Abstract
Replicators are fundamental to the origin of life and evolvability. Their survival depends on the accuracy of replication and the efficiency of growth relative to spontaneous decay. Infrabiological systems are built of two coupled autocatalytic systems, in contrast to minimal living systems that must comprise at least a metabolic subsystem, a hereditary subsystem and a boundary, serving respective functions. Some scenarios prefer to unite all these functions into one primordial system, as illustrated in the lipid world scenario, which is considered as a didactic example in detail. Experimentally produced chemical replicators grow parabolically owing to product inhibition. A selection consequence is survival of everybody. The chromatographized replicator model predicts that such replicators spreading on surfaces can be selected for higher replication rate because double strands are washed away slower than single strands from the surface. Analysis of real ribozymes suggests that the error threshold of replication is less severe by about one order of magnitude than thought previously. Surface-bound dynamics is predicted to play a crucial role also for exponential replicators: unlinked genes belonging to the same genome do not displace each other by competition, and efficient and accurate replicases can spread. The most efficient form of such useful population structure is encapsulation by reproducing vesicles. The stochastic corrector model shows how such a bag of genes can survive, and what the role of chromosome formation and intragenic recombination could be. Prebiotic and early evolution cannot be understood without the models of dynamics.
Keywords: replicator, origin of life, ribozyme, autocatalysis, compartmentation, error threshold
1. Introduction
The replicator, as introduced by Dawkins (1976), has become one of the central concepts in evolutionary theory. He identified two types of replicator with unbounded evolutionary potential, namely genes and memes (memes were meant to be hereditary units of cultural rather than genetic evolution). These ideas have turned out to be extremely fruitful: they have elicited renewed interest in the philosophy of evolution (e.g. Hull 1980) and led to the recognition of other types of replicators with the most important role in evolution (Maynard Smith & Szathmáry 1993, 1995).
A classification of replicators was presented by Maynard Smith & Szathmáry (1995) and it has been refined a number of times (Szathmáry 1995, 2000). Most widely known replicators, including genes, are strongly tied to the world of chemistry: this is obviously not true for memes. Some replicators have only limited heredity (Maynard Smith & Szathmáry 1995), implying that the number of possible types is smaller than or roughly equal to the number of individuals (copies, sequences, etc.) in a plausible (realistic) system. Conversely, in the case of unlimited hereditary replicators, the number of types by far exceeds that of individuals in the population (Szathmáry & Maynard Smith 1997). This shows that a classification of replicators is not naturally hierarchical: there exist molecular and non-molecular replicators with limited or unlimited hereditary potential.
Oparin (1961) defined any system capable of replication and mutation as alive. Most evolutionary biologists would agree with this view. Systems with these properties can evolve complex adaptations (purposeful functions) in the natural world, highly characteristic of living beings. Yet some authors (including Gánti 1971, 1978) have raised doubts concerning such an approach. The acid test is whether viruses are alive or not. Gánti (1971) argued that to regard viruses as living amounts to a conceptual mistake equating programs with computers. In the full analogy, the virus corresponds to a program, written in a decodable language, which says to the computer: ‘Print me again and again, even if you disintegrate as a result of doing so!’ The active part is obviously the computer and not the program. The computer can do many things without such a malign program. In sharp contrast, the program cannot do anything on its own. The living cell is thus analogous to the computer. Since everyone regards the cell in its active state alive, life as such in the example rests with the cell rather than the virus.
Yet viruses evolve. In fact, they have become one of the most accessible test systems for evolutionary hypotheses (e.g. Poon & Chao 2004). Computer programs can also evolve (e.g. Bedau et al. 2000). What is the relationship between units of evolution and units of life? To give a tentative answer, both the concepts must be defined first with sufficient clarity, and only after this the two notions can be compared. Units of evolution must: (i) multiply, (ii) have heredity and (iii) heredity must not be totally accurate (variability). Furthermore, some of the inherited traits must affect the chance of reproduction or of survival of the units. If all these criteria are met, then in a population of such entities, evolution by natural selection can take place (Maynard Smith 1986). Note that this definition does not refer to living systems. Any system satisfying these criteria can evolve in a Darwinian manner.
Units of life as such are less well studied, although cells and organisms are widely known and analysed. Gánti (1971, 1979, 1987, 2003) has refined his ‘life criteria’ that living systems must meet. He observed, correctly, that for the individual living state, reproduction is neither necessary nor sufficient. Many cells and organisms are commonly regarded alive even if they cannot reproduce (any longer). The so-called potential life criteria must be met only if the population of units is to be maintained and evolved. Then, the correct relationship between units of evolution and units of life is that of two partially overlapping sets (Szathmáry 2002).
Some regard the concept of a replicator more informational, detached from real processes of replication, reproduction and development. The elegant concept of a reproducer (Griesemer 2000, 2002) is meant to fill this gap. A reproducer is a unit of multiplication, hereditary variation and development. A reproducer must have at least a minimum developmental capacity required for further multiplication. There is not only an informational link but also material overlap between generations of reproducers. Thus, genes in an organism are replicators but not reproducers. Conversely, an organism is not a replicator but reproducer. In the course of prebiotic and early biological evolution, replicators ganged up to yield reproducers. We shall consider in detail how this could have happened.
2. Survival criteria for informational replicators
Informational replicators, such as genes, have unlimited heredity. The earliest informational replicators must have faced at least two severe constraints. Serious considerations suggest that primordial nucleic acids (or their analogues) must have been rather short molecules owing to excessive noise in their copying. Another consideration emphasizes the fact that replicators must have a growth rate high enough to compensate for spontaneous decay. I consider these two aspects in turn.
(a) The error threshold
Eigen (1971) called attention to the fact that the length of molecules (number of nucleotides) maintained in mutation–selection balance is limited by the copying fidelity. We recapitulate the simplified treatment by Maynard Smith (1983). Imagine two sequences with replication rate constants K and k(<K), respectively. The first sequence mutates into the second with a mutation rate (1−Q). If we assume that they are in a flow reactor where total concentration is kept constant, then the rate equations for growth and competition become
| (2.1a) |
| (2.1b) |
where x and y are concentrations of wild-type and mutant, respectively, Φ=xK+yk and total concentration is (without loss of generality) unity. It is easy to see that in equilibrium, when both templates are present in non-zero concentration, it holds that
| (2.2) |
where it must be true that Q>k/K. If there are ν digits in the sequence, Q=qν can be approximated by e−ν(1−q), where q is the copying fidelity per base per replication. From this we obtain
| (2.3) |
which is Eigen's error threshold of replication. Non-enzymatic replication implies low q, so ν<100 is probable for prebiotic chemistry, which is about the size of a tRNA molecule. Therefore, early genomes must have consisted of independently replicating entities. But they would compete with each other and the one with the highest fitness would win (Eigen 1971). Hence, the ‘Catch-22’ of molecular evolution: no enzymes without a large genome and no genome without enzymes (Maynard Smith 1983).
(b) The decay threshold
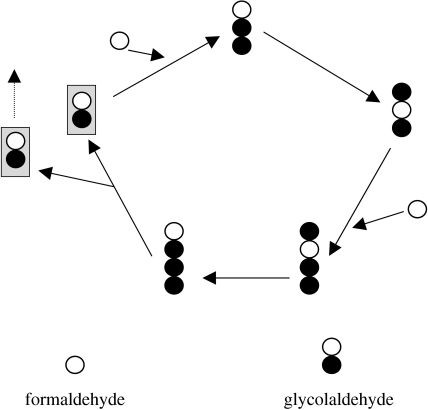
Consider, for a change, a non-informational replicator, such as any intermediate in the formose reaction (figure 1). Note that such an autocatalytic cycle differs markedly from Kauffman's (1993) reflexively autocatalytic protein nets: in the former, each elementary reaction is stoichiometric rather than catalytic. There is a severe problem with the formose reaction: deadly side reactions drain it to such an extent that the intermediates of the cycle disappear ultimately (e.g. Shapiro 1986). This may have been different for cycles on surfaces, but we do not know (yet). As King (1982, 1986) pointed out, the smaller the cycle, the better the chances for its propagation. Suppose that there is a simple autocatalytic cycle of p steps (similar to the system in figure 2, where p=4). At each step, the legitimate reaction leads to the next cycle intermediate, and a number of side reactions drain the system. The latter give rise to all sorts of unwanted by-products. Let the specificity of a reaction at step i be si, which is the rate of legitimate reaction divided by the total rate of all (legitimate+side) reactions. Successful growth of the cycle is guaranteed if
| (2.4) |
or if we calculate with the geometric mean σ of the specificities
| (2.5) |
Figure 1.
The autocatalytic core or seed of the formose reaction (Fernando et al. 2005). Each circle represents a chemical group including one carbon atom.
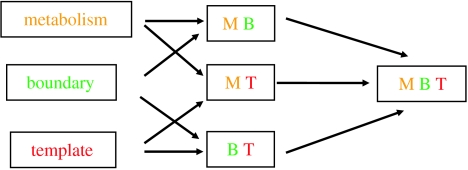
Figure 2.
Elementary combinatorics of infrabiological systems (Fernando et al. 2005). The chemoton is a biological minimal system comprising three qualitatively different subsystems (metabolism, membrane and template).
This shows that the viable system size p increases hyperbolically with specificity. Let us apply Eigen's (1971) full dynamical formalism to this problem (Szathmáry 2002) by assuming that there can be a number of alternative cycles such as the formose reaction that occasionally can produce each other's intermediates:
| (2.6) |
where xi is the concentration of species i; Ri, the rate of replication irrespective of the correctness of the offspring; Qi, the fidelity of replication; Di, the rate of spontaneous decomposition; wij, the mutation rate from species j to species i; and F, an outflow ensuring that the total concentration remains unity. Here, the different ‘species’ mean the catalytic seeds of different alternative cycles (if their existence is feasible, see below), and ‘mutation’ refers to the ‘macromutation’, producing an intermediate of another autocatalytic cycle. Spontaneous decay corresponds to irreversible side reactions; in the case of DNA, it means damage (rather than mutation; damaged DNA is chemically no longer DNA).
When is species i viable? It means that it can increase in concentration when rare. If we forget about selection of, and mutations to, this species for a moment, from equation (2.6) we obtain
| (2.7) |
which after rearrangement yields
| (2.8) |
where it also holds that
| (2.9) |
Lack of enzymatic catalysis implies that the decay rate is rather high. Inequalities (2.8) and (2.9) suggest that copying fidelity must be high. Fortunately, this fits, since mutations are expected to be very rare in the systems composed of cycles of small molecules (most fluctuations cannot propagate their own kind). Thus for autocatalytic cycles, damage is the most severe hurdle (Szathmáry 2000). The same considerations necessarily apply to the fittest cycles. If they coexist, ecology tells us that they must occupy different niches in abstract space, such as requiring different combination of raw materials.
An alternative way of maintaining a variety of cycles is a high mutation rate (low copying fidelity). This is true, but low copying fidelity does not allow the selection for the fittest, because the system gets below the error threshold of replication (see §2a). In such a case, the cycles would cease to be selectable individuals: they would rather form a single, un-evolvable network.
Orgel (1992) called attention to the fact that the intermediates of formose reaction are not informational replicators. In the prebiotic context, Wächtershäuser (1992) called attention to the possibility that there could be, in principle, a limited set of metabolic replicators. These replicators could have limited heredity, allowing some evolution by natural selection. This possibility is intriguing, but it is without any direct experimental support at present: nobody has seen a metabolic replicator, other than the formose reaction, that would run without enzymes. In contemporary systems, such cycles (the Calvin cycle, the reductive citric acid cycle) are well above the damage threshold outlined here, owing to the rate-enhancing effect of evolved enzymes. Thus, the requisite degree of metabolic channelling is one of the biggest (if not the biggest) hurdles of the origin of life.
3. Infrabiological systems and the lipid world scenario
We do not know where RNA came from. Some people think that the first replicators were not even template-based; as we shall see reproducing compartments (vesicles, micelles) are favoured by some. Others see the crucial steps in the linking of different autocatalytic systems that ultimately could evolve into primitive living systems.
(a) Infrabiological systems
Gánti (e.g. 2003) emphasized that contemporary living systems always have: (i) some metabolic subsystem, (ii) some systems for heritable control and (iii) some boundary system to keep the component together. So I consider it unlikely that a chemical system satisfying all the constraints from this abstraction could have appeared just out of chemical chaos. This observation led to the formulation of the concept of infrabiological systems (Szathmáry 2005; Fernando et al. 2005). Infrabiological systems always lack one of the key components just listed. For example, in the original formulation of Ganti (1971), a model of minimal life did not include a boundary system. The combination of a metabolic cycle and a membrane was conceived also by Gánti (1978), and called a self-reproducing microsphere. In contrast, Szostak et al. (2001) conceived a protocell-like entity with a boundary and template replication but no metabolic subsystem. Such systems show a crucial subset of necessary biological phenomena. The three subsystems can be combined to yield three different doublet systems (figure 2).
(b) Composomes and the graded autocatalytic replication domain model
An interesting line of research has been initiated by Doron Lancet with his group, conveniently referred to as the ‘lipid world’ scenario (Segré et al. 2001a). The basic idea is as follows. We know that lipids (more generally, amphiphilic compounds with a hydrophobic tail and a hydrophilic head) tend to form supramolecular structures, such as bilayers, micelles and vesicles. They can grow autocatalytically. Now imagine that we have a mixture of molecules in any one vesicle. Some of them may act as catalysts of certain reactions. It is theoretically possible that some will catalyse their own incorporation (direct autocatalysis), or there will be a gang of molecules each exerting some catalytic function; thus as a net result, the incorporation of all members of the gang is ensured by the gang (reflexive autocatalysis). If this idea holds water, membrane heredity in the lipid world, and natural selection of vesicles without a genetic subsystem, would be feasible. The different, reflexively autocatalytic gangs would constitute compositional genomes or ‘composomes’ (Segré et al. 2001b). Note that the model does not deal with the formation of the lipid constituents: they are assumed to be there in the surrounding soup.
Now, there is nothing mysterious about compositional genomes in the first place. Although relying on direct autocatalysis at the molecular level, the genome of the stochastic corrector (see §7) is also a compositional genome in which the genes are unlinked and the genome is characterized by gene composition. Formally, each protocell can be characterized by a genome vector with entries denoting the number of copies of the ith gene in that vesicle. The change in this number is a stochastic process, which can be characterized by mean and variance. A crucial difference is that, in the stochastic corrector model, we are dealing with a bag of template replicators: there are no genes in Lancet's model.
A similar approach is possible while considering questions in the lipid world; however, the issue is complicated by the fact that we need to tackle the problem of reflexive autocatalysis. This has also precedence in the literature: the reflexively autocatalytic protein networks (e.g. Kauffman 1993) are perhaps the best-known example. I hasten to point out that nobody has seen real reflexively autocatalytic protein sets. Let us see whether one can be more hopeful regarding autocatalytic lipid sets.
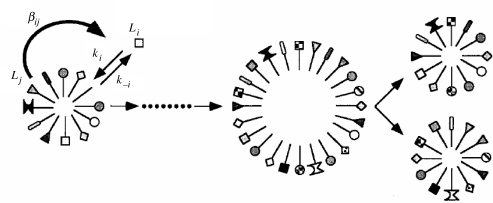
The process imagined is shown in figure 3. It displays a reflexively autocatalytic micelle with many components. The incorporation of amphiphile Li may be catalysed by amphiphile Lj at rate enhancement βij (the ratio of catalysed and uncatalysed reaction rates). The crucial question is this: where can one obtain the values of βij, considering the fact that no such system has been realized so far (the experimental cases are all directly autocatalytic and show no heredity; see Fernando et al. 2005 for review)? The authors suggest translating the model developed for molecular recognition between receptors and ligands (Segré et al. 1998). If catalysis depends on recognition of substrate by catalyst, the reasoning is sound implying that catalysis is a graded phenomenon. From this empirically constrained theoretical distribution, the authors obtain the distribution of βij values in their model.
Figure 3.
The graded autocatalytic replication domain or composome model: catalysed micelle growth and fission (Segré et al. 2001a,b). Li and Lj molecules are different amphiphilic compounds, ki and k−i are rate constants for spontaneous insertion and emigration of amphiphile Li, and βij is the rate enhancement of getting in and out of this molecule from the micelle, catalysed by Lj. Note that the model does not deal with the primary origin of Li molecules per se.
It is imagined that every micelle (or vesicle) is a sample with replacement of a set of possible lipid molecules. Some samples will contain mutually autocatalytic gangs, but not others. The latter ones will not be able to grow. The former will grow and then fragment/divide by some spontaneous process. Micelles containing more efficient gangs (characterized by higher βij values) will take over. Such sets have some heredity; the gangs maintain and propagate their identity by virtue of their mutual catalytic activity.
What are the major concerns apart from the lack of an experimental basis (at this moment) of this model? In the light of the foregoing, I see the following difficulties:
This model works only if the βij values are drawn from a lognormal, rather than a normal distribution. In the latter case, there is no interesting composome population.
The absolute magnitude of the βij values will also matter. Side reactions, as in many other prebiotic models, are neglected in the lipid world scenario. If the catalytic values are too low, then composomes may shrink below the decay threshold, even if without decay very interesting dynamics may unfold.
Even if the decay threshold is not reached, composomal replication may be so inaccurate that fitter composomes cannot be maintained by selection; thus the system may be above the corresponding error threshold.
I hope the fascinating scenario of the lipid world scenario will be complemented by theoretical investigations along these lines. Experimental validation is another formidable problem.
(c) Limited heredity in composomes
Contemporary DNA-based organisms have an unlimited hereditary potential, since the number of types that one can construct from the purely informational point of view greatly exceeds the number of individuals that the Earth can maintain. What is the hereditary potential of composomes? They can have limited heredity only (Szathmáry 2000). First of all, it is only the composition rather than the steric configuration of the system that is maintained. In order to appreciate this point, consider n types of molecules that we use to build our replicator of size k. In the case of template (digital, see later) replication, all possible sequences are potential replicators; Hence, their number is given by
| (3.1) |
as it follows from elementary combinatorics. In the case of ensemble replicators, the positions do not matter and hence the upper bound for the number of possible types is
| (3.2) |
This is clearly an upper bound since every possible subset cannot be realized by the alternative attractors associated with the system. For the same n and k, Ns is always larger than Nc, usually by orders of magnitude. Indeed, by the application of the Stirling formula for factorials, one can deduce an approximate equation for the proportion of the number of types
| (3.3) |
which, for sufficiently large n and k, further approximates to
| (3.4) |
Note that the number of attractors for such collective replicators has not been analytically calculated yet. In any case, the ratio (3.4) showing the advantage of modular template replicators is definitely underestimated. A satisfactory answer must take two considerations into account: (i) the number of attractors in sets of unlimited size (Kauffman 1993) and (ii) finite size k for realistic systems (Segré et al. 1998).
4. Parabolic growth, survival of everybody and the appearance of Darwinian selection
In the field of prebiotic evolution, non-conventional growth laws, such as hyperbolic and parabolic, have been widely discussed. Both represent departures from simple Malthusian growth: hyperbolic and parabolic growth are faster and slower than Malthusian growth, respectively. Hyperbolic growth was thought to be relevant for hypercycles (mutualistic molecular replicators), whereas parabolic growth was experimentally demonstrated to happen with small synthetic replicators. The consequences for selection in a competitive setting are remarkable: survival of the common for hyperbolic growth and survival of everybody for parabolic growth. In this section, I focus mainly on parabolic growth and its consequences.
(a) Growth laws and selection consequences
The simplest reproduction process is the binary fission of the parent object, of which the formal stoichiometry is
where A is a replicator, and S and W are source and waste materials, respectively (here I follow the treatment of Szathmáry & Maynard Smith, 1997). The associated kinetic equation describes a Malthusian growth process
| (4.1) |
which means that growth of x (the concentration of A) is exponential with a per capita rate constant k, provided the concentration of S is kept stationary. When two replicators with different rate constant grow together, the one with larger k will outgrow the other. This is, of course, elementary. For didactic purposes, let us express this outcome through the ratios of the growing concentrations
| (4.2) |
showing that even in a freely growing system, the worse growing population is diluted out in the limit. This is a very simple demonstration of differential survival.
Departures from this simple scheme are easily imaginable. A minimum complication is that two individuals are necessary to produce a third one (akin to sexual reproduction), such as:
and the associated growth equation reads
| (4.3) |
which is called hyperbolic growth, the selection consequences of which are very interesting (Eigen 1971). In order to see this, let us replace the exponent 2 by p and solve the equation by separation to obtain
| (4.4) |
When p>1, defining hyperbolic growth, the system has a finite escape time, i.e. it reaches infinite concentration in finite time. As it is easy to check, for p=2 the asymptote lies at t=1/[x(0)k]. The smaller the time of unbounded explosion, the larger x(0)k. Among the competitors, the one with the highest initial concentration times the growth rate constant wins. Thus, initial conditions also determine the outcome of selection and this phenomenon has been called the ‘survival of the common’, where intrinsic fitness is masked by the growth law (Michod 1983, 1984).
The relevance of hyperbolic growth and survival of the common may be as follows. Eigen (1971) proposed that the hypercycle might have been a link between solitary genes and bacterial genomes. It is a cycle of replicators in which any member catalyses the replication of the next. Each member undergoes a replication cycle as an autocatalyst, and there is the superimposed cyclic network of heterocatalytic aid, hence the term hypercycle. Under simplifying kinetic assumptions, the members of the hypercycle grow coherently and hyperbolically (e.g. Eigen 1971; Eigen & Schuster 1977). Thus, among a set of rival hypercycles, the already common is likely to win. This dynamics was claimed to have been important in the fixation of chirality and the genetic code (e.g. Küppers 1983). Yet this assumption is unwarranted (Szathmáry 1989a), briefly because: (i) parallel simple autocatalytic replication modifies invadability, (ii) stochastic effects allow uncommon, but intrinsically fitter hypercycles to invade and (iii) spatially distinct habitats would have allowed for diversity anyway. Thus, although hypercyclic systems may have played some role in prebiotic evolution, it is unlikely that their hyperbolic growth was very important (cf. Szathmáry et al. 1988).
Parabolic growth ensues when in the equation
| (4.5) |
the solution of which is also given by equation (4.4). When p=1/2, it is reduced to
| (4.6) |
which is why this type of growth is called parabolic.
Parabolic growth entails survival of everybody in a competitive situation. To see this, consider the relative concentration of two parabolically growing replicators in the same environment
| (4.7) |
and in the limit
| (4.8) |
Thus, ‘survival of everybody’ (Szathmáry 1991a) is guaranteed, as shown by selection equations in Szathmáry & Gladkih (1989).
But what kind of molecular mechanism could underlie such an odd type of growth? von Kiedrowski (1986) and Zielinski & Orgel (1987) were the first to show that oligonucleotide analogues follow a square-root growth law in the appropriate medium. The reason, in a simplified form, is as follows. A template molecule A reacts with the source materials whereby a new copy of A is made, which remains associated with the template.
Crucial is the ordering of the rate constants a≫b>c, i.e. association of two template molecules is faster than their dissociation, and replication per se is rate limiting. Note that the immediate product of copying is the replicationally inert AA complex. Thus, replication in this way is self-limiting. The higher the concentration of A, the stronger this self-limitation is. Note also that this type of replication is conservative: there is no material overlap between copy and template, and template and copy are exactly identical as well as complementary (this can be achieved by palindromes).
As it is apparent from the above reaction scheme, the rate of replication is determined by the concentration of free A, and at high enough total concentration of A (denoted by x) and AA (denoted by y), the former is negligible since association is stronger than dissociation. The formation and dissociation of AA are in quasi-equilibrium, thus
| (4.9) |
and therefore,
| (4.10) |
which is formally identical with equation (4.5).
Owing to self-limitation based on molecular complementarity, AA and BB complexes (where A and B are two different replicators) are stronger than AB complexes. Hence, each species limits its own growth more strongly: this condition for joint survival is also found in traditional Lotka–Volterra competitive systems. This is the ultimate cause for survival of the common in parabolic systems (Szathmáry 1991a).
In the meantime, several more replicators obeying the same type of growth dynamics have been constructed among others by Rebek (1994) and Sievers & von Kiedrowski (1995). (In the latter case, the single-stranded templates are not self-complementary.) A detailed kinetic theory for parabolic growth of minimal replicators was worked out by von Kiedrowski (1993). It seems that parabolic growth is a rather robust phenomenon among these replicators, although with the appropriate ‘molecular gymnastics’ nearly exponential growth can be achieved (Kindermann et al. 2005).
One of the important steps of prebiotic evolution must have been the emergence of replicators with exponential growth. Incidentally, this is very likely to have opened up the possibility of a transition from limited to unlimited heredity as well.
(b) A nontrivial consequence of exponential decay
Szathmáry & Gladkih (1989) realized that parabolic growth as expressed in equation (4.5) results in coexistence whenever replicators are in a competitive situation. The system they used was:
| (4.11) |
which implies a constraint of constant total population size (cf. Eigen 1971). The strange result of the analysis of this system was ‘survival of everybody’ (Szathmáry 1991) in contrast to the classical (Darwinian) case of exponential growth (p=1), where survival of the fittest prevails. This result was mathematically confirmed by Varga & Szathmáry (1997) who, by finding an appropriate Liapunov function, demonstrated that there was a single internal, globally stable rest point of the system (4.11).
Lifson & Lifson (1999) recently extended these findings by demonstrating that if single strands decompose by spontaneous (exponential) decay, coexistence is not possible any more and ‘selection of the unfittest’ sets in. Independently, von Kiedrowski (1998) announced that in a simulated chromatographic system of competing self-replicators natural selection could happen, despite the fact that this would not be possible in the spatially homogeneous case, modelled by equation (4.11).
Let us first point out that it is not the system (4.11) that the Lifsons modified. If you introduce decay rates into the model, you get
| (4.12) |
for which survival of everybody is still guaranteed, despite the specific decay rates di. Using essentially the original rationale of Szathmáry & Gladkih (1989) one finds that
| (4.13) |
wh4ich means that the time derivative is positive if the concentration xi is sufficiently low (Scheuring & Szathmáry 2001).
In their model, the Lifsons assume that ‘double strands do not replicate and are resistant to decomposition’ (cf. their equations (3.2) and (4.15)). Their assumption that double strands do not decompose at all is unrealistic. In the following, I review results by von Kiedrowski & Szathmáry (2000) that competitive coexistence is still possible under a range of parameter values for self-replicators with a parabolic growth tendency, even if decay of strands is taken into account.
(c) Theory before experiment: the chromatographized replicator model
A common problem of non-enzymatic artificial replicator systems is product inhibition leading to parabolic instead of exponential amplification. Exponential chemical replication of oligonucleotides was achieved by an iterative stepwise procedure, which employs the surface of a solid support and was called Surface Promoted Replication and Exponential Amplification of DNA analogues (SPREAD; Luther et al. 1998). I review theoretical insights (von Kiedrowski & Szathmáry 2000) into the design of an autonomous variant of the SPREAD procedure. The corresponding program simulates a given set of chemical reactions coupled to a chromatographic process, where the chromatographic column is treated as a series of connected cells. The crucial step is a template-directed reaction occurring at the surface: thus it is assumed that two parabolic replicators compete for their building blocks in the chromatographic column. A simplified semi-analytic treatment confirms that competing parabolic replicators, which spread on mineral surfaces are amenable for Darwinian selection under a wide range of parameter values.
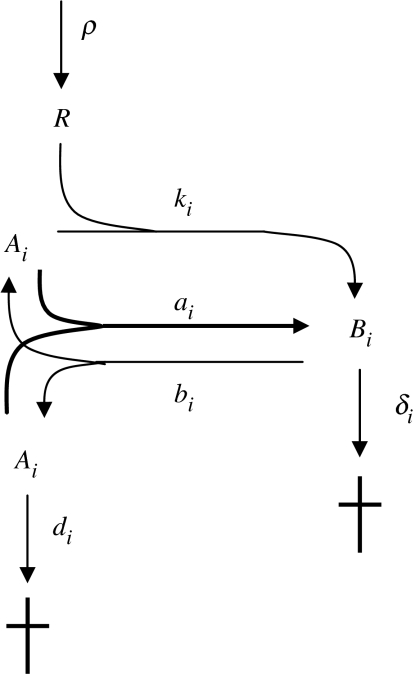
Now my aim is to demonstrate by a semi-analytically soluble simplified model that differential retention can lead to competitive exclusion (von Kiedrowski & Szathmáry 2000). Consider a single compartment with a constant nutrient (raw material) inflow and assume that single strands have a higher decay rate than double strands. This is meant to substitute for the higher retention of double strands on the chromatography column. The scheme of reactions is displayed in figure 4. For two species, we have the following ordinary differential equation system:
| (4.14) |
where R is the common resource and Ai, Bi are the single and double strands of species i, respectively (i=1, 2). We are interested in the conditions under which invasion by the inferior species when rare is not possible, i.e. we have competitive exclusion. A crucial relation is the following:
| (4.15) |
Figure 4.
Stoichiometric scheme of the simplified system with differential decay rates for the double and single strands (von Kiedrowski & Szathma´ry 2000). The resource R is fed into the system at a constant rate ρ. The assumption d≫δ corresponds to that of the more complicated case when the double strand is retained much more strongly than the single strand by the chromatography column.
Thus, when R1 maintained by species 1 alone satisfies condition (4.15), invasion by species 2 is possible, otherwise it is impossible. Obviously, if A2 is to invade, then the rate of its template ligation must be large and that of its decay must be small. A symmetric treatment applies to invasion by species 1 if species 2 is the resident one. The significant fact is that the threshold R1 depends on the decay rates of the single strand (d1) and the double strand (δ1) of the resident species 1 as well.
Competitive exclusion (survival of the fittest) is compatible with
| (4.16) |
but not the other way round. In the chromatographic case, this corresponds to a high retention factor for the double strand and low for the single strand. Note that an increase in δ easily throws the system into the region of coexistence.
I believe that the chromatographized replicator model is relevant to the origin of life on Earth. The chromatographic column is equivalent to a tunnel or a riverbed of minerals in which water containing the resources is continuously running through. Although our model, so far, refers to an isothermal reaction system, it can be easily extended to account for a gradient of increasing temperature along the direction of the column. As long as parabolic replicators need high temperatures whereas short replicators work at low temperatures (von Kiedrowski 1993), long replicators may grow from the consumption of shorter ones synthesized at the entry of the column where the temperature is low. The chromatographized replicator model can be simplified by means of attributing individual desorption rates to individual decay rates. Moreover, the findings from the simplified reaction model, viz. that both selection and coexistence can occur, has been independently confirmed by simulations based on the original model.
The case presented is an unusual one in that theory makes a clear prediction for experiment. Moreover, experimental realization of the model should be relatively straightforward.
5. Real ribozymes and a relaxed error threshold
The error threshold—the critical copying fidelity below which the fittest genotype deterministically disappears—for replication limits the length of the genome that can be maintained by selection; see equation (2.3). Primordial replication must have been error-prone, so early replicators are thought to have been necessarily short (Eigen 1971). The error threshold also depends on the fitness landscape. In an RNA world (Gilbert 1986), there will be many neutral and compensatory mutations that can raise the threshold, below which the functional phenotype, rather than a particular sequence, is still present. A comparative analysis of two extensively mutagenized ribozymes has shown that with a copying fidelity of 0.999 per digit per replication, the phenotypic error threshold rises well above 7000 nucleotides, which permits the selective maintenance of a functionally rich ribo-organism with a genome over 100 different genes the size of a tRNA (Kun et al. 2005a,b). This ‘only’ requires an order of magnitude improvement in the accuracy of in vitro generated polymerase ribozymes (Johnston et al. 2001; Müller & Bartel 2003). Incidentally, this genome size coincides with that estimated for a minimal cell achieved by top-down analysis (comparative analysis of the genomes of reduced organisms: Gil et al. 2004) minus the genes dealing with translation.
Eigen's insight of an error threshold quantifies the problem. Following (2.3), we have
| (5.1) |
where s=K/k is the so-called selective superiority of the fittest (master) sequence. In this simplified treatment, all mutants share the same replication rate, neutral mutations of and back mutations to the master are ignored.
The error threshold was first defined in relation to a particular genotype. However, it is obvious that in an RNA world there will be many neutral and compensatory mutations, which allow the preservation or the restoration of the fittest phenotype rather than of a single genotype. Other things being equal, this will modify the error threshold by increasing it (thus longer genomes will become maintainable). Since in an RNA world the functional ribozymes will have the strongest effect on fitness, one should gather the pertinent data from known ribozymes. As we shall see, there is just enough empirical evidence to formulate an encouraging statement.
To construct a fitness/functionality landscape of a ribozyme: (i) its secondary structure has to be experimentally determined, (ii) this secondary structure cannot contain a pseudo-knot, a special structural element that conventional RNA folding algorithms cannot satisfactorily cope with, (iii) mutagenesis experiments have to reveal all important sites and nucleotides and (iv) the size of the ribozyme should not be very long, otherwise any calculation would be practically unfeasible. The first requirement excludes most of the known ribozymes, since apart from the function only the sequence has been determined. The naturally occurring ribozymes generally fulfil the third requirement, but Hepatitis Delta Virus fails to meet the second requirement and Group I and II introns, as well as RNAase P, fail to meet the fourth. This leaves the hammerhead, the hairpin and the Neurospora VS ribozymes as possible candidates. Kun et al. (2005a) chose the hairpin and the Neurospora VS ribozymes for our study (figure 5). Both are relatively short, naturally occurring self-cleaving ribozymes, which can be divided into a trans-acting enzyme/substrate system where the trans-acting enzyme part does not contain a pseudo-knot.
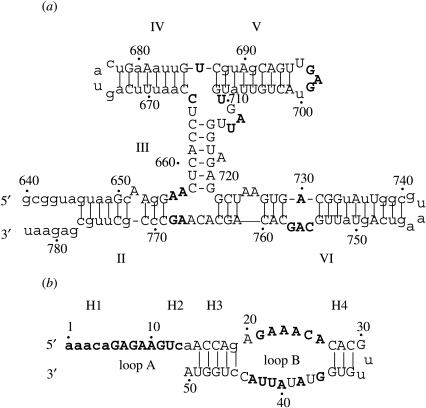
Figure 5.
Secondary structures of (a) Neurospora VS ribozyme and (b) hairpin ribozyme indicating different regions (Kun et al. 2005a,b). Position numbering follows standard convention. Capitalized nucleotides specify those sites that have been subjected to mutagenesis experiments, and enzymatic activities of mutants are available. A total of 183 mutants for the VS ribozyme affecting 83 out of 144 positions, excluding insertions and deletions, were considered. For the hairpin ribozyme, the survey was based on 142 mutants affecting 39 out of 50 positions of the ribozyme and some part of the substrate region. Nucleotides marked in bold are the critical sites.
The construction of the fitness/functionality landscape is based on four general observations: (i) the maintenance of the secondary structure is a major factor in retaining enzymatic activity, but the nature of most individual base pairs is not important and many can be reversed or replaced by a different pair without major loss of activity so long as a base pair is retained at a given position, (ii) the structure can have slight variations which in most cases manifest in some mismatch base pairs and/or some deletions or elongation in a helical region, (iii) there are critical regions in the molecule, where the nature of the base located there is also important and (iv) the effect of multiple mutations is multiplicative, i.e. the product of the activities of single mutants provides the activity of the multiple mutants.
From the fitness/functionality landscapes, the estimated phenotypic error thresholds are and for the VS and hairpin ribozymes, respectively, where is the effective mutation rate per nucleotide per replication. As expected, these figures are substantially higher than those inferred from fitness landscapes that do not take into account the secondary structure of the ribozymes but include information on single mutational effects.
This is the first time that the fitness landscape in terms of functionality has been inferred from real ribozymes (see also Kun et al. 2005b). The phenotypic error threshold thus inferred alleviates Eigen's paradox. This relates to the finding that the fitness landscapes are sufficiently similar. Inequality (5.1) cannot be used to assess the effect of the landscape on the error threshold owing to its restrictive preconditions. A recently derived expression (Takeuchi et al. 2005) offers a much more pertinent approximation:
| (5.2) |
where λ is the fraction of neutral single substitutions. For the VS ribozyme ν=144, q=0.947, λ=0.26; and for the hairpin ribozyme ν=50, q=0.856, λ=0.22. Thus, for ln s we obtain 5.761 and 5.957, respectively.
The fitness values obtained allow us to reconsider Eigen's paradox. Although it was shown that within-gene recombination could raise the error threshold to some extent, it has been unknown until recently what would be the required accuracy of a sufficient replicase ribozyme in a ribo-organism. Substituting an accuracy of q=0.999 in the lower bound of viral RNA replicases into inequality (5.2), and using the two obtained values for λ, we find that ν=7000–8000; namely, such a ribozyme could replicate a genome consisting of more than 100 different genes each of length 70 nucleotides or more than 70 different genes each of length 100. This would be sufficient to run a functionally rich ribo-organism, estimated to harbour about this number of genes (Jeffares et al. 1998). Incidentally, a recent analysis of a core minimal bacterial gene set gives about 200 genes (Gil et al. 2004). This shows that if we take away the genes coding for the whole contemporary translation system, we are again in the same ballpark.
The artificial template-dependent RNA polymerase ribozyme selected by Johnston et al. (2001) has an average fidelity q=0.97. Using formula (5.2) and the fitness/functionality landscape obtained for the VS and hairpin ribozymes (an admitted leap), it was concluded that the accuracy of this ribozyme would allow the maintenance of replicators with length around 250, which means that this ribozyme could replicate itself if other conditions (such as processivity) were favourable. In order to eliminate the burden of Eigen's paradox, a replicase with an error rate of 10−3 per nucleotide per replication might have been sufficient to provide the minimal life requirements in the RNA world.
6. Replicator evolution on the surface
It is a common experience in theoretical ecology and evolutionary biology that population structure promotes coexistence and favours the spread of altruism. Importantly, theoretical investigations in the field of early evolution have paved the way for such investigations to a considerable extent. Without the aim of completeness, I survey some interesting relevant examples.
(a) Metabolic ribozymes coexist on surfaces
Imagine a non-hypercyclic, so-called ‘metabolic’ system (cf. figure 45 in Eigen & Schuster 1978). Undoubtedly, we are here comfortably in the RNA world: we assume that informational replication and selection for enzymatic function has already been achieved. The templates are assumed to contribute to metabolism via enzymatic aid; metabolic products are in turn used up by the templates for replication at different rates. Although all templates contribute to metabolism (‘the common good’), they are able to use it with different efficiency. Thus in a spatially homogenous environment, competitive exclusion follows despite the metabolic coupling (Eigen & Schuster 1978).
Interesting selection dynamics occurs when molecules are bound to the surface without being washed away regularly. This problem was modelled by the use of ‘cellular automata’ (Czárán & Szathmáry 2000). Without becoming too technical, it suffices to say that each square of a grid is assumed to be occupied by a single molecule (template), or be empty. Templates can do two things: to replicate (put an offspring into a neighbouring empty cell if available) and hop away into empty sites nearby. Replication may depend on the composition of the few neighbouring cells. In the case of a hypercycle, for example, the template and a specimen of the preceding cycle member must be present in the same small area if replication of the former is to occur. This of course makes perfect chemical sense.
Boerlijst & Hogeweg (1991) simulated hypercycles on a surface exactly in this way. They found that rotating spirals on the surface appear, provided the hypercycle consists of more than four members. This is linked to the fact that such a hypercycle without population structure shows sustained oscillation in time. Each wing of a rotating spiral looks a bit like the arm of a galaxy, and is dominated by templates of the same membership in the hypercycle. Parasites are unable to kill the hypercycle in that system. This finding was attributed to the dynamics of spirals. Two questions emerge: Are spirals necessary? What happens if one models other systems in the same way (i.e. by cellular automata)?
The dynamics of the non-spatial version of the metabolic system looks as follows.
| (6.1) |
where xi stands for the concentrations of template Ii, and x is the vector of these concentrations. M(x) is a multiplicative function of the concentrations of all the templates, and Φ(x) is an outflow term representing a selection constraint (constant total concentration). This formulation is formally identical to that given by Eigen & Schuster (1978) for a ‘minimum model of primitive translation’. As they noted correctly, the fact that replication of any template is impossible without the presence of all the others does not prohibit the system from undergoing competitive exclusion: M(x) is same in all the equations, hence the system essentially behaves as a collection of Malthusian competitors, whose dynamics are influenced by a common time-dependent factor.
It is assumed that the replicators Ii have dual functionality: as templates they are necessary for their own replication (autocatalysis), and as ‘ribozymes’ (RNAs able to act as enzymes) they contribute to metabolism producing the monomers.
Now we assume that replication takes place on the surface of a mineral (possibly pyrite) substrate. The replicator molecules themselves are of a finite size; therefore the number of replicators bound to a unit area of the substrate is constrained. We consider a two-dimensional square lattice of binding sites as the scene of the replication–diffusion process; each of the sites can harbour a single macromolecule at most. The lattice is toroidal (the opposite edges of the grid are merged in both dimensions) to avoid edge effects.
At t=0, half of the sites are occupied by n different types of macromolecules (we call n the system size). The replicator types are equally abundant in the initial pattern and individual molecules are randomly assigned to sites. The other half of the sites are empty initially. Time is discrete; replication, decay and diffusion take place in each generation of the simulation.
The effect of monomer-producing metabolism is implicit in the model, itself directly acting on the replication process through a local metabolic function. It is local in the sense that its arguments are the copy numbers f(i) of replicator types i (i=1, …, n) within certain localities (neighbourhoods) of the lattice. In accordance with the assumption that the presence of a complete set of replicators is necessary for metabolism to produce monomers for replication, the metabolic function must be a multiplicative form of within-neighbourhood copy numbers f(i). A simple option for the concrete form of the metabolic function M(fs) at a site occupied by a replicator s is the geometric mean of the copy numbers fs(i) within the metabolic neighbourhood of s, i.e.
| (6.2) |
Note that M(fs) is zero if any of the replicator types is missing from the metabolic neighbourhood of s, and that the larger and more uniform the copy numbers of the different replicator types within the metabolic neighbourhood, the more efficient the metabolism at the given locality. By choosing (6.2) as the metabolic function, we assume that the conspecific replicators within the same neighbourhood help replication and that the focal replicator supports its own replication. The first assumption can be interpreted as metabolism being somewhat faster locally in the presence of more catalysts. The actual effect should be rather weak and it should vanish with the copy number increasing; this feature is properly reflected in the metabolic function (6.2): if a replicator type is already present in a replication neighbourhood, then its successive copies do not add too much to the replication chance of the focal template. Implicit in the second assumption is that the time-scale of metabolite diffusion out of the neighbourhood in which it was produced is longer than that of the catalysed reactions of metabolism. The ‘habitat’ of the reaction-diffusion system being an absorptive mineral surface is again straightforward to assume. The size of the metabolically effective neighbourhood is an implicit measure of metabolite and monomer diffusivity: larger neighbourhoods represent faster diffusion of the intermediate metabolites and the monomers.
Czárán & Szathmáry (2000) managed to show that given such a spatial setting, non-hypercyclic systems are once again viable alternatives. The fundamental difference between their model and that of Boerlijst & Hogeweg (1991) is the following: the dynamical link among the replicators is realized through a common metabolism, instead of the direct, intransitive hypercyclic coupling. Using the cellular automaton model of the metabolic system, the aim was to show that
metabolic coupling can lead to coexistence of replicators in spite of an inherent competitive tendency,
parasites cannot easily kill the whole system and
complexity can increase by natural selection.
The result that there is coexistence without any conspicuous pattern (i.e. something like spirals) is robust and counter-intuitive. It is owing to the inherent discreteness (i.e. the corpuscular nature of the replicator molecule populations) and spatial explicitness of the model, which grasp essential features of the living world in general, and macromolecular replicator systems in particular. An inferior (i.e. more slowly replicating) molecule type does not die out since there is an advantage of rarity in the system: a rare template is more likely to be complemented by a metabolically sufficient set of replicators in its neighbourhood than a common one.
(b) Reciprocal altruism on the rocks and the evolution of replicases
Although the question where the first RNA molecules came from is still unsolved, it is nevertheless assumed that catalytic RNA enzymes (ribozymes) with replicase function emerged at some stage of early evolution. Eigen's finding of the error threshold demonstrates that the length of templates maintained by selection is limited by the copying fidelity; therefore, other things being equal, an increase in template length is disadvantageous. On the contrary, longer molecules are expected to be better replicases—a feature not incorporated in the original model. An iterative scenario for longer and longer molecules with better and better replicase function has been suggested (James & Ellington 1999; Poole et al. 1999) and analysed mathematically (Scheuring 2000). A crucial open question is whether parasites (efficient templates that are inefficient replicases) can ruin the system. Absorption to mineral surfaces was hypothesized to help replicases find their useful colleagues in the immediate neighbourhood (Joyce & Orgel 1999). A cellular automaton simulation revealed that copying fidelity, replicase speed and template efficiency could increase by evolution, despite the presence of molecular parasites, essentially owing to reciprocal altruism on the surface, thus making the scenario for a gradual improvement of replicase function more plausible (Szabó et al. 2002).
Consider a population of macromolecules, adsorbed to a surface and built of four different monomers: A, B, C and D. Owing to their catalytic activity, macromolecules located on neighbouring sites of the surface can template-replicate each other, which means building a new macromolecule from free monomers by copying an existing one. In each replication process, two replicator molecules are involved: one is the template and the other acts as a replicase enzyme. We attribute two main properties to replication events, speed and fidelity, which in turn depend on three parameters of the two replicators involved in the process:
replicase activity expresses how fast the molecule can add a monomer to a primer while acting as a replicase,
replicase fidelity measures the accuracy of replication per monomer when the molecule acts as a replicase and
template efficiency defines an average ‘affinity’ of the molecule behaving as a template against others.
The authors assumed that these traits are in a three-way tradeoff: there were no free lunches. Replication speed depends on the activity of the replicase and the quality of the template: higher replicase activity and template efficiency result in faster replication. Given two neighbouring replicator molecules, L and M, on the surface, one of the two different replication events can occur between them: either L as replicase copies and M as a template, or the other way round. Mutations allowed not only point mutations but also additions and deletions of one nucleotide
The outcome was a bimodal population: efficient replicases evolved and short parasites could not ruin the system. This result, together with the chromatographized replicator model, emphasizes the importance of surface dynamics in prebiotic evolution. It also raises the idea that compartmentation offered by vesicles could have been an even more efficient means to evolve more efficient and accurate systems, a possibility to which I now turn.
7. Bags of genes: the stochastic corrector model
It is true that the hypercyclic link ensures indefinite ecological survival of all member replicators. However, problems arise when mutations are taken into account. In order to consider them, it is worthwhile to look at a diagram where auto- and heterocatalytic aids are functionally clearly separate, such as in a hypercycle with protein replicases (figure 6). Mutants providing stronger heterocatalytic aid to the next member are not selected. In contrast, increased autocatalysis is always selected, irrespective of its concomitant effect on heterocatalytic efficiency. This is the well-known problem of parasites in the hypercycle (Maynard Smith 1979). As Eigen et al. (1981) observed, putting hypercycles into reproducing compartments helps, because ‘good’ hypercycles (with efficient heterocatalysis) can be favoured over ‘bad’ ones. The following two questions arise out of this:
Are there other means whereby parasites can be selected against?
Are there non-hypercyclic systems that function well in a compartment context?
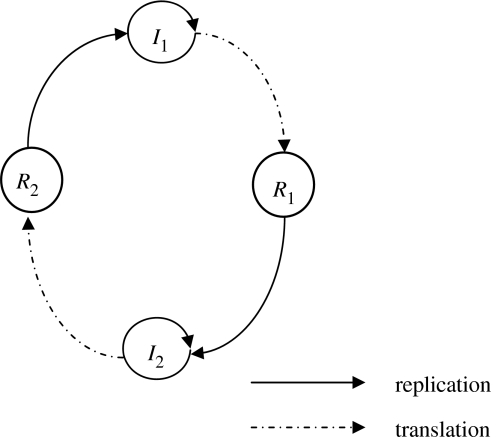
Figure 6.
The hypercycle with translation. Ri is a replicase protein enzyme coded for by gene Ii.
The answers turned out to be ‘yes’ to both of these questions; I discuss them below.
(a) Group selection of early replicators
The phase of evolution just outlined refers to the pre-cellular level. Later in evolution, protocells must have appeared. It turns out that cellularization offers the most natural, and at the same time most efficient, resolution to Eigen's paradox. It also leads to the appearance of linkage, i.e. the origin of chromosomes. The dynamics of genes encapsulated in a reproductive protocell is described by the stochastic corrector model (Szathmáry & Demeter 1987; Szathmáry 1989a,b; Grey et al. 1995; Zintzaras et al. 2002; Fontanari et al. 2006). It rests on the following assumptions (figure 7).
Templates contribute to the fitness of the protocell as a whole and there is an optimal proportion of the genes. Concretely, we assume that the genes encode enzymatic aid given to the intracellular metabolism.
Templates compete with each other within the same protocell. As before, replication rates may differ from gene to gene.
Replication of templates is described by stochastic means. Since the number of genes in any compartment is small (up to a few hundred), their growth is affected by the luck of the draw. Ecologists would express this as demographic stochasticity.
There is no individual regulation of template copy number per protocell.
Templates are assorted randomly into offspring cells upon protocell division.
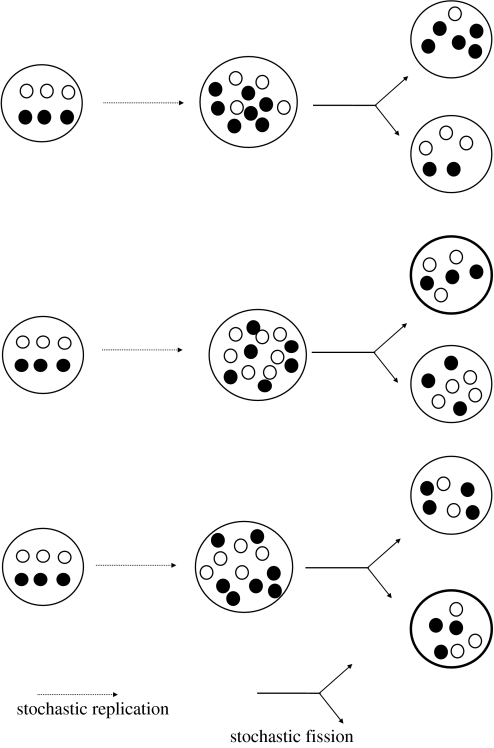
Figure 7.
The stochastic corrector model. Different templates (labelled by open and closed circles) contribute to the well being of the compartments (protocells) in that they catalyse steps of metabolism, for example. During protocell growth, templates replicate at differential expected rates stochastically. Upon division, there is chance assortment of templates into offspring compartments. Stochastic replication and reassortment generate variation among protocells on which natural selection at the compartment level can act and oppose to (correct) internal deterioration owing to within-cell competition.
I must emphasize that in the stochastic corrector model, the templates are not coupled to one another through a reflexive (intransitive) cycle of replicational aid, since it would be a hypercycle. Instead, we assume that they contribute to the ‘common good’ of the protocell by catalysing steps of its metabolism. Within each compartment, the templates are free to compete because they can reap the benefits of a common metabolism differently. (A similar situation can arise among chromosomes and plasmids in contemporary bacteria.) Despite the fact that templates compete, the two sources of stochasticity generate between-cell variation in template copy number on which natural selection (between protocells) can act. This is an efficient means of group selection of templates, since it is the protocells that are the groups obeying the stringent criteria: (i) there are many more groups than templates, (ii) each group has only one ancestor and (iii) there is no migration between the groups (cf. Leigh 1983). Grey et al. (1995) gave a fully rigorous re-examination of the stochastic corrector model. The two mentioned sources of stochasticity effectively lead to the correction of a malign within-protocell trend of harmful competition of the templates. It cannot be too strongly emphasized that the stochastic corrector is not, contrary to common misunderstanding, a hypercyclic system. Hypercycles need compartments but compartments can live without hypercycles. It is interesting to see that genuine group selection is likely to have aided a major transition from naked genes to protocells. Group structure is provided by the physical boundaries of cells.
Within the same context, the origin and establishment of chromosomes (linked genes) in the population have also been analysed (Maynard Smith & Szathmáry 1993). A chromosome consisting of two genes takes about twice as long to be replicated as the single genes. It turns out that chromosomes are strongly selected for at the cellular level even if they have this twofold within-cell disadvantage. Linkage reduces intracellular competition (genes are necessarily replicated simultaneously) as well as the risk of losing one gene by chance upon cell division (a gene is certain to find its complementing partner in the same offspring cell if it is linked to it). The molecular biology of the transition from genes to chromosomes has also been worked out (Szathmáry & Maynard Smith 1993).
(b) Sex and protocells
The results on coexistence leave one (one could say the original) question in the dark: does the error threshold increase or decrease in various systems? Although it was shown that the stochastic corrector model performs better than the compartmentalized hypercycle under a high error rate (Zintzaras et al. 2002), we still do not know the selectively maintainable genome size (or the number of different genes) in the stochastic corrector model. The results on real ribozymes (§5) alleviate, but do not solve, the problem. Lehman (2003) raised the issue that recombination, a frequently ignored player in models of early evolution, could have been crucial to build up primeval genomes of sizeable length. In the article that coined the phrase ‘the RNA world’, Gilbert (1986) already speculated that ‘the RNA molecules evolve in self-replicating patterns, using recombination and mutation to explore new functions and to adapt to new niches’. In this context, Riley & Lehman (2003) have shown that Tetrahymena and Azoarcus ribozymes can promote RNA recombination.
This capability of RNA recombination to potentially reduce the burden imposed by the error threshold has been recently analysed by Santos et al. (2004). They assumed that the recombination in protocells took place via copy-choice means, i.e. the replicase switched between RNA-like templates, as occurs frequently in RNA viruses and is crucial for retroviral replication during reverse transcription. The numerical results showed that there is a quite intricate interplay between mutation, recombination and gene redundancy, but the conclusion from the fitness function they used was that the informational content could have increased by 25% by keeping the same mutational load as that for a population without recombination.
The consequences of imperfect replication in vesicle models are puzzling. For small mutation rates, increased level of polyploidy favours the persistence of protocell lineages since the random loss of essential genes after fission is attenuated. However, for large mutation rates, the situation is reversed: those lineages with low levels of polyploidy are better able to cope with higher mutation rates, particularly when recombination is allowed. This means that gene redundancy was indeed costly. Therefore, selective forces favouring the linkage of genes to make the first chromosomes would eventually outweigh the advantage of faster replicating single genes, because linked genes are less likely to be lost by random assortment when protocells divide (Maynard Smith & Szathmáry 1993).
The role of the number of gene copies in a primitive cell was investigated by Koch (1984), who pointed out the existence of two conflicting forces: (i) higher copy numbers act as a safeguard against random loss of all copies of a gene but (ii) such copy numbers slow down adaptive evolution because a newly arisen favourable mutant is diluted out and cannot be ‘seen’ efficiently by natural selection acting on cells. He further observed that a moderately high (less than 100) copy number per gene is not only optimal, but also confers some additional evolvability by the ‘duplication and divergence’ scenario, as first emphasized by Ohno (1970).
Acknowledgments
This work was supported by the Hungarian Scientific Research Fund (OTKA T 047245) and the National Office for Research and Technology (NAP 2005/KCKHA005). Helpful comments by two anonymous referees are gratefully acknowledged.
Footnotes
One contribution of 19 to a Discussion Meeting Issue ‘Conditions for the emergence of life on the early Earth’.
References
- Bedau M.A, McCaskill J.S, Packard N.H, Rasmussen S, Adami C, Green D.G, Ikegami T, Kaneko K, Ray T.S. Open problems in artificial life. Artif. Life. 2000;6:363–376. doi: 10.1162/106454600300103683. doi:10.1162/106454600300103683 [DOI] [PubMed] [Google Scholar]
- Boerlijst M.C, Hogeweg P. Spiral wave structure in pre-biotic evolution—hypercycles stable against parasites. Physica. 1991;D48:17–28. [Google Scholar]
- Czárán T, Szathmáry E. Coexistence of replicators in prebiotic evolution. In: Dieckmann U, Law R, Metz J.A.J, editors. The geometry of ecological interactions: simplifying spatial complexity. IIASA and Cambridge University Press; Wien, Austria: 2000. pp. 116–134. [Google Scholar]
- Dawkins R. The selfish gene. Oxford University Press; Oxford, UK: 1976. [Google Scholar]
- Eigen M. Self-organization of matter and the evolution of biological macromolecules. Naturwissenschaften. 1971;58:465–523. doi: 10.1007/BF00623322. doi:10.1007/BF00623322 [DOI] [PubMed] [Google Scholar]
- Eigen M, Schuster P. The hypercycle: a principle of natural self-organization. Part A: emergence of the hypercycle. Naturwissenschaften. 1977;64:541–565. doi: 10.1007/BF00450633. doi:10.1007/BF00450633 [DOI] [PubMed] [Google Scholar]
- Eigen M, Schuster P. The hypercycle: a principle of natural self-organization. Part C: the realistic hypercycle. Naturwissenschaften. 1978;65:341–369. doi: 10.1007/BF00450633. doi:10.1007/BF00439699 [DOI] [PubMed] [Google Scholar]
- Eigen M, Schuster P, Gardiner W, Winkler-Oswatitsch R. The origin of genetic information. Sci. Am. 1981;244:78–94. doi: 10.1038/scientificamerican0481-88. [DOI] [PubMed] [Google Scholar]
- Fernando C, Santos M, Szathmáry E. Evolutionary potential and requirements for minimal protocells. Top. Curr. Chem. 2005;259:167–211. [Google Scholar]
- Fontanari J.F, Santos M, Szathmáry E. Coexistence and error propagation in pre-biotic vesicle models: a group selection approach. J. Theor. Biol. 2006;239:247–256. doi: 10.1016/j.jtbi.2005.08.039. doi:10.1016/j.jtbi.2005.08.039 [DOI] [PubMed] [Google Scholar]
- Gánti T. The principle of life (in Hungarian) Gondolat; Budapest, Hungary: 1971. [Google Scholar]
- Gánti T. The principle of life (in Hungarian) Gondolat; Budapest, Hungary: 1978. [Google Scholar]
- Gánti T. University park press; Baltimore, MD: 1979. A theory of biochemical supersystems. [Google Scholar]
- Gánti T. The principle of life. OMIKK; Budapest, Hungary: 1987. [Google Scholar]
- Gánti T. The principles of life. Oxford University Press; Oxford, UK: 2003. [Google Scholar]
- Gil R, Silva F.J, Peretó J, Moya A. Determination of the core of a minimal bacterial gene set. Microbiol. Mol. Biol. Rev. 2004;68:518–537. doi: 10.1128/MMBR.68.3.518-537.2004. doi:10.1128/MMBR.68.3.518-537.2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilbert W. The RNA world. Nature. 1986;319:818. doi:10.1038/319618a0 [Google Scholar]
- Grey D, Hutson V, Szathmáry E. A re-examination of the stochastic corrector model. Proc. R. Soc. B. 1995;262:29–35. [Google Scholar]
- Griesemer J. The units of evolutionary transition. Selection. 2000;1:67–80. doi:10.1556/Select.1.2000.1-3.7 [Google Scholar]
- Griesemer J. What is “epi” about epigenetics? Ann. NY Acad. Sci. 2002;981:97–110. doi: 10.1111/j.1749-6632.2002.tb04914.x. [DOI] [PubMed] [Google Scholar]
- Hull D.L. Individuality and selection. Annu. Rev. Ecol. Syst. 1980;11:311–332. doi:10.1146/annurev.es.11.110180.001523 [Google Scholar]
- James K.D, Ellington A.D. The fidelity of template-directed oligonucleotide ligation and the inevitability of polymerase function. Orig. Life Evol. Biosph. 1999;29:375–390. doi: 10.1023/a:1006544611320. doi:10.1023/A:1006544611320 [DOI] [PubMed] [Google Scholar]
- Jeffares D.C, Poole A.M, Penny D. Relics from the RNA world. J. Mol. Evol. 1998;46:18–36. doi: 10.1007/pl00006280. doi:10.1007/PL00006280 [DOI] [PubMed] [Google Scholar]
- Johnston W.K, Unrau P.J, Lawrence M.S, Glasen M.E, Bartel D.P. RNA-catalyzed RNA polymerization: accurate and general RNA-templated primer extension. Science. 2001;292:1319–1325. doi: 10.1126/science.1060786. doi:10.1126/science.1060786 [DOI] [PubMed] [Google Scholar]
- Joyce G.F, Orgel L.E. Prospects for understanding the origin of the RNA world. In: Gesteland R.F, Cech T.R, Atkins J.F, editors. The RNA world. 2nd edn. Cold Spring Harbor Lab. Press; Plainview, NY: 1999. pp. 49–77. [Google Scholar]
- Kauffman S.A. The origins of order. Oxford University Press; Oxford, UK: 1993. [Google Scholar]
- von Kiedrowski G. A self-replicating hexadeoxy nucleotide. Angew. Chem. Int. Ed. Engl. 1986;25:932–935. doi:10.1002/anie.198609322 [Google Scholar]
- von Kiedrowski G. Mimimal replicator theory I: parabolic versus exponential growth. Bioorg. Chem. Frontiers. 1993;3:113–146. [Google Scholar]
- von Kiedrowski, G. 1998 120. Versammlung Deutscher A¨rtze und Nafurforscher Berlin, 21st September 1998.
- von Kiedrowski G, Szathmáry E. Selection versus coexistence of parabolic replicators spreading on surfaces. Selection. 2000;1:173–179. doi:10.1556/Select.1.2000.1-3.17 [Google Scholar]
- Kindermann M, Stahl I, Reimold M, Pankau W.M, von Kiedrowski G. Systems chemistry: kinetic and computational analysis of a nearly exponential organic replicator. Angew. Chem. 2005;117:6908–6913. doi: 10.1002/anie.200501527. doi:10.1002/ange.200501527 Angew. Chem. Int. Ed. Engl. 44: 6750–6755. [DOI] [PubMed] [Google Scholar]
- King G.A.M. Recycling, reproduction, and life's origin. BioSystems. 1982;15:89–97. doi: 10.1016/0303-2647(82)90022-3. doi:10.1016/0303-2647(82)90022-3 [DOI] [PubMed] [Google Scholar]
- King G.A.M. Was there a prebiotic soup? J. Theor. Biol. 1986;123:493–498. doi:10.1016/S0022-5193(86)80216-8 [Google Scholar]
- Koch A.L. Evolution vs the number of gene copies per primitive cell. J. Mol. Evol. 1984;20:71–76. doi: 10.1007/BF02101988. doi:10.1007/BF02101988 [DOI] [PubMed] [Google Scholar]
- Kun Á, Santos M, Szathmáry E. Real ribozymes suggest a relaxed error threshold. Nat. Genet. 2005a;37:1008–1011. doi: 10.1038/ng1621. doi:10.1038/ng1621 [DOI] [PubMed] [Google Scholar]
- Kun A, Maurel M.-C, Santos M, Szathmáry E. Fitness landscapes, error thresholds, and cofactors in aptamer evolution. In: Klussmann S, editor. The Aptamer handbook. Wiley-Vch; Weinheim, Germany: 2005. pp. 54–92. [Google Scholar]
- Küppers B.-O. Springer; Berlin, Germany: 1983. Molecular theory of evolution. [Google Scholar]
- Lehman N. A case for the extreme antiquity of recombination. J. Mol. Evol. 2003;56:770–777. doi: 10.1007/s00239-003-2454-1. doi:10.1007/s00239-003-2454-1 [DOI] [PubMed] [Google Scholar]
- Leigh E.G. When does the good of the group override the advantage of the individual? Proc. Natl Acad. Sci. USA. 1983;80:2985–2989. doi: 10.1073/pnas.80.10.2985. doi:10.1073/pnas.80.10.2985 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lifson S, Lifson H. Models of prebiotic replication: survival of the fittest versus extinction of the unfittest. J. Theor. Biol. 1999;199:425–433. doi: 10.1006/jtbi.1999.0969. doi:10.1006/jtbi.1999.0969 [DOI] [PubMed] [Google Scholar]
- Luther A, Brandsch R, von Kiedrowski G. Surface-promoted replication and exponential amplification of DNA analogues. Nature. 1998;396:245–248. doi: 10.1038/24343. doi:10.1038/24343 [DOI] [PubMed] [Google Scholar]
- Maynard Smith J. Hypercycles and the origin of life. Nature. 1979;280:445–446. doi: 10.1038/280445a0. doi:10.1038/280445a0 [DOI] [PubMed] [Google Scholar]
- Maynard Smith J. Models of evolution. Proc. R. Soc. B. 1983;219:315–325. [Google Scholar]
- Maynard Smith J. Oxford University Press; Oxford, UK: 1986. The problems of biology. [Google Scholar]
- Maynard Smith J, Szathmáry E. The origin of chromosomes I. Selection for linkage. J. Theor. Biol. 1993;164:437–446. doi: 10.1006/jtbi.1993.1165. doi:10.1006/jtbi.1993.1165 [DOI] [PubMed] [Google Scholar]
- Maynard Smith J, Szathmáry E. The major transitions in evolution. Freeman & Co; Oxford, UK: 1995. [Google Scholar]
- Michod R. Population biology of the first replicators: on the origin of genotype, phenotype, and organism. Am. Zool. 1983;23:5–14. [Google Scholar]
- Michod R. Constraints on adaptation, with special reference to social behaviour. In: Price P.W, Slobodchikoff C.N, Gaud W.S, editors. A new ecology. Wiley; New York, NY: 1984. pp. 253–278. [Google Scholar]
- Müller U.F, Bartel D.P. Substrate 2′-hydroxyl groups required for ribozyme-catalyzed polymerization. Chem. Biol. 2003;10:799–806. doi: 10.1016/s1074-5521(03)00171-6. doi:10.1016/S1074-5521(03)00171-6 [DOI] [PubMed] [Google Scholar]
- Ohno S. Springer; Berlin, Germany: 1970. Evolution by gene duplication. [Google Scholar]
- Oparin A.I. Life, its nature, origin and development. Academic Press; New York, NY: 1961. [Google Scholar]
- Orgel L.E. Molecular replication. Nature. 1992;358:203–209. doi: 10.1038/358203a0. doi:10.1038/358203a0 [DOI] [PubMed] [Google Scholar]
- Poole A, Jeffares D, Penny D. Early evolution: the new kids on the block. BioEssays. 1999;21:880. doi: 10.1002/(SICI)1521-1878(199910)21:10<880::AID-BIES11>3.0.CO;2-P. doi:10.1002/(SICI)1521-1878(199910)21:10<880::AID-BIES11>3.0.CO;2-P [DOI] [PubMed] [Google Scholar]
- Poon A, Chao L. Drift increases the advantage of sex in RNA bacteriophage Phi6. Genetics. 2004;166:19–24. doi: 10.1534/genetics.166.1.19. doi:10.1534/genetics.166.1.19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rebek J. Synthetic self-replicating molecules. Sci. Am. 1994;271:34–40. [Google Scholar]
- Riley C.A, Lehman N. Generalized RNA-directed recombination of RNA. Chem. Biol. 2003;10:1233–1243. doi: 10.1016/j.chembiol.2003.11.015. doi:10.1016/j.chembiol.2003.11.015 [DOI] [PubMed] [Google Scholar]
- Santos M, Zintzaras E, Szathmáry E. Recombination in primeval genomes: a step forward but still a long leap from maintaining a sizeable genome. J. Mol. Evol. 2004;59:507–519. doi: 10.1007/s00239-004-2642-7. doi:10.1007/s00239-004-2642-7 [DOI] [PubMed] [Google Scholar]
- Scheuring I. Avoiding Catch-22 of early evolution by stepwise increase in copying fidelity. Selection. 2000;1:135–145. doi:10.1556/Select.1.2000.1-3.13 [Google Scholar]
- Scheuring I, Szathmáry E. Survival of replicators with parabolic growth tendency and exponential decay. J. Theor. Biol. 2001;212:99–105. doi: 10.1006/jtbi.2001.2360. doi:10.1006/jtbi.2001.2360 [DOI] [PubMed] [Google Scholar]
- Segré D, Lancet D, Kedem O, Pilpel Y. Graded autocatalysis replication domain (GARD): kinetic analysis of self-replication in mutually catalytic sets. Orig. Life Evol. Biosph. 1998;28:501–514. [PubMed] [Google Scholar]
- Segré D, Ben-Eli D, Deamer D.W, Lancet D. The lipid world. Orig. Life Evol. Biosph. 2001a;31:119–145. doi: 10.1023/a:1006746807104. [DOI] [PubMed] [Google Scholar]
- Segré D, Shenhav B, Kafri R, Lancet D. The molecular roots of compositional inheritance. J. Theor. Biol. 2001b;213:481–491. doi: 10.1006/jtbi.2001.2440. [DOI] [PubMed] [Google Scholar]
- Shapiro, R. 1986 A skeptic's guide to the creation of life on Earth New York, NY: Summit Books.
- Sievers D, von Kiedrowski G. Self-replication of complementary nucleotide-based oligomers. Nature. 1995;369:221–224. doi: 10.1038/369221a0. doi:10.1038/369221a0 [DOI] [PubMed] [Google Scholar]
- Szabó P, Scheuring I, Czárán T, Szathmáry E. In silico simulations reveal that replicators with limited dispersal evolve towards higher efficiency and fidelity. Nature. 2002;420:360–363. doi: 10.1038/nature01187. [DOI] [PubMed] [Google Scholar]
- Szathmáry E. The emergence, maintenance, and transitions of the earliest evolutionary units. Oxf. Surv. Evol. Biol. 1989a;6:169–205. [Google Scholar]
- Szathmáry E. The integration of the earliest genetic information. Trends Ecol. Evol. 1989b;4:200–204. doi: 10.1016/0169-5347(89)90073-6. doi:10.1016/0169-5347(89)90073-6 [DOI] [PubMed] [Google Scholar]
- Szathmáry E. Simple growth laws and selection consequences. Trends Ecol. Evol. 1991;6:366–370. doi: 10.1016/0169-5347(91)90228-P. doi:10.1016/0169-5347(91)90228-P [DOI] [PubMed] [Google Scholar]
- Szathmáry E. A classification of replicators and lambda-calculus models of biological organization. Proc. R. Soc. B. 1995;260:279–286. doi: 10.1098/rspb.1995.0092. [DOI] [PubMed] [Google Scholar]
- Szathmáry E. The evolution of replicators. Phil. Trans. R. Soc. B. 2000;355:1669–1676. doi: 10.1098/rstb.2000.0730. doi:10.1098/rstb.2000.0730 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szathmáry E. Units of evolution and units of life. In: Pályi G, Zucchi L, Caglioti L, editors. Fundamentals of life. Elsevier; Paris, France: 2002. pp. 181–195. [Google Scholar]
- Szathmáry E. Life: in search of the simplest cell. Nature. 2005;433:469–470. doi: 10.1038/433469a. doi:10.1038/433469a [DOI] [PubMed] [Google Scholar]
- Szathmáry E, Demeter L. Group selection of early replicators and the origin of life. J. Theor. Biol. 1987;128:463–486. doi: 10.1016/s0022-5193(87)80191-1. [DOI] [PubMed] [Google Scholar]
- Szathmáry E, Gladkih I. Sub-exponential growth and coexistence of non-enzymatically replicating templates. J. Theor. Biol. 1989;138:55–58. doi: 10.1016/s0022-5193(89)80177-8. [DOI] [PubMed] [Google Scholar]
- Szathmáry E, Maynard Smith J. The origin of chromosomes II. Molecular mechanisms. J. Theor. Biol. 1993;164:447–454. doi: 10.1006/jtbi.1993.1166. doi:10.1006/jtbi.1993.1166 [DOI] [PubMed] [Google Scholar]
- Szathmáry E, Maynard Smith J. From replicators to reproducers: the first major transitions leading to life. J. Theor. Biol. 1997;187:555–571. doi: 10.1006/jtbi.1996.0389. doi:10.1006/jtbi.1996.0389 [DOI] [PubMed] [Google Scholar]
- Szathmáry E, Kotsis M, Scheuring I. Limits of hyperbolic growth and selection in molecular and biological populations. In: Hallam T.G, Gross L, Levin S, editors. Mathematical ecology. World Scientific; Singapore, Singapore: 1988. pp. 46–68. [Google Scholar]
- Szostak J.W, Bartel D.P, Luisi P.L. Synthesizing life. Nature. 2001;409:387–390. doi: 10.1038/35053176. doi:10.1038/35053176 [DOI] [PubMed] [Google Scholar]
- Takeuchi N, Poorthuis P.H, Hogeweg P. Phenotypic error threshold; additivity and epistasis in RNA evolution. BMC Evol. Biol. 2005;5:9. doi: 10.1186/1471-2148-5-9. doi:10.1186/1471-2148-5-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varga Z, Szathmáry E. An extremum principle for parabolic competition. Bull. Math. Biol. 1997;59:1145–1154. doi:10.1016/S0092-8240(97)00048-7 [Google Scholar]
- Wächtershäuser G. Groundworks for an evolutionary biochemistry: the iron–sulphur world. Prog. Biophys. Mol. Biol. 1992;58:85–201. doi: 10.1016/0079-6107(92)90022-x. doi:10.1016/0079-6107(92)90022-X [DOI] [PubMed] [Google Scholar]
- Zielinski W.S, Orgel L.E. Autocatalytic synthesis of a tetranucleotide analogue. Nature. 1987;327:346–347. doi: 10.1038/327346a0. doi:10.1038/327346a0 [DOI] [PubMed] [Google Scholar]
- Zintzaras E, Mauro S, Szathmáry E. Living under the challenge of information decay: the stochastic corrector model versus hypercycles. J. Theor. Biol. 2002;217:167–181. doi: 10.1006/jtbi.2002.3026. doi:10.1006/jtbi.2002.3026 [DOI] [PubMed] [Google Scholar]