Abstract
Anthocyanins are secondary metabolites found in higher plants that contribute to the colors of flowers and fruits. In apples (Malus domestica Borkh.), several steps of the anthocyanin pathway are coordinately regulated, suggesting control by common transcription factors. A gene encoding an R2R3 MYB transcription factor was isolated from apple (cv Cripps' Pink) and designated MdMYB1. Analysis of the deduced amino acid sequence suggests that this gene encodes an ortholog of anthocyanin regulators in other plants. The expression of MdMYB1 in both Arabidopsis (Arabidopsis thaliana) plants and cultured grape cells induced the ectopic synthesis of anthocyanin. In the grape (Vitis vinifera) cells MdMYB1 stimulated transcription from the promoters of two apple genes encoding anthocyanin biosynthetic enzymes. In ripening apple fruit the transcription of MdMYB1 was correlated with anthocyanin synthesis in red skin sectors of fruit. When dark-grown fruit were exposed to sunlight, MdMYB1 transcript levels increased over several days, correlating with anthocyanin synthesis in the skin. MdMYB1 gene transcripts were more abundant in red skin apple cultivars compared to non-red skin cultivars. Several polymorphisms were identified in the promoter of MdMYB1. A derived cleaved amplified polymorphic sequence marker designed to one of these polymorphisms segregated with the inheritance of skin color in progeny from a cross of an unnamed red skin selection (a sibling of Cripps' Pink) and the non-red skin cultivar Golden Delicious. We conclude that MdMYB1 coordinately regulates genes in the anthocyanin pathway and the expression level of this regulator is the genetic basis for apple skin color.
Anthocyanins are a class of secondary metabolites that contribute to the red, blue, and purple colors in a range of flowers and fruits. In flowers these pigments attract pollinators and in fruit skin they attract animals to aid in seed dispersal (Regan et al., 2001; Schaefer et al., 2004). Anthocyanins are also found in leaves, particularly during senescence (Feild et al., 2001), in stems, roots, and occasionally in fruit flesh and seeds.
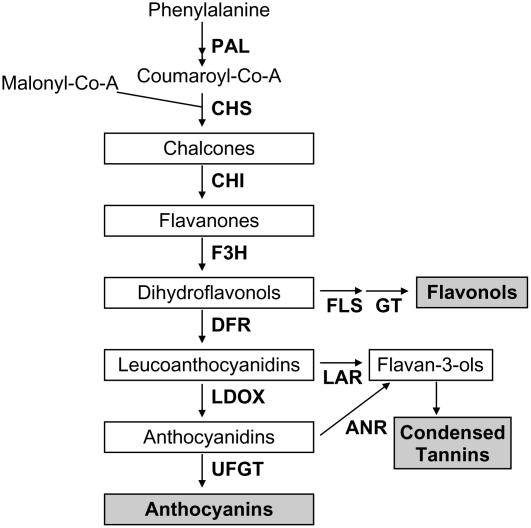
Anthocyanins, condensed tannins (CTs), and flavonols are synthesized via the flavonoid pathway, a branch of the phenylpropanoid pathway. The flavonoid pathway consists of a number of enzymatic steps that each catalyzes a sequential reaction for flavonoid synthesis, as illustrated in Figure 1. The pathway leads to synthesis of anthocyanins with branches for synthesis of flavonols (via flavonol synthase [FLS]) and synthesis of CTs (via leucoanthocyanidin reductase [LAR] and anthocyanidin reductase [ANR]). The genetics and biochemistry of this pathway have been characterized in petunia (Petunia hybrida), maize (Zea mays), snapdragon (Antirrhinum majus), and Arabidopsis (Arabidopsis thaliana; Winkel-Shirley, 2001). Regulation of the flavonoid pathway appears to be primarily at the level of transcription of the structural genes that encode the enzymes for each step of synthesis (Davies and Schwinn, 2003).
Figure 1.
Diagrammatic representation of the flavonoid biosynthetic pathway in apple. Enzymes for each step are shown in bold. Flavonoid intermediates are boxed and principle flavonoid end products are in gray boxes. The Phe ammonia lyase (PAL) enzyme is at a key branch point of the phenylpropanoid pathway that can lead to many classes of compounds. Enzymes required for flavonoid synthesis: CHS, Chalcone synthase; CHI, chalcone isomerase; F3H, flavanone-3β-hydroxylase; DFR, dihydroflavonol-4-reductase; LDOX, leucoanthocyanidin dioxygenase. Enzymes specific for various flavonoid compounds: UFGT, UDP-glycose:flavonoid-3-O-glycosyltransferase for anthocyanin synthesis. The unidentified enzyme encoding a glycosyl transferase for flavonol glycone synthesis is referred to as GT. The flavononoid-3′-hydroxylase enzyme has been omitted for clarity but it may hydroxylate several of the intermediates from the CHI step onwards.
Two families of regulators, the bHLH and MYB proteins, are conserved in the regulation of the anthocyanin and CT pathways in all species analyzed to date (Koes et al., 2005). The bHLH (also called MYC) proteins may have overlapping regulatory targets (Zhang et al., 2003; Zimmermann et al., 2004), but the MYB proteins are the key components providing specificity for the subsets of genes activated. For example, AtPAP1 and AtPAP2, which regulate anthocyanin synthesis, and AtTT2, which regulates CT synthesis, are specific for regulation of these branches of the flavonoid pathway (Borevitz et al., 2000; Nesi et al., 2001).
Arabidopsis has been an excellent model for delineating this fundamental basis of flavonoid synthesis and gene regulation (Winkel-Shirley, 2001). However, Arabidopsis produces dry fruit, similar to legumes, which do not normally accumulate anthocyanin. These fruit mature in a process more like senescence and disperse seeds by dehiscence (Adams-Phillips et al., 2004). To broaden our understanding of regulation of the flavonoid pathway in horticultural crops it will be necessary to conduct studies in plants with fleshy fruit. Grapevine (Vitis vinifera) is probably the best studied crop plant in terms of regulation of anthocyanin synthesis by transcription factors in fruit. In grapevine, two MYB genes, VvMYBA1 and VvMYBA2, which appear to regulate VvUFGT, contain mutations that segregate with white berries (Kobayashi et al., 2004; Walker et al., 2006). Other MYBs may activate a broader range of pathways; for example, when the grape gene VvMYB5a was expressed in tobacco (Nicotiana tabacum) it induced biosynthesis of anthocyanins, CTs, flavonols, and lignin (Deluc et al., 2006). However, regulation of anthocyanin synthesis in grapevine may be atypical in several respects compared to other crops. For example in berry skin only the VvUFGT step is differentially regulated for anthocyanin synthesis (Boss et al., 1996) and in some important cultivars used for wine making, such as Shiraz, anthocyanin synthesis in the fruit does not require direct light (Downey et al., 2004).
In the fruit of many crops, such as apple (Malus domestica), citrus (Citrus sinensis), peach (Prunus persica), pear (Pyrus communis), strawberry (Fragaria spp.), Vaccinium sp. such as cranberry (Vaccinium macrocarpon) and bilberry (Vaccinium myrtillus), eggplant (Solanum melongena), and lychee (Litchi chinensis), multiple flavonoid structural genes are coregulated for anthocyanin synthesis or in many of these fruit anthocyanin synthesis is enhanced by sunlight (Tyas et al., 1998; Matsuzoe et al., 1999; Zhou and Singh, 2000; Li et al., 2001; Jaakola et al., 2002; Kim et al., 2003; Kataoka and Beppu, 2004; Steyn et al., 2004; Tsuda et al., 2004; Jia et al., 2005; Lo Piero et al., 2005; Anttonen et al., 2006; Halbwirth et al., 2006). In none of these cases has a MYB or bHLH regulator specific for induction of anthocyanin synthesis in fruit been isolated. It is not known if either or both of these transcription factors are induced by sunlight or if other transcription factors, for example a bZIP protein (Hartmann et al., 2005) or an inhibitor protein (Hernandez et al., 2004), mediate the response to sunlight. Aside from grapevine, pepper (Capsicum annuum) is the only other fruit in which an anthocyanin MYB regulator, called CaA, has been identified (Borovsky et al., 2004). However, pepper is different from many other crops as anthocyanin is made early in fruit development and is later replaced by carotenoid pigments at ripening. In strawberry a MYB regulator of anthocyanin and flavonol pathways has been identified, called FaMYB1, but based on expression studies in tobacco it is believed to repress these pathways (Aharoni et al., 2001).
In horticultural plants, color is an important consideration in consumer choice. In fruits such as apple, grape, cherry (Prunus avium), strawberry, and others there has been much interest in breeding varieties bearing fruit with altered color, hues, patterns, or total anthocyanin content. In apples, red skin fruit are preferred to other fruit because consumers associate better-colored apples with better taste, ripeness, and flavor (King and Cliff, 2002). Color is also an obvious characteristic that enables product discrimination. For example, the apple cultivar Cripps' Pink, commercially sold as Pink Lady, has been very successful due to its distinct pink hue (Corrigan et al., 1997). Apple skin is also a rich source of compounds that are potent antioxidants, such as anthocyanins, CTs, and flavonols, and intake of such compounds in the human diet has been correlated with lower incidences of cancers and cardiac disease (Gallus et al., 2005; Liu et al., 2005; Rasmussen et al., 2005). For this reason the antioxidative potential of apple skin may become a new quality marker for different apple cultivars (Schirrmacher and Schempp, 2003).
In apple skin, sunlight is the most important external factor regulating anthocyanin synthesis (Saure, 1990; Lancaster, 1992). We recently demonstrated that in the red skin cultivar Cripps' Red, several flavonoid genes required for anthocyanin synthesis were coordinately transcribed in response to light exposure (Takos et al., 2006a). However, the expression of the MdLAR1 and MdANR genes, required for synthesis of CTs, was not light regulated. The coordinated changes in expression of several apple flavonoid genes also have a genetic basis; at fruit ripening transcripts of MdCHS, MdF3H, MdDFR, MdLDOX, and MdUFGT were found to be barely detectable in the non-red skin cultivar Orin but were abundant in the red skin cultivars Fuji and Jonathan (Honda et al., 2002). The data suggest that expression of these genes is controlled by a common regulator that is defective in non-red skin cultivars.
In this study, we report the isolation of a light-induced gene that encodes a MYB regulator of anthocyanin synthesis in apple fruit skin. This gene, called MdMYB1, was characterized in terms of the correlation of its expression with anthocyanin synthesis and function in two heterologous transgene assays. A derived cleaved amplified polymorphic sequence (dCAPS) marker for an allele of MdMYB1 showed segregation with skin color. This marker will be a useful tool for apple breeding programs. Our analysis has delineated the regulation of anthocyanin synthesis in apple fruit skin and will further our understanding of flavonoid regulation in other crops.
RESULTS
Flavonoid Synthesis and Gene Expression in the Skin of Apple Cultivars
Levels of anthocyanins, CTs, and flavonols accumulated in apple fruit skin were measured in several cultivars collected when the fruit was ripening (Fig. 2). Skin from the fruit of the non-red skin cultivars Golden Delicious, Granny Smith, Grandspur, Firm Gold, Shuzaka, and Einscheimer did not contain any detectable anthocyanins (Fig. 2A). Anthocyanin levels ranged from approximately 15 ng mg−1 to approximately 165 ng mg−1 in the red skin cultivars, an unnamed selection with red skin (US), Cripps' Red, Gala, Galaxy, and Hi Early. There was significant variation in the levels of flavonols (Fig. 2B) and CTs (Fig. 2C) among the different cultivars but no clear correlation with color. The non-red skin cultivars accumulated flavonols and CTs in a similar range to that measured in the red skin cultivars.
Figure 2.
Flavonoid concentration in the skin of apple cultivars at fruit ripening. A, Anthocyanin. B, Flavonol. C, CTs. Flavonoids were extracted from pooled samples of peel taken from the entire surface of six to 10 apples for each cultivar. Dashed line separates the non-red skin and red skin cultivars. Data are means of three replicates with error bars indicating ±sds. US, Unnamed selection with red skin.
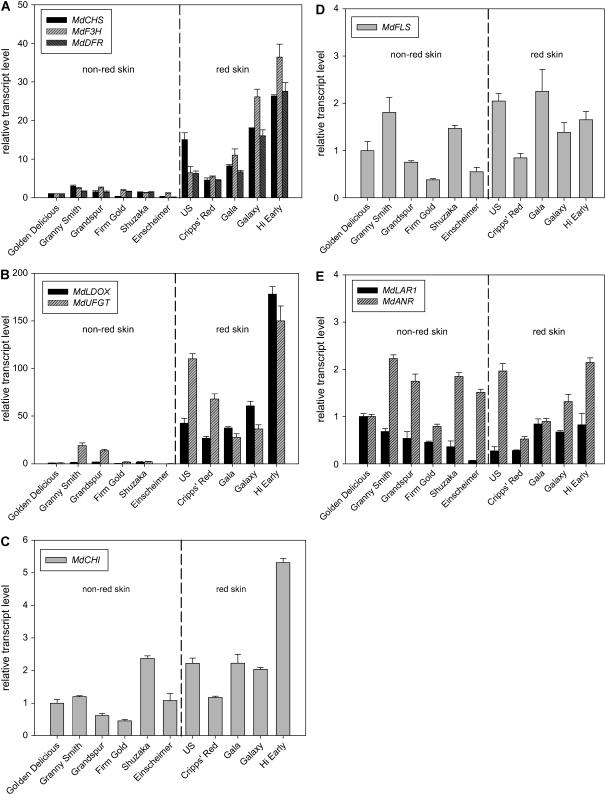
The transcript levels of genes that encode the enzymes of the flavonoid pathway (Fig. 1) were measured in the skin of non-red and red skin apple cultivars by real-time PCR. Transcripts of the early genes of the pathway, required for both anthocyanin and CT synthesis (MdCHS, MdF3H, and MdDFR), were typically 5- to 20-fold higher in red skin cultivars than non-red skin cultivars (Fig. 3A). Transcripts of the late genes of the pathway (MdLDOX and MdUFGT), of which the latter gene is specific for anthocyanin synthesis, ranged from 25- to 180-fold higher in red skin cultivars compared to non-red skin cultivars (Fig. 3B). The exceptions were Granny Smith and Grandspur, which had higher transcript levels of MdUFGT than other non-red skin cultivars although transcript levels were still well below that of the red skin cultivars. Transcripts of the MdCHI gene, which is also required for both anthocyanin and CT synthesis, did not have the same pattern as that observed for the other genes and its expression was at similar levels in all the cultivars analyzed (Fig. 3C). Transcripts of MdFLS, which is specific for the synthesis of flavonols, and MdLAR1 and MdANR, which are specific for the synthesis of CTs, were at similar levels in non-red and red skin cultivars (Fig. 3, D and E).
Figure 3.
Flavonoid structural gene expression in the skin of apple cultivars at fruit ripening. A, MdCHS, MdF3H, and MdDFR. B, MdLDOX and MdUFGT. C, MdCHI. D, MdFLS. E, MdLAR1 and MdANR. Total RNA was extracted from pooled samples of peel described in Figure 2 and reverse transcribed into cDNA. Transcript levels of genes were determined by real-time PCR using gene-specific primers and corrected to the apple UBIQUITIN (MdUBQ) gene. Transcript levels were normalized relative to the transcript level for each gene in the cultivar Golden Delicious. Dashed line separates the non-red skin and red skin cultivars. Data are means of three replicate PCR reactions with error bars indicating ±sds. US, Unnamed selection with red skin.
Isolation of MdMYB1
Anthocyanin synthesis in apple fruit skin is induced by sunlight (Saure, 1990; Lancaster, 1992). The anthocyanin pathway was manipulated artificially in apples by covering fruit on trees with bags early in the season to exclude sunlight. When the bags were removed during the fruit ripening phase, reexposure to sunlight induced anthocyanin synthesis after about 2 to 3 d in Cripps' Red and Cripps' Pink fruit (Takos et al., 2006a; J.-L. Parker, S.P. Robinson, A.R. Walker, and A.M. Takos, personal communication). Coinciding with the reexposure to sunlight, we found increased transcript levels of all the flavonoid structural genes required for anthocyanin synthesis: MdCHS, MdCHI, MdF3H, MdDFR, MdLDOX, and MdUFGT, but synthesis of flavonols and CTs was not induced (Takos et al., 2006a).
One or more transcription factors specific to the anthocyanin pathway should be induced when bags are removed from dark-grown fruit and anthocyanin synthesis is induced by sunlight. To exploit this possibility we prepared cDNA from Cripps' Pink fruit skin in which the anthocyanin pathway had been strongly induced by light and used it as a template for PCR to isolate anthocyanin MYB regulators. Degenerate primers were designed from the conserved regions in the R2R3 domain of MYB transcription factors from other plant species that had been functionally characterized to regulate anthocyanin synthesis. A 246 bp cDNA was isolated that encoded a peptide with approximately 80% sequence identity to the R2R3 region of the petunia MYB transcription factor PhAN2. The 5‘ and 3’ ends of the cDNA were isolated by RACE PCR. The 848 bp full-length cDNA contained a coding region for a deduced amino acid sequence of 243 residues in length and this protein was designated as MdMYB1.
A phylogenetic analysis of the R2R3 region of this deduced amino acid sequence places MdMYB1 in a cluster of MYB proteins that include: Gerbera hybrida GMYB10, snapdragon AmVENOSA, AmROSEA1, and AmROSEA2, pepper CaA, tomato (Lycopersicon esculentum) LeANT1, petunia PhAN2, grape VvMYBA1 and VvMYBA2, and Arabidopsis AtPAP1 and AtPAP2 (Fig. 4A). These MYBs have all been characterized as anthocyanin regulators. The cluster that contains maize ZmPL and ZmC1, which are also specific for regulation of anthocyanin synthesis, and the Arabidopsis AtTT2, which regulates CT synthesis, are more distantly related, as are MYBs that regulate other processes.
Figure 4.
Comparison of MdMYB1 deduced amino acid sequence with other MYB proteins. A, Phylogenetic analysis displaying the similarity of MdMYB1 (red box) with other MYBs that have been characterized as regulators of the anthocyanin pathway. Origins of the various MYBs are given a one or two letter prefix: Md is apple, G is gerbera hybrid, Vv is grapevine, At is Arabidopsis, Ph is petunia, Ca capsicum, Le is tomato, Fa is strawberry, Zm is maize, Am is snapdragon, Dc is carrot, and Nt is tobacco. Functions of the other MYB proteins are indicated. The tree was based on the alignment of the 104 amino acids spanning the R2R3 domain using the CLUSTALW alignment program and default parameters of the Molecular Evolutionary Genetics Analysis package, version 3.1. The tree was then constructed from this file using the neighbor-joining method with a bootstrap test of phylogeny of 10,000 replications and a random seed of 64,238. Bootstrap values are shown at nodes and the scale bar represents 0.05 substitutions per site. Accession numbers of proteins are listed in “Materials and Methods.” B, Protein sequence alignment of the anthocyanin group of MYB regulators and other closely related MYBs identified by the phylogenetic analysis. The R2 and R3 domains are underlined in gray and black, respectively. The bHLH binding motif is boxed in red in the R3 domain. A specific residue of interest common to anthocyanin regulators is indicated with a red arrow within the R2 domain and the number of this residue referred to in the text is that given by Stracke et al. (2001). A red arrow from the box R/S indicates an Arg residue in the amino acid sequence of MdMYB1-1 that is a Ser in MdMYB1-3. Motifs 5 and 6 identified by Stracke et al. (2001) in the C-terminal domain are boxed in red. Protein sequences were aligned with the CLUSTALW program at the Australian National Genome Information Service site (http://www.angis.org.au/) and displayed using the GeneDoc Version 2.6.002 program.
An alignment of the deduced amino acid sequences of some of these proteins shows the high sequence homology within the R2R3 domain at the amino terminus (Fig. 4B). All of the proteins contain the motif [D/E]Lx2[R/K]x3Lx6Lx3R in the R3 domain for interactions with R-like bHLH proteins (Zimmermann et al., 2004). Within the highly conserved R2 domain it is interesting to note that MdMYB1 and the anthocyanin regulators (with the exception of ZmC1) have an Arg at position 39 when a Gly residue is found at this position in ZmC1 and AtTT2 and in 88% of Arabidopsis R2R3 MYBs (Stracke et al., 2001). It is not clear if this residue is critical for the specificity of the anthocyanin MYBs; however, it may be an interesting target for mutagenesis.
There is little homology in the sequence carboxyl terminal to the R2R3 domain; an exception is the motif KPRPR[S/T]F defined by Stracke et al. (2001) as motif 6 in AtPAP1 and PhAN2. This motif is also conserved in CaA and is modified to [K/R]Pxxx[K/T][F/Y] in MdMYB1, GMYB10, AmROSAEA1, LeANT1, and VvMYBA1 but is absent in AtTT2. This motif is also absent in ZmC1, a regulator of anthocyanin synthesis in maize, but it is interesting to note that Baudry et al. (2004) found that ZmC1 could activate the AtANR promoter in Arabidopsis protoplasts, suggesting ZmC1 can activate transcription of at least one CT synthesis gene. Another motif was incorrectly identified as DExWRLxxT in ZmC1 and AtTT2 by Stracke et al. (2001), and designated motif 5, but is actually DEDWLRxxT. This motif is not found in the anthocyanin MYB regulators although there is some homology in this region. When sequencing MdMYB1 cDNAs an A/T polymorphism was found at nucleotide position 573 of the coding sequence that encoded either an Arg (allele MdMYB1-1) or Ser residue (MdMYB1-3), respectively, at amino acid position 191 of the deduced amino acid sequence. MdMYB1-2 is defined later in this report.
MdMYB1 Expression Analysis
Transcript levels of MdMYB1 were determined by real-time PCR using gene-specific primers designed to the region encoding the carboxyl domain. When bags were removed from dark-grown Cripps' Red fruit and the fruit reexposed to sunlight, MdMYB1 transcripts increased by approximately 20-fold after 1 d and then increased to approximately 25-fold higher after 6 d before transcript levels declined again (Fig. 5A). This is similar to the pattern observed for MdUFGT transcript levels and correlates with anthocyanin synthesis when bagged fruit were uncovered (Takos et al., 2006a). In control fruit, which were grown under normal light conditions, MdMYB1 transcripts were approximately 25-fold higher than in the bagged fruit and this level did not fluctuate by more than approximately 2-fold throughout the course of the experiment.
Figure 5.
MdMYB1 gene expression patterns. A, Transcript levels of MdMYB1 in fruit bagging experiment. Fruit from the cultivar Cripps' Red were grown in bags from 48 DAFB until 147 DAFB when the bags were removed and fruit then referred to as unbagged fruit. Control fruit were grown under normal light conditions for the entire period. Total RNA extracted from peel taken from the entire surface of apples on the day that bags were removed from the dark-grown fruit (day 0) and on subsequent days after bag removal. The total RNA was reverse transcribed into cDNA and transcripts were measured by real-time PCR. Data is normalized relative to the transcript levels in the dark-grown fruit at day 0. B, Transcript levels of MdMYB1 in various reproductive tissues of Cripps' Red. Tissues were collected from six to 10 samples at the stages indicated and then pooled for total RNA extraction. The total RNA was reversed transcribed into cDNA for real-time PCR analysis. Data is normalized relative to the transcript levels in flower buds. C, Transcript levels of MdMYB1 in non-red and red skin cultivars. Samples were as described in Figure 3 and data normalized to transcript levels in Golden Delicious. Dashed line separates the non-red skin and red skin cultivars. All data in A to C was corrected to MdUBQ and is the mean of three replicate PCRs with error bars indicating ±sds.
Transcript levels of MdMYB1 in Cripps' Red fruit were approximately 10-fold higher in red skin sectors than in flesh, seeds, flower buds, or young fruit (Fig. 5B). Skin sector samples were taken at 48 and 147 d after full bloom (DAFB), which we found in a previous study coincided with two distinct phases of anthocyanin synthesis in Cripps' Red fruit (Takos et al., 2006b). At 48 DAFB MdMYB1 transcript levels were approximately 24-fold higher in red skin than green skin sectors and at 147 DAFB were approximately 4.5-fold higher.
Transcript levels of MdMYB1 were 15- to 50-fold higher in red skin cultivars than in non-red skin cultivars (Fig. 5C). This correlates with the levels of anthocyanin measured in these cultivars (Fig. 2A) and transcripts of the genes required for anthocyanin synthesis (Fig. 3, A and B). The exceptions were Granny Smith and Grandspur, which had 5-fold higher levels of MdMYB1 transcripts than the other non-red skin cultivars. The data shows a positive correlation between MdMYB1 expression levels and anthocyanin synthesis in response to light, tissue specificity, and genetic background.
Functional Testing of MdMYB1
The function of the MdMYB1 protein was tested in planta by introducing MdMYB1-1 cDNA under the transcriptional control of the cauliflower mosaic virus 35S promoter into two heterologous systems: Arabidopsis plants and cultured grape cells.
Arabidopsis plants, transformed with 35S:MdMYB1-1 cDNA, developed seeds with spots of red pigmentation at the funicular end, indicating anthocyanin synthesis (Fig. 6A). For comparison, plants were also transformed with 35S:AtPAP1 cDNA, the endogenous Arabidopsis MYB anthocyanin regulator. These plants developed a stronger phenotype than for plants transformed with 35S:MdMYB1-1 cDNA as the red pigmentation in seeds was more intense and not restricted to the funicular end (Fig. 6B). For both constructs the phenotype was best observed in immature seeds (less than 10 d after anthesis) before cells of the seed coat die and compress (about 15 d after anthesis) and the seed coat darkens. Seeds of the wild-type plants (ecotype Columbia) had no pigmentation (Fig. 6C) and no pigmentation was observed in any other plant structures for either the wild-type or transformed plants. The data shows that MdMYB1 can function to induce anthocyanin synthesis in Arabidopsis.
Figure 6.
Functional analysis of MdMYB1 in heterologous systems. Photos of developing seeds (less than 10 d after anthesis) from Arabidopsis ecotype Columbia transformed with MdMYB1-1 cDNA (A) or AtPAP1 cDNA (B), while C is seed from untransformed plants. Photos A and B are representative of one of six separate lines of the T2 generation and similar phenotypes were observed in all lines. D, Photo of a cultured grape cell induced to synthesize anthocyanin by bombardment with MdMYB1-1 and AtEGL3 cDNA. E, Stimulation of LUC activity derived from MdDFR and MdUFGT promoters fused to the firefly LUC gene and cobombarded into grape cells with either of two different MdMYB1 cDNAs; MdMYB1-1 encoding an Arg at position 191 and MdMYB1-3 encoding a Ser at position 191. Data is the mean of the ratio of P. pyralis LUC activity to R. formis LUC activity normalized to bombardment in which the MYB cDNA was absent. Error bars are ±sds of three bombardments from three separate experiments.
Cultured grape cells bombarded with gold particles coated with MdMYB1-1 cDNA and the bHLH partner AtEGL3, both driven by the cauliflower mosaic virus 35S promoter, were induced to synthesize anthocyanin (Fig. 6D). No pigmented cells were observed when transformed with AtEGL3 cDNA alone. The MdMYB1-1 and MdMYB1-3 cDNAs, which encoded either an Arg or Ser at position 191, respectively (Fig. 4B), were tested for their efficacy to induce expression from the promoters of apple flavonoid structural genes in the grape cell assay. Both these MdMYB1 cDNAs were able to induce an approximately 25-fold and approximately 10-fold increase in luciferase (LUC) enzyme activity when cobombarded with constructs containing the MdDFR and MdUFGT promoters, respectively, fused to the firefly LUC gene (Fig. 6E). Similar numbers of pigmented cells were also observed after bombardment with both MdMYB1 cDNAs (data not shown). These results show that MdMYB1 can activate expression of two apple anthocyanin pathway genes and indicate that either an Arg or a Ser at position 191 does not affect MdMYB1 function.
MdMYB1 Genomic Sequence Analysis
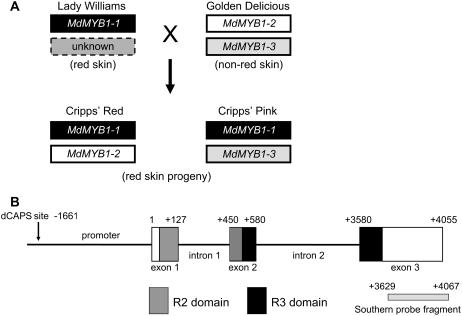
Genomic fragments encompassing approximately 6 kb of MdMYB1 and spanning from approximately 2 kb of the promoter to the stop codon were isolated from genomic DNA of Golden Delicious and the red skin siblings Cripps' Pink and Cripps' Red. These siblings are progeny of a cross of the red skin cultivar Lady Williams with the non-red skin cultivar Golden Delicious. Sequence analysis revealed the presence of three MdMYB1 alleles in these cultivars (Fig. 7A). The MdMYB1-1 allele was inherited by both Cripps' Pink and Cripps' Red but was not present in the parent Golden Delicious. Therefore, MdMYB1-1 must have been inherited from the red skin parent Lady Williams and thus was an excellent candidate single dominant gene controlling anthocyanin synthesis in apple skin.
Figure 7.
Analysis of MdMYB1 genomic DNA. A, Diagrammatic representation of the inheritance of MdMYB1 alleles in the cross of Lady Williams with Golden Delicious as determined by sequencing. MdMYB1 alleles were sequenced in Golden Delicious, Cripps' Red, and Cripps' Pink. The presence of the MdMYB1-1 allele in Lady Williams is inferred since this allele is not present in Golden Delicious. The identity of the other MdMYB1 allele present in Lady Williams is unknown. B, Genomic organization of MdMYB1-1 gene (not drawn to scale). Numbers refer to position relative to the first nucleotide of the start codon. Coding sequence is shown in boxes and noncoding sequence is shown as black line. Location of R2 and R3 domains is indicated by gray and black, respectively. Exons, introns, and the promoter region are labeled. Location of dCAPS forward primer binding site is indicted in the promoter (it is this primer sequence that is restricted by BstEII in the PCR fragment derived from MdMYB1-1). The probe fragment used for Southern analysis is indicated by a pale gray box.
The genomic organization of MdMYB1-1 consists of three exons and two introns (Fig. 7B). The R2 domain is split over exons 1 and 2, while the R3 domain is split over exons 2 and 3. This is similar to the structure of other R2R3 MYBs (Jiang et al., 2004). Intron 1 is 323 bp in length while intron 2 is 3 kb in length. Southern blotting under stringent conditions using a DNA probe made to the region encoding the carboxyl-terminal domain indicated there is a single copy of MdMYB1 in the genome of Lady Williams, Golden Delicious, Cripps' Red, and Cripps' Pink (data not shown). The other two alleles of MdMYB1 have the same basic gene structure as MdMYB1-1; one of these was designated MdMYB1-2 and the other corresponded to MdMYB1-3 that was functional in the grape cell bombardment (Fig. 6E).
A comparison of the nucleotide sequences of the three alleles shows that they are identical in exons 1 and 2 but in MdMYB1-1 and MdMYB1-2 there are a small number of nucleotide differences in the promoter and a single nucleotide difference in intron 2 (Table I). The MdMYB1-3 allele is more different from the other alleles with many nucleotide differences in the promoter and introns and a nucleotide change in exon 3 that results in the Arg-to-Ser change at position 191 of the deduced amino acid sequence (Fig. 4B).
Table I.
Comparison of features of genomic sequence of MdMYB1 alleles
| Allele | Sequence Comparison with MdMYB1-1a
|
|||
|---|---|---|---|---|
| Promoter | Intron 1 | Intron 2 | Exon 3 | |
| MdMYB1-1 | 1,865 bp | 323 bp | 3,000 bp | 475 bp |
| MdMYB1-2 | Eight nucleotide differences | Identical | One nucleotide difference | Identical |
| MdMYB1-3 | >25 nucleotide differences | 256 bp (93.5% homology) | 2,995 bp (98.3% homology) | A → T (Arg → Ser position 191) |
Exons 1 and 2 are identical in the three alleles.
MdMYB1-1 Cosegregation with Skin Color
A single nucleotide polymorphism (SNP) in the promoter of the MdMYB1-1 allele is different from the SNP in the MdMYB1-2 and MdMYB1-3 alleles. This SNP was used as the basis for designing primers for a dCAPS PCR assay to test the segregation of MdMYB1 alleles with the inheritance of fruit skin color. The forward primer was designed with the SNP incorporated in the primer sequence and its location in the promoter is shown in Figure 7B. After amplification, PCR products that contained the MdMYB1-1 promoter sequence were restricted by the enzyme BstEII to a size of 263 bp, while products that did not contain this sequence, such as derived from the MdMYB1-2 and MdMYB1-3 alleles, were not digested and were 291 bp in size.
The 263 bp PCR fragment was detected in all of the red skin cultivars we tested, indicating they contained the MdMYB1-1 allele (Fig. 8A). Most of these cultivars appeared to be heterozygous for MdMYB1-1 as an undigested PCR product of 291 bp was also detected, indicating the presence of another allele. For two of the red skin cultivars, Fuji and Hi Early, only the 263 bp PCR fragment was detected, which indicates they could be homozygous for MdMYB1-1 or that they contain an allele whose sequence could either not be amplified or distinguished from MdMYB1-1 by the dCAPS assay. The latter possibility seems most likely since Fuji is heterozygous for skin color inheritance as when it is crossed with a non-red skin cultivar it produces progeny with a 1:1 segregation of skin color.
Figure 8.
Analysis of segregation of MdMYB1-1 allele by dCAPS assay. A, Fragments from dCAPS assay of genomic DNA extracted from the leaves of non-red and red skin cultivars. B, Photos of fruit from progeny of the cross of a red selection with Golden Delicious and below fragments from dCAPS assay of genomic DNA from leaves of these progeny. PCR fragments were digested with BstEII and separated on 3% (w/v) NuSieve GTG Agarose gel. Migration of molecular mass markers is not shown but the top fragment is 291 bp and the bottom fragment is 263 bp.
For most of the non-red skin cultivars only the 291 bp PCR product was detected, indicating they do not contain the MdMYB1-1 allele. In the cases of Granny Smith and Grandspur, which are genetically related, the dCAPS assay indicated the presence of the MdMYB1-1 allele, yet these are non-red skin cultivars. Sequence analysis of this allele confirmed that, while it had the same sequence as MdMYB1-1 in the region where the dCAPS primers were designed, there were numerous other differences compared to the MdMYB1-1 promoter. Therefore these cultivars contain at least one unidentified allele of MdMYB1.
The segregation of MdMYB1-1 with color inheritance was tested in progeny from the cross of an unnamed red skin selection (Red Selection, from a cross of Lady Williams by Golden Delicious and a sibling of Cripps' Pink and Cripps' Red) and Golden Delicious. The utility of the dCAPS assay for predicting skin color inheritance is shown in a small number of the progeny, which is typical of the results we obtained for a total of 136 progeny (Fig. 8B). Apples in which the 263 bp PCR fragment was detected, indicating the presence of MdMYB1-1, had varying levels of pigmentation perhaps reflecting the degree of exposure to sunlight. Apples in which only the 291 bp PCR fragment was detected had none or very poor pigmentation. In total, 93 progeny were scored for red skin color and in all of these the MdMYB1-1 allele was detected by the dCAPS assay, while for the 43 progeny scored as non-red the MdMYB1-1 allele was not detected (Table II). The MdMYB1-1 allele shows segregation with red skin color inheritance, indicating it is probably the color locus for apple fruit skin in these cultivars.
Table II.
Segregation of the MdMYB1-1 allele in apple progeny
| Red Selection (MdMYB1-1, MdMYB1-2)a | ||
|---|---|---|
| × | ||
| Golden Delicious (MdMYB1-2, MdMYB1-3)
| ||
| Scoring of Progeny | Inheritance of MdMYB1-1b | |
| Red skin | 93 | 93/93 |
| Non-red skin | 43 | 0/43 |
The selection is a sibling of Cripps' Pink and Cripps' Red.
Predicted by the dCAPS marker.
DISCUSSION
Activity of the Flavonoid Pathway in Apple Cultivars
To determine the nature of the genetic basis of apple fruit skin color we analyzed flavonoid synthesis and structural gene transcription in several non-red and red-skin cultivars. We found that non-red skin cultivars, despite being deficient in anthocyanin synthesis, were still able to synthesize CTs and flavonols in fruit skin to a similar level as found in red skin cultivars. Our results are in agreement with the previous analysis of flavonoid accumulation in the fruit skin of several apple cultivars at ripening (Lata et al., 2005; Vanzani et al., 2005). The study by Lister et al. (1994) also showed that similar composition and concentrations of flavonols and CTs accumulated in the fruit skin of the red skin cultivar Splendour and the non-red skin cultivar Granny Smith throughout fruit maturation.
Using real-time PCR we determined that transcript levels of MdCHS, MdF3H, MdDFR, MdLDOX, and MdUFGT genes required for anthocyanin synthesis were lower in fruit skin of non-red than red skin cultivars. This result is in agreement with the enzyme activity data of Lister et al. (1996) and gene transcription data of Honda et al. (2002) and Kondo et al. (2002) that indicated multiple anthocyanin pathway steps were deficient in the fruit skin of non-red skin cultivars. Unlike the anthocyanin pathway genes, transcripts of genes specific to the flavonol and CT pathways were at similar levels in all the cultivars tested.
Since transcription of at least five structural flavonoid genes was affected, the genetic basis of apple skin color is probably due to the activity of a common regulator of these genes. This would be analogous to AtPAP1, which is a specific regulator of the anthocyanin pathway in Arabidopsis (Borevitz et al., 2000). Since the flavonol and CT pathways are not perturbed in non-red apples it is likely there are separate regulators for these pathways. In Arabidopsis, specific MYB regulators of these pathways have been found, for example AtMYB12 is a regulator of AtCHS and AtFLS, which are required for flavonol synthesis (Mehrtens et al., 2005), while AtTT2 regulates AtDFR, AtLDOX, and AtANR, which are required for CT synthesis (Nesi et al., 2001). In apple, there could be orthologs of these MYB regulators that can stimulate transcription from promoters of some of the early flavonoid pathway genes so as to generate the precursor compounds required for flavonol and CT synthesis. These regulators would be expressed early in fruit development when these compounds are made, and in particular transcripts of the genes required for flavonol and CT synthesis are abundant (Takos et al., 2006b).
The MdCHI gene was the only anthocyanin pathway gene that did not seem to have reduced expression in non-red skin apple cultivars at fruit ripening. This suggests that MdCHI is not coregulated with the other pathway steps. However, both Lister et al. (1996) and Li et al. (2002) found that MdCHI enzyme activity was much lower in non-red than red skin cultivars at fruit ripening. We analyzed one isoform of MdCHI but Southern analysis has indicated that there are at least two sequences similar to MdCHI in the apple genome (Takos et al., 2006b). It is possible that another MdCHI gene isoform we have not analyzed is involved in anthocyanin synthesis and so analysis of other putative MdCHI genes is needed to clarify whether this step is coregulated.
Characterization of MdMYB1
We isolated a gene that encoded an R2R3 MYB family transcription factor we named MdMYB1. Phylogenetic analysis placed MdMYB1 in a group of MYB proteins that have been functionally characterized to regulate anthocyanin synthesis and separate from MYBs that regulate other processes.
Consistent with a role in regulating anthocyanin synthesis, MdMYB1 gene transcripts increased within 1 d when bags were removed from apple fruit exposing skin to sunlight. This increase in MdMYB1 transcripts was accompanied by the accumulation of anthocyanin and flavonoid structural gene transcripts in the skin (Takos et al., 2006a). Other anthocyanin MYB regulators have also been shown to respond to light; for example, in Arabidopsis anthocyanin synthesis and expression of both AtPAP1 and AtPAP2 respond to light under stress conditions (Vanderauwera et al., 2005). Correlation has been found between light, anthocyanin synthesis, and transcript levels of MYB anthocyanin regulators in maize seed aleurone and seedling mesocotyl and in Perilla frutescens leaves (Procissi et al., 1997; Gong et al., 1999; Piazza et al., 2002). However, MdMYB1 is the first MYB regulator of the anthocyanin pathway to show light induction in the skin of a fleshy fruit. This is different from grapevine (Shiraz) berries where anthocyanin synthesis is unperturbed by bunch shading (Downey et al., 2004), presumably indicating the VvMYBA1 and VvMYBA2 genes are regulated differently compared to MdMYB1. There is some variability in the response of grapevine cultivars to light. For example, compared to those grown in full sun, Cabernet Sauvignon berries contain less anthocyanin when shaded from sunlight; however, VvMYBA1 expression seems to be only slightly affected (Jeong et al., 2004).
The preferential expression of MdMYB1 in red skin sectors of apple fruit is consistent with the model of a specific regulator of the anthocyanin branch of the pathway. This is because CT synthesis and MdLAR and MdANR transcripts are distributed almost equally between red and green skin sectors of the same fruit (Takos et al., 2006b) and so would probably require a regulator that is expressed in both these tissue types. Furthermore, transcripts of MdMYB1 were more abundant in the red skin compared to the non-red skin apple cultivars (Fig. 5C), which are still able to accumulate flavonols and CTs and express the flavonoid pathway genes specific to their synthesis. Similarly, in grapevine (Shiraz) berries CT and flavonol synthesis is temporally separated from anthocyanin synthesis, or is induced by different environmental conditions, and so these pathways are also likely to be controlled by separate regulatory factors (Downey et al., 2003a, 2003b, 2004; Bogs et al., 2005). Specific MYB regulators for the flavonol and CT pathways are yet to be isolated from apple and grapevine fruit or any other fleshy fruit.
We demonstrated that MdMYB1 can function as an anthocyanin regulator by introducing the coding region of the cDNA into Arabidopsis plants and cultured grape cells. In both these systems anthocyanin synthesis was induced (Fig. 6, A–D). The Arabidopsis seed coat is rich in CTs, but when the AtANR gene is disrupted anthocyanins can accumulate (Devic et al., 1999). The plants transformed with MdMYB1 cDNA developed partially red seeds, suggesting this regulator was able to divert some of the flux of intermediates through the flavonoid pathway away from synthesis of CTs and toward synthesis of anthocyanin, presumably by activation of an endogenous UFGT gene (Lee et al., 2005). The AtPAP1 cDNA produced a stronger phenotype than MdMYB1 cDNA and this may be because of a better interaction of AtPAP1 with other endogenous regulatory cofactors or flavonoid structural gene promoters or because of posttranscriptional or posttranslational regulation.
Mechanism of Light Induction of Flavonoid Structural Genes by MdMYB1
We have shown that MdMYB1 can activate transcription from the apple MdDFR and MdUFGT promoters when cobombarded into grape cells (Fig. 6E). The promoters of MdDFR and MdUFGT contain putative light regulatory units, consisting of an ACGT element and MYB-recognition element (MRE), or MRE-like sequences, which have been identified by Hartmann et al. (2005) to be necessary for light responsiveness of several structural genes required for flavonol synthesis in Arabidopsis. The ACGT elements are bound by bZIP factors or bHLH factors and the MRE/MRE-like sequence elements are bound by MYB factors. It was proposed that a combination of a MYB and bZIP factor conferred the light responsiveness while a MYB and bHLH factor conferred tissue specificity. In Perilla a model has been proposed in which a MYB factor, MYB-P1, and two bHLH factors are required for anthocyanin synthesis and the genes encoding the MYB and one of the bHLH factors, MYC-F3G1, are both light induced (Yamazaki et al., 2003). In maize the bHLH gene Sn and the MYB genes C1 and Pl are light induced but it is the expression of the MYB genes that are considered to be rate limiting for anthocyanin synthesis (Procissi et al., 1997). We have shown in apple that the MYB component for regulation of anthocyanin structural genes is encoded by a light-induced gene, MdMYB1, but the apple bHLH and/or bZIP cofactors required for anthocyanin synthesis have not been isolated and so their contribution to light regulation remains to be determined.
Genetic Segregation of MdMYB1 Alleles
We isolated and sequenced three alleles of MdMYB1 in cultivars descended from Lady Williams crossed with Golden Delicious (Fig. 7A). Using a dCAPS PCR assay we found that one of these alleles, MdMYB1-1, segregated with skin color in most of the cultivars we tested. The exceptions were the non-red skin cultivars Granny Smith and Grandspur; however, these cultivars contain a different unidentified allele of MdMYB1 that is presumably nonfunctional, probably due to low expression (Fig. 5C). Our data is in agreement with the random-amplified polymorphic DNA markers developed by Cheng et al. (1996), which also predict skin color inheritance to be determined by a single dominant gene. The random-amplified polymorphic DNA markers were also unable to correctly predict color inheritance for the cultivar Granny Smith but with our sequence data dCAPS markers could be designed to test the segregation of MdMYB1 alleles from this cultivar. The presence of a different allele in Granny Smith and Grandspur correlates with the expression of MdMYB1 and flavonoid structural genes that were slightly higher in these cultivars compared to other non-red skin cultivars (Figs. 5C and 3B).
The perfect segregation of the MdMYB1-1 allele with color in 136 progeny of Red Selection (a sibling of Cripps' Pink) backcrossed to Golden Delicious demonstrated that this allele is tightly linked to the color locus in these cultivars. However, the segregation ratio of greater than 2:1 of red-skinned to green-skinned progeny is different from the expected 1:1 ratio. There are a number of possible reasons why this has occurred. Apples have a self incompatibility system that might reduce pollination by one allele since the parents are genetically related (Schneider et al., 2005). The population of 136 progeny is from a selection trial where individuals have already been culled; if germination was incomplete or slow, the seedling did not grow in a strong and characteristic fashion or if the young tree did not survive and thrive after transplanting into the selection block. Only trees that have started to bear fruit could be scored and the red-fruited trees might establish more quickly than trees with non-red fruit. Mutations in linked genes affecting these processes could distort the segregation ratio. Although this unexpected segregation ratio is interesting, it does not detract from our observation that skin color and MdMYB1-1 cosegregate.
The two other alleles sequenced, MdMYB1-2 and MdMYB1-3, encode, respectively, an identical protein to MdMYB1-1 and a protein with an Arg-to-Ser exchange in the C-terminal domain (Fig. 4B), but this change does not perturb functional activity of the MdMYB1 cDNA in the grape cell assay (Fig. 6E). Therefore it seems likely these alleles are deficient in regulating anthocyanin synthesis due to altered gene regulation, as evidenced by the poor expression in non-red skin cultivars (Fig. 5C). This would explain why non-red skin cultivars can still develop a faint blush under high light (Reay, 1999; Kondo et al., 2002). Presumably these conditions increase the levels of a regulatory factor of MdMYB1 or activate alternate regulatory pathways to induce some MdMYB1 expression.
The MdMYB1-1 dCAPS marker could be a useful tool for apple breeding programs as it would allow the prediction of fruit skin color formation long before immature trees are able to bare fruit. This would be of even greater significance if MdMYB1 also controls later steps of pigment formation that could affect skin hue. For example, overexpression of the MYBs LeANT1 and AtPAP1 in tomato and Arabidopsis, respectively, induces genes involved in flavonoid modification and transport into vacuoles (Mathews et al., 2003; Tohge et al., 2005). In Arabidopsis, novel anthocyanins were produced. If MdMYB1 has a similar spectrum of action, then alleles that control fine differences in color intensity or shading could be identified and used for breeding.
There has been extensive testing of growth conditions to improve anthocyanin synthesis in orchard-grown apples, including chemical sprays and plant hormones, the use of reflecting films on the canopy floor to alter light conditions and manipulation of the crop load, the canopy, and temperature conditions (Arakawa, 1988; Reay et al., 1998; Ju et al., 1999; Reay, 1999; Awad et al., 2001a, 2001b; Kondo et al., 2001; Reay and Lancaster, 2001; Iglesias et al., 2002; Layne et al., 2002; Embree and Nichols, 2005). The identification of an anthocyanin regulatory gene MdMYB1 provides a molecular tool to study the impact of these processes and conditions on anthocyanin synthesis and may provide a genetic solution to improving the consistency of color in apples.
CONCLUSION
In apple fruit the flavonoid pathway is spatially and temporally regulated but it is also responsive to light. We have identified a MYB regulatory gene, MdMYB1, that has an expression pattern correlating with that of the structural genes during fruit development and in response to light and coincides with anthocyanin synthesis. The MYB regulator can activate the promoters of two of these structural genes in a transient assay and one allele of this gene segregates with skin color in progeny of a cross of two apples with different-colored skins. This suggests that MdMYB1 regulates structural genes across the flavonoid pathway in apple fruit, determining skin color and providing the potential to modulate fruit color by altering expression of this regulatory gene. It will be of great interest to determine how the response to light is also mediated through this key regulator in apple fruit.
MATERIALS AND METHODS
Apple Fruit Samples
Samples of fruit skin, flesh, seeds, and early fruit development stages of apple (Malus domestica Borkh. cv Cripps' Red) were collected at the Primary Industries and Resources South Australia Site at Lenswood in the Adelaide Hills, South Australia from November, 2002 until April, 2003 as described by Takos et al. (2006b). Fruit bagging experiments were conducted on cultivars Cripps' Red in April, 2003 and Cripps' Pink in April, 2004 at the Lenswood site as described by Takos et al. (2006a). All other apple samples were collected at Manjimup, western Australia, including fruit skin samples from various cultivars collected in the mid to late stages of ripening in 2003. Samples were pooled from six to 10 individual fruit. Young leaves for genomic DNA extraction from a red-skinned selection (Red Selection, from a cross between Lady Williams and Golden Delicious) × Golden Delicious progeny were collected in 2003 and 2005. All samples were immediately frozen in liquid nitrogen and stored at −80°C.
Flavonoid Analysis
Anthocyanins and flavonols were extracted from 0.1 g of finely ground plant material in 1 mL 1% (v/v) HCL-methanol for 1 h at room temperature on a rotating wheel in darkness. Samples were clarified by centrifugation at 13,000g for 15 min at room temperature and 10 μL of supernatant was analyzed by HPLC, as described by Takos et al. (2006a).
CTs were extracted from 0.1 g of finely ground plant material in 1 mL 70% (v/v) acetone. The extract was processed in a tannin-protein binding assay as according to Downey and Adams (2005).
Preparation of Nucleic Acids and cDNA Synthesis
Total RNA was isolated from 1 g apple tissue by a hot borate method (Wan and Wilkins, 1994). Genomic DNA was isolated from 20 mg of young leaf material using a DNeasy plant mini kit (QIAGEN).
First-strand cDNA was synthesized from 4 μg total RNA except for flesh samples where 2 μg was used due to an inhibition of synthesis at higher concentrations. The reaction was primed by oligo dT(18) and synthesis of cDNA catalyzed by SuperScriptIII RNase H-Reverse Transcriptase with RNAseOUT (Invitrogen) included in the reaction to inhibit RNA degradation.
Expression Analysis
For real-time PCR, cDNA was diluted 1:20 with water and 5 μL placed into a reaction containing 266 nm of each primer and Absolute QPCR SYBR Green buffer (ABgene). Primers for flavonoid structural genes and thermocycling conditions are described in Takos et al. (2006b). Primers for MdMYB1 real-time PCR were: forward, 5′-AGACCAATGTGATAAGACCTCAG-3′ and reverse, 5′-AACCAAAAACTTGTGAAGAGTTC-3′, and reaction conditions were as described for flavonoid structural genes. The PCR reactions were carried out in a RotorGene2000 real-time PCR instrument (Corbett Research). The relative gene copy number for each cDNA was calculated against standards of purified PCR fragments from each gene as described in Takos et al. (2006b). For all analysis the signal obtained for a gene of interest was normalized against the signal obtained for the apple UBIQUITIN (MdUBQ) gene.
Isolation of MdMYB1 cDNA
First-strand cDNA was synthesized as described above from 4 μg total RNA extracted from dark-grown Cripps' Pink fruit that had been exposed to sunlight for 2 d. Degenerate PCR primers for isolation of MYB genes were: forward, 5′-TGYATHRAYAARTAYGGIGARGGIAARTGG-3′ and reverse, 5′-GTRTTCCARTARTTYTTIACRTCRTTNGC-3′. Standard PCR conditions were used for Platinum Taq DNA Polymerase (Invitrogen) with the addition of betaine to 1 m. Thermocycling conditions were: initial denaturation of 94°C for 2 min; 35 cycles of 94°C for 30 s, 45°C for 30 s, 72°C for 1 min; and final extension of 72°C for 5 min. A single 246 bp PCR product was ligated to pDrive (QIAGEN) and several clones sequenced. A single sequence was obtained and designated MdMYB1. Nested primers were designed for 3′ and 5′ RACE PCR. The cDNA template primed with oligo dT(18) described above was used for the 3′ RACE PCR while for 5′ RACE PCR fresh first-strand cDNA was made from the Cripps' Pink total RNA sample above using a GeneRacer kit (Invitrogen). Using the sequence information obtained by RACE PCR primers were designed to amplify a PCR fragment containing the full-length coding sequence of MdMYB1 and were: forward, 5′-GCGGTACCGGTAGCAGGCAAAAGAATAGCTAAGC-3′ and reverse, 5′-GCGGATCCCACATTTACAAGCAAGGAAAATA-3′. An 848 bp PCR fragment was obtained from the Cripps' Pink cDNA template used above in a reaction catalyzed by PfuTurbo (Stratagene) following the manufacturer's reaction conditions. The PCR fragment was tailed with dATP using a QIAGEN A-Addition kit and was ligated to pDrive (QIAGEN). Several clones were sequenced and two MdMYB1 cDNAs, which differed in nucleotide 573 of the coding sequence resulting in an Arg (MdMYB1-1) or Ser (MdMYB1-3) in position 191 of the deduced amino acid sequence, were identified.
Generation of MdMYB1 Transgenic Arabidopsis Plants
The full-length coding region sequence MdMYB1-1 or AtPAP1 was ligated to the binary vector pART27 or pBART27, a derivative of pART27, respectively (Gleave, 1992), and used to transform Agrobacterium tumefaciens Agl1. Arabidopsis (Arabidopsis thaliana) plants, ecotype Columbia, were transformed with the Agrobacterium by the floral dipping method (Clough and Bent, 1998). T1 MdMYB1 transgenic plants were selected on one-half-strength Murashige and Skoog media containing 8 g/L agar and 350 mg/L kanamycin. Kanamycin-resistant T1 seedlings were transferred to soil and grown at 20°C in a growth chamber (Phoenix Biosystems) with a 16 h day length and a light intensity of 150 mmol m−2 s−2. Plants transformed with AtPAP1 were grown on soil and selected by resistance to basta. Seeds of individual self-fertilized T2 lines were collected and single-copy insertion lines were selected based on a Mendelian segregation ratio.
Functional Assay Using Grape Cells
Two MdMYB1 ORFs, MdMYB1-1 and MdMYB1-3, were ligated to the pART7 vector (Gleave, 1992). Fragments containing promoters of MdDFR and MdUFGT genes, which were approximately 2 kb and approximately 1.8 kb in size, respectively, were isolated with a Universal GenomeWalker kit (BD Biosciences CLONTECH) following the manufacturer's instructions and using Cripps' Pink genomic DNA as a template. These fragments were ligated to the vector pLuc containing the Photinus pyralis LUC gene (Horstmann et al., 2004). Gold particles were coated by the method in Ramsay et al. (2003) with a mixture of 150 ng each of the constructs; MdMYB1 in pART7, AtEGL3 in pFF19 (Ramsay et al., 2003), either MdDFR or MdUFGT promoter fragments in pLuc, and 3 ng of the Renilla reniformis LUC plasmid pRluc (Horstmann et al., 2004). A suspension culture of grapevine (Vitis vinifera) Chardonnay petiole callus was grown in Grape Cormier medium (Do and Cormier, 1991) to a log phase and filtered onto sterile Whatman1 discs (5.5 cm) on the surface of Grape Cormier-agar plates. The Chardonnay cells were bombarded with DNA-coated gold particles at 350 kPa helium in a vacuum of 75 kPa and a distance of 14 cm as described by Torregrosa et al. (2002). Cells were harvested 48 h after transfection and lysed by grinding on ice in 150 μL of Passive Lysis buffer (Promega). Lysates were clarified by centrifugation for 2 min at 500g and the activity of P. pyralis and R. reniformis LUC enzymes determined using the Dual-Luciferase Reporter Assay system (Promega). Light emission was measured using a TD-20/20 Luminometer (Turner Biosystems).
Cloning of MdMYB1 Genomic Sequence
Primers for the PCR of the full-length MdMYB1 cDNA (above) were also used to amplify the full-length genomic sequence from Cripps Pink, Cripps' Red, and Golden Delicious genomic DNA templates. Fragments, approximately 4 kb in length, were amplified using Platinum Taq DNA Polymerase High Fidelity (Invitrogen) following the manufacturer's recommended reaction conditions. The PCR products were ligated to pDrive (QIAGEN) and several clones sequenced for each reaction.
An approximately 2 kB fragment of the promoter sequence of MdMYB1 was isolated with a Universal GenomeWalker kit (BD Biosciences) as described above. Primers specific to each MdMYB1 allele, based upon differences in intron 1, were then used to amplify the promoter sequence that corresponded to each allele of MdMYB1 with Platinum Taq DNA Polymerase High Fidelity (Invitrogen) following the manufacturer's conditions. The PCR products were ligated to pDrive (QIAGEN) and several clones sequenced for each reaction.
MdMYB1-1 dCAPS Assay
Primers for dCAPS PCR were: forward, 5′-CCTGAACACGTGGGAACCGGCCCGTTGGTAAC-3′ and reverse, 5′-GTGAAGGTTGTCTTTATTAGTGACGTG-3′. The approximately 250 bp PCR product was amplified with Platinum Taq DNA Polymerase (Invitrogen) following the manufacturer's conditions in a reaction of 50 μL. Thermocycling conditions were: initial denaturation of 94°C for 2 min; 35 cycles of 94°C for 30 s, 55°C for 30 s, 72°C for 30 s; and final extension of 72°C for 5 min. After PCR, 15 μL of the reaction was digested with 10 units of BstEII (Promega) at 65°C for 4 h. Digested DNA fragments were resolved on a 3% (w/v) NuSieve GTG Agarose gel (FMC BioProducts).
Accession Numbers of Sequences
Sequences referred to in this article are deposited at the National Center for Biotechnology Information under the following accession numbers: apple, MdMYB1-1 (DQ886414), MdMYB1-2 (DQ886415), MdMYB1-3 (DQ886416), promoter of MdDFR (DQ886412), promoter of MdUFGT (DQ886413); gerbera hybrid, GMYB10 (CAD87010); tomato (Lycopersicon esculentum), LeANT1 (AAQ55181); petunia (Petunia hybrida), PhAN2 (AAF66727), PhPH4 (AAY52377), and PhODO1 (AAV98200); capsicum (Capsicum annuum) CaA (CAE75745); Arabidopsis, AtPAP1 (ABB03879), AtTT2 (Q9FJA2), AtMYB12 (ABB03913), AtWER (AAF18939), and AtGL1 (AAC97387); strawberry (Fragaria spp.), FaMYB1 (AAK84064); grapevine, VvMYBA1 (BAD18977), VvMYBA2 (AB097924), and VvMYB5a (AAS68190); maize (Zea mays), ZmC1 (AAA33482) and ZmPl (AAA19821); snapdragon (Antirrhinum majus), AmMIXTA (CAA55725), AmROSEA1 (ABB83826), AmROSEA2 (ABB83827), and AmVENOSA (ABB83828); carrot (Daucus carota), DcMYB1 (BAE54312); and tobacco (Nicotiana tabacum), NtMYB2 (BAA88222).
Note Added in Proof
A similar gene, MdMYB10 (DQ267896–DQ267898), has been isolated from apple and shown to regulate anthocyanin production (Espley RV, Hellens RP, Putterill J, Stevenson DE, Kutty-Amma S, Allan AC [2006] Red colouration in apple fruit is due to the activity of MYB transcription factor, MdMYB10. Plant J [in press]).
Acknowledgments
We thank Commonwealth Scientific and Industrial Research Organization colleagues Nicole Cordon for help with HPLC analysis, and Karin Sefton and Debra McDavid for excellent technical assistance; staff from Primary Industries and Resources South Australia for allowing us to sample apple fruits at Lenswood in the Adelaide Hills; Eva Decker, the University of Freiburg, for luciferase expression vectors; Mark Downey, Department of Primary Industries Victoria, for developing methods of analysis of flavonoids; and the Department of Agriculture and Food Western Australia and Horticulture Australia Ltd. for a grant to support this work.
This work was supported by a grant from the Department of Agriculture and Food Western Australia and Horticulture Australia Ltd.
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (www.plantphysiol.org) is: Amanda R. Walker (mandy.walker@csiro.au).
References
- Adams-Phillips L, Barry C, Giovannoni J (2004) Signal transduction systems regulating fruit ripening. Trends Plant Sci 9: 331–338 [DOI] [PubMed] [Google Scholar]
- Aharoni A, De Vos CHR, Wein M, Sun ZK, Greco R, Kroon A, Mol JNM, O'Connell AP (2001) The strawberry FaMYB1 transcription factor suppresses anthocyanin and flavonol accumulation in transgenic tobacco. Plant J 28: 319–332 [DOI] [PubMed] [Google Scholar]
- Anttonen MJ, Hoppula KI, Nestby R, Verheul MJ, Karjalainen RO (2006) Influence of fertilization, mulch color, early forcing, fruit order, planting date, shading, growing environment, and genotype on the contents of selected phenolics in strawberry (Fragaria x ananassa Duch.) fruits. J Agric Food Chem 54: 2614–2620 [DOI] [PubMed] [Google Scholar]
- Arakawa O (1988) Characteristics of color development in some apple cultivars—changes in anthocyanin synthesis during maturation as affected by bagging and light quality. J Jpn Soc Hortic Sci 57: 373–380 [Google Scholar]
- Awad MA, De Jager A, Dekker M, Jongen WMF (2001. a) Formation of flavonoids and chlorogenic acid in apples as affected by crop load. Sci Hortic (Amst) 91: 227–237 [Google Scholar]
- Awad MA, Wagenmakers PS, de Jager A (2001. b) Effects of light on flavonoid and chlorogenic acid levels in the skin of “Jonagold” apples. Sci Hortic (Amst) 88: 289–298 [Google Scholar]
- Baudry A, Heim MA, Dubreucq B, Caboche M, Weisshaar B, Lepiniec L (2004) TT2, TT8, and TTG1 synergistically specify the expression of BANYULS and proanthocyanidin biosynthesis in Arabidopsis thaliana. Plant J 39: 366–380 [DOI] [PubMed] [Google Scholar]
- Bogs J, Downey MO, Harvey JS, Ashton AR, Tanner GJ, Robinson SP (2005) Proanthocyanidin synthesis and expression of genes encoding leucoanthocyanidin reductase and anthocyanidin reductase in developing grape berries and grapevine leaves. Plant Physiol 139: 652–663 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borevitz JO, Xia YJ, Blount J, Dixon RA, Lamb C (2000) Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell 12: 2383–2393 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borovsky Y, Oren-Shamir M, Ovadia R, De Jong W, Paran I (2004) The A locus that controls anthocyanin accumulation in pepper encodes a MYB transcription factor homologous to Anthocyanin2 of Petunia. Theor Appl Genet 109: 23–29 [DOI] [PubMed] [Google Scholar]
- Boss PK, Davies C, Robinson SP (1996) Expression of anthocyanin biosynthesis pathway genes in red and white grapes. Plant Mol Biol 32: 565–569 [DOI] [PubMed] [Google Scholar]
- Cheng FS, Weeden NF, Brown SK (1996) Identification of co-dominant RAPD markers tightly linked to fruit skin color in apple. Theor Appl Genet 93: 222–227 [DOI] [PubMed] [Google Scholar]
- Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16: 735–743 [DOI] [PubMed] [Google Scholar]
- Corrigan VK, Hurst PL, Boulton G (1997) Sensory characteristics and consumer acceptability of “Pink Lady” and other late-season apple cultivars. New Zeal J Crop Hortic Sci 25: 375–383 [Google Scholar]
- Davies KM, Schwinn KE (2003) Transcriptional regulation of secondary metabolism. Funct Plant Biol 30: 913–925 [DOI] [PubMed] [Google Scholar]
- Deluc L, Barrieu F, Marchive C, Lauvergeat V, Decendit A, Richard T, Carde JP, Merillon JM, Hamdi S (2006) Characterization of a grapevine R2R3-MYB transcription factor that regulates the phenylpropanoid pathway. Plant Physiol 140: 499–511 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devic M, Guilleminot J, Debeaujon I, Bechtold N, Bensaude E, Koornneef M, Pelletier G, Delseny M (1999) The BANYULS gene encodes a DFR-like protein and is a marker of early seed coat development. Plant J 19: 387–398 [DOI] [PubMed] [Google Scholar]
- Do CB, Cormier F (1991) Effects of high ammonium concentrations on growth and anthocyanin formation in grape (Vitis-Vinifera L) cell-suspension cultured in a production medium. Plant Cell Tissue Organ Cult 27: 169–174 [Google Scholar]
- Downey M, Adams D (2005) Practical use of a total tannin assay for red wine grapes. Aust Vitic 9: 68–71 [Google Scholar]
- Downey MO, Harvey JS, Robinson SP (2003. a) Analysis of tannins in seeds and skins of Shiraz grapes throughout berry development. Aust J Grape Wine Res 9: 15–27 [Google Scholar]
- Downey MO, Harvey JS, Robinson SP (2003. b) Synthesis of flavonols and expression of flavonol synthase genes in the developing grape berries of Shiraz and Chardonnay (Vitis vinifera L.). Aust J Grape Wine Res 9: 110–121 [Google Scholar]
- Downey MO, Harvey JS, Robinson SP (2004) The effect of bunch shading on berry development and flavonoid accumulation in Shiraz grapes. Aust J Grape Wine Res 10: 55–73 [Google Scholar]
- Embree CG, Nichols DS (2005) Bio-regulators improve fruit size and colour and reduce crop-load and annual bearing of Honeycrisp (TM) apples. Can J Plant Sci 85: 453–455 [Google Scholar]
- Feild TS, Lee DW, Holbrook NM (2001) Why leaves turn red in autumn: the role of anthocyanins in senescing leaves of red-osier dogwood. Plant Physiol 127: 566–574 [PMC free article] [PubMed] [Google Scholar]
- Gallus S, Talamini R, Giacosa A, Montella M, Ramazzotti V, Franceschi S, Negri E, La Vecchia C (2005) Does an apple a day keep the oncologist away? Ann Oncol 16: 1841–1844 [DOI] [PubMed] [Google Scholar]
- Gleave AP (1992) A versatile binary vector system with a T-DNA organizational-structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol Biol 20: 1203–1207 [DOI] [PubMed] [Google Scholar]
- Gong ZZ, Yamazaki M, Saito K (1999) A light-inducible Myb-like gene that is specifically expressed in red Perilla frutescens and presumably acts as a determining factor of the anthocyanin formation. Mol Gen Genet 262: 65–72 [DOI] [PubMed] [Google Scholar]
- Halbwirth H, Puhl I, Haas U, Jezik K, Treutter D, Stich K (2006) Two-phase flavonoid formation in developing strawberry (Fragaria x ananassa) fruit. J Agric Food Chem 54: 1479–1485 [DOI] [PubMed] [Google Scholar]
- Hartmann U, Sagasser M, Mehrtens F, Stracke R, Weisshaar B (2005) Differential combinatorial interactions of cis-acting elements recognized by R2R3-MYB, BZIP, and BHLH factors control light-responsive and tissue-specific activation of phenylpropanoid biosynthesis genes. Plant Mol Biol 57: 155–171 [DOI] [PubMed] [Google Scholar]
- Hernandez JM, Heine GF, Irani NG, Feller A, Kim MG, Matulnik T, Chandler VL, Grotewold E (2004) Different mechanisms participate in the R-dependent activity of the R2R3 MYB transcription factor C1. J Biol Chem 279: 48205–48213 [DOI] [PubMed] [Google Scholar]
- Honda C, Kotoda N, Wada M, Kondo S, Kobayashi S, Soejima J, Zhang ZL, Tsuda T, Moriguchi T (2002) Anthocyanin biosynthetic genes are coordinately expressed during red coloration in apple skin. Plant Physiol Biochem 40: 955–962 [Google Scholar]
- Horstmann V, Huether CM, Jost W, Reski R, Decker EL (2004) Quantitative promoter analysis in Physcomitrella patens: a set of plant vectors activating gene expression within three orders of magnitude. BMC Biotechnol 4: 1–13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iglesias I, Salvia J, Torguet L, Cabus C (2002) Orchard cooling with overtree microsprinkler irrigation to improve fruit colour and quality of “Topred Delicious” apples. Sci Hortic (Amst) 93: 39–51 [Google Scholar]
- Jaakola L, Maatta K, Pirttila AM, Torronen R, Karenlampi S, Hohtola A (2002) Expression of genes involved in anthocyanin biosynthesis in relation to anthocyanin, proanthocyanidin, and flavonol levels during bilberry fruit development. Plant Physiol 130: 729–739 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeong ST, Goto-Yamamoto N, Kobayashi S, Esaka A (2004) Effects of plant hormones and shading on the accumulation of anthocyanins and the expression of anthocyanin biosynthetic genes in grape berry skins. Plant Sci 167: 247–252 [Google Scholar]
- Jia HJ, Araki A, Okamoto G (2005) Influence of fruit bagging on aroma volatiles and skin coloration of “Hakuho” peach (Prunus persica Batsch). Postharvest Biol Technol 35: 61–68 [Google Scholar]
- Jiang CZ, Gu X, Peterson T (2004) Identification of conserved gene structures and carboxy-terminal motifs in the Myb gene family of Arabidopsis and Oryza sativa L. ssp. indica. Genome Biol 5: R46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ju ZQ, Duan YS, Ju ZG (1999) Effects of covering the orchard floor with reflecting films on pigment accumulation and fruit coloration in “Fuji” apples. Sci Hortic (Amst) 82: 47–56 [Google Scholar]
- Kataoka I, Beppu K (2004) UV irradiance increases development of red skin color and anthocyanins in “Hakuho” peach. HortScience 39: 1234–1237 [Google Scholar]
- Kim SH, Lee JR, Hong ST, Yoo YK, An G, Kim SR (2003) Molecular cloning and analysis of anthocyanin biosynthesis genes preferentially expressed in apple skin. Plant Sci 165: 403–413 [Google Scholar]
- King MC, Cliff MA (2002) Development of a model for prediction of consumer liking from visual attributes of new and established apple cultivars. J Am Pom Soc 56: 223–229 [Google Scholar]
- Kobayashi S, Goto-Yamamoto N, Hirochika H (2004) Retrotransposon-induced mutations in grape skin color. Science 304: 982. [DOI] [PubMed] [Google Scholar]
- Koes R, Verweij W, Quattrocchio F (2005) Flavonoids: a colorful model for the regulation and evolution of biochemical pathways. Trends Plant Sci 10: 236–242 [DOI] [PubMed] [Google Scholar]
- Kondo S, Maeda M, Kobayashi S, Honda C (2002) Expression of anthocyanin biosynthetic genes in Malus sylvestris L. “Mutsu” non-red apples. J Hortic Sci Biotechnol 77: 718–723 [Google Scholar]
- Kondo S, Tsukada N, Niimi Y, Seto H (2001) Interactions between jasmonates and abscisic acid in apple fruit, and stimulative effect of jasmonates on anthocyanin accumulation. J Jpn Soc Hortic Sci 70: 546–552 [Google Scholar]
- Lancaster JE (1992) Regulation of skin color in apples. CRC Crit Rev Plant Sci 10: 487–502 [Google Scholar]
- Lata B, Przeradzka M, Binkowska M (2005) Great differences in antioxidant properties exist between 56 apple cultivars and vegetation seasons. J Agric Food Chem 53: 8970–8978 [DOI] [PubMed] [Google Scholar]
- Layne DR, Jiang ZW, Rushing JW (2002) The influence of reflective film and ReTain on red skin coloration and maturity of “Gala” apples. Horttechnology 12: 640–645 [Google Scholar]
- Lee Y, Yoon HR, Paik YS, Liu JR, Chung W, Choi G (2005) Reciprocal regulation of Arabidopsis UGT78D2 and BANYULS is critical for regulation of the metabolic flux of anthocyanidins to condensed tannins in developing seed coats. J Plant Biol 48: 356–370 [Google Scholar]
- Li YJ, Sakiyama R, Maruyama H, Kawabata S (2001) Regulation of anthocyanin biosynthesis during fruit development in “Nyoho” strawberry. J Jpn Soc Hortic Sci 70: 28–32 [Google Scholar]
- Li ZH, Sugaya S, Gemma H, Iwahori S (2002) Flavonoid biosynthesis and accumulation and related enzyme activities in the skin of “Fuji” and “Oorin” apples during their development. J Jpn Soc Hortic Sci 71: 317–321 [Google Scholar]
- Lister CE, Lancaster JE, Sutton KH, Walker JRL (1994) Developmental changes in the concentration and composition of flavonoids in skin of a red and a green skin apple cultivar. J Sci Food Agric 64: 155–161 [Google Scholar]
- Lister CE, Lancaster JE, Walker JRL (1996) Developmental changes in enzymes of flavonoid biosynthesis in the skins of red and green apple cultivars. J Sci Food Agric 71: 313–320 [Google Scholar]
- Liu RH, Liu JR, Chen BQ (2005) Apples prevent mammary tumors in rats. J Agric Food Chem 53: 2341–2343 [DOI] [PubMed] [Google Scholar]
- Lo Piero AR, Puglisi I, Rapisarda P, Petrone G (2005) Anthocyanins accumulation and related gene expression in red orange fruit induced by low temperature storage. J Agric Food Chem 53: 9083–9088 [DOI] [PubMed] [Google Scholar]
- Mathews H, Clendennen SK, Caldwell CG, Liu XL, Connors K, Matheis N, Schuster DK, Menasco DJ, Wagoner W, Lightner J, et al (2003) Activation tagging in tomato identifies a transcriptional regulator of anthocyanin biosynthesis, modification, and transport. Plant Cell 15: 1689–1703 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuzoe N, Yamaguchi M, Kawanobu S, Watanabe Y, Higashi H, Sakata Y (1999) Effect of dark treatment of the eggplant on fruit skin color and its anthocyanin component. J Jpn Soc Hortic Sci 68: 138–145 [Google Scholar]
- Mehrtens F, Kranz H, Bednarek P, Weisshaar B (2005) The Arabidopsis transcription factor MYB12 is a flavonol-specific regulator of phenylpropanoid biosynthesis. Plant Physiol 138: 1083–1096 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nesi N, Jond C, Debeaujon I, Caboche M, Lepiniec L (2001) The Arabidopsis TT2 gene encodes an R2R3 MYB domain protein that acts as a key determinant for proanthocyanidin accumulation in developing seed. Plant Cell 13: 2099–2114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piazza P, Procissi A, Jenkins GI, Tonelli C (2002) Members of the c1/pl1 regulatory gene family mediate the response of maize aleurone and mesocotyl to different light qualities and cytokinins. Plant Physiol 128: 1077–1086 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Procissi A, Dolfini S, Ronchi A, Tonelli C (1997) Light-dependent spatial and temporal expression of pigment regulatory genes in developing maize seeds. Plant Cell 9: 1547–1557 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramsay NA, Walker AR, Mooney M, Gray JC (2003) Two basic-helix-loop-helix genes (MYC-146 and GL3) from Arabidopsis can activate anthocyanin biosynthesis in a white-flowered Matthiola incana mutant. Plant Mol Biol 52: 679–688 [DOI] [PubMed] [Google Scholar]
- Rasmussen SE, Frederiksen H, Krogholm KS, Poulsen L (2005) Dietary proanthocyanidins: occurrence, dietary intake, bioavailability, and protection against cardiovascular disease. Mol Nutr Food Res 49: 159–174 [DOI] [PubMed] [Google Scholar]
- Reay PF (1999) The role of low temperatures in the development of the red blush on apple fruit (“Granny Smith”). Sci Hortic (Amst) 79: 113–119 [Google Scholar]
- Reay PF, Fletcher RH, Thomas VJG (1998) Chlorophylls, carotenoids and anthocyanin concentrations in the skin of “Gala” apples during maturation and the influence of foliar applications of nitrogen and magnesium. J Sci Food Agric 76: 63–71 [Google Scholar]
- Reay RF, Lancaster JE (2001) Accumulation of anthocyanins and quercetin glycosides in “Gala” and “Royal Gala” apple fruit skin with UV-B-visible irradiation: modifying effects of fruit maturity, fruit side, and temperature. Sci Hortic (Amst) 90: 57–68 [Google Scholar]
- Regan BC, Julliot C, Simmen B, Vienot F, Charles-Dominique P, Mollon JD (2001) Fruits, foliage and the evolution of primate colour vision. Philos Trans R Soc Lond B Biol Sci 356: 229–283 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saure MC (1990) External control of anthocyanin formation in apple. Sci Hortic (Amst) 42: 181–218 [Google Scholar]
- Schaefer HM, Schaefer V, Levey DJ (2004) How plant-animal interactions signal new insights in communication. Trends Ecol Evol 19: 577–584 [Google Scholar]
- Schirrmacher G, Schempp H (2003) Antioxidative potential of flavonoid-rich extracts as new quality marker for different apple varieties. J Appl Bot 77: 163–166 [Google Scholar]
- Schneider D, Stern RA, Goldway M (2005) A comparison between semi- and fully compatible apple pollinators grown under suboptimal pollination conditions. HortScience 40: 1280–1282 [Google Scholar]
- Steyn WJ, Holcroft DM, Wand SJE, Jacobs G (2004) Regulation of pear color development in relation to activity of flavonoid enzymes. J Am Soc Hortic Sci 129: 6–12 [Google Scholar]
- Stracke R, Werber M, Weisshaar B (2001) The R2R3-MYB gene family in Arabidopsis thaliana. Curr Opin Plant Biol 4: 447–456 [DOI] [PubMed] [Google Scholar]
- Takos AM, Robinson SP, Walker AR (2006. a) Transcriptional regulation of the flavonoid pathway in the skin of dark-grown “Cripps' Red” apples in response to sunlight. J Hortic Sci Biotechnol 81: 735–744 [Google Scholar]
- Takos AM, Ubi BE, Robinson SP, Walker AR (2006. b) Condensed tannin biosynthesis genes are regulated separately from other flavonoid biosynthesis genes in apple fruit skin. Plant Sci 170: 487–499 [Google Scholar]
- Tohge T, Nishiyama Y, Hirai MY, Yano M, Nakajima J, Awazuhara M, Inoue E, Takahashi H, Goodenowe DB, Kitayama M, et al (2005) Functional genomics by integrated analysis of metabolome and transcriptome of Arabidopsis plants over-expressing an MYB transcription factor. Plant J 42: 218–235 [DOI] [PubMed] [Google Scholar]
- Torregrosa L, Verries C, Tesniere C (2002) Grapevine (Vitis vinifera L.) promoter analysis by biolistic-mediated transient transformation of cell suspensions. Vitis 41: 27–32 [Google Scholar]
- Tsuda T, Yamaguchi M, Honda C, Moriguchi T (2004) Expression of anthocyanin biosynthesis genes in the skin of peach and nectarine fruit. J Am Soc Hortic Sci 129: 857–862 [Google Scholar]
- Tyas JA, Hofman PJ, Underhill SJR, Bell KL (1998) Fruit canopy position and panicle bagging affects yield and quality of “Tai So” lychee. Sci Hortic (Amst) 72: 203–213 [Google Scholar]
- Vanderauwera S, Zimmermann P, Rombauts S, Vandenabeele S, Langebartels C, Gruissem W, Inze D, Van Breusegem F (2005) Genome-wide analysis of hydrogen peroxide-regulated gene expression in Arabidopsis reveals a high light-induced transcriptional cluster involved in anthocyanin biosynthesis. Plant Physiol 139: 806–821 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vanzani P, Rossetto M, Rigo A, Vrhovsek U, Mattivi F, D'Amato E, Scarpa M (2005) Major phytochemicals in apple cultivars: contribution to peroxyl radical trapping efficiency. J Agric Food Chem 53: 3377–3382 [DOI] [PubMed] [Google Scholar]
- Walker AR, Lee E, Bogs J, McDavid DAJ, Thomas MR, Robinson SP (2006) White grapes arose through mutation of two similar and adjacent regulatory genes. Plant J (in press) [DOI] [PubMed]
- Wan C, Wilkins TA (1994) A modified hot borate method significantly enhances the yield of high quality RNA from cotton (Gossypium hirsutum L.). Anal Biochem 223: 7–12 [DOI] [PubMed] [Google Scholar]
- Winkel-Shirley B (2001) Flavonoid biosynthesis: a colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126: 485–493 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamazaki M, Makita Y, Springob K, Saito K (2003) Regulatory mechanisms for anthocyanin biosynthesis in chemotypes of Perilla frutescens var. crispa. Biochem Eng J 14: 191–197 [Google Scholar]
- Zhang F, Gonzalez A, Zhao MZ, Payne CT, Lloyd A (2003) A network of redundant bHLH proteins functions in all TTG1-dependent pathways of Arabidopsis. Development 130: 4859–4869 [DOI] [PubMed] [Google Scholar]
- Zhou Y, Singh BR (2000) Effects of light on the anthocyanin synthesis in cranberry fruits. FASEB J 14: A1416–A1416 [Google Scholar]
- Zimmermann IM, Heim MA, Weisshaar B, Uhrig JF (2004) Comprehensive identification of Arabidopsis thaliana MYB transcription factors interacting with R/B-like BHLH proteins. Plant J 40: 22–34 [DOI] [PubMed] [Google Scholar]