Abstract
Mechanical wounding not only damages plant tissues, but also provides pathways for pathogen invasion. To understand plant responses to wounding at a genomic level, we have surveyed the transcriptional response of 8,200 genes in Arabidopsis plants. Approximately 8% of these genes were altered by wounding at steady-state mRNA levels. Studies of expression patterns of these genes provide new information on the interactions between wounding and other signals, including pathogen attack, abiotic stress factors, and plant hormones. For example, a number of wound-responsive genes encode proteins involved in pathogen response. These include signaling molecules for the pathogen resistance pathway and enzymes required for cell wall modification and secondary metabolism. Many osmotic stress- and heat shock-regulated genes were highly responsive to wounding. Although a number of genes involved in ethylene, jasmonic acid, and abscisic acid pathways were activated, many in auxin responses were suppressed by wounding. These results further dissected the nature of mechanical wounding as a stress signal and identified new genes that may play a role in wounding and other signal transduction pathways.
Wounding is a common damage that occurs to plants as a result of abiotic stress factors such as wind, rain, hail, and of biotic factors, especially insect feeding. Wounding presents a constant threat to plant survival because it not only physically destroys plant tissues, but also provides a pathway for pathogen invasion. To cope with wounding effectively, plants must prepare for pathogen attack while defending against insect predators. Therefore, it is hypothesized that plants may have evolved mechanisms that integrate the pathogen and wounding response. In support of this idea, studies have shown that wounding regulates a number of genes that are also regulated by or play a role in pathogen response (Reymond and Farmer, 1998; Durrant et al., 2000; Reymond et al., 2000). Wounding and pathogen responses also share a number of components in their signaling pathways (Maleck and Dietrich, 1999). For example, studies have shown that several plant hormones are important for wounding and pathogen responses. These include jasmonic acid (JA), salicylic acid (SA), and ethylene (O'Donnell et al., 1996; Creelman and Mullet, 1997; Dong, 1998; Penninckx et al., 1998; Reymond and Farmer, 1998; Thomma et al., 1998). In particular, JA as a typical wounding hormone is essential for certain pathogen responses (Dong, 1998; Penninckx et al., 1998; Thomma et al., 1998; Rojo et al., 1999). Signaling pathways initiated by these hormones and pathogen infection have been further addressed by a recent study using cDNA arrays to identify genes commonly regulated by these hormones and an avirulent pathogen (Schenk et al., 2000). These studies indicate the existence of a substantial network of regulatory interactions and coordination during pathogen and wounding responses. Another study using microarray approach focused on transcriptional profiling of genes after pathogen infection and identified regulons that are involved in systemic acquired resistance (Maleck et al., 2000). This study also identified genes that are reported to be regulated by wounding. In addition to pathogen resistance, wounding pathways may also interact with other signaling processes. Several studies show that some genes are induced by osmotic stress and wounding treatment (Yamaguchi-Shinozaki and Shinozaki, 1994; Kudla et al., 1999; Reymond et al., 2000), suggesting possible interaction between wounding and abiotic stress responses.
Wounding may elicit pathways that interact with pathogen resistance and possibly other signaling pathways. Dissecting crosstalk between these pathways is critical to the understanding of the plant response to environmental cues in general and to wounding in particular. To map the interaction between the wound response and other signaling pathways in a comprehensive manner at the genomic level, and to identify novel genes involved in these processes, it is necessary to survey global gene expression pattern following wounding. Although several previous studies (Durrant et al., 2000; Reymond et al., 2000) have taken different high-throughput methods to identify wounding-responsive genes, our study extends the throughput to a new level by using an Affymetrix Arabidopsis Genome GeneChip array representing 8,200 genes of the genome (Zhu and Wang, 2000). Hundreds of genes were identified to be activated or suppressed by wounding. In particular, a large number of genes encoding signaling molecules were identified as early wound-responsive genes. Extensive overlap between pathogen- and wound-response genes was further demonstrated. Novel interactions between wounding response and responses to several other signals have been identified by this study. Of particular interest is that wounding activates genes encoding heat shock proteins and that auxin-responsive genes are negatively regulated by wounding.
RESULTS AND DISCUSSION
Wounding Alters the mRNA Levels of Approximately 8% of the Total Number of Genes Assayed, a Large Number of Which Encode Signaling Molecules
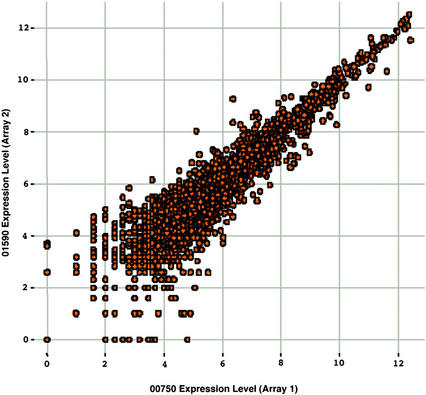
Gene expression patterns of 8,200 genes representing one-third of the Arabidopsis genome were surveyed among parallel control and wounded plants at 30 min and 6 h after wounding. The two time points were designed to identify early and late wounding response genes. Because the two time points were 5.5 h apart, parallel control plants were included to eliminate those genes that are regulated by the circadian clock (Harmer et al., 2000). For example, if a gene is regulated by circadian clock only, this gene should be regulated in the same manner in the wound-treated or control sample at the same time point. As a result, the gene will not be selected as “differentially expressed” when comparing its expression level in the parallel control and wounded samples. Duplicate GeneChip experiments were conducted to select commonly induced or repressed genes. Highly correlated results with average false positive rates less than 1% were obtained (r = 0.86% ± 0.24%, Fig. 1). The noise level of expression (average difference) was arbitrarily determined as 25, which is the scaling target selected (see “Materials and Methods”). With such confidence, genes with an expression level above 25 that are altered more than 2.0-fold in the wounded plants as compared with the unwounded plants at the same time point were defined as differentially expressed genes regulated by wounding. These genes were selected for further analysis (see Supplemental table, which can be viewed at www.plantphysiol.org). Nearly all previously reported wound-responsive genes that were represented on the genechip were identified in this study, demonstrating the internal consistency of the analysis. In addition, a separate study has validated the Genechip technology in profiling gene expression in Arabidopsis (Zhu et al., 2001a). This genomic approach significantly extended our knowledge on wound-responsive genes (from less than 100 reported using conventional methods to 657 identified here).
Figure 1.
A representative scatter plot of the expression level of 8,200 genes, showing the reproducibility between GeneChip experiments. Total RNA was prepared from the control sample at the 0 time point, and labeled samples were independently synthesized and hybridized to the arrays. Note that a high correlation is observed, indicating the results are reproducible.
Of particular interest is the identification of a large number of genes that encode regulatory proteins (Table I). Among all the wounding-regulated genes, about 20% of them encode proteins that play a role in signal transduction and regulation of gene expression. These include 41 protein kinases, six phosphatases, five GTP-binding proteins, seven calcium-binding proteins, four enzymes involved in PI or inositol phosphate metabolism, and 65 transcription factors. It is generally expected that most signaling components should be present in the cell in preparation for extracellular stimuli to initiate signaling processes. Here, we show that these signaling components are transcriptionally regulated by signals such as wounding, implicating feedback control at the mRNA level in the regulation of the signaling pathways.
Table I.
Signaling and regulatory proteins encoded by wound-responsive genes
| Gene Family | Accession No. | Folda
|
|
|---|---|---|---|
| W0.5b | W6 | ||
| Phosphatases | |||
| MP2C-like protein phosphatase 2C | CAB45796 | 37.16 | |
| Putative protein phosphatase 2C | AAC31850 | 18.01 | |
| Protein phosphatase 2C-like protein | CAA19874 | 6.15 | |
| Protein phosphatase 2C-like protein | CAA16879 | 2.83 | |
| Protein phosphatase 2C | BAA07287 | 2.18 | |
| Protein phosphatase 2C | CAA05875 | 2.12 | |
| Receptor-like protein kinases (RLKs) | |||
| Putative receptor protein kinase | AAB80675 | 9.26 | |
| Similar to RLK | AAB65477 | 8.15 | |
| Similar to RLK3 | CAA18469 | 7.42 | |
| Similar to S-receptor kinase (BRLK) | CAA20204 | 6.71 | |
| RLK3 | CAA09731 | 5.61 | |
| Similar to RLK5 | AAD40144 | 4.76 | |
| Similar to S-receptor kinase 8 precursor | AAC18787 | 4.61 | |
| Putative protein kinase | AAF07814 | 4.61 | −2.07 |
| Putative receptor protein kinase | AAF07816 | 4.27 | |
| Similar to RLK3 | CAB45516 | 4.27 | |
| Similar to wall-associated kinase 1 | CAB41191 | 4.21 | |
| Protein kinase catalytic domain, AK2 | CAA60510 | 3.58 | |
| Similar to RLK5 | AAD32284 | 2.88 | |
| RLK | CAA18465 | 2.41 | |
| Similar to S-receptor kinase (ARK1) | AAC13905 | 2.26 | |
| Putative receptor-like protein kinase | AAD28318 | 2.07 | |
| S-Like receptor protein kinase | AAC13903 | 2.76 | |
| Similar to RLK | AAC06160 | −3.64 | |
| Putative RLK | AAB82629 | −2.05 | |
| Similar to ARK3 | CAB44328 | −2.24 | |
| Similar to CLV1 receptor kinase | CAA16688 | −2.44 | |
| Putative receptor protein kinase | CAA19724 | −5.25 | |
| Non-Receptor protein kinases | |||
| Similar to NPK1 protein kinase | AAC31848 | 8.23 | |
| Similar NAK protein kinase | AAC95171 | 6.03 | |
| Ribosomal-protein S6 kinase (AtPK19) | BAA07661 | 5.62 | |
| Protein kinase-like protein | CAB44327 | 5.51 | 3.38 |
| Protein kinase (APK2a) | BAA24694 | 5.31 | |
| Protein kinase 6-like protein | CAB51172 | 5.21 | |
| Putative protein kinase | AAF26067 | 3.31 | |
| Similar to Pti1 kinase | AAC34243 | 3.09 | |
| Similar to APK1 protein kinase | CAA20030 | 3.05 | |
| Similar to protein kinase NAK | AAB97121 | 3.02 | |
| Similar to CBL-interacting protein kinase | AAC16938 | 2.59 | 3.81 |
| Calcium-dependent protein kinase 1 | AAF27092 | 2.05 | |
| Hypothetical Ser-Thr protein kinase | AAD25942 | 2.03 | |
| Similar to CTR1, MAP3K family | AAD30219 | 4.64 | |
| Similar to EDR1, MAP3K family | CAA19821 | 4.37 | |
| Putative protein kinase | AAC09037 | −2.71 | |
| Casein kinase I (CKI2) | CAA55397 | −2.94 | |
| AK23, protein kinase catalytic domain | CAA60531 | −2.08 | |
| Putative protein kinase | CAB10237 | −2.33 | |
| Contain protein kinase catalytic domain | AAD26873 | −3.35 | |
| Calcium-binding proteins | |||
| Similar to CCD-1 | CAA19722 | 22.18 | |
| Calmodulin (CAM)-related protein (TCH2) | AAB82713 | 7.52 | |
| Putative calmodulin like, MSS3 | AAB64310 | 4.71 | |
| Calmoudulin-like protein, CALL_ARATH | AAC78532 | 4.15 | |
| Calmodulin-like protein | CAB42906 | 3.01 | 4.31 |
| Putative calcium-binding protein (RD20) | AAB80656 | 3.36 | |
| Putative calmodulin (CAM9) | CAB41312 | −2.38 | |
| GTP-binding proteins | |||
| Extra-large G-protein like | CAB36716 | 3.66 | |
| Similar to GTP-binding protein | CAB38902 | 3.55 | |
| Similar to GTP exchanger factor | AAB61488 | 3.47 | |
| SGP1 monomeric G-protein | CAB54517 | 3.18 | |
| GTP-binding protein Rab7 | AAC25512 | −2.77 | |
| Phosphatidylinositol (PI) signaling molecules | |||
| PI 5-phosphatase like | AAD10829 | 2.95 | |
| Inositol trisphosphate 5-phosphatase | CAB59428 | 2.68 | |
| Similar to PI 5-phosphatase | AAD46036 | 13.42 | |
| PI synthase (PIS1) | AAF16562 | −2.02 | |
| AP2/EREB-type transcription factors | |||
| Putative EREBP protein | AAC31840 | 53.77 | |
| DREB1B/CBF1 | BAA33435 | 38.89 | |
| EREBP homolog | CAB36718 | 26.11 | |
| Similar to AtERF-4 | AAF16756 | 20.61 | |
| AtERF-5 | BAA32422 | 6.42 | |
| AtERF-4 | AAF34830 | 6.05 | |
| EREBP-4 like protein | CAB10530 | 5.77 | |
| AtERF-1 | BAA32418 | 5.71 | |
| Similar to TINY isolog | AAC25505 | 4.78 | |
| RAV2 | BAA34251 | 3.74 | −2.31 |
| CRT/DRE binding factor 2 (CBF2) | AAD15976 | 2.14 | |
| AP2 domain containing protein RAP2.6 | AAC36019 | 105.25 | |
| AP2 domain transcription factor | AAD20907 | 14.33 | |
| AP2 domain containing protein RAP2.9 | AAC49775 | 3.61 | |
| AP2 domain containing protein RAP2.4 | AAC49770 | 2.81 | |
| AP2 domain transcription factor | AAD20668 | −2.18 | |
| AP2 domain containing protein RAP2.7 | AAD21489 | −2.74 | |
| Putative AP2 domain protein | CAA23041 | −2.85 | |
| Putative AP2 transcription factor | AAD32841 | −3.11 | |
| WRKY-type transcription factors | |||
| AtWRKY40 | AAF14671 | 25.52 | |
| AtWRKY33 | AAC67339 | 15.12 | |
| AtWRKY53 | CAA23047 | 13.24 | |
| AtWRKY22 | AAB61016 | 4.18 | |
| AtWRKY11 | CAB45914 | 3.01 | |
| AtWRKY15 | AAB87100 | 2.81 | |
| AtWRKY60 | AAD23013 | 2.63 | |
| MYB-type transcription factors | |||
| AtMYB15 | CAA74603 | 17.88 | |
| AtMYB51 | CAB09206 | 8.42 | |
| AtMYB77/AtMYBR2 | CAA74604 | 7.91 | |
| AtMYB96 | AAD53106 | 4.18 | 2.52 |
| AtMYB73 | CAB16756 | 3.36 | −3.01 |
| AtMYB44/AtMYBR1 | CAB09200 | 2.48 | |
| AtMYB20 | AAC83591 | −2.01 | −2.42 |
| AtMYB28 | CAB09183 | −2.14 | |
| AtMYBL2 | CAA92280 | −2.68 | |
| AtMYB106 | AAF26160 | −3.32 | |
| AtMYB16 | CAB09174 | −3.62 | |
| AtMYB3 | AAC83581 | −4.14 | |
| AtMYB4 | AAC83582 | −5.67 | |
| Zinc finger-type transcription factors | |||
| Zinc finger protein (AtRZAT7) | CAA67234 | 52.01 | |
| Putative C2H2-type zinc finger protein | AAC98070 | 18.91 | |
| Putative CCCH-type zinc finger protein | AAF18728 | 15.96 | −2.33 |
| C2H2 zinc finger protein (AtZat12) | CAA67232 | 11.64 | 2.17 |
| Putative Cys-3-His zinc finger protein | AAD25930 | 11.17 | −2.71 |
| Salt-tolerance STZ/ZAT10 protein | AAF24959 | 10.21 | |
| RING zinc finger protein (ATL2) | AAC77829 | 7.08 | |
| Similar to salt tolerance STZ/ZAT10 | AAB80922 | 7.65 | 2.72 |
| RING-H2 finger protein (RHA3b) | AAC68674 | 4.46 | 22.21 |
| Putative zinc finger protein (PMZ) | AAD37511 | 4.05 | 3.31 |
| Similar to zinc-binding protein (PWA33) | AAC24052 | 3.75 | 3.18 |
| Similar to GATA transcription factor 3 | CAB41103 | 3.01 | |
| RING-H2 zinc finger protein (ATL6) | AAD33584 | 2.92 | |
| Putative DOF zinc finger protein | AAD21486 | 2.17 | |
| Putative CCCH-type zinc finger protein | AAC62135 | −2.04 | |
| RING zinc finger protein (RMA1) | AAD11593 | −2.26 | |
| Putative CCCH-type zinc finger protein | AAC42256 | −2.32 | |
| C2H2-type zinc finger protein (ZFP8) | AAA87304 | −3.22 | |
| Other-type transcription factors | |||
| Putative bHLH transcription factor | AAAC63587 | 10.69 | |
| Similar to bHLH transcription factor | AAD22130 | 2.14 | |
| Similar to bHLH factor GBOF-1 | CAB16839 | −2.03 | −2.55 |
| Heat shock transcription factor 4 | AAC31756 | 6.67 | 4.58 |
| Heat shock transcription factor 21 | AAC31792 | 5.68 | |
| Homeodomain zipper protein (AtHB-7) | AAC69925 | 2.89 | |
| Homeodomain zipper protein (AtHB-12) | AAC39462 | 2.28 | |
| Homeodomain zipper protein (AtHB-16) | AAD46064 | −2.33 | |
Average channel intensity ratio of wounding-treated samples over control samples over control samples from two independent experiments.
W0.5 and W6 indicate 0.5 and 6 h after wounding.
Wounding Response Overlaps Extensively with Pathogen Response
Among the wound-responsive genes identified in this study, a large fraction of them are known pathogen responsive genes or are postulated to play a role in pathogen resistance. We categorized these genes into two major groups: genes for signaling/regulatory components and those for effector proteins. The signaling/regulatory components include RLKs, non-receptor protein kinases, protein phosphatases, calcium-binding proteins, G-proteins, PI pathway enzymes, and transcription factors (Table I). The effector proteins include defense-related proteins and enzymes involved in the secondary metabolism (Table II). Here, we discuss several categories of newly discovered wound-responsive genes that may encode pathogen response-related proteins.
Table II.
Wound-regulated genes involved in putative defense mechanisms
| Putative Function | Accession No. | Folda
|
|
|---|---|---|---|
| W0.5b | W6 | ||
| Pathogenesis-related genes | |||
| Similar to NDR1/HIN1-like protein 3 | AAC77866 | 18.02 | |
| Similar to Hs1pro-1 | AAB95285 | 12.08 | |
| Similar to regulatory protein NPR1 | CAA19683 | 5.07 | |
| Similar to Mlo proteins | AD25552 | 4.51 | |
| Putative lectin like protein | AAC27148 | 3.88 | |
| Non-race-specific disease resistance protein (NDR1) | AAB95208 | 3.71 | |
| Similar to Avr9/Cf-9 rapidly elicited protein 137 | AAD25764 | 3.69 | |
| Resistance protein RPP5-like | CAA16929 | 2.47 | |
| Disease resistance protein RPP1-WsC | AAC72979 | 2.13 | |
| Similar to Mlo proteins | AAC28997 | 14.03 | |
| Similar to disease resistance protein Cf-2.2 | AAC12833 | 4.88 | |
| Putative disease resistance protein | AAD26901 | 4.44 | |
| AIG1, disease resistance protein | AAC49282 | 3.66 | |
| Disease resistance RPP5 like protein | CAB10461 | −3.77 | |
| Putative β-1,3-glucanase | AAD15611 | 3.61 | |
| β-1,3-glucanase-like protein | CAA16806 | 2.12 | |
| Putative β-1,3-glucanase | CAA18827 | −2.61 | |
| Putative endochitinase | AAB64044 | 16.35 | |
| Class IV chitinase | CAA74930 | 8.42 | |
| Putative endochitinase | AAB64049 | 2.79 | |
| Similar to class I chitinases | AAC72865 | 2.26 | |
| Putative antifungal protein, AFP3_ARATH | AAC31244 | −2.78 | −2.15 |
| Thaumatin-like protein, PR-5 | AAF21072 | −2.08 | |
| Putative protease inhibitor | AAC79626 | 4.59 | |
| Putative trypsin inhibitor | AAB64325 | 4.13 | |
| Oxidative burst | |||
| Peroxidase ATP24a | CAA72484 | 15.41 | |
| Peroxidase ATP15a | CAA67551 | 5.25 | 11.75 |
| Respiratory burst oxidase protein D | AAC39479 | 4.81 | |
| Putative l-ascorbate oxidase | CAA18769 | 2.71 | |
| CTF2B, similar to monooxygenase 2 | AAD09952 | 2.51 | |
| GA 2-oxidase | CAB41008 | 2.07 | 2.31 |
| Similar to glutathione S-transferase | AAD32888 | 5.91 | |
| Putative glutathione S-transferase | AAC95194 | 5.45 | |
| Peroxidase ATP2a | CAA66863 | 3.46 | |
| Putative p-hydroxyphenylpyruvate dioxygenase | AAF24813 | 3.33 | |
| Putative glutathione S-transferase | AAC95192 | 3.28 | |
| Similar to aldehyde reductase | AAC62136 | 3.22 | |
| Glutathione S-transferase | CAA74639 | 2.72 | |
| Putative amine oxidase | AAD22129 | 2.67 | |
| Putative copper amine oxidase | AAD23730 | 2.23 | |
| Putative ferredoxin | AAD15602 | 2.21 | |
| Putative glutathione S-transferase | AC95196 | 2.11 | |
| Glutathione reductase | AAB67841 | 2.07 | |
| Catalase 3 | AAC49807 | −3.27 | |
| Cytochrome P450 | |||
| Putative cytochrome P450 | AAC02748 | 6.31 | |
| Putative cytochrome P450 | AAB87109 | 6.25 | |
| Putative cytochrome P450 | AAC06158 | 4.22 | |
| Cytochrome P450 monooxygenase | BAA28534 | −2.21 | |
| Phenylpropanoid biosynthesis | |||
| Putative leucoanthocyanidin dioxygenase | AAC27173 | 13.66 | |
| Cinnamyl-alcohol dehydrogenase ELI3-2 | CAB37539 | 11.75 | |
| Similar to cinnamyl alcohol dehydrogenase | AAC33210 | 8.89 | |
| Cinnamyl alcohol dehydrogenase-like protein | CAA17549 | 8.31 | |
| Putative cinnamoyl-CoA reductase | AAB80681 | 2.49 | |
| Putative chorismate mutase/prephenate dehydratase | AAC73018 | −2.13 | |
| Flavonoid 3′,5′-hydroxylase-like protein | CAB45977 | −6.83 | |
| TRP pathway | |||
| Phosphoribosylanthranilate isomerase | AAD38141 | 2.72 | |
| Strictosidine synthase | AAB40594 | 2.67 | |
| TRP synthase α-chain | AAC49117 | 2.11 | |
| Anthranilate synthase, α-subunit | AAC35228 | 2.05 | |
| Anthranilate N-benzoyltransferase-like protein | AAB95283 | −2.14 | 2.01 |
| Putative chorismate mutase/prephenate dehydratase | AAC73018 | −2.13 | |
| Berberine biosynthesis | |||
| Putative berberine bridge enzyme | AAD25759 | 6.51 | 9.25 |
| Berberine bridge enzyme-like protein | CAB42588 | 2.61 | 6.14 |
| Berberine bridge enzyme-like protein | CAB45881 | 2.32 | 3.27 |
| Tyr transaminase-like protein | CAA23026 | 2.55 | |
| Putative berberine bridge enzyme | AAC12821 | 2.54 | |
| Putative Tyr transaminase | AAD23027 | 2.08 | |
| Cell wall modification | |||
| Xyloglucan endotransglycosylase-related protein | AAB18367 | 10.50 | |
| Xyloglucan transglycosylase-related protein (TCH4) | AAC05572 | 4.08 | 2.18 |
| Similar to TCH4 gene | AAC27142 | 4.06 | |
| Similar to xyloglucan fucosyltransferase FT1 | AAD22285 | 3.91 | |
| Glycosyltransferase-like protein | CAB62445 | 2.46 | |
| Putative glucosyltransferase | AAC16958 | 3.13 | |
| Putative glucosyltransferase | AAB64024 | 2.96 | |
| Putative xyloglucan endotransglycosylase | AAD31572 | 2.19 | |
| Similar to pectinesterase | AAC19280 | 3.37 | 3.41 |
| Putative pectin methylesterase | AAC02974 | 3.11 | 4.01 |
| Pectinesterase 1 | AAF02857 | 2.39 | |
| Similar to pectinesterase | CAB51210 | 16.91 | |
| Putative pectinesterase | AAB82640 | 3.35 | |
| Putative pectin methylesterase | AAC02973 | 2.26 | |
| Putative pectate lyase | CAA22985 | −2.28 | |
| Cellulose synthase isolog | AAB63623 | 4.88 | |
| Cellulose synthase like protein | CAB10434 | −2.01 | 2.23 |
| UDP-Glc 4-epimerase-like protein | CAB45812 | 11.61 | |
| Putative UDP-galactose-4-epimerase | AAD25749 | 2.18 | 2.15 |
| Extensin-like protein | CAA22152 | 13.41 | |
| Extensin like protein, pEARLI 1 | CAB41718 | −2.03 | 4.74 |
| pEARLI 1-like protein | CAB41717 | 6.11 | |
| Expansin At-EXP1 | AAB38070 | −2.16 | |
| Putative expansin | AAB87577 | −3.29 | |
| Gly-rich protein | AAA32722 | 12.70 | |
| Putative Pro-rich protein | AAD29802 | 2.62 | |
| Gly-rich RNA-binding protein AtGRP2 like | CAB36849 | −2.44 | |
| Gly-rich protein, atGRP-2 | AAB24074 | −3.26 | |
Average channel intensity ratio of wounding-treated samples over control samples from two independent experiments.
W0.5 and W6 indicate 0.5 and 6 h after wounding.
In the protein kinase group, several plant-specific protein kinases, including the RLK family, NPK1-like, EDR1-like, and Pti1-like proteins, which are particularly relevant to pathogen response, were found to be regulated by wounding. Among the 22 wound-regulated genes encoding RLKs (Table I), two new RLKs are highly homologous to RLK3 and RLK5, respectively. RLK3, 4, 5, and 6 have been previously shown to be regulated by pathogen and SA in recent studies (Czernic et al., 1999; Du and Chen, 2000). Possible function of some RLKs identified here is further supported by their elevated expression in pathogen-treated samples from independent microarray experiments (Maleck et al., 2000; Schenk et al., 2000). RLK proteins are believed to perceive external stimuli and transduce the signals across the plasma membrane through phosphorylation cascades that ultimately lead to expression of appropriate target genes. During the activation of plant defense responses, pathogen- or host-derived elicitor molecules may serve as ligands for these RLKs (Song et al., 1995). Furthermore, at least one typical RLK, Xa21, encodes a disease resistance protein in rice (Oryza sativa; Song et al., 1995). We speculate that some of wound-responsive RLKs identified in this study may be involved in pathogen resistance. Further functional analyses will test this hypothesis.
Several genes for Ca2+-binding proteins were found to be transcriptionally activated by wounding, including the CCD-1-like protein and CaM-like proteins (Table I). Similar genes also respond to pathogen signals and function in pathogen resistance responses (Heo et al., 1999; Takezawa, 2000). Ca2+-binding proteins, including CaM, CaM-like proteins, calcineurin B-like protein, and calcium-dependent protein kinase are calcium sensors that specify the cellular response to different extracellular signals (Trewavas and Malho, 1998; Kudla et al., 1999). The wound- and pathogen-responsive CaM-like proteins and CCD-1 homolog may play a role in defense response signaling pathways against pathogen and insects.
Wounding induces expression of a large number of genes encoding transcription factors. Among them are members in the AP2, WRKY, and MYB families. We found several genes encoding AP2 domain-containing proteins that are strongly activated upon wounding. Similar AP2 gene families have been reported to be induced by SA, JA, ethylene, and pathogen attack (Maleck et al., 2000; Schenk et al., 2000). Wounding also induces seven genes encoding WRKY transcription factors, including AtWRKY40, AtWRKK33, AtWRKY53, AtWRKY22, AtWRKY11, AtWRKY15, and AtWRKY60 (Table I). WRKY proteins were originally defined as transcription factors that target genes regulated by pathogen attack, fungal elicitors, and SA (Rushton et al., 1996; for review, see Eulgem et al., 2000). A recent study (Du and Chen, 2000) reported that AtWRKY18 regulates the expression of RLKs including RLK4, which in turn is induced by bacterial infection and salicylate treatment. Several wound-responsive WRKY genes reported here are also regulated by pathogen infection (Maleck et al., 2000; Schenk et al., 2000; Chen et al., 2002), further supporting the notion that wound-inducible expression of WRKY genes shown in this study may function in pathogen and wounding defense mechanisms.
Wounding alters the gene expression of many MYB transcription factors that regulate the transcription of a number of flavonoid genes (Meissner et al., 1999; Borevitz et al., 2000). However, different MYB transcription factors may show diverse expression pattern following wounding. We found that AtMYB15 and AtMYB51 are rapidly up-regulated by wounding, whereas MYB3 and MYB4 are down-regulated by wounding. The AtMYB15 gene is highly similar to the NtMYB2 gene from tobacco (Nicotiana tabacum) that is also inducible by wounding and elicitors and is known to positively regulate PR gene expression (Sugimoto et al., 2000). AtMYB51 is highly similar to the ATR1 protein that activates Trp gene expression in Arabidopsis (Bender and Fink, 1998). Mutant analysis shows that AtMYB3 and AtMYB4 are repressors of phenylpropanoid biosynthesis gene expression (Jin et al., 2000). It is significant that wounding up-regulates AtMYB15 and 51, positive regulators of secondary metabolism, and down-regulates AtMYB3 and 4, repressors of secondary metabolism. This finding further supports the hypothesis that wounding positively regulates the secondary metabolic pathways involved in activation of defense response.
A number of genes encoding putative components of disease resistance pathways were also shown to be wound inducible in this study (Table II). These genes include a NPR1-like gene, NDR1, and a NDR1-like gene, glucanase-like genes, chitinase homologs, and several putative disease resistance genes (R-gene), including Mlo-like genes, an RPP5-like gene, RPP1, and a Cf2.2-like gene. The products of R-genes in plants and their direct or indirect interaction with avirulent gene products mediate specific pathogen-host interactions (Ryals et al., 1996; Glazebrook et al., 1997; Wilson et al., 1997; Nurnberger and Scheel, 2001). NDR1, which encodes a protein with two predicted transmembrane domains, is required for the action of most CC-NBS-LRR class of disease resistance genes. Its expression is also induced in response to pathogen attack and may function to integrate various pathogen recognition signals (Century et al., 1997). The systemic acquired resistance pathway is mediated, at least in part, by SA that activates the expression of PR genes, such as PR1, PR2 and PR5 (Ryals et al., 1996; Nurnberger and Scheel, 2001). This pathway requires the function of the NPR1 gene product (Cao et al., 1997). NPR1 interacts with several members of the TGA subclass of basic domain/Leu zipper transcription factors that bind to the SA-responsive element found in the PR-1 gene promoter (Zhang et al., 1999). The fact that NPR1-like and NDR1-like genes, together with several R-genes, are up-regulated by wounding as shown in Table II clearly indicates that wounding enhances the levels of a variety of defense response genes.
Another interesting wound-induced gene from our study is the Mlo-like gene (Table II). The barley (Hordeum vulgare) Mlo gene shows homology to the G-protein-coupled receptors. Lack of the wild-type Mlo protein leads to broad spectrum disease resistance against the pathogenic powdery mildew fungus and deregulated leaf cell death (Buschges et al., 1997). A genome-wide search for Mlo sequence-related genes in Arabidopsis revealed approximately 35 family members that have not been previously studied. Our study showed that at least one of these genes was regulated by wounding and provided a starting point toward further functional analysis of Mlo family genes in plant defense mechanisms.
Among the late response genes regulated by wounding are a number of genes involved in secondary metabolism and cell wall modifications (Table II). Their respective gene products were defined as effector proteins involved in late defense responses. Most striking is the activation of the phenylpropanoid pathway that produces many secondary metabolites, which act as anti-pathogen and anti-insect agents (De Luca and St-Pierre, 2000). Some of them are important intermediates for lignin biosynthesis during cell wall thickening. A number of genes encoding enzymes in this pathway are activated by wounding. These include genes for a putative leucoanthocyanidine dioxygenases, three putative cinnamyl-alcohol dehydrogenases, and a putative cinnamoyl-coenzyme A reductase (Table II). The coordinate expression of these genes suggests that they may be controlled by one or more common regulatory factors. As discussed earlier, regulation of MYB family transcription factors by wounding provides a potential link between the signaling pathway and the secondary metabolism (Jin et al., 2000).
Enzymes involved in the Trp and alkaloid biosynthetic pathways are also activated at the transcriptional level. Several genes encoding enzymes in the Trp biosynthetic pathway, such as anthranilate synthase α-subunit, phosphoribosylanthranilate isomerase, and Trp synthase α-subunit, are induced by wounding (Table II). The Arabidopsis Trp pathway can lead to the biosynthesis of the phytoalexin camalexin and other secondary compounds (Radwanski and Last, 1995). Alkaloids, one of the largest groups of natural products, provide many pharmacologically active compounds. Among them, berberine, a benzylisoquinolone alkanoid isolated from Captis (Ranunculaceae), is used as an anti-pathogen agent by plants. The biosynthetic pathway from l-Tyr to berberine has 13 different enzymatic reactions. It is not known whether Arabidopsis produces berberine and related alkaloids. Our study reveals several wound-responsive genes encoding berberine biosynthetic enzymes. Among them, two Tyr transaminase like genes and four berberine bridge enzyme-like genes are late wound-response genes (Table II). This finding suggests that the berberine biosynthesis pathway may occur in Arabidopsis and may participate in defense responses.
Genes whose products are involved in cell wall biosynthesis and modifications were shown to be regulated by wounding (Table II), and a large fraction of these are also known to be regulated by pathogens (Maleck et al., 2000; Schenk et al., 2000). These genes encode enzymes involved in biosynthesis of cell wall structure components, such as cellulose, hemicellulose, pectin, and proteins. Some wound-regulated genes encode enzymes such as endoglucanases, xyloglucan endotransglycosylases, and a number of glycosyl transferases, which alter carbohydrate linkages and modify secreted cell wall components. Our study further confirms the idea that defense mechanism against microbial pathogens and insects involves cell wall modifications.
Our results also confirmed that the oxidative burst, an essential component of the pathogen response, is highly regulated by wounding. A number of genes for the enzymes involved in hydrogen peroxide production and processing are activated by wounding treatments. These include genes encoding peroxidase ATP21, ATP15a, and ATP24a, respiratory burst oxidase protein D, putative l-ascorbate oxidase, glutathione reductase, and glutathione S-transferase (Table II). Hydrogen peroxide can act as a local signal for hypertensive cell death and can also serve as a diffusible signal for the induction of defense genes in adjacent cells (Alvarez et al., 1998). Wounding stress also induces hydrogen peroxide accumulation locally and systemically in the leaves of several plant species (Orozco-Cardenas and Ryan, 1999). It has recently been shown that hydrogen peroxide acts as a second messenger for induction of defense genes in tomato (Lycopersicon esculentum) plants in response to wounding, systemin, and methyl jasmonate (Orozco-Cardenas et al., 2001). This could partially explain why a large number of genes are commonly regulated by pathogen attack and wounding in plants. This extensive overlap between pathogen and wounding response as shown in this study further supports the hypothesis that plants have evolved mechanisms that integrate pathogen defense into wounding response.
Interaction between Wounding and Abiotic Stress Responses
In addition to pathogen response genes, our study shows that a number of genes regulated by abiotic stress response pathways are activated by wounding (Table III). These include genes responsive to drought, cold, high salt, heat shock, and others. These so-called “stress genes” are normally silent and rapidly induced by stress conditions (Shinozaki and Yamaguchi-Shinozaki, 2000). Several transcription factors such as DREB1B/CBF1 and DREB1C/CBF2 that govern the stress regulations are among the early responsive genes after wounding (Table III). These proteins contain the AP2/EREBP domain and bind to the DRE/CRT motif in drought-, high salt-, and cold stress-responsive gene promoters (Jaglo-Ottosen et al., 1998; Liu et al., 1998). DREB1/CBFs are known to be induced by low-temperature stress, indicating that it may function in cold-responsive gene expression, whereas DREB2 genes are implicated in drought-responsive gene expression because its mRNA level is induced by drought but not by cold (Liu et al., 1998). A recent study (Seki et al., 2001) shows that many drought-responsive genes such as Rd29A/Iti78/Cor78, Kin1, Kin2/Cor6.6, Cor15a, and Rd17/Cor47 are also induced by cold stress in a DREB1A-dependent manner. We show in this study that wounding not only induces transcription factor DREB1B/CBF, but also activated many downstream stress genes such as Rd29A/cor78, Kin1, and Kin2. This result suggests that mechanical wounding may activate drought and cold response pathways.
Table III.
Wound-regulated genes related to abiotic stress responses
| Putative Function | Accession No. | Folda
|
|
|---|---|---|---|
| W0.5b | W6 | ||
| Cold/drought responsive | |||
| DREB1B/CBF1 | BAA33435 | 38.89 | |
| CRT/DRE binding factor 2 (CBF2) | AAD15976 | 2.14 | |
| LEA D113 homolog type 1 | CAA63008 | 36.76 | |
| LEA76 homolog type 1 | CAA63012 | 36.45 | |
| Dehydrin | AAB00374 | 15.36 | |
| Cor78/Rd29A | AAA32775 | 12.01 | |
| Cor15b precursor | AAD23000 | 6.63 | |
| Cor47/Rd17 | BAA23547 | 5.54 | |
| Lti29 | CAA62448 | 4.88 | |
| Similar to LEA14 | AAC62908 | 4.87 | |
| Rd20 | AAB80656 | 3.36 | |
| Homeodomain transcription factor (AtHB-7) | AAC69925 | 2.89 | |
| Putative dessication-related protein LEA14 | CAA71174 | 2.72 | |
| Rd22 | BAA01546 | 2.44 | |
| Homeodomain transcription factor (AtHB-12) | AAC39462 | 2.28 | |
| Kin1 | CAA35838 | 2.23 | |
| Cold-regulated cor6.6/kin2 | CAA38894 | 2.05 | |
| Salt/abscisic acid responsive | |||
| Similar to protein phosphatase MP2C | CAB45796 | 37.17 | |
| Salt tolerance zinc finger protein (STZ/ZAT10) | AAF24959 | 10.21 | |
| Similar to salt tolerance factor STZ/ZAT10 | AAB80922 | 7.65 | 2.72 |
| Ribosomal protein S6 kinase homolog (AtPK19) | BAA07661 | 5.63 | |
| Putative Suc-proton symporter (AtSUC3) | AAC32907 | 2.42 | |
| Trehalose-6-phosphate phosphatase-like | CAA22164 | 2.11 | |
| Calcium-dependent protein kinase 1 (CDPK1) | AAF27092 | 2.05 | |
| Aquaporin/MIP-like protein | CAB41102 | 4.92 | |
| Putative Pro transporter | AAD24641 | 3.13 | |
| Pro transporter (AAP4) | CAA54631 | 2.85 | |
| Suc transport protein (SUC1) | AAF34305 | 2.56 | |
| Aquaporin (PIP2a) | AAD18142 | 2.51 | |
| Aquaporin (PIP1a) | CAA53475 | 2.39 | |
| Δ1-Pyrroline-5-carboxylate synthetase (ATP5CS) | BAA06864 | 2.29 | |
| Similar to trehalose-6-phosphate synthase | AAC24048 | 2.28 | |
| Suc synthase | CAA50317 | 2.24 | |
| Putative ABI1 and ABI2 homolog | CAA05875 | 2.12 | |
| Heat shock responsive | |||
| HSF-like protein | CAA16745 | 7.25 | |
| HSF 4 | AAC31756 | 6.67 | 4.58 |
| HSF 21 | AAC31792 | 5.68 | |
| Heat shock protein (HSP) 17.6-II | CAA45039 | 11.01 | |
| HSP 70 | CAA05547 | 4.51 | |
| HSP 17.6A | CAA74399 | 3.72 | |
| Hsc70-G8 protein | CAA70105 | 2.81 | |
| DnaJ-like protein | CAA18498 | 2.45 | |
| Hsc70-G7 protein | CAA70111 | 2.18 | |
| HSP 83 | AAA32822 | 2.15 | |
Average channel intensity ratio of wounding-treated samples over control samples from two independent experiments.
W0.5 and W6 indicate 0.5 and 6 h after wounding.
Wounding treatment also activated a number of genes related to the high salinity response. These include genes for ATPK19, MP2C-like protein phosphatase, AtCDPK1, STZ/ZAT10, water channels (PIP1a/2a), sugar synthetic enzymes (ATP5CS and Suc synthase), and Pro transporters (Table III). Although the mechanism underlying the plant response to high salinity is not clear, gene activation has been widely documented to be an important response (Hasegawa et al., 2000). The salt-responsive genes encode proteins that are often involved in synthesis of osmolytes (e.g. Pro and Suc), water flux control, and membrane transport of ions (Hasegawa et al., 2000). In addition to previously characterized salt-responsive genes, a Pro transporter gene has also been identified by this study, suggesting that transport of osmolyte may be one of the limiting factors in the plant osmotic stress response. Further analysis indicates that this gene is also activated in the osmotic stress-treated samples (T. Zhu, unpublished data).
HSPs are thought to facilitate growth and survival of plants under conditions of severe heat stress whereby lethal temperatures can be tolerated for short periods (Gurley, 2000; Lee and Vierling, 2000). Major classes of HSPs present in plants include the small HSPs (ranging in Mr from 15–28 kD), HSP60, HSP70, and a constitutively expressed heat shock cognate protein, HSC70, HSP90, and HSP100 (Gurley, 2000). As in other organisms, these HSPs play an essential part in protein folding and other cellular processes under normal and stress conditions in plants (Lee and Vierling, 2000). Expression of heat shock genes is regulated by HSFs. Wounding activated several genes encoding HSFs and HSPs (Table III). This is rather unexpected because HSPs have not been studied in the context of the wound response. It is interesting that wound activation of heat shock response genes follows a time frame consistent with the normal heat shock response. Two HSF genes (HSF4 and HSF21) are early wounding-response genes that were activated within 30 min after wounding. This activation is followed by transcription of a number of HSP genes, including HSP17.6II, HSP17.6A, HSP70, Hsc70-G8, Hsc70-G7, HSP83, and a Dna-J-like gene. Therefore, it is tempting to speculate that HSF4 and HSF21 are the transcription factors that control the expression of these HSPs. To determine if other stress conditions or signals (besides heat shock and wounding) also regulate the expression of HSPs, we performed a systematic database search on four genes including HSF21, HSF4, HSP70, and HSP17.6A in the Arabidopsis gene expression database of Torrey Mesa Research Institute. As shown in Table IV, HSF4, HSP70, and HSP17.6A are activated by a number of stress conditions and signals including osmotic stress, electric shock, pathogen attack, light, and several plant hormones. HSF21 is specifically activated by wounding and pathogen elicitor, but not by other signals. This result suggests that different HSPs may play distinct roles in various stress responses. Some such as HSF4, HSP70, and HSP17.6A may be important for a range of stress conditions, whereas others such as HSF21 may be involved in more specific stress responses.
Table IV.
Response of heat shock-responsive proteins upon various environmental stimuli
| Response | Putative Function
|
||
|---|---|---|---|
| Classification | Treatment | Response | |
| Early (30 min) | Heat shock transcription factor-like protein (HSF21: CAA16745) | ||
| 1 = Stress | Wounding | 1 h after treatment | |
| 2 = Pathogen | Pseudomonas syringae | 3 h after treatment | |
| 3 = Virus | Tobacco rattle virus | ? | |
| 4 = Others | Senescent | ? | |
| Constitutive (30 min and 6 h) | Heat shock transcription factor 4 (HSF4: AAC31756) | ||
| 1 = Stress | Wounding | 0.5 and 1 h after treatment | |
| Detachment | 12 h after treatment | ||
| Osmotic stress | 3 and 27 h after treatment | ||
| Electroporation shock | 17 h after treatment | ||
| 2 = Pathogen | P. syringae | 3, 6, 9, 12, 27, 30, and 48 h after treatment | |
| Peronospora parasitica | 12 and 48 h after treatment | ||
| 3 = Virus | Turnip mosaic virus | 28 and 92 h after treatment | |
| Tobacco rattle virus | 92 h after treatment | ||
| Cucumber mosaic virus | 92 h after treatment | ||
| 4 = Chemical | Benzothiadiazole (BTH) | 48 and 72 h after treatment | |
| Jasmonic acid | 40 h after treatment | ||
| β-Aminobutyric acid (BABA) | 72 h after treatment | ||
| 5 = Others | Light | 4, 8, 12, 16, 20, 24, and 40 h after treatment | |
| Cold | 3 and 27 h after treatment | ||
| Late (6 h) | Heat shock protein 70 and 17.6A (HSP70 and 17.6A CAA05547 and CAA74399) | ||
| 1 = Stress | Osmotic | 3 and 27 h after treatment | |
| Electroporation shock | ? | ||
| 2 = Pathogen | P. parasitica | 48 h after treatment | |
| 3 to 48 h after treatment | |||
| 3 = Virus | Oil seed virus | 28 h after treatment | |
| Tobacco rattle virus | 52 h after treatment | ||
| Turnip vein clearing virus | 52 h after treatment | ||
| Turnip mosaic virus | 92 h after treatment | ||
| 4 = Chemical | γ-Aminobutyric acid (GABA) | 24 h after treatment | |
| Jasmonic acid | 40 h after treatment | ||
| Salicylic acid | 2 h after treatment | ||
| BTH | 48 and 72 h after treatment | ||
| 5 = Others | Light | 24 h after treatment | |
| Cold | 3 and 27 h after treatment | ||
Interaction of Wounding and Hormonal Signaling Pathways
As discussed earlier, wounding and pathogen responses involve a number of plant hormones, including JA, SA, and ethylene (Reymond and Farmer, 1998). JA accumulates in wounded plants and activates expression of various defense genes such as proteinase inhibitors, thionin, and enzymes involved in secondary metabolism (Creelman and Mullet, 1997). We found that a number of genes, such as a putative lipase, putative lipoxygenase, putative allene oxide synthase, and 12-oxophytodienoate reductase, which are involved in biosynthesis of JA, are up-regulated by wounding (Table V). Most of these JA synthesis genes are early wounding-response genes and were activated within 30 min. This result is consistent with earlier studies (Reymond and Farmer, 1998; Reymond et al., 2000) and suggests that wounding may trigger the JA biosynthesis upon direct activation of genes encoding the relevant biosynthetic enzymes.
Table V.
Wound-regulated genes related to hormonal responses
| Putative Function | Accession No. | Folda
|
|
|---|---|---|---|
| W0.5b | W6 | ||
| Jasmonic acid pathway | |||
| Putative allene oxide synthase | AAD03574 | 2.72 | 3.94 |
| Phospholipase like protein | AAC37472 | 2.45 | |
| Putative phospholipase | AAC27833 | 2.06 | |
| Putative GDSL motif lipase/hydrolase | AAD25823 | 3.98 | |
| Lipoxygenase | CAB56692 | 3.68 | |
| 12-Oxophytodienoate reductase (OPR1) | AAC78440 | 2.16 | |
| Ethylene pathway | |||
| Putative EREBP protein | AAC31840 | 53.77 | |
| EREBP homolog | CAB36718 | 26.11 | |
| ACC synthase (AtACS-6) | CAB51412 | 23.66 | |
| ACC synthase | AAC63850 | 21.26 | |
| Similar to AtERF-4 | AAF16756 | 20.61 | |
| AtERF-5 | BAA32422 | 6.42 | |
| AtERF-4 | AAF34830 | 6.05 | |
| EREBP-4 like protein | CAB10530 | 5.77 | |
| AtERF-1 | BAA32418 | 5.71 | |
| Similar to senescense-associated protein | AAF18611 | 4.31 | |
| Senescense-associated protein | AAD20613 | 8.72 | |
| Auxin pathway | |||
| Putative auxin-regulated protein | AAB82641 | 6.25 | |
| Auxin-responsive GH3-like protein | AAD23040 | 2.42 | |
| Putative auxin-regulated protein | AAC69926 | −2.55 | |
| Putative GH3-like protein | AAD14468 | −3.29 | |
| Auxin-induced protein-like | CAB38618 | −3.83 | −2.29 |
| SAUR-AC1= small auxin up RNA | AAB30527 | −4.15 | −2.31 |
| Putative auxin-induced protein | CAB38620 | −7.13 | −3.86 |
| Auxin-responsive protein IAA2 | AAB97164 | −2.12 | |
| Match to Arabidopsis IAA3 | AAB70452 | −2.12 | |
| Auxin-induced protein (IAA20) | AAC34236 | −2.16 | |
| Similar to auxin-induced protein | AAB62852 | −2.18 | |
| Putative auxin-regulated protein | AAD29795 | −2.54 | |
Average channel intensity ratio of wounding-treated samples over control samples from two independent experiments.
W0.5 and W6 indicate 0.5 and 6 h after wounding.
Ethylene plays a highly pleiotropic role in plant growth and development and is involved in a number of processes, including germination, senescence, abscission, and fruit ripening. Ethylene also participates in a variety of defense responses and abiotic stress responses (Ecker, 1995). Exogenous application of ethylene induces the transcription of genes encoding class I basic chitinases, glucanases, and other PR genes. It has been also well documented that wounding induces ethylene production that affects the downstream wound response (O'Donnell et al., 1996). Two mechanisms may be involved in the interaction between ethylene signaling and wound response according to results from this study. First, several 1-aminocyclopropane-1-carboxylic acid (ACC) synthase genes, which are involved in ethylene biosynthesis, have been identified as early wound-response genes (Ecker, 1995; Reymond and Farmer, 1998). Following the induction of ACC synthase genes, we observed the significant induction of ethylene response genes such as senescence-associated genes, chitinases, and glucanases as late wound-response genes (Tables II and V). Second, many of the ethylene response transcription factors such as EREBPs are rapidly induced by wounding. These transcriptional factors may directly participate in the activation of ethylene-responsive genes. Wounding-mediated ethylene synthesis implicates ethylene as a chemical messenger for wounding as reported previously (O'Donnell et al., 1996). However, early induction of EREBPs suggests that crosstalk may occur between wounding and ethylene signaling pathway at the level of transcriptional regulation.
Perhaps the most surprising finding is the interaction between wounding and the auxin signaling pathway. In almost all cases, genes that are positively responsive to auxin signaling pathway are down-regulated by wounding (Table V). Although auxins, primarily indole-3-acetic acid (IAA), control many important developmental processes in plants (Bartel, 1997; Normanly and Bartel, 1999), little is known on auxin function in stress or defense responses. Now we show that wounding negatively regulates IAA responsive genes, revealing a new level of crosstalk between wounding and auxin response in plants. Negative regulation of auxin response by wounding can happen through at least two possible pathways. One is by reducing production of endogenous IAA, and the other is negative crosstalk of the signaling pathways. We addressed the first possibility by compiling the expression pattern of genes encoding IAA biosynthetic enzymes. The starting point for IAA synthesis is in the Trp biosynthetic pathway, and in recent years, it has become generally accepted that there exists a “Trp-independent” pathway branching off from the Trp biosynthetic pathway at a Trp precursor such as indole as well as a Trp-dependent pathway (Normanly and Bartel, 1999). One gene encoding nitrilase was shown to be down-regulated by wounding (Table V). This enzyme catalyzes the conversion of indole-3-acetonitrile to IAA (Normanly et al., 1997). It is interesting that two genes encoding IAA glucosyltransferases are also highly induced. As these enzymes are responsible for the formation of IAA-sugar conjugates, this finding may provide a mechanism for wound-induced IAA conjugation, thereby reducing the endogenous level of active IAA. Our study also provides evidence for an alternative mechanism of negative regulation of IAA response by wounding. As discussed earlier, a NPK1-like gene may be a homolog of ANP1 that negatively regulates IAA responsive genes (Kovtun et al., 1998). The expression of the NPK1-like gene is up-regulated by wounding (Table I), implicating activation of the NPK1-like gene in down-regulation of IAA signaling. Although our result does not elucidate the mechanism of down-regulation of IAA response by wounding, finding antagonism between wounding and IAA responses sheds new light on IAA function and on the interaction of these pathways in Arabidopsis.
Kinetics of Gene Activation and a Novel Method for Pathway Finding
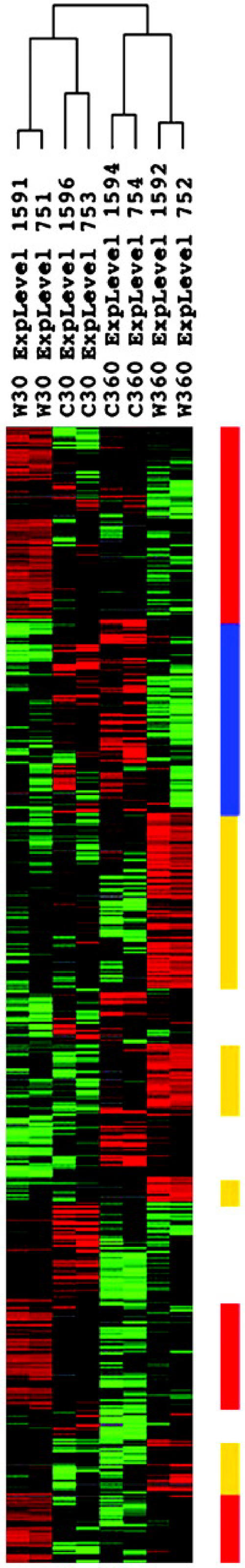
In this study, we surveyed global gene expression at two time points after wounding. Those genes with significantly altered mRNA levels within 30 min are considered early response genes, whereas those responsive after 6 h are considered late response genes. Two interesting phenomena have been found when comparing these two groups of genes. First, more than 90% of the genes are early or late response genes (Fig. 2). Only a very small number of genes (10%) are in both groups. This finding indicates that induction of most early response genes is transient, but not sustained. Second, the time frame of induction of a particular gene by a signal often reflects the position of the gene product in the response pathway. For example, the early response genes are often those that encode “signaling or regulatory components” such as protein kinases and transcription factors (Table I). The late response genes are often those that encode “effector proteins” such as enzymes in the metabolism (Fig. 2). In several cases, it is clear that the protein products of early response genes function to regulate the expression of late response genes, suggesting a cascade of gene regulation. There are certainly exceptions to this cascade of regulation. For example, early response genes encoding several RLKs have been shown to be targets for WRKY transcriptional factors (Du and Chen, 2000). Several examples of possible gene regulation cascades are illustrated in Figure 3, which may help elucidate the specific signaling pathways involved. In one case, the HSFs are activated within 30 min, which is followed by induction of a number of HSPs in the 6-h samples. It will be interesting to determine if accumulation of HSFs is a prerequisite for activation of downstream HSP genes (Gurley, 2000; Lee and Vierling, 2000). Second, abiotic transcription factors such as CBFs/DREBs are encoded by early wounding response genes, whereas the stress defense genes such as RD29A, Kin1 and Kin2, and Cor15b are all late response genes. It is known that CBFs/DREBs are the transcription factors that activate the target genes including Cor/RD/Kin (Jaglo-Ottosen et al., 1998; Liu et al., 1998; Seki et al., 2001). In the third example, MYB transcriptional factors are early response genes and their target genes such as those encoding secondary metabolic enzymes are late response genes (Meissner et al., 1999; Borevitz et al., 2000). Furthermore, WRKY and AP2/EREBP-type transcription factors are also early response genes and their target genes encoding enzymes such as chitinase, glutathione S-transferase, and strictosidine synthase are late response genes (Rushton et al., 1996; Du and Chen, 2000; Memelink et al., 2001). These examples demonstrate that one can build hypothetical models for gene regulation pathways based on transcriptional profiling. It is certain that further studies by genetic and biochemical approaches are required to test the model pathways.
Figure 2.
Hierarchical cluster of the wounding-induced and -reduced genes using Spearman rank correlation, showing the early “signal gene clusters” and late “effector gene clusters.” Up-regulated gene clusters are indicated by red bars (signal gene clusters) or orange bars (effector gene clusters). Down-regulated gene clusters are indicated by the blue bar.
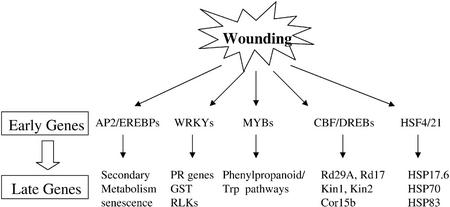
Figure 3.
A hypothetical model for early wound-responsive transcription factors to activate late responsive genes in wounding signal transduction pathways.
We conclude that this study has not only identified a large number of novel genes that are regulated by wounding, but it also provides new insights into the interaction and overlap between signal transduction pathways initiated by wounding and other stimuli. In addition, the expression kinetics of wounding-inducible genes provides useful information for pathway finding in plants.
MATERIALS AND METHODS
Plant Treatment and RNA Isolation
Four-week-old Arabidopsis (ecotype Columbia) seedlings grown under short-day conditions (8 h of light at 500 μmol m−2 s−2 at 21°C–23°C in 75% humidity at day and night) were used for the wounding stress treatments. Wounding was performed by puncturing leaves with a hemostat. Approximately 50% of the leaves in the treated plants were wounded. One sample was collected at each time point from one pot containing 20 rosette plants. Two sets of potted plants were prepared for the same treatment and were harvested separately for RNA extraction. The parallel control plants were grown under the same conditions and were harvested at the same time points except that the control plants were not wounded. To prevent the subsequent pathogen infection, plants were grown in a disinfected growth chamber and hemostats were sterilized.
Total RNAs were isolated from the parallel control and wounded leaf material using the RNAwiz isolation regent (Ambion, Austin, TX) and were then purified using RNeasy columns (Qiagen, Valencia, CA). The RNA samples were prepared separately from the two sets of plant materials and were pooled for microarray analysis.
Microarray Experiments and Data Analyses
Microarray experiments were conducted in duplicates according to Zhu et al. (2001). Briefly, double-stranded cDNA was synthesized from 5 μg of total RNA samples using a combination of oligo dT(24) primer containing a 5′ T7 RNA polymerase promoter sequence, SuperScript II reverse transcriptase (Invitrogen, Carlsbad, CA), and Escherichia coli DNA polymerase and ligase. The synthesized products were purified by phenol/chloroform extraction and ethanol precipitation. Synthesized cDNAs (approximately 0.1 μg) were used as templates to produce biotinylated cRNA probes by in vitro transcription using T7 RNA Polymerase (BioArray High-Yield RNA Transcript Labeling kit; Enzo Diagnostics, New York). Labeled cRNAs were purified using affinity resin (RNeasy Spin Columns; Qiagen) and were randomly fragmented to produce molecules of approximately 35 to 200 bases.
Arabidopsis Genome GeneChip arrays (Affymetrix, Santa Clara, CA) were used for the gene expression detection. Fragmented cRNAs were denatured and hybridized to the probe array cartridge. Hybridization was carried out at 45°C for 16 h with mixing on a rotisserie at 60 rpm. After hybridization, the array cartridge was washed and stained with streptavidin/phycoerythrin in a fluidics station (Affymetrix). The probe array was scanned twice and the intensities were averaged with a GeneArray Scanner (Hewlett-Packard, Palo Alto, CA). Scanned images were processed and quantified using GeneChip Suite 3.2 (Affymetrix).
The expression level of the genes was measured by the average difference of the perfect match probes and mismatch probes. The expression levels for all genes presented on the microarrays were globally normalized across different experiments, so that all of the values are directly comparable. This was done by adjusting the mean of the global average difference in each array to 100. The reproducibility of the microarray experiments was characterized by comparing each set of data generated from the duplicated experiments. Synthesis of cDNA and cRNA from each set of biological samples was performed independently. The labeled cRNA samples were then hybridized to two different GeneChip microarrays. A correlation coefficient was calculated between the duplicate experiments. Genes with an average difference above 25 and showing a difference of 2-fold or above between the duplicate data set were identified as false positives. A false positive rate was also calculated based on the percentage of genes showing significant changes in duplicated experiments. Any gene with an average difference value above 25 and with a “present” call is considered as an expressed gene. These genes were further filtered to select those that were differentially expressed in response to wound treatments. The differentially expressed genes were selected by pair-wise comparison of the control and wounded samples from the same time point. These differentially expressed genes are defined as expression level of 25 or above, have a present call in at least one sample, and showed more than 2-fold induction or reduction in expression level. Only those genes that were altered 2-fold in both chip replicates were selected. Average values of fold changes from the duplicate chips were presented. Selected genes were clustered according to their expression level using self-organizing map and by hierarchical cluster analysis using the Cluster program, and displayed by TreeView program (Eisen et al., 1998). The Spearman rank correlation was used in the hierarchical cluster of the wounding-induced and -reduced genes. In addition, genes were also categorized according to their functions as described in the “Results and Discussion.”
Supplementary Material
ACKNOWLEDGMENTS
We thank Drs. Jane Glazebrook, John Grant, and Liang Shi for critical reading of the manuscript, and Dr. Wenqiong Chen for sharing unpublished stress-responsive gene expression results.
Footnotes
This work was supported in part by Syngenta Research and Technology.
The online version of this article contains Web-only data. The supplemental material is available at www.plantphysiol.org.
Article, publication date, and citation information can be found at www.plantphysiol.org/cgi/doi/10.1104/pp.002857.
LITERATURE CITED
- Alvarez ME, Pennell RI, Meijer P-J, Ishikawa A, Dixon RA, Lamb C. Reactive oxygen intermediates mediate a systemic signal network in the establishment of plant immunity. Cell. 1998;92:773–784. doi: 10.1016/s0092-8674(00)81405-1. [DOI] [PubMed] [Google Scholar]
- Bartel B. Auxin biosynthesis. Annu Rev Plant Physiol Plant Mol Biol. 1997;48:51–66. doi: 10.1146/annurev.arplant.48.1.51. [DOI] [PubMed] [Google Scholar]
- Bender J, Fink GR. A Myb homolog, ATR1, activates tryptophan gene expression in Arabidopsis. Proc Natl Acad Sci USA. 1998;95:5655–5660. doi: 10.1073/pnas.95.10.5655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borevitz JO, Xia Y, Blount J, Dixon RA, Lamb C. Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell. 2000;12:2383–2393. doi: 10.1105/tpc.12.12.2383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buschges R, Hollricher K, Panstruga R, Simons G, Wolter M, Frijters A, Van Daelen R, Van Der Lee T, Diergarde P, Groenendijk J et al. The barley Mlo gene: a novel control element of plant pathogen resistance. Cell. 1997;88:695–705. doi: 10.1016/s0092-8674(00)81912-1. [DOI] [PubMed] [Google Scholar]
- Cao H, Glazebrook J, Clarke JD, Volko S, Dong X. The Arabidopsis NPR1 gene that controls systemic acquired resistance encodes a novel protein containing ankyrin repeats. Cell. 1997;88:57–63. doi: 10.1016/s0092-8674(00)81858-9. [DOI] [PubMed] [Google Scholar]
- Century KS, Shapiro AD, Repetti PP, Dahlbeck D, Holub E, Staskawicz BJ. NDR1, a pathogen-induced component required for Arabidopsis disease resistance. Science. 1997;278:1963–1965. doi: 10.1126/science.278.5345.1963. [DOI] [PubMed] [Google Scholar]
- Chen W, Provart N, Glazebrook J, Katagiri F, Chang HS, Zou G, Whitham SA, Budworth PR, Tao Y, Xie Z et al. Expression profile matrix of Arabidopsis transcription factor genes suggests their putative functions in response to environmental stresses. Plant Cell. 2002;14:559–574. doi: 10.1105/tpc.010410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Creelman RA, Mullet JE. Oligosaccharins, brassinolides, and jasmonates: nontraditional regulators of plant growth, development, and gene expression. Plant Cell. 1997;9:1211–1223. doi: 10.1105/tpc.9.7.1211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Czernic P, Visser B, Sun W, Savoure A, Deslandes L, Marco Y, Van Montagu M, Verbruggen N. Characterization of an Arabidopsis thalianareceptor-like protein kinase gene activated by oxidative stress and pathogen attack. Plant J. 1999;18:321–327. doi: 10.1046/j.1365-313x.1999.00447.x. [DOI] [PubMed] [Google Scholar]
- De Luca V, St-Pierre B. The cell and developmental biology of alkaloid biosynthesis. Trends Plant Sci. 2000;5:168–173. doi: 10.1016/s1360-1385(00)01575-2. [DOI] [PubMed] [Google Scholar]
- Dong X. SA, JA, ethylene, and disease resistance in plants. Curr Opin Plant Biol. 1998;1:316–323. doi: 10.1016/1369-5266(88)80053-0. [DOI] [PubMed] [Google Scholar]
- Du L, Chen Z. Identification of genes encoding receptor-like protein kinases as possible targets of pathogen- and salicylic acid-induced WRKY DNA-binding proteins in Arabidopsis. Plant J. 2000;24:837–847. doi: 10.1046/j.1365-313x.2000.00923.x. [DOI] [PubMed] [Google Scholar]
- Durrant WE, Rowland O, Piedras P, Hammond-Kosack KE, Jones JDG. cDNA-AFLP reveals a striking overlap in race-specific resistance and wound response gene expression profiles. Plant Cell. 2000;12:963–977. doi: 10.1105/tpc.12.6.963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ecker JR. The ethylene signal transduction pathway in plants. Science. 1995;268:667–675. doi: 10.1126/science.7732375. [DOI] [PubMed] [Google Scholar]
- Eisen MB, Spellman PT, Brown PO, Botstein D. Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA. 1998;95:14863–14868. doi: 10.1073/pnas.95.25.14863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eulgem T, Rushton PJ, Robatzek S, Somssich IE. The WRKY superfamily of plant transcription factors. Trends Plant Sci. 2000;5:199–206. doi: 10.1016/s1360-1385(00)01600-9. [DOI] [PubMed] [Google Scholar]
- Glazebrook J, Rogers EE, Ausubel FM. Use of Arabidopsis for genetic dissection of plant defense responses. Annu Rev Genet. 1997;31:547–569. doi: 10.1146/annurev.genet.31.1.547. [DOI] [PubMed] [Google Scholar]
- Gurley WB. HSP101: a key component for the acquisition of thermotolerance in plants. Plant Cell. 2000;12:457–460. doi: 10.1105/tpc.12.4.457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harmer SL, Hogenesch JB, Straume M, Chang H-S, Han B, Zhu T, Wang X, Kreps JA, Kay SA. Orchestrated transcription of key pathways in Arabidopsis by the circadian clock. Science. 2000;290:2110–2113. doi: 10.1126/science.290.5499.2110. [DOI] [PubMed] [Google Scholar]
- Hasegawa PM, Bressan RA, Zhu J-K, Bohnert HJ. Plant cellular and molecular responses to high salinity. Annu Rev Plant Physiol Plant Mol Biol. 2000;51:463–499. doi: 10.1146/annurev.arplant.51.1.463. [DOI] [PubMed] [Google Scholar]
- Heo WD, Lee SH, Kim MC, Kim JC, Chung WS, Chun HJ, Lee KJ, Park CY, Park HC, Choi JY et al. Involvement of specific calmodulin isoforms in salicylic acid-independent activation of plant disease resistance responses. Proc Natl Acad Sci USA. 1999;96:766–771. doi: 10.1073/pnas.96.2.766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jaglo-Ottosen KR, Gilmour SJ, Zarka DG, Schabenberger O, Thomashow MF. Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science. 1998;280:104–106. doi: 10.1126/science.280.5360.104. [DOI] [PubMed] [Google Scholar]
- Jin H, Cominelli E, Bailey P, Parr A, Mehrtens F, Jones J, Tonelli C, Weisshaar B, Martin C. Transcriptional repression by AtMYB4 controls production of UV-protecting sunscreens in Arabidopsis. EMBO J. 2000;19:6150–6161. doi: 10.1093/emboj/19.22.6150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kovtun Y, Chiu W-L, Zeng W, Sheen J. Suppression of auxin signal transduction by a MAPK cascade in higher plants. Nature. 1998;395:716–720. doi: 10.1038/27240. [DOI] [PubMed] [Google Scholar]
- Kudla J, Xu Q, Harter K, Gruissem W, Luan S. Genes for calcineurin B-like proteins in Arabidopsis are differentially regulated by stress signals. Proc Natl Acad Sci USA. 1999;96:4718–4723. doi: 10.1073/pnas.96.8.4718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee GJ, Vierling E. A small heat shock protein cooperates with heat shock protein 70 systems to reactivate a heat-denatured protein. Plant Physiol. 2000;122:189–197. doi: 10.1104/pp.122.1.189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Q, Kasuga M, Sakuma Y, Abe H, Miura S, Yamaguchi-Shinozaki K, Shinozaki K. Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought- and low temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell. 1998;10:1391–1406. doi: 10.1105/tpc.10.8.1391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maleck K, Dietrich RA. Defense on multiple fronts: How do plants cope with diverse enemies? Trends Plant Sci. 1999;4:215–219. doi: 10.1016/s1360-1385(99)01415-6. [DOI] [PubMed] [Google Scholar]
- Maleck K, Levine A, Eulgem T, Morgan A, Schmid J, Lawton KA, Dangl JL, Dietrich RA. The transcriptome of Arabidopsis thaliana during systemic acquired resistance. Nat Genet. 2000;26:403–409. doi: 10.1038/82521. [DOI] [PubMed] [Google Scholar]
- Meissner RC, Jin H, Cominelli E, Denekamp M, Fuertes A, Greco R, Kranz HD, Penfield S, Petroni K, Urzainqui A et al. Function search in a large transcription factor gene family in Arabidopsis: assessing the potential of reverse genetics to identify insertional mutations in R2R3 MYB genes. Plant Cell. 1999;11:1827–1840. doi: 10.1105/tpc.11.10.1827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Memelink J, Verpporte R, Kijne JW. ORCAnization of jasmonate-responsive gene expression in alkaloid metabolism. Trends Plant Sci. 2001;6:212–219. doi: 10.1016/s1360-1385(01)01924-0. [DOI] [PubMed] [Google Scholar]
- Normanly J, Bartel B. Redundancy as a way of life: IAA metabolism. Curr Opin Plant Biol. 1999;2:207–213. doi: 10.1016/s1369-5266(99)80037-5. [DOI] [PubMed] [Google Scholar]
- Normanly J, Grisafi P, Fink GR, Bartel B. Arabidopsis mutants resistant to the auxin effects of indole-3-acetonitrile are defective in the nitrilase encoded by the NIT1 gene. Plant Cell. 1997;9:1781–1790. doi: 10.1105/tpc.9.10.1781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nurnberger T, Scheel D. Signal transmission in the plant immune response. Trends Plant Sci. 2001;6:372–379. doi: 10.1016/s1360-1385(01)02019-2. [DOI] [PubMed] [Google Scholar]
- O'Donnell PJ, Calvert C, Atzorn R, Wasternack C, Leyser HMO, Bowles DJ. Ethylene as a signal mediating the wound response of tomato plants. Science. 1996;274:1914–1917. doi: 10.1126/science.274.5294.1914. [DOI] [PubMed] [Google Scholar]
- Orozco-Cardenas M, Narvaez-Vasquez J, Ryan C. Hydrogen peroxide acts as a second messenger for the induction of defense genes in tomato plants in response to wounding, systemin, and methyl jasmonate. Plant Cell. 2001;13:179–191. [PMC free article] [PubMed] [Google Scholar]
- Orozco-Cardenas M, Ryan CA. Hydrogen peroxide is generated systemically in plant leaves by wounding and systemin via the octadecanoid pathway. Proc Natl Acad Sci USA. 1999;96:6553–6557. doi: 10.1073/pnas.96.11.6553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Penninckx IAMA, Thomma BPHJ, Buchala A, Metraux J-P, Broekaert WF. Concomitant activation of jasmonate and ethylene response pathways is required for induction of a plant defensin gene in Arabidopsis. Plant Cell. 1998;10:2103–2113. doi: 10.1105/tpc.10.12.2103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Radwanski ER, Last RL. Tryptophan biosynthesis and metabolism: biochemical and molecular genetics. Plant Cell. 1995;7:921–934. doi: 10.1105/tpc.7.7.921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reymond P, Farmer EE. Jasmonate and salicylate as global signals for defense gene expression. Curr Opin Plant Biol. 1998;1:404–411. doi: 10.1016/s1369-5266(98)80264-1. [DOI] [PubMed] [Google Scholar]
- Reymond P, Weber H, Damond M, Farmer EE. Differential gene expression in response to mechanical wounding and insect feeding in Arabidopsis. Plant Cell. 2000;12:707–719. doi: 10.1105/tpc.12.5.707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rojo E, Leon J, Sanchez-Serrano JJ. Cross-talk between wound signalling pathways determines local versus systemic gene expression in Arabidopsis thaliana. Plant J. 1999;20:135–142. doi: 10.1046/j.1365-313x.1999.00570.x. [DOI] [PubMed] [Google Scholar]
- Rushton PJ, Torres JT, Parniske M, Wernert P, Hahlbrock K, Somssich IE. Interaction of elicitor-induced DNA binding proteins with elicitor response elements in the promoters of parsley PR1 genes. EMBO J. 1996;15:5690–5700. [PMC free article] [PubMed] [Google Scholar]
- Ryals JA, Neuenschwander UH, Willits MG, Molina A, Steiner H-Y, Hunt MD. Systemic acquired resistance. Plant Cell. 1996;8:1809–1819. doi: 10.1105/tpc.8.10.1809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schenk PM, Kazan K, Wilson I, Anderson JP, Richmond T, Somerville SC, Manners JM. Coordinated plant defense responses in Arabidopsis revealed by microarray analysis. Proc Natl Acad Sci USA. 2000;97:11655–11660. doi: 10.1073/pnas.97.21.11655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seki M, Narusaka M, Abe H, Kasuga M, Yamaguchi-Shinozaki K, Carninci P, Hayashizaki Y, Shinozaki K. Monitoring the expression pattern of 1300 Arabidopsis genes under drought and cold stresses by using a full-length cDNA microarray. Plant Cell. 2001;13:61–72. doi: 10.1105/tpc.13.1.61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shinozaki K, Yamaguchi-Shinozaki K. Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signaling pathways. Curr Opin Plant Biol. 2000;3:217–223. [PubMed] [Google Scholar]
- Song W-Y, Wang G-L, Chen L-L, Kim H-S, Pi L-Y, Holsten T, Gardner J, Wang B, Zhai W-X, Zhu L-H et al. A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science. 1995;270:1804–1806. doi: 10.1126/science.270.5243.1804. [DOI] [PubMed] [Google Scholar]
- Sugimoto K, Takeda S, Hirochika H. MYB-related transcription factor NtMYB2 induced by wounding and elicitors is a regulator of the tobacco retrotransposon Tto1 and defense-related genes. Plant Cell. 2000;12:2511–2527. doi: 10.1105/tpc.12.12.2511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takezawa D. A rapid induction by elicitors of the mRNA encoding CCD-1, a 14-kDa Ca2+-binding protein in wheat cultured cells. Plant Mol Biol. 2000;42:807–817. doi: 10.1023/a:1006431724090. [DOI] [PubMed] [Google Scholar]
- Thomma BPHJ, Eggermont K, Penninckx IAMA, Mauch-Mani B, Vogelsang R, Cammue BPA, Broekaert WF. Separate jasmonate-dependent and salicylate-dependent defense-response pathways in Arabidopsis are essential for resistance to distinct microbial pathogens. Proc Natl Acad Sci USA. 1998;95:15107–15111. doi: 10.1073/pnas.95.25.15107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trewavas AJ, Malho R. Ca2+signalling in plant cells: the big network. Curr Opin Plant Biol. 1998;1:428–433. doi: 10.1016/s1369-5266(98)80268-9. [DOI] [PubMed] [Google Scholar]
- Wilson I, Vogel J, Somerville S. Signalling pathways: a common theme in plants and animals. Curr Biol. 1997;7:R175–R178. doi: 10.1016/s0960-9822(97)70082-4. [DOI] [PubMed] [Google Scholar]
- Yamaguchi-Shinozaki K, Shinozaki K. A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress. Plant Cell. 1994;6:251–264. doi: 10.1105/tpc.6.2.251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y, Fan W, Kinkema M, Li X, Dong X. Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene. Proc Natl Acad Sci USA. 1999;96:6523–6528. doi: 10.1073/pnas.96.11.6523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu T, Budworth P, Han B, Brown D, Chang H-S, Zou G, Wang X. Toward elucidating the global gene expression patterns of developing Arabidopsis: parallel analysis of 8300 genes by a high-density oligonucleotide probe array. Plant Physiol Biochem. 2001;39:221–242. [Google Scholar]
- Zhu T, Chang H-S, Schmeits J, Gil P, Shi L, Budworth P, Zou G, Chen X, Wang X. Gene expression microarrays: improvements and applications towards agricultural gene discovery. J Assoc Lab Automat. 2001a;6:95–98. [Google Scholar]
- Zhu T, Wang X. Large-scale profiling of the Arabidopsis transcriptome. Plant Physiol. 2000;124:1472–1476. doi: 10.1104/pp.124.4.1472. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.