Abstract
The iProClass database provides comprehensive, value-added descriptions of proteins and serves as a framework for data integration in a distributed networking environment. The protein information in iProClass includes family relationships as well as structural and functional classifications and features. The current version consists of about 830 000 non-redundant PIR-PSD, SWISS-PROT, and TrEMBL proteins organized with more than 36 000 PIR superfamilies, 145 000 families, 4000 domains, 1300 motifs and 550 000 FASTA similarity clusters. It provides rich links to over 50 database of protein sequences, families, functions and pathways, protein–protein interactions, post-translational modifications, protein expressions, structures and structural classifications, genes and genomes, ontologies, literature and taxonomy. Protein and superfamily summary reports present extensive annotation information and include membership statistics and graphical display of domains and motifs. iProClass employs an open and modular architecture for interoperability and scalability. It is implemented in the Oracle object-relational database system and is updated biweekly. The database is freely accessible from the web site at http://pir.georgetown.edu/iproclass/ and searchable by sequence or text string. The data integration in iProClass supports exploration of protein relationships. Such knowledge is fundamental to the understanding of protein evolution, structure and function and crucial to functional genomic and proteomic research.
INTRODUCTION
The completion of the draft human genome sequences marked the beginning of a new era of biological research, in which scientists have begun systematically to explore gene functions and other complex regulatory processes by studying organisms at the global scale of genomes, transcriptomes and proteomes. With the accelerated accumulation of molecular data, advanced bioinformatics infrastructures must be developed in order to fully explore these valuable data and to generate new hypotheses and derive scientific knowledge. One major challenge lies in the volume, complexity and dynamic nature of the data, which are being collected and maintained in heterogeneous and distributed sources. The iProClass database (1) was designed to offer a comprehensive, integrated view of protein information to facilitate knowledge discovery.
OVERVIEW AND CURRENT CONTENTS
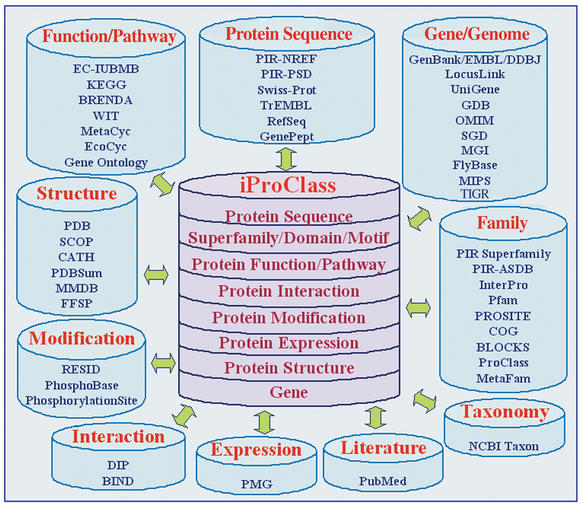
The iProClass database (Fig. 1) contains value-added descriptions of proteins, including family relationships at both global (superfamily/family) and local (domain, motif, site) levels, as well as structural and functional classifications and features. The database was first released in October 2000 and contained about 200 000 proteins from the PIR Protein Sequence Database (PIR-PSD) (2) and SWISS-PROT (3). It is updated biweekly and currently consists of about 830 000 non-redundant protein sequences from the PIR-PSD, SWISS-PROT, and TrEMBL (3) databases. The protein entries are organized with more than 36 000 PIR superfamilies (4), 145 000 families, 3700 Pfam (5) and PIR homology domains, 1300 ProSite (6) motifs, 550 000 FASTA (7) similarity clusters, and links to over 50 molecular biology databases.
Figure 1.
iProClass database overview.
Database cross-references in iProClass are represented by rich links, which include both the links and related summary information. This approach effectively combines data warehouse and hypertext navigation methods for data integration to provide timely information from distributed sources. iProClass collects information from and links to databases for protein sequences (PIR-PSD, PIR-NREF, SWISS-PROT, TrEMBL, GenPept, RefSeq), families (InterPro, Pfam, ProSite, Blocks, Prints, COG, MetaFam, PIR-ASDB, ProClass), functions and pathways (EC-IUBMB, KEGG, BRENDA, WIT, MetaCyc, EcoCyc), interactions (DIP, BIND), post-translational modifications (RESID, PhosphoSite DB), protein expression and proteomes (PMG), structures and structural classifications (PDB, PDBSum, SCOP, CATH, FSSP, MMDB), genes and genomes (GenBank, EMBL, DDBJ, LocusLink, TIGR, SGD, FlyBase, MGI, GDB, OMIM, MIPS, GenProtEC), ontologies (GO), literature (PubMed) and taxonomy (NCBI Taxonomy). The information content is continually enhanced by: (i) adding links to more databases, (ii) adding executive summary information from the linked databases and (iii) increasing the number of occurrences of links to the databases that iProClass already links to. The composite annotations collected from multiple sources are presented with attribution to the underlying databases.
iProClass presents comprehensive views for protein sequences and superfamilies in two types of summary reports. The protein sequence report covers information on family, structure, function, gene, genetics, disease, ontology, taxonomy and literature, with cross-references to relevant molecular databases and executive summary lines, as well as a graphical display of domain and motif sequence regions and a link to related sequences in pre-computed FASTA clusters. The superfamily report provides PIR superfamily membership information with length, taxonomy and keyword statistics, complete member listing separated into major kingdoms, family relationships at the whole protein and domain and motif levels with direct mapping to other classifications, structure and function cross-references, graphical display of domain and motif architecture of members, and a link to dynamically generated multiple sequence alignments and phylogenetic trees for superfamilies with curated seed members.
DATABASE ACCESS AND USAGE
The iProClass database employs an open and modular database architecture to provide a framework for data integration in a distributed networking environment. The modular structure makes the system scalable, customizable, and extendable for adding new components. The database is implemented in the Oracle object-relational system and freely accessible from our web site at http://pir.georgetown.edu/iproclass/. Direct report retrieval is based on unique identifiers such as PIR or SWISS-PROT sequence ID (e.g. http://pir.georgetown.edu/cgi-bin/ipcEntry?id=A31997) or PIR superfamily ID (e.g. http://pir.georgetown.edu/cgi-bin/ipcSF?id=SF000130). Matching lists of proteins or superfamilies are retrievable by sequence search [BLAST (8) or peptide match] or combinations of text strings (from more than 50 searchable fields of unique identifiers and annotations). The lists are displayed with line-summaries that contain protein IDs, matched fields, protein name, taxonomy, superfamily, domain, motif and matched sequence region (for BLAST and peptide search), with hypertext links to the full protein and superfamily reports.
With up-to-date information from many sources, iProClass provides much richer protein annotation than is found in any single database. The data integration allows interesting relationships to be revealed between objects in different databases. This can facilitate discovery of functional associations beyond sequence homology. For example, searches of PIR superfamilies sharing the adenylylsulfate kinase (EC 2.7.1.25) domain have led to the identification of functionally related domains for sulfate adenylyltransferase (EC 2.7.7.4) that bear no detectable sequence similarity via their association in multi-domain proteins (9). Knowledge of such relationship is fundamental to the understanding of protein evolution, structure and function and crucial to functional genomic and proteomic research.
Acknowledgments
ACKNOWLEDGEMENTS
This study is supported by grant DBI-9974855 and DBI-0138188 from National Science Foundation, and grant P41 LM05798 from National Library of Medicine.
REFERENCES
- 1.Wu C.H., Xiao,C., Hou,Z., Huang,H. and Barker,W.C. (2001) iProClass: an integrated and comprehensive protein classification database. Nucleic Acids Res., 29, 52–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Wu C.H., Huang,H., Arminski,L., Castro-Alvear,J., Chen,Y., Hu,Z., Ledley,R.S., Lewis,K.C., Mewes,H.-W., Orcutt,B.C., Suzek,B., Tsugita,A., Vinayaka,C.R., Yeh,L.-S., Zhang,J. and Barker,W.C. (2002) The Protein Information Resource: an integrated public resource of functional annotation of proteins. Nucleic Acids Res., 30, 35–37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bairoch A. and Apweiler,R. (2000) The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res., 28, 45–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Barker W.C., Pfeiffer,F. and George,D.G. (1996) Superfamily classification in PIR-International Protein Sequence Database. Methods Enzymol., 266, 59–71. [DOI] [PubMed] [Google Scholar]
- 5.Bateman A., Birney,E., Cerruti,L., Durbin,R., Etwiller,L., Eddy,S.R., Griffiths-Jones,S., Howe,K.L., Marshall,M. and Sonnhammer,E. (2002) The Pfam Protein Families Database. Nucleic Acids Res., 30, 276–280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Falquet L., Pagni,M., Bucher,P., Hulo,N., Sigrist,C.J.A., Hofmann,K. and Bairoch,A. (2002) The PROSITE database, its status in 2002. Nucleic Acids Res., 30, 235–238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Pearson W.R. and Lipman,D.J. (1988) Improved tools for biological sequence comparison. Proc. Natl Acad. Sci. USA, 85, 2444–2448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Altschul S.F., Madden,T.L., Schaffer,A.A., Zhang,J., Zhang,Z., Miller,W. and Lipman,D.J. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res., 25, 3389–3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wu C.H. and Barker,W.C. (2003) Functional annotation of proteins: a family classification approach. In Wang,L. (ed.), The Practical Bioinformatician. World Scientific Inc., in press.