Abstract
Green leaf volatiles (GLVs) are commonly emitted by green plants, and their production is drastically enhanced when they are under biotic stress. To clarify the ecological function of naturally emitted GLVs, we studied the response of Arabidopsis, whose GLV biosynthesis had been modified, when subjected to herbivory or a pathogenic infection. There was a significant increase in GLV production after herbivory by cabbage white butterfly larvae and pathogen (gray mold) infection in hydroperoxide lyase (HPL) sense Arabidopsis compared with WT controls. The HPL sense modification resulted in the plant being more attractive to the parasitic wasp Cotesia glomerata, leading to higher mortality of the herbivores. The HPL sense modification also resulted in greater inhibition of growth of the fungus. By contrast, HPL antisense Arabidopsis produced fewer GLVs, attracted fewer parasitoids, and was more susceptible to the pathogens than the WT control. These data show that (i) one of the ecological functions of GLV biosynthesis related to resistance against both herbivores and pathogens, and (ii) the genetic modification of GLV biosynthesis could be a unique approach for improving plant resistance against such biotic stresses.
Keywords: Arabidopsis, hydroperoxide lyase, tritrophic interactions, Cotesia glomerata, Botrytis cinerea
Green plants emit C6–aldehydes, C6-alcohols, and their acetates, and these so-called green leaf volatiles (GLVs) are biosynthesized via the lipoxygenase/hydroperoxide lyase (HPL) pathway (1). It has been reported that GLVs negatively affect herbivore performance (2, 3) and play an important role in the recruitment of the carnivorous natural enemies of herbivores (4–8). Further, GLVs are known to have antifungal activity toward several microorganisms (9–11). These findings suggest that GLVs have the potential to control both pest insects and plant pathogens.
Intact plants produce only trace amounts of GLVs, whereas greater amounts are rapidly emitted after wounding (12). In Arabidopsis, for example, the production of one GLV, (Z)-3-hexenal, increased ≈30-fold within 5 min of leaves being mechanically damaged (13). It is important to evaluate the defensive functions of GLVs actually emitted by plants when infested by herbivores or infected by pathogens, as previous studies have exposed test plants to synthetic GLVs. Vancanneyt et al. (3) reported that antisense mediated deletion of HPL in transgenic potato (Solanum tuberosum) led to an increase in aphid performance, suggesting that the GLV biosynthetic pathway plays an important role in induced direct defensive responses of plants against herbivores. However, no studies have reported the role of naturally occurring GLVs on recruitment of the herbivores' natural enemies and induced direct defensive response against pathogens.
The focus of this study is twofold: to test whether the increase in naturally emitted GLVs in response to biotic stress is actually multifunctional and to determine whether genetically manipulating the GLV biosynthetic pathway could be a useful approach for improving the resistance of plants against various biotic stresses. To do so, we tested the responses of Arabidopsis plants, whose GLV biosynthesis had been modified, to attack by a herbivore, the larvae of the cabbage white butterfly, Pieris rapae, and a fungal pathogen, the gray mold Botrytis cinerea. We chose these systems as we have shown that Cotesia glomerata, a parasitic wasp that attacks P. rapae larvae, was attracted to at least two GLVs, (E)-2-hexenal and (Z)-3-hexenyl acetate, which were emitted by Arabidopsis in response to herbivore damage (4). Furthermore, we have shown that GLVs induced several defense genes in Arabidopsis, resulting in higher resistance to gray mold (14). Thus, our working hypothesis was that the transgenic manipulation of GLV biosynthesis in Arabidopsis could (i) alter the response of C. glomerata to host-infested plants, and thus modify the incidence of P. rapae larval parasitism, and (ii) affect the susceptibility of the plant to B. cinerea infection.
Results and Discussion
Response of HPL Sense and Antisense Arabidopsis to Damage Made by Herbivory and Pathogen Infection.
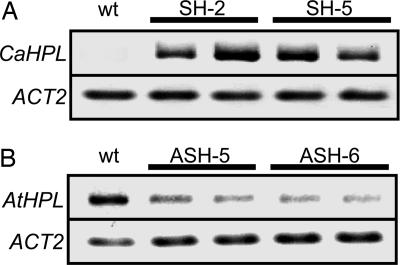
As the lipoxygenase/HPL pathway produces the GLVs, we created two HPL sense Arabidopsis lines (SH-2 and SH-5) in which HPL enzymatic activity was enhanced to approximately twice that of WT plants (Fig. 1A) (P < 0.05, Tukey's test), by the expression of CaHPL (Fig. 2A). We also created two HPL antisense lines (ASH-5 and ASH-6) with HPL enzymatic activity ≈25% of that in controls (Fig. 1A) (P < 0.05, Tukey's test), by the expression of antisense AtHPL that reduced AtHPL expression (Fig. 2B).
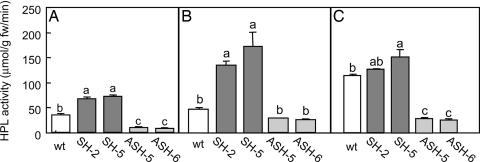
Fig. 1.
Enzymatic activity of HPL in WT, SH, and ASH plants that were either intact (A), infested by P. rapae larvae (B), or infected by B. cinerea (C). See Materials and Methods for details on methods of infestation and infection. Within any given treatment, bars with different letters indicate significant differences between plant types (P < 0.05, Tukey's test).
Fig. 2.
Accumulation of CaHPL mRNA in SH plants (A) and AtHPL mRNA in ASH plants (B) determined by RT-PCR, with ACT2 as the control.
When infested with P. rapae larvae, the HPL enzymatic activity of the two SH lines was significantly enhanced relative to WT plants, whereas ASH lines did not differ from controls (Fig. 1B) (P < 0.05, Tukey's test). HPL activity of WT plants increased about two times after infected by B. cinerea (Fig. 1 A and B). The activity in the infected SH-5 line was significantly higher than in infected WT plants (Fig. 1C) (P < 0.05, Tukey's test), but that of infected SH-2 was not (Fig. 1C). In the infected ASH lines, HPL activity was less than that of infected WT plants (Fig. 1C) (P < 0.05, Tukey's test).
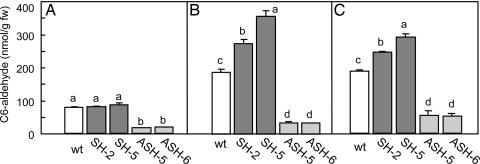
To evaluate GLV biosynthesis in HPL sense and antisense Arabidopsis we measured (Z)-3-hexenal, the first GLV component that is then converted into other GLVs such as (E)-2-hexenal, (Z)-3-hexenol, and (Z)-3-hexenyl acetate within the lipoxygenase/HPL pathway (1, 12, 13). The levels of (Z)-3-hexenal were not significantly different in intact SH lines and WT plants (Fig. 3A) (P > 0.05, Tukey's test), but were significantly lower in ASH lines (Fig. 3A) (P < 0.05, Tukey's test). The lack of difference between SH lines and WT plants was probably caused by them all having similar levels of (Z)-3-hexenal precursors, i.e., linolenic acid, whereas the lower amount in intact ASH plants were probably caused by lower levels of HPL activity. Feeding by P. rapae larvae resulted in significantly higher levels of (Z)-3-hexenal in the two SH lines than the WT plants (P < 0.05, Tukey's test), and they were significantly lower than controls in the two ASH lines (P < 0.05, Tukey's test) (Fig. 3B). The same trends were seen when SH, ASH, and WT plants were infected by B. cinerea (Fig. 3C) (P < 0.05, Tukey's test). These results show that HPL sense and antisense Arabidopsis plants have up-regulated and down-regulated GLV biosynthesis, respectively. There were significant differences between SH-2 and SH-5 in the production of (Z)-3-hexenal, whether in response to herbivory or pathogenic infection. This difference would be caused by separate gene-insertion events.
Fig. 3.
Concentrations of (Z)-3-hexenal in WT, SH, and ASH plants that were either intact (A), infested by P. rapae larvae (B), or infected by B. cinerea (C). See Materials and Methods for details on methods of infestation and infection. Within any given treatment, bars with different letters indicate significant differences between plant types (P < 0.05, Tukey's test).
Response of C. glomerata to HPL Sense and Antisense Arabidopsis Infested by P. rapae Larvae.
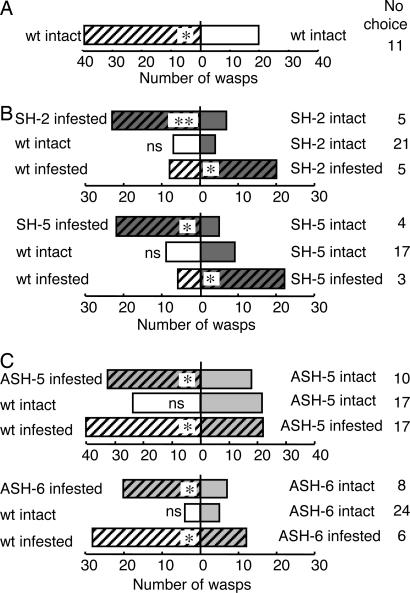
Two SH lines, two ASH lines, and WT plants infested with P. rapae larvae all attracted more C. glomerata females than their intact counterparts (Fig. 4). Furthermore, the wasps preferred infested SH plants over infested WT ones (Fig. 4B). As the levels of damage by P. rapae larvae were the same for all three types of infested plants (≈5% of total leaf area per plant), the higher attractiveness of SH lines would be explained by the higher GLV biosynthetic activity. In contrast, infested ASH plants were significantly less attractive than infested WT plants (Fig. 4C), supporting the hypothesis that the attraction of C. glomerata females to plants was positively associated with GLV biosynthetic activity. The amounts of (E)-2-hexenal, one of the GLVs emitted from P. rapae-infested Arabidopsis that attracts C. glomerata (4), were significantly higher from both P. rapae-infested SH lines than from infected WT plants (data not shown). These increases would, at least in part, explain the increased attractiveness of infested SH lines over infested controls.
Fig. 4.
Preference of C. glomerata females for one of two alternatively treated Arabidopsis plants [WT (A), SH (B), and ASH (C)] in a small cage. Plants were either subjected to P. rapae larval infestation for 2 h (with stripes) or not (without stripes). WT plants in B and C were used as the control. ∗∗, 0.001 < P < 0.01; ∗, 0.01 < P < 0.05; ns, not significantly different (binomial test).
To test whether the increased attractiveness of SH plants to C. glomerata resulted in a significantly higher incidence of parasitism in P. rapae larvae, we conducted an experiment in a climate-controlled room with SH-2 and WT plants. The experiment was replicated five times, and significantly more parasitized P. rapae larvae were found on SH-2 than WT plants (Fig. 5) (paired t test, P = 0.0006). These results suggest that naturally emitted GLVs play an important role in attracting C. glomerata, and that enhancing GLV biosynthesis is one means of ameliorating the attractiveness of plants to the carnivorous natural enemies of herbivores.
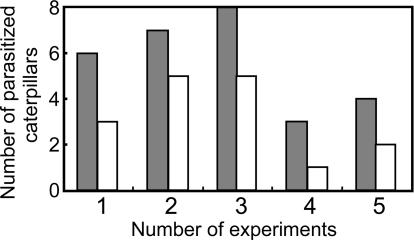
Fig. 5.
Incidence of parasitism of P. rapae larvae by C. glomerata on WT (empty bars) and SH-2 (filled bars) plants. A paired t test was used to compare the numbers of parasitized larvae on the two plant groups.
Response of B. cinerea to HPL Sense and Antisense Arabidopsis.
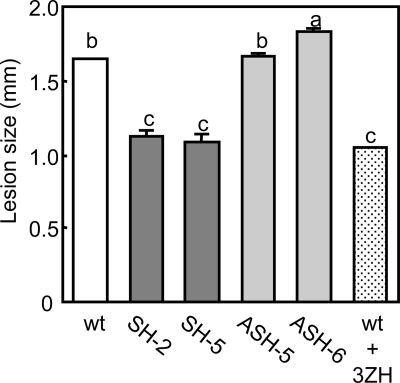
To test whether naturally emitted GLVs could make the plants more resistance to fungal disease, we inoculated two SH lines, two ASH lines, and WT plants with B. cinerea. After 4 days the mean diameter of the necrotic area was significantly smaller on the leaves of SH lines than WT plants (P < 0.05, Tukey's test), whereas on ASH-6 leaves they were significantly larger than those of the controls (P < 0.05, Tukey's test) (Fig. 6). However, there was no difference in lesion diameter between ASH-5 and control leaves (Fig. 6). We believe that the reduced susceptibility in HPL sense Arabidopsis can be explained, in part, by the increased concentrations of GLVs emitted by plants upon infection. This idea was supported by the fact that when (Z)-3-hexenal was added to infected WT leaves, the diameter of lesions were significantly smaller than on untreated WT leaves and similar to those on infected SH-2 and SH-5 ones (Fig. 6) (P < 0.05, Tukey's test). The actual mechanism/mode of action of GLVs is unknown, but may be involved with the reinforcement of plant cell walls and/or the accumulation of antimicrobial proteins and phytoalexins in a plant (15). (E)-2-hexenal, (E)-2-hexenol, and (Z)-3-hexenol have been shown to up-regulate defense-related genes in Arabidopsis, lima beans, and citrus (16–18) and affect phytoalexin accumulation in cotton plants (Gossypium hirsutum L.) (19) and Arabidopsis (15). Furthermore, GLVs themselves have antifungal properties, which would directly delay fungal growth (20).
Fig. 6.
Comparison of the diameters of necrotic lesions caused by B. cinerea on (Z)-3-hexenal-treated and control WT plants and on SH and ASH plants. Different letters indicate significant differences between treatments (P < 0.05, Tukey's test). 3ZH, (Z)-3-hexenal.
GLVs are commonly found in small amounts in the headspace of intact green plants but increase dramatically when plants are under biotic stress. In this study, using genetically modified Arabidopsis we were able to show how such natural changes in GLVs play a role in plant resistance to herbivores through the attraction of natural enemies and against fungi through the suppression of their growth. To date, the majority of work on genetically modified crops for pest management has focused on the introduction of toxic compounds in plants (21); however, our findings suggest an alternative direction that would involve genetic transformations to increase plants' natural defense systems through the enhanced production of GLVs. However, it is clear that the efficacy of such an approach will have to be tested on a larger spatial scale under natural conditions.
Materials and Methods
Preparation of Transgenic Plants.
To enhance HPL enzymatic activity, full-length bell pepper (Capsicum annuum L.) HPL cDNA (22) was inserted into a pCGN5138 vector, along with the enhanced promoter of the cauliflower mosaic virus 35S (CaMV35S) ribosomal RNA promoter (p-e35S), the leader sequence of HSP70, and the nopaline synthase 3′ terminator sequences (23). The pCGN5138 vector contained the nptII gene between the CaMV35S promoter and tml 3′ terminator sequences. These constructs were transferred to Agrobacterium tumefaciens LBA4404. Transgenic Arabidopsis plants (ecotype No-0) were created by using the vacuum infiltration method (24). Homozygotes of the transgenic lines were selected, based on kanamycin resistance in the T2 generation, and further confirmation of the genotype was obtained by PCR amplification of the transgene. This approach gave us two lines (SH-2 and SH-5) from transformed bell pepper HPL cDNA. Using the same protocol, we produced transgenic plants expressing antisense AtHPL by inserting full-length AtHPL cDNA (25) into a pCGN5138 vector in antisense orientation. We obtained two lines (ASH-5 and ASH-6) with transformed AtHPL cDNA in antisense orientation and reduced HPL enzymatic activity. T3 to T6 generations of transgenic and WT Arabidopsis plants were grown in plastic pots (five plants per pot) in a climate-controlled room (24 ± 2°C, 16-h:8-h light/dark photoperiod) for 4–5 weeks before being used in the experiments.
RT-PCR.
Total RNA was extracted from Arabidopsis leaves (≈200 mg fresh weight) with Trizol Reagent (Invitrogen, Carlsbad, CA); remaining DNA was degraded by treating it with a DNA-free kit (Ambion, Austin, CA). cDNA was synthesized from the total RNA (4 μg) by a Thermoscript RT-PCR system (Invitrogen), with oligo(dT)20 as a primer, according to the manufacturer's instructions. RT-PCR was performed by using Takara ExTaq polymerase (Takara Biomedicals, Otsu, Japan). The reaction mixture contained 0.1 μmg of cDNA, 0.5 μmM each of forward and reverse primers, 1× ExTaq buffer, 250 nM dNTPs, and 0.5 units of ExTaq polymerase in a 20-μl solution.
Primers for CaHPL were: 5′-CATTTTCTTGCCAACCCTTCT-3′ (forward primer) and 5′-AGCAACCTTTAGTACCTACCA-3′ (reverse primer).
Primers for AtHPL were: 5′-AACCCTAACATCGTCGCCGTTCT-3′ (forward primer) and 5′-ATTTTGAATAGCCTCATCTCGGG-3′ (reverse primer).
Primers for ACT2 (At3g18780) were the same as those used previously (12). The PCR was carried out on a Perkin-Elmer 9700 (Applied Biosystems, Foster City, CA) at 94°C for 2 min followed by 23–24 cycles of 94°C for 45 s, 57°C for 45 s, and 72°C for 45 s, followed by a final 72°C extension for 8 min.
Infestation with P. rapae Larvae.
Two third-stadium P. rapae larvae were placed on each plant (five plants per pot) and allowed to feed for 2 h before the measurement of GLV production or to test the flight response of parasitic wasps.
Inoculation of Plants with B. cinerea.
To measure resistance to B. cinerea [strain IuRy-1 (14)], a conidial suspension (3 μl; 3 × 104 cfu·ml−1 in 2.5% glucose) was placed on the center of Arabidopsis rosette leaves. The inoculated leaves were incubated at 22°C under a 16-hr light/8-hr dark photoperiod for 4 days, and then the diameter of the necrotic lesion was measured. To confirm the effect of GLVs on resistance to B. cinerea, WT plants were exposed to (Z)-3-hexenal, and 24 h later, a conidial suspension was placed on the center of the rosette leaves. These plants were incubated for 4 days, when the diameters of the necrotic lesions were measured and compared with those on control infected WT plants.
Enzyme Assay.
The 13-HPL activity in leaves was determined by using 100 μM of 13-HPO linoleic acid in 100 mM sodium phosphate (pH 6.3), using a modified version of a method described (21). For herbivory, each plant (five plants per pot) was infested with two third-stadium P. rapae larvae for 2 h. For pathogenic infections, the leaves of Arabidopsis plants were spotted with 5 μl of B. cinerea conidial suspension in 2.5% glucose (1 × 105 cfu·ml−1) and incubated in a growth chamber (22°C; 16 h light:8 h dark) for 12 h.
Crude enzyme solutions, prepared from Arabidopsis leaves, were reacted with 13-hydroperoxide of linoleic acid for 1 min at 25°C. The resultant aldehydes were quantified by HPLC after converting them to 2,4-dinitrophenylhydrazones (22).
Analysis of (Z)-3-Hexenal.
Arabidopsis leaves (0.5 g) were soaked in 5 ml of 80% methanol, containing 5 nmol of n-heptanal, and incubated for 1 h at 60°C. The solution was then mixed with 2.5 ml of 0.1% 2,4-dinitrophenylhydrazine (in ethanol containing 2% acetic acid) and incubated at 25°C for 1 h. Hydrazones were extracted with 2.5 ml of n-hexane, reconstituted with 50 μl of acetonitrile, and analyzed by HPLC (22).
Two-Choice Tests.
C. glomerata wasps were obtained from parasitized host larvae collected in the field. The flight responses of female wasps, at 25 ± 2°C, were observed in a cage (25 × 35 × 30 cm) with three windows (covered with nylon gauze) and one door. Five wasps were released at the midpoint between two groups of potted Arabidopsis plants (five plants per pot) and observed for 30 min. The transgenic and WT plants used in this study were matched for both shape and color. The first plant on which each wasp landed was recorded as its choice. If the wasp did not land on any plant during the assay, it was classified as a no-choice response. All herbivores and herbivore products were removed from the plants before bioassay, and the wasps used were mated but naïve with respect to oviposition.
Incidence of Parasitism of P. rapae Larvae on Transgenic Plants.
Five SH-2 and five WT plants were each infested with 10 second-stadium P. rapae larvae for 1 day. The plant groups were then placed 2 m apart in a climate-controlled chamber (3.2 × 2.7 × 2.2 m; 25 ± 2°C), and 10 naïve C. glomerata females were released at the center of the room. After 6 h the larvae were collected, and the incidence of parasitism was determined by keeping them until either the wasp larvae egressed or the caterpillar molted to form a normal crysalid.
Statistics.
Tukey's HSD (honestly significant difference) test was used to determine significant differences among plant types in experiments for HPL enzymatic activity (n = 6–9 replicates), the concentration of (Z)-3-hexenal (n = 6–9), and the diameter of necrotic lesions (n = 3). An alpha level of 0.05 was used for statistical significance.
Two-choice tests were analyzed by using a binomial probability function, with the null hypothesis that wasps would have a 1:1 distribution on the two groups of plants. Wasps that did not make a choice (data in Fig. 4) were excluded from the analyses. Paired t tests were used to compare the numbers of parasitized larvae on WT and transgenic plants (n = 5).
Acknowledgments
We thank J. McNeil for his critical reading of the manuscript. This study was supported by Core Research for Evolutional Science and Technology of the Japan Science and Technology Agency.
Abbreviations
- GLV
green leaf volatile
- HPL
hydroperoxide lyase
Footnotes
The authors declare no conflict of interest.
References
- 1.Matsui K. Curr Opin Plant Biol. 2006;9:274–280. doi: 10.1016/j.pbi.2006.03.002. [DOI] [PubMed] [Google Scholar]
- 2.Zhang QH, Schlyer R, Anderson P. J Chem Ecol. 1999;25:2847–2861. [Google Scholar]
- 3.Vancanneyt G, Sanz C, Farmaki T, Paneque M, Ortego F, Castanera P, Sanchez-Serrano JJ. Proc Natl Acad Sci USA. 2001;98:8139–8144. doi: 10.1073/pnas.141079498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Shiojiri K, Ozawa R, Matsui K, Kishimoto K, Kugimiya S, Takabayashi J. J Chem Ecol. 2006;32:969–979. doi: 10.1007/s10886-006-9047-6. [DOI] [PubMed] [Google Scholar]
- 5.Takabayashi J, Noda T, Takahashi S. Appl Entomol Zool. 1991;26:237–243. [Google Scholar]
- 6.Whitman DW, Eller FJ. J Chem Ecol. 1992;18:1743–1753. doi: 10.1007/BF02751099. [DOI] [PubMed] [Google Scholar]
- 7.Brikett MA, Chamberlain K, Guerrieri E, Pickett JA, Wadhams LJ, Yashuda T. J Chem Ecol. 2003;29:1589–1600. doi: 10.1023/a:1024218729423. [DOI] [PubMed] [Google Scholar]
- 8.Shimoda T, Ozawa R, Sano K, Yano E, Takabayashi J. J Chem Ecol. 2005;31:2019–2032. doi: 10.1007/s10886-005-6075-6. [DOI] [PubMed] [Google Scholar]
- 9.Croft KPC, Juttner F, Slusarenko A. Plant Physiol. 1993;101:13–24. doi: 10.1104/pp.101.1.13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kubo I, Fujita K. J Agric Food Chem. 2001;49:5750–5754. doi: 10.1021/jf010728e. [DOI] [PubMed] [Google Scholar]
- 11.Nakamura S, Hatanaka A. J Agric Food Chem. 2002;50:7639–7644. doi: 10.1021/jf025808c. [DOI] [PubMed] [Google Scholar]
- 12.Hatanaka A. Phytochemistry. 1993;34:1201–1218. [Google Scholar]
- 13.Matsui K, Kurishita S, Hisamitsu A, Kajiwara T. Biochem Soc Trans. 2000;28:857–860. [PubMed] [Google Scholar]
- 14.Kishimoto K, Matsui K, Ozawa R, Takabayashi J. Plant Cell Physiol. 2005;46:1093–1102. doi: 10.1093/pcp/pci122. [DOI] [PubMed] [Google Scholar]
- 15.Kishimoto K, Matsui K, Ozawa R, Takabayashi J. Plant Sci. 2006;170:715–723. doi: 10.1016/j.plantsci.2006.05.004. [DOI] [PubMed] [Google Scholar]
- 16.Bate NJ, Rothstein SJ. Plant J. 1998;16:561–569. doi: 10.1046/j.1365-313x.1998.00324.x. [DOI] [PubMed] [Google Scholar]
- 17.Arimura G, Ozawa R, Horiuchi J, Nishioka T, Takabayashi J. Biochem Syst Ecol. 2001;29:1049–1061. [Google Scholar]
- 18.Gomi K, Yamasaki Y, Yamamoto H, Akimitsu K. J Plant Physiol. 2003;160:1219–1231. doi: 10.1078/0176-1617-01177. [DOI] [PubMed] [Google Scholar]
- 19.Zeringue HJ., Jr Phytochemistry. 1992;31:2305–2308. [Google Scholar]
- 20.Matsui K, Minami A, Hornung E, Shibata H, Kishimoto K, Ahnert V, Kindl H, Kajiwara T, Feussner I. Phytochemistry. 2006;67:649–657. doi: 10.1016/j.phytochem.2006.01.006. [DOI] [PubMed] [Google Scholar]
- 21.High SM, Cohen MB, Shu QY, Altossar I. Trends Plants Sci. 2004;9:286–292. doi: 10.1016/j.tplants.2004.04.002. [DOI] [PubMed] [Google Scholar]
- 22.Matsui K, Shibutani M, Hase T, Kajiwara T. FEBS Lett. 1996;394:21–24. doi: 10.1016/0014-5793(96)00924-6. [DOI] [PubMed] [Google Scholar]
- 23.McBirde KE, Summerfelt KR. Plant Mol Biol. 1990;14:269–276. doi: 10.1007/BF00018567. [DOI] [PubMed] [Google Scholar]
- 24.Bechtold N, Pelletier G. Methods Mol Biol. 1998;82:259–266. doi: 10.1385/0-89603-391-0:259. [DOI] [PubMed] [Google Scholar]
- 25.Matsui K, Wilkinson J, Hiatt B, Knauf V, Kajiwara T. Plant Cell Physiol. 1999;40:477–481. doi: 10.1093/oxfordjournals.pcp.a029567. [DOI] [PubMed] [Google Scholar]