” and optic fibers, indicative of the onset of retinal ganglion cell genesis, does not occur until approximately the 6th postovulatory week. Consistent with this finding is the fact that the gene expression profile of the hES cells after 3 weeks in RD conditions resembles that of the 91-day (after conception) fetal retina (albeit with a reduced expression of photoreceptor genes). The hES cells are therefore accelerated by 3–4 weeks over the normal human developmental time course.
The degeneration of neurons and photoreceptors in the retina that occur in a number of disorders are common causes of blindness for which there are currently few therapies. The repair of the retina with replacement cells derived from human ES cells could thus provide the basis for new approaches for treating a wide variety of retinal degenerations. In this study, we report a method for directing pluripotent hES cells to a retinal progenitor identity. The hES-derived retinal progenitors are highly similar to retinal progenitors derived from human fetal stages, using both quantitative PCR and immunofluorescence labeling. The retinal progenitor cells are able to integrate with retinas from a mouse model of Leber’s congenital amaurosis caused by a mutation in the Aipl1 gene that causes photoreceptor degeneration, and cells expressing photoreceptor markers differentiate from the hES cells in these cocultures. Thus, hES cells may provide an excellent source of new neurons and photoreceptors for retinal repair.
Materials and Methods
Cell Culture and Retinal Induction.
The H-1 (WA-01) human embryonic stem cell line was obtained from Wicell Research Institute. The cells were cultured and passaged on a feeder layer made of irradiated mouse embryonic fibroblasts. Embryoid bodies were formed by treating undifferentiated hES colonies with 1 mg/ml type IV collagenase (Invitrogen, Carlsbad, CA) and resuspending ≈150 100-cell clumps per ml in a six-well ultra-low attachment plate (VWR) in the presence of media containing DMEM:F12 (Gibco, Grand Island, NY), 10% knockout serum (Invitrogen), B-27 supplement (Invitrogen), 1 ng/ml mouse noggin (R & D Systems, Minneapolis, MN), 1 ng/ml human recombinant Dkk-1 (R & D Systems) and 5 ng/ml human recombinant insulin-like growth factor-1 (IGF-1) (R & D Systems). The cells were cultured as embryoid bodies for 3 days. On the fourth day, 25–35 embryoid bodies were plated onto each poly(d-lysine)-Matrigel (Collaborative Research, Waltham, MA) coated 35-mm plates and cultured in the presence of DMEM:F12, B-27 supplement, N-2 Supplement (Invitrogen), 10 ng/ml mouse noggin, 10 ng/ml human recombinant Dkk-1, 10 ng/ml human recombinant IGF-1, and 5 ng/ml human recombinant basic fibroblast growth factor (bFGF) (R & D Systems). The media was changed every 2–3 days. Hsf6 hES cell line experiments were carried out in a similar manner at the stem cell core facility at the University of Washington.
Immunocytochemistry and Immunohistochemistry.
Cells and eyes were fixed with 4% paraformaldehyde and analyzed with the following antibodies: rabbit anti-recoverin (a generous gift from Jim Hurley, University of Washington, Seattle, WA), mouse anti-Tuj-1 (Covance, Austin, TX), mouse anti-Hu C/D (Molecular Probes, Eugene, OR), rabbit anti-neurofilament-M (Chemicon, Billerica, MA), mouse anti-NCAM (DHSB, Iowa City, IA), mouse anti-Rho-4D2 (gift from R. S. Molday, University of British Columbia, Vancouver, BC, Canada), mouse anti-Pax6 (DHSB), rabbit anti-Pax6 (CRP, Denver, PA), mouse anti-human nuclear antigen (Chemicon), mouse anti-human mitochondrial antigen (Chemicon), rabbit anti-GFP (University of Alberta), mouse anti-GFP (Molecular Probes), goat anti-Sox2 (Santa Cruz Biotechnology, Santa Cruz, CA), rabbit anti-Chx10 (gift from T. M. Jessel, Columbia University, New York, NY), rabbit anti-Prox1 (Chemicon), rabbit anti-S-opsin (gift from J. Nathans, Johns Hopkins University, Baltimore, MD), rabbit anti-Nrl (gift from A. Swaroop, University of Michigan, Ann Arbor, MI), rabbit anti-Crx (gift from C. Gregory-Evans, Imperial College, London, U.K.), rabbit anti-internexin (Chemicon), rabbit anti-PKCα (Research Diagnostics, Concord, MA) and mouse anti-synaptophysin (Chemicon). Secondary antibody staining was done using the corresponding Alexa Fluor 488, Alexa Fluor 568, and Alexa Fluor 350 fluorescent-tagged antibodies (Molecular Probes).
Real-Time Quantitative PCR Analysis (QPCR).
Total RNA was extracted from the cultures by using TriZol (Invitrogen) followed by chloroform extraction as per manufacturer’s instructions. This was followed by DNase-1 (Qiagen, Valencia, CA) treatment followed by RNA cleanup using Qiagen RNA mini cleanup kit. cDNA was reverse transcribed by using Superscript II RT kit (Invitrogen) as per the manufacturer’s instructions. QPCR was performed for various genes (Table 1, which is published as supporting information on the PNAS web site), and results were normalized to β-actin levels.
Animals.
All experiments were done in accordance with approved protocols and the animals were housed and bred in the Department of Comparative Medicine at the University of Washington. Wild-type and Aipl1−/− mice (26) bred onto a global GFP expressing background (The Jackson Laboratory, Bar Harbor, ME) were used for in vitro explant coculture experiments.
In Vitro Explant Coculture Experiments.
Whole retinas were cultured on a nitrocellulose membrane for up to two weeks in vitro, in a method modified from Caffe et al. (27). Briefly, retinas from mice were dissected free from the lens, pigmented epithelium, and extra-ocular tissue in Hanks’s balanced salt solution (HBSS), four small incisions were made in the peripheral retina to allow better flattening, and retinas were placed photoreceptor side down on a Millicell-CM 0.4-μm filter insert. Filters were placed into a six-well plate containing 1 ml of explant media [DMEM:F12 (Gibco), 0.6% glucose, 5 nM Hepes, 0.11% NaHCO3, 25 μg/ml insulin, 100 μg/ml transferrin, 60 μM putrescine, 30 nM selenium, 20 nM progesterone, 800 nM l-glutamine, penicillin and streptomycin (Gibco), N2 supplement, and 10% FBS (Gibco)]. Explants were cultured at the gas-liquid interface at 37°C, 5% CO2 and media was replaced every other day. The day after placement of explants onto the nitrocellulose membrane, ≈20 100-cell ES cell clumps were gently dropped onto the surface of the explant. The explants were maintained for 6 days on the nitrocellulose membrane. For immunohistochemical analysis, the explants were fixed with 4% PFA and gently lifted off the membrane. They were subsequently embedded in OCT and cryosectioned.
Calcium Imaging of Glutamate Responses.
Intracellular calcium ([Ca2+]i) transients in hES cells after >3 weeks in RD conditions were studied. Cells were kept serum-free for 1 h and then loaded with 8 μM acetomethoxy (AM) ester form of Oregon green 488 BAPTA-1 (Molecular Probes) in bath solution (see below) with 0.05 mg/ml pluronic acid for 30 min at 37°C (95% O2/5% CO2). Cells were transferred to a recording chamber (1-ml volume) and perfusated with bath solution (adapted from the cell culture media containing 119 mM NaCl, 4.16 mM KCl, 2.5 mM CaCl2, 0.3 mM MgCl2, 0.4 mM MgSO4, 0.5 mM Na2HPO4, 0.45 mM NaH2PO4, 20 mM Hepes, and 19 mM glucose) at rate of 1 ml/min at 35–37°C and pH of 7.4. Fluorescence was monitored by using a laser-scanning confocal microscope (Zeiss LSM5/Axioskop2 MOT, Jena, Germany). Fluorescence images (excitation at 488 nm; emission >505 nm; maximal pinhole opening) were acquired at 2 Hz and 387 ms exposure time per image at a resolution of 256 × 256 pixels. Images were processed and analyzed by using ImageJ software (http://rsb.info.nih.gov/ij). The fluorescence changes (%ΔF/F0) for individual cells were calculated by using the formula % ΔF/F0 = (F1 − F0) × 100/F0, where F1 was the fluorescence averaged over the pixel of a cell soma after a stimulus, whereas F0 was the average fluorescence of that cell before stimulus application, averaged over three images. Background intensity was zero. Before drug application, cells were perfused for >10 min. For stimulation experiments, a glass electrode was filled with either bath solution (control) or bath solution containing 1 mm of glutamate. The electrode was connected to a PicoSpritzer (PLI-100; Medical Systems, Greenvale, NY), which delivered 100-ms air puffs that propelled 10 μl of solution over the area of interest during continuous perfusion. All data presented were analyzed by using Student′s t test.
Human Fetal Eyes.
Eyes from 78- 95-days-postconception fetuses, without identifiers, were obtained from therapeutic abortions through the fetal tissue bank at the University of Washington. Individual eyes were rinsed with sterile HBSS; retinas were then dissected from other ocular tissue. The retinas were then either cultured on Matrigel-coated coverslips or fixed in 4% paraformaldehyde for subsequent freezing in OCT or used for RNA extraction using TriZol.
Supplementary Material
Acknowledgments
We thank Drs. Tony Blau and C. Murray for advice in human ES cell culture, the Stem Cell Core technical expertise of Angelique Nelson, without whom this work would not have been possible; members of the Reh laboratory for critical discussions over the course of the experiments; Drs. Jim Hurley and Visvanathan Ramamurthy (University of Washington) for the Aipl1−/− mice; Dr. Rachel Wong (University of Washington) for help with the calcium imaging; Dr. Olivia Bermingham-McDonogh (University of Washington) for helpful comments on the manuscript; and Paige Etter for technical assistance and manuscript editing. This work was supported by National Institutes of Health Grants P20GM69983, and NS28308 (to T.A.R.).
Abbreviations
- hES
human embryonic stem
- IGF-I
insulin-like growth factor I
- RD
retinal determination.
Footnotes
Conflict of interest statement: No conflicts declared.
This paper was submitted directly (Track II) to the PNAS office.
References
- 1.Moshiri A., Close J., Reh T. A. Int. J. Dev. Biol. 2004;48:1003–1014. doi: 10.1387/ijdb.041870am. [DOI] [PubMed] [Google Scholar]
- 2.Meyer J. S, Katz M. L, Maruniak J. A., Kirk M. D. Stem Cells. 2006;24:274–283. doi: 10.1634/stemcells.2005-0059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ikeda H., Osakada F., Watanabe K., Mizuseki K., Haraguchi T., Miyoshi H., Kamiya D., Honda Y., Sasai N., Yoshimura N., et al. Proc. Natl. Acad. Sci. USA. 2005;102:11331–11336. doi: 10.1073/pnas.0500010102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bjorklund L. M., Sanchez-Pernaute R., Chung S., Andersson T., Chen I. Y., McNaught K. S., Brownell A. L., Jenkins B. G., Wahlestedt C., Kim K. S., Isacson O. Proc. Natl. Acad. Sci. USA. 2002;99:2344–2349. doi: 10.1073/pnas.022438099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kim J. H., Auerbach J. M., Rodriguez-Gomez J. A., Velasco I., Gavin D., Lumelsky N., Lee S. H., Nguyen J., Sanchez-Pernaute R., Bankiewicz K., McKay R. Nature. 2002;418:50–56. doi: 10.1038/nature00900. [DOI] [PubMed] [Google Scholar]
- 6.Nishimura F., Yoshikawa M., Kanda S., Nonaka M., Yokota H., Shiroi A., Nakase H., Hirabayashi H., Ouji Y., Birumachi J., et al. Stem Cells. 2003;21:171–180. doi: 10.1634/stemcells.21-2-171. [DOI] [PubMed] [Google Scholar]
- 7.Takagi Y., Takahashi J., Saiki H., Morizane A., Hayashi T., Kishi Y., Fukuda H., Okamoto Y., Koyanagi M., Ideguchi M., et al. J. Clin. Invest. 2005;115:102–109. doi: 10.1172/JCI21137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sanchez-Pernaute R., Studer L., Ferrari D., Perrier A., Lee H., Vinuela A., Isacson O. Stem Cells. 2005;23:914–922. doi: 10.1634/stemcells.2004-0172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.McDonald J. W., Liu X. Z., Qu Y., Liu S., Mickey S. K., Turetsky D., Gottlieb D. I., Choi D. W. Nat. Med. 1999;5:1410–1412. doi: 10.1038/70986. [DOI] [PubMed] [Google Scholar]
- 10.Nistor G. I., Totoiu M. O., Haque N., Carpenter M. K., Keirstead H. S. Glia. 2005;49:385–396. doi: 10.1002/glia.20127. [DOI] [PubMed] [Google Scholar]
- 11.Liu S., Qu Y., Stewart T. J., Howard M. J., Chakrabortty S., Holekamp T. F., McDonald J. W. Proc. Natl. Acad. Sci. USA. 2000;97:6126–6131. doi: 10.1073/pnas.97.11.6126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Mueller D., Shamblott M. J., Fox H. E., Gearhart J. D., Martin L. J. J. Neurosci. Res. 2005;82:592–608. doi: 10.1002/jnr.20673. [DOI] [PubMed] [Google Scholar]
- 13.Zaghloul N. A., Yan B., Moody S. A. Biol. Cell. 2005;97:321–337. doi: 10.1042/BC20040521. [DOI] [PubMed] [Google Scholar]
- 14.Mukhopadhyay M., Shtrom S., Rodriguez-Esteban C., Chen L., Tsukui T., Gomer L., Dorward D. W., Glinka A., Grinberg A., Huang S. P., et al. Dev. Cell. 2001;1:423–434. doi: 10.1016/s1534-5807(01)00041-7. [DOI] [PubMed] [Google Scholar]
- 15.Anderson R. M., Lawrence A. R., Stottmann R. W., Bachiller D., Klingensmith J. Development (Cambridge, U.K.) 2002;129:4975–4987. doi: 10.1242/dev.129.21.4975. [DOI] [PubMed] [Google Scholar]
- 16.Bachiller D., Klingensmith J., Kemp C., Belo J. A., Anderson R. M., May S. R., McMahon J. A., McMahon A. P., Harland R. M., Rossant J., De Robertis E. M. Nature. 2000;403:658–661. doi: 10.1038/35001072. [DOI] [PubMed] [Google Scholar]
- 17.Lamb T. M., Knecht A. K., Smith W. C., Stachel S. E., Economides A. N., Stahl N., Yancopolous G. D., Harland R. M. Science. 1993;262:713–718. doi: 10.1126/science.8235591. [DOI] [PubMed] [Google Scholar]
- 18.Smith W. C., Knecht A. K., Wu M., Harland R. M. Nature. 1993;361:547–549. doi: 10.1038/361547a0. [DOI] [PubMed] [Google Scholar]
- 19.Hemmati-Brivanlou A., Kelly O. G., Melton D. A. Cell. 1994;77:283–295. doi: 10.1016/0092-8674(94)90320-4. [DOI] [PubMed] [Google Scholar]
- 20.Pera E. M., Wessely O., Li S. Y., De Robertis E. M. Dev. Cell. 2001;1:655–665. doi: 10.1016/s1534-5807(01)00069-7. [DOI] [PubMed] [Google Scholar]
- 21.Glinka A., Wu W., Delius H., Monaghan A. P., Blumenstock C., Niehrs C. Nature. 1998;391:357–362. doi: 10.1038/34848. [DOI] [PubMed] [Google Scholar]
- 22.Zuber M. E., Gestri G., Viczian A. S., Barsacchi G., Harris W. A. Development (Cambridge, U.K.) 2003;130:5155–5167. doi: 10.1242/dev.00723. [DOI] [PubMed] [Google Scholar]
- 23.Itsykson P., Ilouz N., Turetsky T., Goldstein R. S., Pera M. F., Fishbein I., Segal M., Reubinoff B. E. Mol. Cell. Neurosci. 2005;30:24–36. doi: 10.1016/j.mcn.2005.05.004. [DOI] [PubMed] [Google Scholar]
- 24.Watanabe K., Kamiya D., Nishiyama A., Katayama T., Nozaki S., Kawasaki H., Watanabe Y., Mizuseki K., Sasai Y. Nat. Neurosci. 2005;8:288–296. doi: 10.1038/nn1402. [DOI] [PubMed] [Google Scholar]
- 25.O’Rahilly R. Anat Embryol. (Berlin) 1983;168:87–99. doi: 10.1007/BF00305401. [DOI] [PubMed] [Google Scholar]
- 26.Ramamurthy V., Niemi G. A., Reh T. A., Hurley J. B. Proc. Natl. Acad. Sci. USA. 2004;101:13897–13902. doi: 10.1073/pnas.0404197101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Caffe A. R., Visser H., Jansen H. G., Sanyal S. Curr. Eye Res. 1989;8:1083–1092. doi: 10.3109/02713688908997401. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.