Abstract
The ORF29 gene of varicella-zoster virus encodes a single-stranded DNA binding protein that is predominantly nuclear during lytic infection but appears to be restricted to the cytoplasm of latently infected neurons. Following reactivation, ORF29p accumulates in the nuclei of neurons, suggesting that its confinement to the cytosol may be critical for maintaining quiescence. When autonomously expressed, ORF29p accumulates in the nuclei of fibroblasts and the cytoplasm of cells (guinea pig enteric neurons) and cell lines (U373MG) of neuronal origin. Inhibition of the 26S proteasome redirects the accumulation of ORF29p to the nucleus in cells of neuronal origin. Here, we show that ORF29p is ubiquitinated and sumoylated in 293T cells and subsequently degraded from the N terminus. Ubiquitinated ORF29p accumulates in both the nuclei and the cytoplasm of fibroblasts, but degradation products are seen primarily in the cytoplasm. Modification and degradation of ORF29p occurs in 293T, U373MG, and MeWo cells. Therefore, these processes are ubiquitous; however, the robustness of the degradation process is cell type specific. The proteasome-mediated mechanism of nuclear exclusion in U373MG cells is an active process that is not specific for the endogenous ORF29p nuclear localization signal but can be saturated by protein stabilization or overexpression, which leads to nuclear accumulation of ORF29p. The evidence for ORF29p ubiquitination and previous data regarding the effect of proteasome inhibitors on the abundance and distribution of ORF29p implicate the 26S proteasome in influencing the protein's cell type-specific localization.
Primary infection of cutaneous epithelial cells by the human alphaherpesvirus varicella-zoster virus (VZV) results in the chicken pox rash (varicella). Following this lytic infection, VZV establishes latency in the sensory ganglia and can reactivate later in the host's life to cause shingles (zoster) (1). During latency, viral DNA replication, late gene expression, and virion assembly do not occur. It has been reported that VZV proteins encoded by immediate-early and early open reading frames (ORFs) 4, 21, 29, 62, 63, and 66 are expressed during latency and that their subcellular distribution correlates with whether the virus is undergoing latent or lytic infection (3-5, 10, 15, 21). These proteins are detained in the cytoplasm during latency until the block to productive infection is relieved, after which they accumulate in the nucleus. Thus, the confinement of ORF29p and other VZV latency-associated proteins (LAPs) in the cytosol may be a hallmark of latency and play a role in maintaining quiescence. However, while the LAPs appear to be cytoplasmic during latency, the possibility that these proteins accumulate below the levels of detection in the nucleus or shuttle between the nucleus and cytoplasm cannot be ruled out. An understanding of the subcellular distribution of the VZV LAPs during latency and reactivation is important for identifying the cell- and virus-specified proteins that govern VZV infection.
The in vivo cell type dependency of ORF29p localization can be recapitulated in vitro. In fibroblasts infected with an adenovirus vector expressing ORF29p (AdORF29), the protein localizes primarily to the nucleus with diffuse cytoplasmic staining. In contrast, in astrocytoma-derived U373MG cells and cultured guinea pig enteric ganglia (EG) infected with AdORF29, ORF29p is cytoplasmic (33). Previous investigations by our laboratory revealed a role for the 26S proteasome in the cellular localization of ORF29p. Inhibition of the proteasome with MG132 led to the accumulation of autonomously expressed ORF29p in the nuclei of U373MG cells and EG, where it is normally detected only in the cytoplasm. Reversal of MG132 resulted in the disappearance of ORF29p from the nucleus and its reappearance in the cytoplasm. If proteasome activity was restored in the presence of cycloheximide, the signal corresponding to ORF29p disappeared (33). This demonstrated that nuclear translocation of ORF29p does not guarantee its survival, as the protein's distribution and stability were reset when inhibition of proteasome activity was reversed. The disappearance of ORF29p from the nucleus upon the removal of the drug also suggested that ORF29p is either destroyed in the nucleus or exported and rapidly degraded (33).
Pulse-chase studies show that the half-life of ORF29p is shorter in U373MG cells, where it is excluded from the nucleus, than in MeWo cells, where nuclear accumulation of the protein occurs. Thus, depending on its cellular localization, ORF29p is differentially targeted for degradation (33). Moreover, the increase in the stability of ORF29p following the inhibition of the proteasome correlated with its accumulation in the nucleus, thus further supporting a role for this pathway in the subcellular localization of this protein. The same studies also revealed the presence of slower- and more-rapidly migrating immunoreactive species at early times in U373MG cells and at later times in MeWo cells. These aberrantly migrating species might represent modified or degraded forms of ORF29p. Of note, the alternative species were less abundant in samples treated with the proteasome inhibitor MG132.
A protein destined for destruction is often marked by the covalent attachment of multiple ubiquitin moieties before it is escorted to the proteasome for rapid hydrolysis. In addition to ubiquitination, regulating the targeting of some proteins to the 26S proteasome involves the covalent attachment of other small, ubiquitin-like molecules to substrates. One ubiquitin-like molecule, SUMO, is a small moiety that posttranslationally forms an isopeptide bond with an internal lysine of a substrate, just as ubiquitin does. SUMO may regulate its substrates by competing with ubiquitin and precluding substrate degradation or by controlling the subcellular localization of proteins (9).
In the present study, we investigated the posttranslational modification of ORF29p to attempt to unravel the basis for the subcellular targeting of this protein. Biochemical analyses reveal that ORF29p is ubiquitinated and sumoylated. Whereas ubiquitinated ORF29p accumulates in both the nucleus and the cytoplasm of fibroblasts, degradation products appear primarily in the cytoplasm. ORF29p modification and degradation occur in 293T, U373MG, and MeWo cells; thus, these processes are ubiquitous. However, the fate and subsequent localization of ORF29p are cell type specific and we therefore believe that it is the robustness of the degradation process that dictates localization. The proteasome-dependent mechanism involved in the nuclear exclusion of ORF29p in U373MG cells is an active process that can be saturated by protein overexpression, which leads to nuclear accumulation of ORF29p. The evidence for ORF29p ubiquitination and degradation implicates the proteasome as one of the determinants of the protein's cell type-specific localization. The differential compartmentalization of ORF29p and the other LAPs correlates with whether VZV infection is lytic or latent, suggesting that the pathways involved in the distribution of these proteins may also contribute to the maintenance of latency.
MATERIALS AND METHODS
Mammalian cells.
Human 293T fibroblast cells, astrocytoma-derived U373MG cells, and human melanoma (MeWo) cells were maintained as monolayer cultures as previously described (33, 34). Twenty-four hours prior to infection or transfection, the cells were seeded onto coverslips in six-well tissue culture dishes for fluorescence microscopy assays or 100-mm dishes for all other experiments. During virus infection, MeWo and U373MG cells were maintained in Dulbecco's modified Eagle's medium (GIBCO-BRL, Grand Island, NY) supplemented with 2% fetal bovine serum, 100 U/ml penicillin, and 100 μg/ml streptomycin.
Virus.
Jones VZV, a wild-type clinical isolate, was propagated in MeWo cell monolayers by serial passage of infected cells onto uninfected cells as previously described (11). Cell-free Jones VZV was obtained as previously described (34), and titers were determined by plaque assay on MeWo cells. Cell-free virus stocks were stored in 0.5-ml aliquots at −80°C. Adenovirus AdORF29 expresses ORF29p from a mouse cytomegalovirus (mCMV) promoter (33).
Transfections.
All transfections were performed using Lipofectamine PLUS in Opti-MEM media (Invitrogen, Carlsbad, CA). Forty-eight hours following transfection, cells were processed for analysis by fluorescence microscopy.
Antibodies.
Rabbit polyclonal antibodies against amino acids 1086 to 1201 of ORF29p were previously described (21). Mouse monoclonal antibody to FLAG M2 and rabbit polyclonal antibodies to human c-Jun were purchased from Stratagene (La Jolla, CA). Mouse monoclonal antibody to bovine erythrocyte ubiquitin was purchased from Calbiochem (San Diego, CA). Mouse monoclonal antibody to α-tubulin was obtained from Sigma (St. Louis, MO). Mouse monoclonal antibody to bromodeoxyuridine (BrdU) was purchased from Roche (Indianapolis, IN). Mouse monoclonal antibody to β-galactosidase (β-Gal) was obtained from Promega (Madison, WI). Alexa Fluor 488 conjugated goat anti-mouse and Alexa Fluor 546 goat anti-rabbit antibodies were purchased from Molecular Probes (Carlsbad, CA). Goat anti-rabbit and anti-mouse antibodies conjugated to horseradish peroxidase for immunoblotting were purchased from KPL (Gaithersburg, MD).
Drug treatment.
Cells were treated with 20 μM MG132 (EMD Biosciences, La Jolla, CA) from a 10 mM stock in dimethyl sulfoxide (DMSO; Sigma) and 50 μg/ml cycloheximide (Sigma) from a 10-mg/ml stock in water.
Indirect immunofluorescence (IF) microscopy.
Cells on glass coverslips were prepared and processed as previously described (34) except for the following modifications made for BrdU detection. Cells analyzed for BrdU incorporation during heterokaryon assays were permeabilized by incubation in 0.2% Triton X-100 in phosphate-buffered saline (PBS) for 12 min. After several washes with PBS, the cells were treated for 10 min at room temperature with 4 N HCl to expose the BrdU residues for staining. The acid was removed, and the cells were washed quickly several times in PBS, followed by two 10-min washes in PBS on a shaking platform.
All samples were visualized with a Zeiss Axiovert 200 M inverted microscope, and images were acquired with a Zeiss Axiocam (Carl Zeiss Microimaging Inc., Thornwood, NY) using Openlab 4.1 software (Improvision, Lexington, MA). Images were merged using OpenLab software, assembled with Photoshop, and labeled in Illustrator (CS Adobe Systems Inc., San Jose, CA).
Plasmids and cloning.
p29-12 (34) and pDC516-29 (33) express ORF29p under the chicken actin promoter and the mCMV promoter, respectively, and were previously described. pFLAGORF29 was constructed by digesting p29-12 with EcoRI and NotI and cloning the ORF29 fragment into the same restriction sites of pCF2HN. pCF2HN expresses transgenes with an N-terminal FLAG tag under a human CMV (hCMV) promoter and was derived from pCF2H (30). p29SV40NLS was constructed by PCR amplification with oligonucleotides ORF29ES5′PRM.ECO (5′-GGGAATTCGATGGAAAATACTCAGAAGACTG-3′) (GIBCO-BRL) and 3′SV40NLS (5′-GGGCGGCCGCTTATACTTTTCGCTTCTTCTTAGGCATTTCCATTGTAATGTTCCCATG-3′) (Proligo LLC, Boulder, CO), using p29-12 as a template. The primer 3′SV40NLS contains the coding sequence for the simian virus 40 (SV40) T-antigen nuclear localization signal (NLS) (PKKKRKV) followed by a stop codon. The amplification product was digested with EcoRI and NotI and cloned into the pTriEx-1 vector (Novagen, Madison, WI). pHM829 was provided by Christian Shindler's laboratory at Columbia University and expresses β-Gal and green fluorescent proteins (32). To express a β-Gal SV40 NLS fusion protein, pβgalSV40NLS was constructed by inserting the linker formed by the annealing of oligonucleotides sv40nlsstop (5′-CTAGCCCTAAGAAGAAGCGAA AAGTATAAC-3′) and sv40nlsbot (5′-TCGAGTTATACTTTTCGCTTCTTCTTAGGG-3′) into linearized pHM829 at the Nhe1 and XhoI sites. pCMV-Flag-53 and pHis6-HA-SUMO were provided by Jan-Philipp Kruse and the Wei Gu laboratory at Columbia University, and pCMV-HA-UB was kindly contributed by Atish Choudhury and the Richard Baer laboratory at Columbia University (26).
Metabolic labeling.
Cell proteins were radiolabeled as previously described (33) and the cells were then washed three times with PBS before harvesting.
Cell fractionation.
Cytoplasmic and nuclear fractions were prepared as previously described (19) and then subjected to immunoprecipitation with anti-hemagglutinin (HA) affinity matrix (Roche).
Immunoprecipitation.
Cells were harvested and immunoprecipitated as previously described (33). Proteins immunoprecipitated with ORF29p antiserum conjugated to GammaBind Plus Sepharose beads were released from beads by boiling in 50 ml PBS and 10 ml 6× sodium dodecyl sulfate (SDS) sample buffer (300 mM Tris-HCl [pH 6.8], 12% SDS, 0.6% bromophenol blue, 60% glycerol, 600 mM β-mercaptoethanol) for 10 min. The released protein was then subjected to SDS-polyacrylamide gel electrophoresis (PAGE). Tagged proteins purified by binding to anti-FLAG and anti-HA beads were eluted overnight at 4°C with 50 ml of 150 mg/ml FLAG peptide (Sigma) in radioimmunoprecipitation assay (RIPA) buffer or 500 mg/ml HA peptide (Roche) in RIPA buffer, respectively. Ten milliliters of 6× SDS sample buffer was added to the eluates before they were boiled for 10 min and subjected to SDS-PAGE.
Preparation of whole cell lysates.
Infected MeWo and U373MG cells were harvested for protein preparation and Western blot analysis as described above, except the cells were resuspended in only 100 μl of RIPA buffer.
Western blot analysis.
Proteins were transferred from the acrylamide gels to nitrocellulose membranes (Schleicher and Schuell, Keene, NH) with a Bio-Rad Trans-Blot Semi-Dry apparatus and analyzed by Western blotting as previously described (33, 34).
Coomassie blue staining of proteins.
Twenty-five microliters of each eluate was subjected to SDS-PAGE on a NuPAGE 4 to 12% bis-Tris gel (Invitrogen) in 1× MES (morpholineethanesulfonic acid) running buffer (Invitrogen). The gel was rinsed twice for 5 min each in water, fixed in a 50% methanol and 7% acetic acid solution for 15 min, and washed twice more in water. Twenty milliliters of GelCode blue stain reagent (Pierce, Rockford, IL) was added to the gel and incubated overnight at room temperature. The stained gel was washed several times with water before the bands were excised for matrix-assisted laser desorption (MALDI) mass spectrophotometry analysis.
Northern blot analysis.
Total RNA was isolated from transfected 293T cells by using the TRIzol reagent (Invitrogen) and analyzed for ORF29 and GAPDH RNA as previously described (33).
Heterokaryon assays.
MeWo or U373MG cells were infected with AdORF29 at a multiplicity of infection (MOI) of 50. At 48 hours postinfection, 5.0 × 105 infected cells were mixed with 1.0 × 106 uninfected cells of the other cell type, which had been incubated for 24 h in normal growth media supplemented with 50 μM BrdU (Sigma) and washed repeatedly in PBS before they were mixed with the infected cells. The BrdU-labeled DNA of the uninfected cells allowed for identification of these nuclei in heterokaryons. The mixed cultures were allowed to adhere to glass coverslips in six-well culture dishes for 12 h. The medium was removed, and the cells were incubated for 90 s in either PBS or 50% polyethylene glycol (PEG) with a molecular weight of 1,450 (Sigma) in PBS at 37°C. The cells were then washed four times in PBS and incubated for 12 h in normal growth media before ORF29p localization and BrdU incorporation were visualized by indirect IF microscopy.
RESULTS
Expression profile of ORF29p in MeWo and U373MG cells.
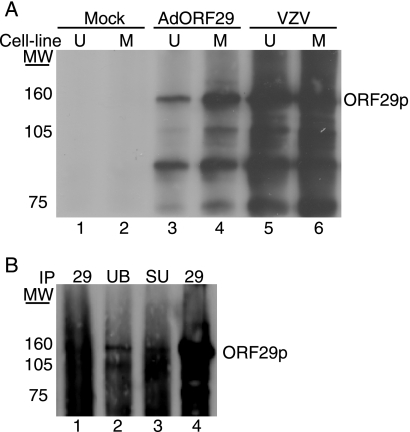
Inhibition of the proteasome with MG132 or epoxomicin results in the redistribution of ORF29p from the cytoplasm to the nucleus in U373MG cells infected with AdORF29 or in guinea pig enteric neurons latently infected with VZV (33). This proteasome-dependent relocalization hints at ubiquitination as a modification that might explain the differences in the cellular partitioning of ORF29p. To determine whether alternative species of ORF29p accumulated when the protein was localized to the cytoplasm in comparison to the nucleus, the SDS-PAGE profile of ORF29p expressed during VZV infection of MeWo and U373MG cells was compared with that of the protein generated during AdORF29 infection of these cell lines. Infected cell proteins were metabolically labeled prior to harvesting and lysing them in RIPA buffer. The cell lysates were immunoprecipitated with an ORF29p-specific antiserum, and the bound proteins were subjected to SDS-PAGE. The ORF29p that accumulates predominantly in the nuclei of MeWo cells and that which accumulates in the cytoplasm of U373MG cells migrate with the same mobility (Fig. 1A, lanes 3 and 4). Note that there is less ORF29p in U373MG cells where the protein is unstable (34). These proteins also comigrate with ORF29p expressed during lytic VZV infection of MeWo and U373MG cells (Fig. 1A, lanes 5 and 6). These data, however, do not rule out cell type-specific protein modifications that may not be detectable by SDS-PAGE.
FIG. 1.
Analysis of ORF29p electrophoretic mobility. (A) MeWo (M) (lanes 2, 4, and 6) and U373MG (U) (lanes 1, 3, and 5) cells were either infected with cell-free Jones VZV at an MOI of 1 (lanes 5 and 6), infected with AdORF29 at an MOI of 50 (lanes 3 and 4), or mock infected. At 3 days postinfection, cells were labeled with 500 μCi/ml Tran35S-label for 24 h, harvested, and lysed in RIPA buffer. ORF29p was immunoprecipitated from the labeled cell lysates by using an ORF29p-specific antiserum, and bound proteins were subjected to SDS-PAGE analysis on a 5% gel. Proteins were visualized by autoradiography. (B) MeWo cells were either mock infected (lane 1) or infected with cell-free VZV at an MOI of 1 (lanes 2 to 4). At 5 days postinfection, the cells were harvested, lysed in RIPA buffer, and immunoprecipitated (IP) with ubiquitin-specific antibody (UB) (lane 2), SUMO-specific antibody (SU) (lane 3), or ORF29p-specific antiserum (lanes 1 and 4) (29). The bound proteins were subjected to SDS-PAGE and analyzed by Western blotting with antiserum specific for ORF29p. MW, molecular weights in thousands.
ORF29p migrates as at least two distinct species.
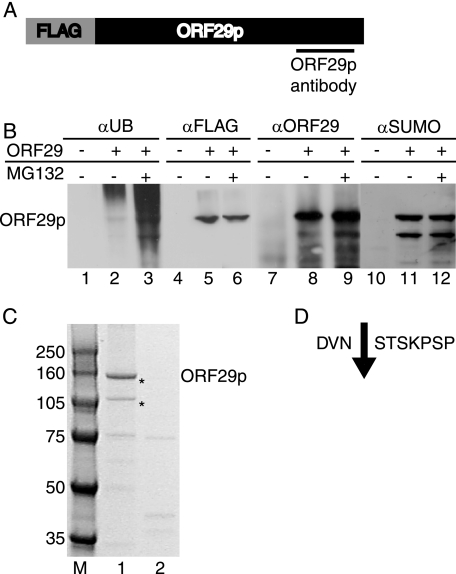
MeWo and U373MG cells infected with either VZV or AdORF29 accumulate additional protein species that immunoprecipitated with the ORF29p-specific antiserum (Fig. 1A, lanes 3 to 6). These polypeptides migrated to the same position as those observed during pulse chase experiments with ORF29p (33). To determine whether the smaller peptides represented specific truncations of ORF29p, a FLAG-tagged ORF29p expression construct (pFLAGORF29) was prepared and transfected into 293T cells (Fig. 2A). Transfected 293T cells were either mock treated or treated with 20 μM MG132 and lysed, and the cell lysates were reacted with an anti-FLAG M2 agarose matrix. The bound material was eluted with FLAG peptide and subjected to Western blot analyses. Antiserum specific to the C terminus of ORF29p reacted with peptides of the same molecular weights as those observed during labeling experiments (Fig. 1 and 2B, lanes 8 to 9). The FLAG-specific antiserum recognized only the band corresponding to full-length ORF29p (Fig. 2B, lanes 5 to 6). Thus, the N terminus was absent from the faster-migrating species. The observation that peptides lacking the N-terminal FLAG tag copurified with those bound by the FLAG affinity matrix demonstrates that the more-rapidly migrating ORF29p species interacts with the full-length FLAG-tagged ORF29p molecule and provides evidence that ORF29p can exist as a multimer. We cannot rule out the possibility that the truncated version of ORF29p is derived not from degradation but from a specific cleavage event that occurs to one of the peptide chains when they are multimerized. This would also result in a failure to detect the N terminus in our SDS-PAGE analysis. Ubiquitin-specific antiserum also reacted with ORF29p expressed in VZV and AdORF29-infected MeWo and U373MG cells (Fig. 1A and B).
FIG. 2.
Analysis of two species of ORF29p. (A) The N-terminal, FLAG-tagged ORF29 expression cassette from pFLAGORF29 is diagrammed (not to scale), with the region recognized by the ORF29p-specific antiserum underlined. 293T cells were transiently transfected with either pFLAGORF29 (B, lanes 2, 3, 5, 6, 8, 9, 11, and 12; C, lane 1) or empty pCF2HN (B, lanes 1, 4, 7, and 10; C, lane 2). At 48 h posttransfection, cells were either treated with DMSO (B, lanes 1, 2, 4, 5, 7, 8, 10, and 11; C) or treated with 20 μM MG132 (B, lanes 3, 6, 9, and 12) for 6 h. Cells were lysed in RIPA buffer and incubated with an anti-FLAG M2 agarose matrix. Bound proteins were eluted with FLAG peptide and resolved by SDS-PAGE. (B) Western blot analysis of the eluates with antiserum specific to ubiquitin (αUB) (lanes 1 to 3), FLAG peptide (αFLAG) (lanes 4 to 6), ORF29p (αORF29) (lanes 7 to 9), or SUMO (αSUMO) (lanes 10 to 12). (C) Coomassie blue staining of bound proteins. Asterisks indicate the bands that were sequenced by MALDI mass spectrophotometry. MW, molecular weights in thousands. (D) Sequence of ORF29p amino acids 301 to 310 and the cleavage site (↓) that produces the 105-kDa ORF29p peptide as determined by MALDI mass spectrophotometry.
Antibody to ubiquitin and SUMO recognized the full-length and at least one truncated version of ORF29p, which demonstrates that ORF29p is modified and that the smaller ORF29p-related peptides retain the ubiquitin and SUMO moieties (Fig. 2B, lanes 2 to 3 and 11 to 12). The levels of ubiquitinated ORF29p increased following MG132 treatment, indicating that the ubiquitinated ORF29p peptides were stabilized (Fig. 2B, lane 3).
To verify that the two most prominent species obtained by immunoprecipitation were forms of ORF29p, Coomassie blue-stained bands corresponding to the presumed full-length ORF29p (130 kDa) and an associated 105-kDa peptide were analyzed by MALDI mass spectrophotometry (Fig. 2C). Mass spectrophotometry sequencing verified that the 130-kDa species contained the full-length, FLAG-tagged ORF29p sequence, while the 105-kDa sample was a fragment of ORF29p whose N terminus was amino acid 304. This slower-migrating species lacks the ORF29p NLS (34). The sequence surrounding the N terminus of the ORF29p truncation, with an arrow indicating the location of the presumed cleavage site, is shown in Fig. 2D. The predicted truncation site was determined by the detection of a unique peptide ion at an m/z of 1,336.6 that was identified to be 304-STSKPSPSGGFER-316, from the lower band. Mass spectrophotometry sequencing also reported that peptides corresponding to the FLAG epitope or the N terminus of ORF29p were not present in the 105-kDa species sample. These data and those derived from Western blot analysis reveal that ORF29p accumulates in expressing cells as a full-length species and as smaller peptides generated by cleavage near the N terminus.
One characteristic of the proteasome is that not all substrates are hydrolyzed to completion. Some substrates are truncated, and these products can serve functions that are distinct from the full-length protein. This proteolysis step can serve as a potent regulatory tool for transforming a protein from one form to another. It is, therefore, possible that one or more of the smaller isoforms of ORF29p function during VZV infection. However, in cotransfection experiments, an ORF29p N-terminal truncation that is missing the ORF29p NLS does not affect the localization of full-length ORF29p or vice versa, thus arguing against either peptide having a dominant negative effect on localization (data not shown). These results also demonstrate that although the full-length ORF29p and the N-terminal deletion appear to interact, this multimer does not accumulate in the nucleus.
ORF29p is ubiquitinated and sumoylated in transfected 293T cells.
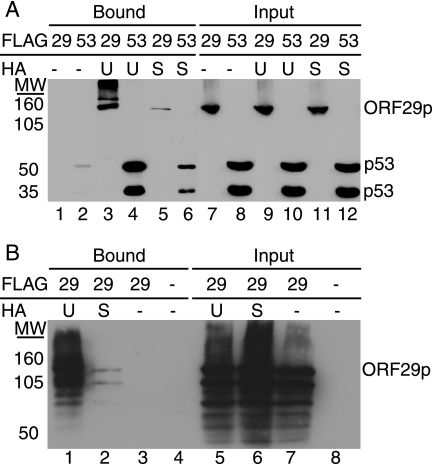
The biochemical approach of Xirodimas et al. (36) was employed to provide further evidence that the ORF29p isoforms were ubiquitinated and sumoylated. Briefly, pFLAGORF29 was transfected into 293T cells either alone or in combination with a construct expressing HA-tagged ubiquitin (pCMV-HA-UB) or HA-tagged SUMO (pHis6-HA-SUMO). A FLAG-tagged p53 expression vector (pCMV-Flag-53) provided a positive substrate control for both modifications (12, 18-20). Cell lysates were prepared and incubated with an HA affinity matrix to remove the ubiquitinated or sumoylated molecules. Figure 3 illustrates that ORF29p, like p53, bound to the HA affinity columns when coexpressed with HA-UB or HA-SUMO but not when expressed alone. Western blots with FLAG-specific antiserum confirmed that ORF29p was ubiquitinated and sumoylated, and in agreement with previous data, the FLAG antibody reacted only with the full-length or more-slowly migrating forms of ORF29p (Fig. 3A). When the same eluates were subjected to Western blot analysis and probed with ORF29p-specific antiserum, the full-length ORF29p molecule was detected along with higher- and lower-molecular-weight species (Fig. 3B). Thus, the lower-molecular-weight ORF29p species retain the ubiquitin and SUMO moieties and interact with the modified full-length ORF29p. The signal representing sumoylation of ORF29p was reproducibly lower than that for ubiquitination, which may reflect the level of modification. These data demonstrate that ORF29p is ubiquitinated and sumoylated.
FIG. 3.
Analysis of ORF29p modifications. 293T cells were transiently transfected with either pFLAGORF29 (A, lanes 1, 3, 5, 7, 9, and 11; B, lanes 1 to 3 and 5 to 7), pCMV-Flag-53 (A, lanes 2, 4, 6, 8, 10, and 12), or empty pCF2HN (B, lanes 4 and 8) either alone (A, lanes 1, 2, 7, and 8; B, lanes 3, 4, 7, and 8) or in conjunction with pCMV-HA-UB (UB) (A, lanes 3, 4, 9, and 10; B, lanes 1 and 5) or pHis6-HA-SUMO (SU) (A, lanes 5, 6, 11, and 12; B, lanes 2 and 6). At 48 h posttransfection, cells were lysed in RIPA buffer and reacted with an anti-HA matrix. Bound proteins were eluted with HA peptide. Bound and input fractions were resolved by SDS-PAGE, and ORF29p and p53 were detected by Western blot analysis with FLAG-specific antiserum (A) or ORF29p-specific antiserum (B). MW, molecular weights in thousands.
Degradation products of ORF29p accumulate predominantly in the cytoplasm.
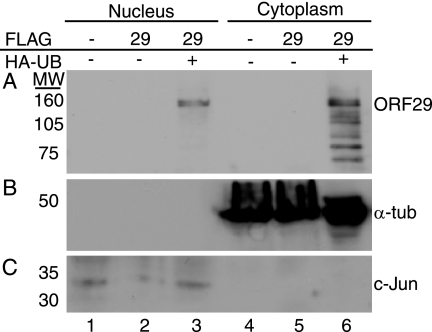
We next asked whether isoforms of ORF29p were the same in the cytoplasm and nucleus. Fractionated cell extracts were prepared from 293T cells transfected with pFLAGORF29, subjected to HA affinity chromatography, and assayed by Western blot analyses with an ORF29p-specific antiserum. Full-length, ubiquitinated ORF29p was abundant in the nucleus (Fig. 4A), whereas full-length ORF29p and other, more rapidly migrating related peptides were present in the cytoplasmic fraction. Thus, ORF29p is predominantly degraded in the cytoplasm, as the lower-molecular-mass products were not detected in the nuclear fraction. The distributions of c-Jun and α-tubulin were used to monitor the efficiency of the cell fractionation procedure (Fig. 4B). These data suggest that some ORF29p may be inhibited from accumulating in the nucleus because it is rapidly degraded in the cytoplasm by the 26S proteasome or immediately exported from the nucleus following degradation, as the smaller products were not detected in the nuclear fraction. Metabolic labeling and Western blot analyses demonstrated that ORF29p ubiquitination and degradation occur in both fibroblast- and astrocytoma-derived cells regardless of the different localization patterns of ORF29p in these contexts (Fig. 1). Therefore, it appears to be the efficiency of this process that is cell type specific.
FIG. 4.
Subcellular distribution of ubiquitinated ORF29p. 293T cells were transiently transfected with either empty pCF2HN (lanes 1 and 4) or pFLAGORF29 (lanes 2, 3, 5, and 6) either alone (lanes 1, 2, 4, and 5) or in conjunction with pCMV-HA-UB (lanes 3 and 6). At 48 h posttransfection, cells were harvested, separated into nuclear and cytoplasmic fractions, and incubated with an anti-HA matrix. Bound proteins were eluted with HA peptide. Eluates (A) and input fractions (B and C) were resolved by SDS-PAGE. ORF29p (ORF29) (A), c-Jun (B), and α-tubulin (α-tub) (C) were detected by Western blotting. MW, molecular weights in thousands.
The SV40 large-T-antigen NLS is not able to target ORF29p to the nucleus.
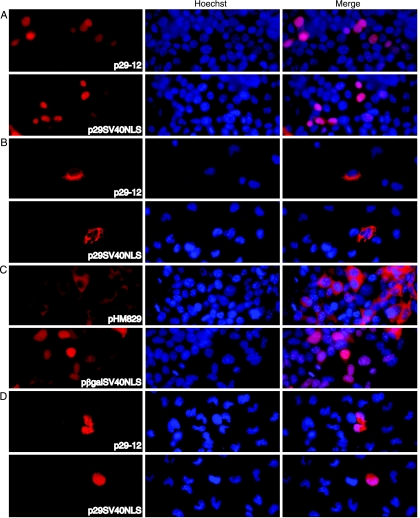
Inhibition of the nuclear import of some proteins is achieved by masking the nuclear targeting signal (2, 13, 25, 38). To determine whether the block to nuclear import of ORF29p in U373MG cells was NLS specific, ORF29p was expressed as a C-terminal SV40 NLS fusion. The SV40 NLS is a well-characterized signaling domain that can effectively direct other cytoplasmic proteins to the nucleus (14). Fusion of the SV40 NLS to the C terminus of the β-Gal protein, which typically resides in the cytoplasm, allows for nuclear targeting of the resulting chimeric protein (Fig. 5C). To test the effect of the same SV40 NLS sequence on the subcellular distribution of ORF29p, 293T and U373MG cells were transfected with either p29-12 or p29SV40NLS, which express wild-type ORF29p or an ORF29p-SV40 NLS fusion, respectively, and the distributions of these proteins were analyzed by indirect IF (Fig. 5). As expected, both proteins were present in the nuclei of transfected 293T cells (Fig. 5A). However, both the wild-type ORF29p and the SV40 NLS fusion were detected only in the cytoplasm of U373MG cells (Fig. 5B). This suggests that the block to nuclear import is an active process and cannot be overridden by the presence of a canonical NLS.
FIG. 5.
Localization of an ORF29p SV40 NLS fusion protein. 293T (A and C) and U373MG (B and D) cells were transfected with p29-12, p29SV40NLS, pHM829, or pβgalSV40NLS. At 48 h posttransfection, cell cultures were treated with either DMSO (A to C) or 20 μM MG132 in DMSO (D) for 6 h prior to fixing them onto glass coverslips. ORF29p, β-galactosidase, and the SV40 NLS fusions of these proteins were detected by indirect immunofluorescence microscopy after reacting them with specific antisera. The nuclei were visualized by counterstaining with Hoechst.
To determine whether similar mechanisms detain both the wild-type ORF29p and the SV40 NLS fusion in the cytoplasm, transfected U373MG cells were treated with MG132 for 6 h before processing (Fig. 5D). Both proteins accumulated in the nuclei of expressing U373MG cells following inhibition of the proteasome with MG132 treatment. Thus, the proteasome degradation pathway is involved in the nuclear exclusion of these proteins in U373MG cells.
ORF29p overexpression results in its nuclear accumulation in U373MG cells.
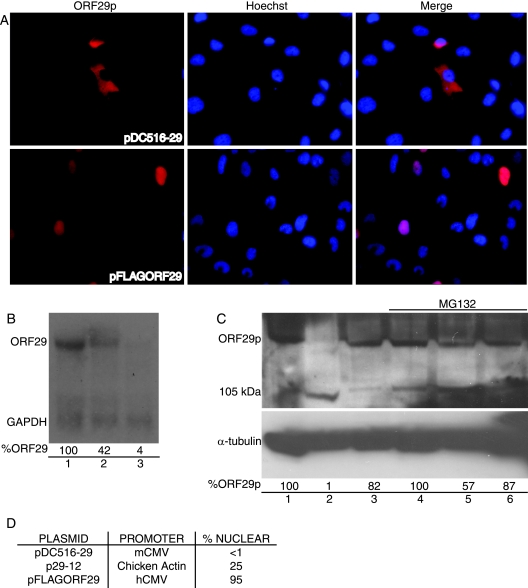
Because the cumulative evidence suggested that cytoplasmic retention of ORF29p is an active process, we were interested in determining whether overexpression of ORF29p in U373MG cells would saturate the system and result in nuclear localization. To test this hypothesis, we transfected U373MG cells with plasmids wherein ORF29p expression was regulated from either a mouse CMV (pDC516-29) or a human CMV (pFLAGORF29) promoter. The hCMV promoter directs the synthesis of higher levels of protein than the mCMV promoter in human cells. In U373MG cells transfected with pDC516-29, ORF29p levels are low and the protein accumulates in the cytoplasm (Fig. 6A). In contrast, expression of ORF29p from the hCMV promoter of pFLAGORF29 results in accumulation of ORF29p in both the nucleus and the cytoplasm in over 95% of the cells observed (Fig. 6A).
FIG. 6.
Overexpression of ORF29p in U373MG cells. U373MG cells were transfected with pDC516-29 (B, lane 3; C lanes 2 and 5), pFLAGORF29 (B, lane 1; C, lanes 1 and 4), or p29-12 (B, lane 2; C, lanes 3 and 6), which express ORF29p from mCMV, hCMV, or chicken actin promoters, respectively. At 48 h posttransfection, the cells were treated with either DMSO (A, B, and C, lanes 1 to 3) or 20 μM MG132 (C, lanes 4 to 6) for 6 h before fixing them onto glass coverslips (A) or harvesting them for Northern (B) and Western (C) blot analyses. (A) ORF29p was detected by indirect immunofluorescence microscopy after it was reacted with specific antiserum. (B) Total RNA was isolated, separated by gel electrophoresis, and transferred to a nylon membrane. The mRNA was probed with radiolabeled probes specific for ORF29 and GAPDH (glyceraldehyde-3-phosphate dehydrogenase). Following hybridization, these RNAs were visualized by exposure to film. The percentages of ORF29 transcript were calculated in comparison to transcript levels from the hCMV promoter (100%) after normalization to GAPDH levels by using the ImageJ program. (C) Infected U373MG cells were lysed in RIPA buffer and subjected to SDS-PAGE. After the transfer of proteins to a nitrocellulose membrane, Western blot analysis was done using ORF29p- and α-tubulin-specific antisera. The percentages of ORF29p were calculated in comparison to protein levels from the hCMV promoter (100%) after normalization to α-tubulin levels by using the ImageJ program. (D) The percentages of ORF29p-expressing cells that exhibited nuclear staining when reacted with antiserum specific for ORF29p during IF experiments, according to the promoter.
To confirm that transcription from the hCMV promoter resulted in the accumulation of more RNA than that from its mouse counterpart in human cells, a Northern blot analysis was performed. Total RNA was isolated from 293T cells transfected with pDC516-29, pFLAGORF29, or p29-12. ORF29p expression from p29-12 is driven by the chicken actin promoter and results in nuclear exclusion of ORF29p in 75% of expressing U373MG cells (Fig. 6D). This situation represents an intermediate intracellular localization phenotype compared to expression driven by the mouse or human CMV promoters (Fig. 5B and 6D). Northern blot analysis demonstrated that of the three promoters tested, ORF29 had the highest mRNA levels when driven by the hCMV promoter and the lowest in cells transfected with the mCMV construct (Fig. 6B). The presence of a single ORF29 RNA species also confirms that the truncated forms of ORF29p detected during metabolic labeling and Western blot experiments are a consequence of posttranslational events.
We next asked whether the protein levels of ORF29p correlated with the RNA levels. Accordingly, whole-cell protein extracts from transfected U373MG cells were subjected to SDS-PAGE and Western blot analysis. Full-length ORF29p expressed in U373MG cells driven by the mCMV promoter was barely detectable compared to ORF29p levels in cells transfected with pFLAGORF29 (Fig. 6C, lanes 1 and 3). In addition, the 105-kDa ORF29p species was more abundant in U373MG cells transfected with pDC516-29 than in cultures transfected with the other constructs. In other cultures, transfected U373MG cells were also treated with MG132 before analysis. Inhibition of the proteasome increased the levels of full-length ORF29p in cells transfected with pDC516-29, bringing the amounts closer to those seen in cells transfected with pFLAGORF29 (Fig. 6C, lanes 4 and 5). Inhibition of the proteasome did not affect the levels of full-length protein expressed from the hCMV promoter, at which time ORF29p was nuclear in both treated and untreated cultures (Fig. 6). The levels of ORF29 RNA and protein that accumulated from the chicken actin promoter-driven construct were intermediate compared to those seen with the mouse and human CMV promoters (Fig. 6). Therefore, Northern and Western blot results correlated the levels of protein expression with the subcellular distribution patterns observed during IF experiments. These data suggest that a threshold for the inhibition of nuclear transport exists, as overexpression or stabilization of ORF29p allows for its nuclear accumulation.
ORF29p localization in MeWo and U373MG heterokaryons is dependent on cell type ratios.
Our data demonstrate that ORF29p is less stable in U373MG cells, where it resides in the cytoplasm, than in MeWo cells, where the protein accumulates in the nucleus. Therefore, differences that make ORF29p more susceptible to breakdown in one environment than the other exist between the cell types. Heterokaryon assays have previously been used to characterize the shuttling properties of proteins (6, 7, 29). Accordingly, we used such an assay to ask whether the cytoplasm of either cell type would arrest or enhance the compartmentalization of ORF29p.
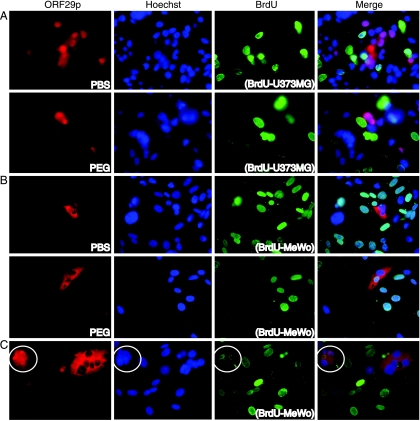
MeWo cells expressing ORF29p for 48 h were cocultured with U373MG cells that had been labeled with BrdU. Twelve hours later, the cells were incubated for 90 s with either PBS or 50% PEG to induce cell fusion. The cells were washed and allowed to recover for 12 h before ORF29p localization and BrdU incorporation were visualized by indirect IF microscopy (Fig. 7). U373MG nuclei were distinguished from MeWo nuclei by their reactivities with antiserum to BrdU. In mixed cultures treated with PBS only, isolated MeWo cells expressed ORF29p (Fig. 7A). The subcellular distribution of ORF29p within these cells was predominantly nuclear with diffuse cytoplasmic staining as previously described (33). In PEG-treated cultures, numerous multinucleated cells were present. In 75% of expressing heterokaryons, ORF29p was present only in nuclei that did not contain BrdU-labeled DNA, indicating that the protein expressed in MeWo cells did not translocate into U373MG nuclei (Fig. 7A). Thus, the presence of cytoplasm from MeWo cells did not permit ORF29p accumulation in U373MG nuclei in the majority of expressing heterokaryons and the ORF29p present in the nuclei of MeWo cells was also not affected by the presence of U373MG cytoplasm.
FIG. 7.
ORF29p localization in U373MG and MeWo cell heterokaryons. MeWo (A) and U373MG (B and C) cells were infected with AdORF29 at an MOI of 50. At 48 hours postinfection, infected cells were mixed with U373MG (A) or MeWo (B and C) cells that were labeled by incubation with BrdU and seeded onto coverslips for 12 h. The mixed cultures were incubated with either PBS or 50% PEG in PBS for 90 s at 37°C, washed four times, and allowed to recover in normal growth media for 12 h at 37°C before they were fixed and analyzed by indirect immunofluorescence with antisera specific for ORF29p and BrdU. For each panel, the cell line containing BrdU is indicated in parentheses. The white circle surrounds the heterokaryon containing a 5:1 MeWo-to-U373MG nucleus ratio.
Similar experiments were done in the presence of 50 μg/ml of cycloheximide to inhibit de novo protein synthesis. Cells were treated with cycloheximide for 20 min, fused, and allowed to recover for 12 h in the presence of the drug. In both PBS- and PEG-treated cultures, there was no loss of nuclear ORF29p signal after cycloheximide treatment, which is characteristic of expression in homogenous MeWo cell cultures (data not shown) (33).
The protocol was then reversed so that ORF29p was expressed in U373MG cells before they were mixed with nonexpressing MeWo cells labeled with BrdU. As expected, ORF29p was cytoplasmic in U373MG cells when mixed cultures were treated with PBS (Fig. 7B). Following cell fusion, ORF29p was excluded from nuclei of both cell types in 80% of expressing heterokaryons (Fig. 7B). These data indicate that the presence of MeWo cytosol is not usually sufficient for nuclear translocation of ORF29p expressed in U373MG cells. However, ORF29p accumulated in the nuclei of both cell types in a small percentage of multinucleated heterokaryons. From an analysis of this small percentage of heterokaryons that allowed for ORF29p nuclear targeting, we deduced that a relationship between the ratio of MeWo to U373MG nuclei and the ability of ORF29p to target to U373MG nuclei existed. ORF29p accumulated in the nuclei of heterokaryons, regardless of whether they were of MeWo or U373MG origin, that contained a 4:1 or higher ratio of MeWo to U373MG nuclei. In Fig. 7C, the circled heterokaryon in the upper left-hand corner harbors a 5:1 ratio of MeWo to U373MG nuclei and ORF29p is found in the nuclei of both cell types, while the heterokaryon on the right contains more U373MG nuclei and ORF29p remained in the cytoplasm. The ability of the more abundant cell type in a heterokaryon to influence the subcellular distribution of ORF29p suggests that it is the ratio of the cytoplasms that determines the cellular localization and abundance of ORF29p.
When ORF29p was expressed in U373MG cells, cycloheximide treatment ablated ORF29p signal in both PBS- and PEG-treated cells (data not shown). This is not surprising, as the half-life of ORF29p in U373MG cells is short (33). Therefore, in the absence of de novo protein synthesis, most of the ORF29p was eliminated during the cycloheximide pretreatment period.
DISCUSSION
The subcellular distribution of ORF29p changes according to whether VZV infection is lytic or latent. ORF29p is predominantly nuclear during lytic infection of the dermis and epidermis but appears to be restricted to the cytoplasm of latently infected neurons (3, 21, 33). In reactivated enteric neurons in culture or in neurons examined at autopsy from patients with active zoster, ORF29p accumulates in the nucleus (21, 33). When expressed autonomously, ORF29p displays a similar pattern of cell type-specific localization (33, 34). Therefore, it is the cell type-specific environment that governs the segregation patterns of ORF29p.
We have previously shown that ORF29p has a novel NLS signal that uses the classical nuclear transport pathway to reach the nucleus (34). In this report, we demonstrate that a canonical NLS cannot target ORF29p to the nuclei of U373MG cells and, therefore, does not override the retention of ORF29p in the cytoplasm. This supports the hypothesis that selective nuclear exclusion involves recognition of an amino acid sequence within the protein that is distinct from the NLS and that retention of ORF29p in the cytoplasm of U373MG cells is an active process. Another cellular pathway that affects ORF29p localization is the proteasome degradation pathway. Inhibition of the proteasome leads to stabilization and nuclear accumulation of ORF29p in U373MG cells and cultured guinea pig EG (33). The ORF29p cell type-specific subcellular distribution patterns may result from the differential targeting or accessibility of the 26S proteasome.
Most proteins targeted to the proteasome for degradation are tagged by the covalent attachment of multiple ubiquitin moieties. We have demonstrated by Western blot and biochemical analyses that ORF29p is ubiquitinated, sumoylated, and degraded in 293T, MeWo, and U373MG cells and that multiple truncated ORF29p isoforms accumulate in expressing cells (Fig. 2 and 3). The predominant truncated ORF29p species migrates at 105 kDa and represents a peptide containing ORF29p amino acids 304 to 1203. Ubiquitin- and SUMO-specific antisera also recognize the 105-kDa band, demonstrating that this peptide is also modified. The role of these modifications in the regulation of the intracellular localization of ORF29p is unknown. It is possible that the level and/or sites of either of these modifications determine the fate of ORF29p.
The origin of the 105-kDa ORF29p isoform remains undefined. Amino acids 303 and 304 both contain uncharged polar-side chains that constitute an atypical target for all three proteolytic sites (trypsin like, chymotrypsin like, and caspase like) used by the 26S proteasome (8, 17, 27, 28). There are no methionine residues near the N terminus of the 105-kDa truncated species. This argues against the possibility of an alternative translational start site, and caspase-mediated cleavage is unlikely because the truncation does not occur after an aspartic acid residue (35).
One characteristic of the proteasome is that not all substrates are hydrolyzed to completion. Some substrates are truncated, and these products can serve functions that are distinct from those of the full-length protein. This proteolysis step can provide a potent regulatory tool for transforming a protein from one form to another. Many viruses encode polyproteins that are specifically cleaved into smaller polypeptides that serve distinct functions during infection. While it is possible that one or more of the smaller isoforms of ORF29p function during VZV infection, cotransfection experiments suggested that an ORF29p N-terminal truncation does not affect the localization of full-length ORF29p or vice versa. Moreover, the multimeric form of the protein containing the N-terminal truncation and full-length polypeptide was not found in the nucleus.
ORF29p that accumulates in the nuclei of U373MG cells after MG132 treatment is immediately turned over when proteasome activity is restored (33). Therefore, stabilization leads to nuclear accumulation but not vice versa. In contrast, ORF29p in MeWo cell nuclei is stable, even after cycloheximide treatment and in the presence of U373MG cytoplasm when heterokaryons are formed. Biochemical analyses of 293T subcellular fractions demonstrated that while ubiquitinated ORF29p is found in both the nucleus and the cytoplasm of 293T cells, degradation products are detected only in the cytoplasm. Thus, ORF29p may either be targeted for degradation before entering the nucleus or be rapidly exported from the nucleus and degraded in the cytoplasm. It appears that in MeWo and 293T cells, the bulk of ORF29p escapes degradation and is transported to the refuge of the nuclei, while in U373MG cells, ORF29p has no escape from proteasome-directed destruction.
Pulse chase experiments reveal that ORF29p degradation occurs more slowly in MeWo cells than in U373MG cells (33). The numerous species that copurify with ORF29p appear immediately in U373MG cells but not until later time points in MeWo cells. We have demonstrated that shortly after synthesis in U373MG cells, a truncated polypeptide containing ORF29p amino acids 304 to 1203 appears. The resulting polypeptide lacks an NLS and thus cannot enter the nucleus on its own. Since degradation is delayed in MeWo cells, ORF29p can be imported into the sanctuary of the nucleus. In both cell types, ORF29p that remains in the cytoplasm is susceptible to proteasome-directed degradation.
The degradation system responsible for inhibiting ORF29p nuclear translocation is saturated by overexpression (Fig. 6). Thus, there is a threshold for achieving nuclear translocation that may be overcome by a change in expression levels or abundance. This characteristic creates the caveat that in the heterokaryon assays, we were investigating not only the localization of ORF29p but also its abundance. In most heterokaryons, the presence of MeWo cytosol had no effect on the exclusion of ORF29p from U373MG nuclei and U373MG cytoplasm had no effect on ORF29p residing in MeWo nuclei. However, in heterokaryons that contained a 4:1 or higher ratio of MeWo to U373MG cell nuclei, ORF29p accumulated in the nuclei of both cell types. In this context, the active degradation process in U373MG cells is probably diluted and/or overcome by the presence of abundant MeWo cell cytosol.
Our studies show that the degradation pathway is also active, although less efficient, in those cell lines where ORF29p accumulates in the nucleus. This observation suggests that during lytic infection, this mechanism is also conserved. Like that of other alphaherpesviruses, VZV gene expression is temporally regulated during a productive infection (31). The ORF29 promoter is stimulated by IE62, which may allow for accumulation of ORF29p at peak times of VZV DNA replication (22, 23, 37). At later times during infection, when virions are being packaged, ORF62 is downregulated and the protein accumulates in the cytoplasm (16). In response to the relocalization of ORF62p, ORF29p expression levels may drop below the threshold required for nuclear import. A higher proportion of ORF29p will, therefore, be degraded in the cytoplasm. Therefore, during productive infection, the proteasome in conjunction with the expression of ORFs 61p and 62p may assist in regulating the amount of cellular ORF29p during viral replication. ORF29p resides in small quantities in the cytoplasm during lytic infection but is not packaged in the VZV virion, indicating that it does not gain access to the assembly machinery. We speculate that association with the proteasome degradation components sequesters ORF29p away from the virion packaging machinery.
ORF29p degradation and nuclear exclusion during latency may be necessary to maintain viral DNA in a quiescent state. During latency, IE62 appears to be retained in the cytoplasm with ORF29p. The apparent inaccessibility of IE62 during latency might serve to limit expression from the ORF29 promoter (22-24, 37). With only low levels of ORF29p being expressed, degradation predominates and the protein cannot accumulate in the nucleus. Following reactivation, ORF29p levels are increased, which could saturate the proteasome-mediated degradation pathway and permit nuclear import.
The elucidation of the interplay between cellular pathways and virus-specified proteins in the regulation of the differential compartmentalization of ORF29p and the other LAPs may expand our understanding of VZV latency and reactivation.
Acknowledgments
These studies were supported by a grant from the Public Health Service, AI-024021, to S.J.S.
Footnotes
Published ahead of print on 6 September 2006.
REFERENCES
- 1.Arvin, A. M. 1996. Varicella-zoster virus. Clin. Microbiol. Rev. 9:361-381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Blank, V., P. Kourilsky, and A. Israel. 1991. Cytoplasmic retention, DNA binding and processing of the NF-kappa B p50 precursor are controlled by a small region in its C-terminus. EMBO J. 10:4159-4167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chen, J. J., A. A. Gershon, Z. S. Li, O. Lungu, and M. D. Gershon. 2003. Latent and lytic infection of isolated guinea pig enteric ganglia by varicella zoster virus. J. Med. Virol. 70(Suppl. 1):S71-S78. [DOI] [PubMed] [Google Scholar]
- 4.Cohrs, R. J., D. H. Gilden, P. R. Kinchington, E. Grinfeld, and P. G. Kennedy. 2003. Varicella-zoster virus gene 66 transcription and translation in latently infected human ganglia. J. Virol. 77:6660-6665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Croen, K. D., and S. E. Straus. 1991. Varicella-zoster virus latency. Annu. Rev. Microbiol. 45:265-282. [DOI] [PubMed] [Google Scholar]
- 6.Davidson, R. L., and P. S. Gerald. 1976. Improved techniques for the induction of mammalian cell hybridization by polyethylene glycol. Somatic Cell Genet. 2:165-176. [DOI] [PubMed] [Google Scholar]
- 7.Davidson, R. L., K. A. O'Malley, and T. B. Wheeler. 1976. Polyethylene glycol-induced mammalian cell hybridization: effect of polyethylene glycol molecular weight and concentration. Somatic Cell Genet. 2:271-280. [DOI] [PubMed] [Google Scholar]
- 8.Dick, T. P., A. K. Nussbaum, M. Deeg, W. Heinemeyer, M. Groll, M. Schirle, W. Keilholz, S. Stevanovic, D. H. Wolf, R. Huber, H. G. Rammensee, and H. Schild. 1998. Contribution of proteasomal beta-subunits to the cleavage of peptide substrates analyzed with yeast mutants. J. Biol. Chem. 273:25637-25646. [DOI] [PubMed] [Google Scholar]
- 9.Glickman, M. H., and A. Ciechanover. 2002. The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol. Rev. 82:373-428. [DOI] [PubMed] [Google Scholar]
- 10.Grinfeld, E., and P. G. Kennedy. 2004. Translation of varicella-zoster virus genes during human ganglionic latency. Virus Genes 29:317-319. [DOI] [PubMed] [Google Scholar]
- 11.Grose, C., and P. A. Brunel. 1978. Varicella-zoster virus: isolation and propagation in human melanoma cells at 36 and 32 degrees C. Infect. Immun. 19:199-203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Haupt, Y., R. Maya, A. Kazaz, and M. Oren. 1997. Mdm2 promotes the rapid degradation of p53. Nature 387:296-299. [DOI] [PubMed] [Google Scholar]
- 13.Henkel, T., U. Zabel, K. van Zee, J. M. Muller, E. Fanning, and P. A. Baeuerle. 1992. Intramolecular masking of the nuclear location signal and dimerization domain in the precursor for the p50 NF-kappa B subunit. Cell 68:1121-1133. [DOI] [PubMed] [Google Scholar]
- 14.Kalderon, D., W. D. Richardson, A. F. Markham, and A. E. Smith. 1984. Sequence requirements for nuclear location of simian virus 40 large-T antigen. Nature 311:33-38. [DOI] [PubMed] [Google Scholar]
- 15.Kennedy, P. G., E. Grinfeld, and J. E. Bell. 2000. Varicella-zoster virus gene expression in latently infected and explanted human ganglia. J. Virol. 74:11893-11898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kinchington, P. R., and S. E. Turse. 1998. Regulated nuclear localization of the varicella-zoster virus major regulatory protein, IE62. J. Infect. Dis. 178(Suppl. 1):S16-S21. [DOI] [PubMed] [Google Scholar]
- 17.Kisselev, A. F., T. N. Akopian, V. Castillo, and A. L. Goldberg. 1999. Proteasome active sites allosterically regulate each other, suggesting a cyclical bite-chew mechanism for protein breakdown. Mol. Cell 4:395-402. [DOI] [PubMed] [Google Scholar]
- 18.Kubbutat, M. H., S. N. Jones, and K. H. Vousden. 1997. Regulation of p53 stability by Mdm2. Nature 387:299-303. [DOI] [PubMed] [Google Scholar]
- 19.Li, M., C. L. Brooks, F. Wu-Baer, D. Chen, R. Baer, and W. Gu. 2003. Mono- versus polyubiquitination: differential control of p53 fate by Mdm2. Science 302:1972-1975. [DOI] [PubMed] [Google Scholar]
- 20.Lohrum, M. A., D. B. Woods, R. L. Ludwig, E. Balint, and K. H. Vousden. 2001. C-terminal ubiquitination of p53 contributes to nuclear export. Mol. Cell. Biol. 21:8521-8532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lungu, O., C. A. Panagiotidis, P. W. Annunziato, A. A. Gershon, and S. J. Silverstein. 1998. Aberrant intracellular localization of varicella-zoster virus regulatory proteins during latency. Proc. Natl. Acad. Sci. USA 95:7080-7085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Meier, J. L., X. Luo, M. Sawadogo, and S. E. Straus. 1994. The cellular transcription factor USF cooperates with varicella-zoster virus immediate-early protein 62 to symmetrically activate a bidirectional viral promoter. Mol. Cell. Biol. 14:6896-6906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Meier, J. L., and S. E. Straus. 1995. Interactions between varicella-zoster virus IE62 and cellular transcription factor USF in the coordinate activation of genes 28 and 29. Neurology 45:S30-S32. [DOI] [PubMed] [Google Scholar]
- 24.Meier, J. L., and S. E. Straus. 1993. Varicella-zoster virus DNA polymerase and major DNA-binding protein genes have overlapping divergent promoters. J. Virol. 67:7573-7581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Nigg, E. A. 1997. Nucleocytoplasmic transport: signals, mechanisms and regulation. Nature 386:779-787. [DOI] [PubMed] [Google Scholar]
- 26.Nikolaev, A. Y., M. Li, N. Puskas, J. Qin, and W. Gu. 2003. Parc: a cytoplasmic anchor for p53. Cell 112:29-40. [DOI] [PubMed] [Google Scholar]
- 27.Nussbaum, A. K., T. P. Dick, W. Keilholz, M. Schirle, S. Stevanovic, K. Dietz, W. Heinemeyer, M. Groll, D. H. Wolf, R. Huber, H. G. Rammensee, and H. Schild. 1998. Cleavage motifs of the yeast 20S proteasome beta subunits deduced from digests of enolase 1. Proc. Natl. Acad. Sci. USA 95:12504-12509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Orlowski, M. 1990. The multicatalytic proteinase complex, a major extralysosomal proteolytic system. Biochemistry 29:10289-10297. [DOI] [PubMed] [Google Scholar]
- 29.Pinol-Roma, S., and G. Dreyfuss. 1992. Shuttling of pre-mRNA binding proteins between nucleus and cytoplasm. Nature 355:730-732. [DOI] [PubMed] [Google Scholar]
- 30.Relaix, F., X. J. Wei, X. Wu, and D. A. Sassoon. 1998. Peg3/Pw1 is an imprinted gene involved in the TNF-NFkappaB signal transduction pathway. Nat. Genet. 18:287-291. [DOI] [PubMed] [Google Scholar]
- 31.Shiraki, K., and R. W. Hyman. 1987. The immediate early proteins of varicella-zoster virus. Virology 156:423-426. [DOI] [PubMed] [Google Scholar]
- 32.Sorg, G., and T. Stamminger. 1999. Mapping of nuclear localization signals by simultaneous fusion to green fluorescent protein and to beta-galactosidase. BioTechniques 26:858-862. [DOI] [PubMed] [Google Scholar]
- 33.Stallings, C. L., G. J. Duigou, A. A. Gershon, M. D. Gershon, and S. J. Silverstein. 2006. The cellular localization pattern of varicella-zoster virus ORF29p is influenced by proteasome-mediated degradation. J. Virol. 80:1497-1512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Stallings, C. L., and S. Silverstein. 2005. Dissection of a novel nuclear localization signal in open reading frame 29 of varicella-zoster virus. J. Virol. 79:13070-13081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Talanian, R. V., C. Quinlan, S. Trautz, M. C. Hackett, J. A. Mankovich, D. Banach, T. Ghayur, K. D. Brady, and W. W. Wong. 1997. Substrate specificities of caspase family proteases. J. Biol. Chem. 272:9677-9682. [DOI] [PubMed] [Google Scholar]
- 36.Xirodimas, D. P., C. W. Stephen, and D. P. Lane. 2001. Cocompartmentalization of p53 and Mdm2 is a major determinant for Mdm2-mediated degradation of p53. Exp. Cell Res. 270:66-77. [DOI] [PubMed] [Google Scholar]
- 37.Yang, M., J. Hay, and W. T. Ruyechan. 2004. The DNA element controlling expression of the varicella-zoster virus open reading frame 28 and 29 genes consists of two divergent unidirectional promoters which have a common USF site. J. Virol. 78:10939-10952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Zabel, U., T. Henkel, M. S. Silva, and P. A. Baeuerle. 1993. Nuclear uptake control of NF-kappa B by MAD-3, an I kappa B protein present in the nucleus. EMBO J. 12:201-211. [DOI] [PMC free article] [PubMed] [Google Scholar]