Abstract
As millions of single-nucleotide polymorphisms (SNPs) have been identified and high-throughput genotyping technologies have been rapidly developed, large-scale genomewide association studies are soon within reach. However, since a genomewide association study involves a large number of SNPs it is therefore nearly impossible to ensure a genomewide significance level of 0.05 using the available statistics, although the multiple-test problems can be alleviated, but not sufficiently, by the use of tagging SNPs. One strategy to circumvent the multiple-test problem associated with genome-wide association tests is to develop novel test statistics with high power. In this report, we introduce several nonlinear tests, which are based on nonlinear transformation of allele or haplotype frequencies. We investigate the power of the nonlinear test statistics and demonstrate that under certain conditions, some nonlinear test statistics have much higher power than the standard  -test statistic. Type I error rates of the nonlinear tests are validated using simulation studies. We also show that a class of similarity measure-based test statistics is based on the quadratic function of allele or haplotype frequencies, and thus they belong to nonlinear tests. To evaluate their performance, the nonlinear test statistics are also applied to three real data sets. Our study shows that nonlinear test statistics have great potential in association studies of complex diseases.
-test statistic. Type I error rates of the nonlinear tests are validated using simulation studies. We also show that a class of similarity measure-based test statistics is based on the quadratic function of allele or haplotype frequencies, and thus they belong to nonlinear tests. To evaluate their performance, the nonlinear test statistics are also applied to three real data sets. Our study shows that nonlinear test statistics have great potential in association studies of complex diseases.
WITH the imminent completion of the HapMap Project providing a comprehensive catalog of common genetic variations in human populations (Altshuler and Clark 2005) and rapid development of technologies enabling efficient and economical genotyping of a large number of variants (Borsting et al. 2005), genomewide association studies will become practically feasible in the near future. However, a limit, which may keep genomewide association studies from realization, pertains to problems of a statistical nature. Considering the adjustment for millions of statistical tests, a stringent P-value of 10−6– has been suggested to ensure a genomewide significance level of 0.05 (Freimer and Sabatti 2004; Neale and Sham 2004; Wang et al. 2005). Although this problem can be alleviated by selecting and typing tag SNPs (Halldorsson et al. 2004; Ahmadi et al. 2005), the effect of such a strategy on the significance level is still limited. Therefore, developing novel test statistics with high power requires immediate consideration.
has been suggested to ensure a genomewide significance level of 0.05 (Freimer and Sabatti 2004; Neale and Sham 2004; Wang et al. 2005). Although this problem can be alleviated by selecting and typing tag SNPs (Halldorsson et al. 2004; Ahmadi et al. 2005), the effect of such a strategy on the significance level is still limited. Therefore, developing novel test statistics with high power requires immediate consideration.
The primary assumption for association studies is that a mutation (a disease allele) increases disease susceptibility. Under this assumption, one expects that the disease allele will occur more frequently in the affected individuals (cases) than in the unaffected ones (controls) (Pritchard and Donnelly 2001). The standard  -test for association studies is to identify the disease locus by comparing the differences in allele or haplotype frequencies between the affected and unaffected individuals. More precisely, the
-test for association studies is to identify the disease locus by comparing the differences in allele or haplotype frequencies between the affected and unaffected individuals. More precisely, the  -statistic is a quadratic form of difference of allele or haplotype frequencies between the affected and unaffected individuals (Chapman and Wijsman 1998; Akey et al. 2001). A natural way to amplify differences in frequency is to conduct linear transformation of allele or haplotype frequencies in the currently used statistics for association studies. However, any statistics arising from linear transformation will not change the values of pretransformation statistics. We propose to use nonlinear transformations of allele or haplotype frequencies in cases (PA) and in controls (P), i.e.,
-statistic is a quadratic form of difference of allele or haplotype frequencies between the affected and unaffected individuals (Chapman and Wijsman 1998; Akey et al. 2001). A natural way to amplify differences in frequency is to conduct linear transformation of allele or haplotype frequencies in the currently used statistics for association studies. However, any statistics arising from linear transformation will not change the values of pretransformation statistics. We propose to use nonlinear transformations of allele or haplotype frequencies in cases (PA) and in controls (P), i.e.,  and f(P), with the expectation that statistics based on the difference
and f(P), with the expectation that statistics based on the difference  will be more powerful than those based on the difference
will be more powerful than those based on the difference  . For example, the case–control differential may be enhanced with some nonlinear transformations of allele or haplotype frequencies. Association tests with such nonlinear transformation are referred to as nonlinear association tests hereafter.
. For example, the case–control differential may be enhanced with some nonlinear transformations of allele or haplotype frequencies. Association tests with such nonlinear transformation are referred to as nonlinear association tests hereafter.
The main purpose of this report is to develop a general statistical framework of nonlinear association tests and to present several nonlinear test statistics for association studies. To accomplish this, we first study the properties of nonlinear transformations of allele or haplotype frequencies in cases and controls. We then study how to construct test statistics on the basis of the nonlinear transformations of allele or haplotype frequencies and to derive asymptotic distributions of the nonlinear test statistics under null and alternative hypotheses. Alternative to comparing differences in allele or haplotype frequencies, a recently developed class of association tests compares similarities of a genome region between affected and unaffected individuals (Tzeng et al. 2003; Zhang et al. 2003). Under the general statistical framework for nonlinear association tests, we show that many similarity measure-based test statistics are nonlinear association tests with quadratic transformation of allele or haplotype frequencies. Thus, we can unify the allele or haplotype frequency-based association tests and similarity measure-based association tests. Since different nonlinear tests may have different power, selection of nonlinear statistics is critical to a successful application of nonlinear tests to association studies. We compare the power of several nonlinear test statistics and uncover the relationship between the power of the nonlinear test statistics and the strength of nonlinearity used in the test statistics (Bates and Watts 1980). To demonstrate that amplification of the differences in allele or haplotype frequencies by nonlinear test statistics will not cause false positive problems, we study the type I error rates of the nonlinear test statistics by simulations. Finally, to evaluate the performance of the nonlinear test statistics for association studies, the presented nonlinear test statistics are applied to three real data examples.
METHODS
Nonlinear transformations of allele or haplotype frequencies:
The principle behind the standard  -test in case–control studies is to compare the difference in allele or haplotype frequencies between cases and controls. We expect that amplifying such a difference may improve the power to detect disease susceptibility genes. One strategy to amplify the difference is to nonlinearly transform the frequencies. The difference in the values of nonlinear function of allele or haplotype frequencies between cases and controls should be larger than the difference in original allele or haplotype frequencies. Therefore, our goal is to search for nonlinear transformations that meet this requirement. To achieve this goal, we first investigate the factors that would affect the difference in values of nonlinear function of allele or haplotype frequencies between the two populations. For convenience of presentation, we study only haplotypes. The results can be adapted easily for the alleles.
-test in case–control studies is to compare the difference in allele or haplotype frequencies between cases and controls. We expect that amplifying such a difference may improve the power to detect disease susceptibility genes. One strategy to amplify the difference is to nonlinearly transform the frequencies. The difference in the values of nonlinear function of allele or haplotype frequencies between cases and controls should be larger than the difference in original allele or haplotype frequencies. Therefore, our goal is to search for nonlinear transformations that meet this requirement. To achieve this goal, we first investigate the factors that would affect the difference in values of nonlinear function of allele or haplotype frequencies between the two populations. For convenience of presentation, we study only haplotypes. The results can be adapted easily for the alleles.
Consider two alleles D and d at the disease locus. Let D denote the disease allele and  , and
, and  be the penetrance of genotypes DD, Dd, and dd, respectively. Let
be the penetrance of genotypes DD, Dd, and dd, respectively. Let  be the prevalence of disease. Define
be the prevalence of disease. Define
 |
where  and
and  are the frequencies of alleles D and d, respectively. Suppose that K marker loci span m haplotypes
are the frequencies of alleles D and d, respectively. Suppose that K marker loci span m haplotypes  . Let
. Let  and
and  be the overall measures of linkage disequilibrium (LD) between haplotype
be the overall measures of linkage disequilibrium (LD) between haplotype  and disease allele D and allele d, respectively, and define
and disease allele D and allele d, respectively, and define
 |
where  and
and  are the frequencies of the haplotypes
are the frequencies of the haplotypes  and
and  , respectively (Xiong et al. 2003). It is known that
, respectively (Xiong et al. 2003). It is known that  , where
, where  and
and  are the frequencies of the haplotype
are the frequencies of the haplotype  in the cases and controls, respectively, and
in the cases and controls, respectively, and  (Zhao et al. 2005). Let
(Zhao et al. 2005). Let  be a nonlinear function of the haplotype frequency
be a nonlinear function of the haplotype frequency  . We now calculate the difference between the nonlinear transformation of the haplotype frequency in the affected individuals
. We now calculate the difference between the nonlinear transformation of the haplotype frequency in the affected individuals  and the nonlinear transformation of the haplotype frequency in the general population
and the nonlinear transformation of the haplotype frequency in the general population  . By Taylor's expansion, we can obtain
. By Taylor's expansion, we can obtain
 |
where  and
and  are the first and second derivatives of the function
are the first and second derivatives of the function  with respect to
with respect to  . This equation still holds if the haplotype frequencies are replaced by allele frequencies.
. This equation still holds if the haplotype frequencies are replaced by allele frequencies.
From the above equation, the difference between the nonlinear functions of the frequencies in cases and controls depends on the first and second derivatives of the function  with respect to
with respect to  as well as the overall measure of the LD between the haplotype
as well as the overall measure of the LD between the haplotype  and the disease allele D. If
and the disease allele D. If  , then we have
, then we have  , which implies that the absolute value of the difference in nonlinear functions of the haplotype frequencies between cases and controls is larger than that of the original frequency difference under this condition.
, which implies that the absolute value of the difference in nonlinear functions of the haplotype frequencies between cases and controls is larger than that of the original frequency difference under this condition.
Test statistics:
Assume that  affected individuals and
affected individuals and  unaffected individuals are sampled. Let
unaffected individuals are sampled. Let  and
and  be the estimators of frequencies of haplotype
be the estimators of frequencies of haplotype  in cases and controls, respectively. The allele or haplotype frequencies are asymptotically distributed as multivariate normal distributions
in cases and controls, respectively. The allele or haplotype frequencies are asymptotically distributed as multivariate normal distributions  and
and  , respectively, where
, respectively, where  ,
,  ,
,  , and
, and  .
.
Let  be a continuously differentiable nonlinear function with a nonzero differential at x. Let
be a continuously differentiable nonlinear function with a nonzero differential at x. Let  for
for  ,
,  ,
,  , and
, and  . Then, the random vectors X and Y are asymptotically distributed as multivariate normal distributions
. Then, the random vectors X and Y are asymptotically distributed as multivariate normal distributions  and
and  , respectively (Serfling 1980), where
, respectively (Serfling 1980), where  ,
,  ,
,  ,
,  ,
,  , and
, and  .
.
Define the matrix
 |
Let  be an estimator of the matrix
be an estimator of the matrix  . We propose the test statistic
. We propose the test statistic  to test the association of the alleles or haplotypes with disease,
to test the association of the alleles or haplotypes with disease,
 |
where  is the generalized inverse of matrix
is the generalized inverse of matrix  . The null hypothesis is that there is no association of alleles or haplotypes with the disease; i.e.,
. The null hypothesis is that there is no association of alleles or haplotypes with the disease; i.e.,  . Let
. Let  . Under the null hypothesis,
. Under the null hypothesis,  is asymptotically distributed as a central
is asymptotically distributed as a central  with r degrees of freedom (Greenwood and Nikulin 1996; Serfling 1980). The test statistic TN defines a class of nonlinear tests. Various nonlinear functions with some regularity can be used to construct the test statistic. Table 1 lists some of the nonlinear functions used in this study and their corresponding derivatives.
with r degrees of freedom (Greenwood and Nikulin 1996; Serfling 1980). The test statistic TN defines a class of nonlinear tests. Various nonlinear functions with some regularity can be used to construct the test statistic. Table 1 lists some of the nonlinear functions used in this study and their corresponding derivatives.
TABLE 1.
Some of the nonlinear transformations for allele or haplotype frequencies
| Function | Derivative |
|---|---|
| Entropy | |

|
 |
| Exponential | |

|
 |
| Quadratic | |

|
 |
| Reciprocal | |

|
 |
Similarity measure-based statistics are special cases of the nonlinear tests:
We often observe that affected individuals share common haplotypes in the region surrounding disease mutations more often than unaffected individuals (Fan and Lange 1998; Jorde 2000). There are two ways to quantify the excessive sharing of common haplotypes among affected individuals. One way is to measure differences in allele or haplotype frequencies between affected and unaffected individuals (Akey et al. 2001). Another way is to measure differences in similarity of the genome region between affected and unaffected individuals (Bourgain et al. 2001; Tzeng et al. 2003). In appendix b, we show that the similarity measure of the genome region is a quadratic function of allele or haplotype frequencies. Therefore, similarity measure-based statistics are nonlinear test statistics.
Analytic formulas for power calculation of the nonlinear tests:
To evaluate the performance of the nonlinear test for association studies, we need to calculate its power. The alternative hypothesis is that there is at least one allele or haplotype associated with the disease; i.e.,  . Under the alternative hypothesis, the test statistic
. Under the alternative hypothesis, the test statistic  is asymptotically distributed as a noncentral
is asymptotically distributed as a noncentral  with noncentrality parameter
with noncentrality parameter  , where
, where  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  , and
, and  .
.
The noncentrality parameter  can be approximated by
can be approximated by
 |
(appendix c), where
 |
and
 |
The matrix S measures the strength of the nonlinearity of the nonlinear transformation  (appendix c). Note that under the same alternative hypothesis, the traditional
(appendix c). Note that under the same alternative hypothesis, the traditional  -test statistic, which is defined as
-test statistic, which is defined as
 |
is a noncentral  -distribution with the noncentrality parameter
-distribution with the noncentrality parameter
 |
Comparing the noncentrality parameters  and
and  , we can see that the noncentrality parameter
, we can see that the noncentrality parameter  involves one more term S than the noncentrality parameter
involves one more term S than the noncentrality parameter  . The matrix S characterizes the nonlinearity of the nonlinear function. The power of the nonlinear test statistics depends on the strength of the nonlinearity of the nonlinear function through the matrix S. The matrix S is referred to as the strength matrix of the nonlinearity of the nonlinear function.
. The matrix S characterizes the nonlinearity of the nonlinear function. The power of the nonlinear test statistics depends on the strength of the nonlinearity of the nonlinear function through the matrix S. The matrix S is referred to as the strength matrix of the nonlinearity of the nonlinear function.
If the product terms of the haplotype frequencies in the variance–covariance matrices  and
and  are ignored, the matrices
are ignored, the matrices  and
and  can be approximated by
can be approximated by  and
and  . Then the noncentrality parameters
. Then the noncentrality parameters  and
and  will be further reduced to
will be further reduced to
 |
where  . The parameter
. The parameter  is proportional to the curvature of a nonlinear function (Bates and Watts 1980) and influences the noncentrality parameter
is proportional to the curvature of a nonlinear function (Bates and Watts 1980) and influences the noncentrality parameter  .
.
From the above formulas, we can see that both noncentrality parameters  and
and  depend on the frequencies of the allele or haplotypes, penetrance, the measure of the LD between the marker alleles or haplotypes, and the disease allele as well as sample size. In addition, the noncentrality parameter of nonlinear test
depend on the frequencies of the allele or haplotypes, penetrance, the measure of the LD between the marker alleles or haplotypes, and the disease allele as well as sample size. In addition, the noncentrality parameter of nonlinear test  also depends on the curvature, which measures the degree of nonlinearity of nonlinear function.
also depends on the curvature, which measures the degree of nonlinearity of nonlinear function.
RESULTS
Distribution of the nonlinear test statistics:
In the previous sections, we have shown that when the sample size is large enough to apply large sample theory, the nonlinear test statistics under the null hypothesis of no association are asymptotically distributed as a central  -distribution. To examine the validity of this statement, we performed a series of simulation studies. The computer program SNaP (Nothnagel 2002) was used to generate haplotypes of the sample individuals. Two data sets with a single haplotype block each were simulated. The first data set has two marker loci that generated four haplotypes with frequencies 0.2952, 0.2562, 0.1957, and 0.2529. The second data set has six marker loci that generated eight haplotypes with frequencies 0.1820, 0.1461, 0.1406, 0.1291, 0.1211, 0.1107, 0.0817, and 0.0887. For each data set, 20,000 individuals who were equally divided into cases and controls were generated in the general population.
-distribution. To examine the validity of this statement, we performed a series of simulation studies. The computer program SNaP (Nothnagel 2002) was used to generate haplotypes of the sample individuals. Two data sets with a single haplotype block each were simulated. The first data set has two marker loci that generated four haplotypes with frequencies 0.2952, 0.2562, 0.1957, and 0.2529. The second data set has six marker loci that generated eight haplotypes with frequencies 0.1820, 0.1461, 0.1406, 0.1291, 0.1211, 0.1107, 0.0817, and 0.0887. For each data set, 20,000 individuals who were equally divided into cases and controls were generated in the general population.
To examine whether the asymptotic results of the nonlinear test statistics still hold for small sample size under the null hypothesis of no association, 100–500 individuals were randomly sampled from each of the cases and controls. Ten thousand simulations were repeated for each of the nonlinear test statistics. In each simulation, the nonlinear test statistics were calculated. Table 2 shows that the estimated type I error rates (at the significance level 0.05) of the nonlinear test statistics were not appreciably different from the nominal level  .
.
TABLE 2.
Estimated type I error rates for the nonlinear test statistics (10,000 simulations)
| Sample size | Entropy | Exponential | Quadratic | Reciprocal |
|---|---|---|---|---|
Two-SNP haplotypes ( = 0.05) = 0.05) | ||||
| 100 | 0.0460 | 0.0514 | 0.0548 | 0.0544 |
| 200 | 0.0510 | 0.0508 | 0.0546 | 0.0544 |
| 300 | 0.0560 | 0.0486 | 0.0532 | 0.0490 |
| 400 | 0.0570 | 0.0476 | 0.0508 | 0.0538 |
| 500 | 0.0540 | 0.0500 | 0.0496 | 0.0524 |
Six-SNP haplotypes ( = 0.05) = 0.05) | ||||
| 100 | 0.0450 | 0.0530 | 0.0544 | 0.0522 |
| 200 | 0.0490 | 0.0508 | 0.0476 | 0.0490 |
| 300 | 0.0502 | 0.0478 | 0.0518 | 0.0508 |
| 400 | 0.0508 | 0.0488 | 0.0476 | 0.0508 |
| 500 | 0.0500 | 0.0498 | 0.0462 | 0.0512 |
Power of nonlinear test statistics and standard χ2-test statistic:
Power of a test statistic for association studies depends on the allele or haplotype frequencies at the marker loci and the frequency of the disease allele, measure of LD between the alleles or haplotypes at the marker loci and the disease allele, sample size, the disease model, and the measure of nonlinearity of the nonlinear function. To evaluate the performance of nonlinear tests, we compare the power of several nonlinear test statistics with that of the standard  -test statistic by both analytical method and simulation. The results are very similar. In this report, we present only the power calculation by analytical method.
-test statistic by both analytical method and simulation. The results are very similar. In this report, we present only the power calculation by analytical method.
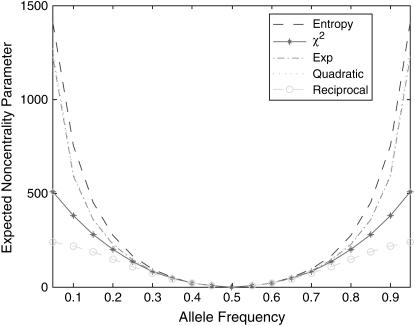
We first investigate the expected noncentrality parameters of nonlinear test statistics at the disease locus. We assume that frequencies of two alleles at the disease locus in controls are both equal to 0.5. Figure 1 plots the expected noncentrality parameters of the nonlinear test statistics and the standard  -test statistic as a function of frequency of disease allele in cases. From Figure 1 we can see three remarkable features. First, the expected noncentrality parameters of all test statistics increase as the difference in frequency of disease allele between cases and controls increases. Second, except for the reciprocal-based statistic that uses reciprocal function as nonlinear transformation of allele/haplotype frequencies, expected noncentrality parameters for all the other nonlinear test statistics are larger than that of the standard
-test statistic as a function of frequency of disease allele in cases. From Figure 1 we can see three remarkable features. First, the expected noncentrality parameters of all test statistics increase as the difference in frequency of disease allele between cases and controls increases. Second, except for the reciprocal-based statistic that uses reciprocal function as nonlinear transformation of allele/haplotype frequencies, expected noncentrality parameters for all the other nonlinear test statistics are larger than that of the standard  -test statistic. Third, except for the reciprocal-based statistic, expected noncentrality parameters for all the other nonlinear test statistics are almost indistinguishable.
-test statistic. Third, except for the reciprocal-based statistic, expected noncentrality parameters for all the other nonlinear test statistics are almost indistinguishable.
Figure 1.—
Expected noncentrality parameters of the nonlinear test statistics and the standard  -test statistic as a function of the frequency of the disease allele in cases, assuming that the frequencies of two alleles at the disease locus in the controls are both equal to 0.5.
-test statistic as a function of the frequency of the disease allele in cases, assuming that the frequencies of two alleles at the disease locus in the controls are both equal to 0.5.
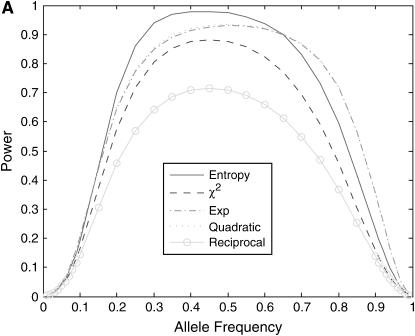
We then investigate the power of nonlinear test statistics at the disease locus. Figure 2 plots the power of the nonlinear test statistics and the standard  -test statistic as a function of disease allele frequency under three different disease models: (i) disease model with penetrance
-test statistic as a function of disease allele frequency under three different disease models: (i) disease model with penetrance  ,
,  , and
, and  ; (ii) disease model with penetrance
; (ii) disease model with penetrance  ,
,  , and
, and  ; and (iii) genotype relative risk model for r = 4, in which the genotype relative risk for genotypes Dd and DD is r and
; and (iii) genotype relative risk model for r = 4, in which the genotype relative risk for genotypes Dd and DD is r and  times greater than that for the genotype dd (Risch and Merikangas 1996). Several features emerge from Figure 2. First, power for most of the nonlinear test statistics is higher than that of the standard
times greater than that for the genotype dd (Risch and Merikangas 1996). Several features emerge from Figure 2. First, power for most of the nonlinear test statistics is higher than that of the standard  -test statistic, but power of the reciprocal-based test statistic is lower than that of the standard
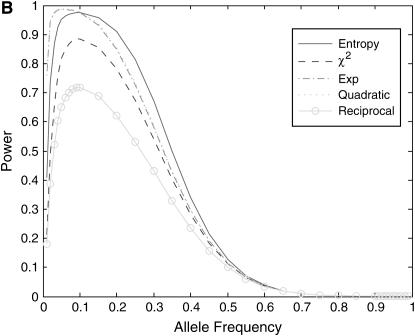
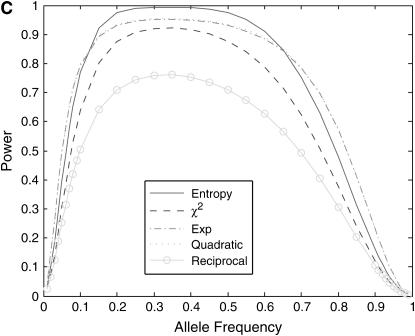
-test statistic, but power of the reciprocal-based test statistic is lower than that of the standard  -test statistic. The power curves of the exponential and quadratic functions are similar. Second, power of the nonlinear test statistics is influenced by disease models. Shapes of the nonlinear test statistics in disease model ii are different from those of the test statistics in disease models i and iii. Third, power of the test statistics depends on disease allele frequency. Shapes of the power curves in disease models i and iii are roughly bell; however, shapes of the power curves in disease model ii are skewed to the left.
-test statistic. The power curves of the exponential and quadratic functions are similar. Second, power of the nonlinear test statistics is influenced by disease models. Shapes of the nonlinear test statistics in disease model ii are different from those of the test statistics in disease models i and iii. Third, power of the test statistics depends on disease allele frequency. Shapes of the power curves in disease models i and iii are roughly bell; however, shapes of the power curves in disease model ii are skewed to the left.
Figure 2.—
(A) Power of the nonlinear test statistics and the standard  -test statistic at the disease locus with a significance level of 0.001 for a disease model with penetrance
-test statistic at the disease locus with a significance level of 0.001 for a disease model with penetrance  , and
, and  as a function of the disease allele frequency, assuming equal sample size (n =100) in both cases and controls. (B) Power of the nonlinear test statistics and the standard
as a function of the disease allele frequency, assuming equal sample size (n =100) in both cases and controls. (B) Power of the nonlinear test statistics and the standard  -test statistic at the disease locus with a significance level of 0.001 for a disease model with penetrance
-test statistic at the disease locus with a significance level of 0.001 for a disease model with penetrance  , and
, and  as a function of the disease allele frequency, assuming equal sample size (n = 100) in both cases and controls. (C) Power of the nonlinear test statistics and the standard
as a function of the disease allele frequency, assuming equal sample size (n = 100) in both cases and controls. (C) Power of the nonlinear test statistics and the standard  -test statistic at the disease locus with a significance level of 0.001 for a genotype relative risk disease model (r = 4) as a function of the disease allele frequency, assuming equal sample size (n = 100) in both cases and controls.
-test statistic at the disease locus with a significance level of 0.001 for a genotype relative risk disease model (r = 4) as a function of the disease allele frequency, assuming equal sample size (n = 100) in both cases and controls.
Real data examples:
Nonlinear test statistics are also applied to three real examples. The first example is a test of association of COMT haplotypes with schizophrenia. P-values of the nonlinear tests for testing associations of two-SNP haplotypes (generated from two SNP markers) and three-SNP haplotypes (generated from three SNP markers) with schizophrenia are presented in Table 3. Table 3 also includes P-values of the standard  -tests by Shifman et al. (2002). Improvement of the nonlinear tests over the standard
-tests by Shifman et al. (2002). Improvement of the nonlinear tests over the standard  -test varies among nonlinear tests and among haplotypes. The quadratic-based test has the largest improvement over the standard
-test varies among nonlinear tests and among haplotypes. The quadratic-based test has the largest improvement over the standard  -test when it is applied to three-SNP haplotypes. The P-value of the quadratic-based test is 4.0 ×
-test when it is applied to three-SNP haplotypes. The P-value of the quadratic-based test is 4.0 ×  , which is much smaller than the
, which is much smaller than the  obtained by the standard
obtained by the standard  -test.
-test.
TABLE 3.
Association tests for COMT haplotypes with schizophrenia
| Two-SNP haplotypesa
|
Three-SNP haplotypea: H4 | |||
|---|---|---|---|---|
| P-values for | H1 | H2 | H3 | |
| Entropy | 1.9e-009 | 2.7e-006 | 2.9e-006 | 1.5e-012 |
| Exponential | 1.2e-013 | 9.5e-009 | 1.8e-005 | 8.0e-014 |
| Quadratic | 7.5e-013 | 1.5e-008 | 9.5e-006 | 4.0e-014 |
| Reciprocal | 1.4e-010 | 8.3e-008 | 2.4e-006 | 2.9e-013 |
| χ2 b | 1.4e-004 | 5.7e-003 | 1.1e-003 | 4.5e-004 |
All data (including males and females) are used in the analysis.
H1, rs737865–rs165599; H2, rs737865–rs165688; H3, rs165599–rs165688; H4, rs165688–rs737865–rs165599.
P-values reported by Shifman et al. (2002).
The second example is a test of association of functional haplotypes in the promoter of the matrix metalloproteinase-2 (MMP-2) gene with esophageal cancer in the Chinese Han population (Yu et al. 2004). Two SNPs in the MMP-2 gene were typed in 527 esophageal cancer patients and 777 controls. P-values of the nonlinear tests are given in Table 4. We can see that P-values for most of the nonlinear tests are 10–100 times smaller than that of the standard  -test, whereas the P-value of the reciprocal-based test is almost the same as that of the standard
-test, whereas the P-value of the reciprocal-based test is almost the same as that of the standard  -test.
-test.
TABLE 4.
P-values of nonlinear tests for the MMP-2 gene with esophageal cancer
| Nonlinear transformation | P-value |
|---|---|
| Entropy | 3.2e-008 |
| Exponential | 2.3e-007 |
| Quadratic | 1.9e-007 |
| Reciprocal | 5.1e-006 |
| χ2 | 7.0e-006 |
To examine whether nonlinear test statistics show significant association or not when the standard  -test shows no significance, the proposed nonlinear test statistics were also applied to test association of a functional SNP in ZDHHC8 with schizophrenia in a Japanese case–control population (Saito et al. 2005). The results are summarized in Table 5. The data demonstrate that when the
-test shows no significance, the proposed nonlinear test statistics were also applied to test association of a functional SNP in ZDHHC8 with schizophrenia in a Japanese case–control population (Saito et al. 2005). The results are summarized in Table 5. The data demonstrate that when the  -test shows no association of the functional SNP in the ZDHHC8 gene with schizophrenia, nonlinear test statistics also show no evidence of association. P-values of the nonlinear test statistics are the same as that of the standard
-test shows no association of the functional SNP in the ZDHHC8 gene with schizophrenia, nonlinear test statistics also show no evidence of association. P-values of the nonlinear test statistics are the same as that of the standard  -test.
-test.
TABLE 5.
Association tests of a functional SNP in the ZDHHC8 gene with schizophrenia
| Sample
|
|||
|---|---|---|---|
| Nonlinear transformation | Male | Female | All |
| Entropy | 0.2586 | 0.7134 | 0.6177 |
| Exponential | 0.2586 | 0.7134 | 0.6177 |
| Quadratic | 0.2586 | 0.7134 | 0.6177 |
| Reciprocal | 0.2604 | 0.7135 | 0.6178 |
| χ2 | 0.2603 | 0.7135 | 0.6178 |
DISCUSSION
In the near future, genomewide association studies performing millions of statistical tests will be conducted. To ensure a genomewide significance level of 0.05, a stringent P-value is required for the statistical test. There is crucial need for increased efforts in developing new statistical methods that can achieve small P-values. As an attempt toward this direction, in this report, we present nonlinear tests for association studies.
The traditional  -test statistic is a quadratic function of the difference (
-test statistic is a quadratic function of the difference ( ) in allele or haplotype frequencies between the affected and unaffected individuals. Although the
) in allele or haplotype frequencies between the affected and unaffected individuals. Although the  -test statistic itself is a nonlinear function of allele or haplotype frequencies, its basic unit (
-test statistic itself is a nonlinear function of allele or haplotype frequencies, its basic unit ( ) is a linear transformation of allele or haplotype frequencies. If the difference in nonlinear transformation of allele or haplotype frequencies is larger than the difference in allele or haplotype frequencies, i.e.,
) is a linear transformation of allele or haplotype frequencies. If the difference in nonlinear transformation of allele or haplotype frequencies is larger than the difference in allele or haplotype frequencies, i.e.,  , where
, where  denotes a norm of the vector, then the statistics based on
denotes a norm of the vector, then the statistics based on  may have higher power than the statistics based on (
may have higher power than the statistics based on ( ). On the basis of this simple idea, we have developed a general statistical framework for nonlinear tests that provides basic procedures about how to construct test statistics using nonlinear transformations of allele or haplotype frequencies. We have showed that, in general, similarity measure-based statistics can be formulated as the differences in quadratic forms of allele or haplotype frequencies. Therefore, using the proposed statistical framework for nonlinear tests, we can derive many similarity measure-based statistics. As a by-product, nonlinear test theory can unify two classes of association tests: tests of the difference in allele or haplotype frequencies and tests based on a similarity measure of the genome region being tested.
). On the basis of this simple idea, we have developed a general statistical framework for nonlinear tests that provides basic procedures about how to construct test statistics using nonlinear transformations of allele or haplotype frequencies. We have showed that, in general, similarity measure-based statistics can be formulated as the differences in quadratic forms of allele or haplotype frequencies. Therefore, using the proposed statistical framework for nonlinear tests, we can derive many similarity measure-based statistics. As a by-product, nonlinear test theory can unify two classes of association tests: tests of the difference in allele or haplotype frequencies and tests based on a similarity measure of the genome region being tested.
The distributions of nonlinear test statistics are based on the asymptotic statistical theory of nonlinear transformations. We investigate the distributions of several nonlinear test statistics under the null hypothesis by simulation studies. Even with moderate sample size ( , distributions of the proposed nonlinear statistics are still close to central
, distributions of the proposed nonlinear statistics are still close to central  -distribution (data not shown). To validate the test statistics, we calculate the type I error rates of the presented nonlinear statistics by simulations. This showed that the type I error rates of nonlinear statistics were close to the nominal significance levels, which implies that the nonlinear tests for association study are valid in a single homogeneous population.
-distribution (data not shown). To validate the test statistics, we calculate the type I error rates of the presented nonlinear statistics by simulations. This showed that the type I error rates of nonlinear statistics were close to the nominal significance levels, which implies that the nonlinear tests for association study are valid in a single homogeneous population.
To evaluate the performance of the nonlinear test statistics, we compare the power of the nonlinear test statistics with that of the standard  -test statistic. To reveal the relationships between the power of the nonlinear test statistics and the measure of nonlinearity of nonlinear transformations, we developed analytical tools for calculations of the power of the test statistics. Power of the nonlinear statistics depends on several parameters such as disease model, allele or haplotype frequencies, measure of LD between the allele or haplotype and disease allele, and the measure of nonlinearity of the nonlinear transformations of the allele or haplotype frequencies. We showed that, in many cases, most of the studied nonlinear test statistics have higher power than the standard
-test statistic. To reveal the relationships between the power of the nonlinear test statistics and the measure of nonlinearity of nonlinear transformations, we developed analytical tools for calculations of the power of the test statistics. Power of the nonlinear statistics depends on several parameters such as disease model, allele or haplotype frequencies, measure of LD between the allele or haplotype and disease allele, and the measure of nonlinearity of the nonlinear transformations of the allele or haplotype frequencies. We showed that, in many cases, most of the studied nonlinear test statistics have higher power than the standard  -test statistic, with the exception of the reciprocal transformation whose power, in general, is lower than that of the standard
-test statistic, with the exception of the reciprocal transformation whose power, in general, is lower than that of the standard  -test statistic. However, since the power of a statistic is a complex issue, there is not one statistic that is uniformly most powerful. Forms of nonlinear transformation are crucial for developing nonlinear test statistics. Our preliminary results showed that the larger the measure of nonlinearity of the nonlinear transformation is, the higher the power of its corresponding nonlinear test statistic. Power of nonlinear test statistics is a complicated function of the measure of nonlinearity of the nonlinear transformation and other genetic and population parameters, particularly allele/haplotype frequencies. Our experience shows that when the frequencies of alleles/haplotypes are <0.05, nonlinear test statistics may not be a good choice for association analysis. We suggest using nonlinear test statistics when the frequencies of alleles/haplotypes are >0.05; i.e., we use nonlinear test statistics for association analysis of common diseases with common alleles. A clear and consistent pattern of power of the nonlinear test statistics depends on the measure of nonlinearity of the nonlinear transformation and is difficult to obtain. More investigations are needed.
-test statistic. However, since the power of a statistic is a complex issue, there is not one statistic that is uniformly most powerful. Forms of nonlinear transformation are crucial for developing nonlinear test statistics. Our preliminary results showed that the larger the measure of nonlinearity of the nonlinear transformation is, the higher the power of its corresponding nonlinear test statistic. Power of nonlinear test statistics is a complicated function of the measure of nonlinearity of the nonlinear transformation and other genetic and population parameters, particularly allele/haplotype frequencies. Our experience shows that when the frequencies of alleles/haplotypes are <0.05, nonlinear test statistics may not be a good choice for association analysis. We suggest using nonlinear test statistics when the frequencies of alleles/haplotypes are >0.05; i.e., we use nonlinear test statistics for association analysis of common diseases with common alleles. A clear and consistent pattern of power of the nonlinear test statistics depends on the measure of nonlinearity of the nonlinear transformation and is difficult to obtain. More investigations are needed.
To further evaluate the performance of the nonlinear test statistics, the proposed nonlinear test statistics were also applied to three real data examples. The results showed that when the standard  -test detected association of the COMT gene with schizophrenia, all nonlinear test statistics demonstrated strong association of the COMT gene with schizophrenia and when the standard
-test detected association of the COMT gene with schizophrenia, all nonlinear test statistics demonstrated strong association of the COMT gene with schizophrenia and when the standard  -test detected no association of the gene ZDHHC8 with schizophrenia, all nonlinear test statistics with almost the same P-values as that of the standard
-test detected no association of the gene ZDHHC8 with schizophrenia, all nonlinear test statistics with almost the same P-values as that of the standard  -test also showed no association.
-test also showed no association.
The results in this report are very limited. Theoretical and empirical studies should be conducted to compare and investigate the relative strengths and weaknesses of nonlinear tests and other existing association tests. The properties of the nonlinear test statistics should be further investigated both by theoretical studies and by empirical simulations. In this report, we studied only very limited nonlinear functions. It is worth developing general theory for searching optimal nonlinear functions with the highest power. Nonlinear tests are a new concept for developing test statistics, which will open new ways for developing powerful statistics in genetic studies of complex diseases. Theory for nonlinear tests is at its infancy. Many theoretical works and empirical evaluations are needed in the future.
Acknowledgments
We thank Sagiv Shifman and Ariel Darvasi for providing the detailed data information for schizophrenic haplotype analyses. We thank two anonymous reviewers for helpful comments on the manuscript, which led to its improvement. We also thank Ranjan Deka for his constructive comments. M. M. Xiong is supported by the National Institutes of Health (NIH)–National Institute of Arthritis and Musculoskeletal and Skin Diseases grants IP50AR44888 and HL74735 and by NIH grant ES09912. J. Y. Zhao is supported by NIH grant ES09912.
APPENDIX A
In the following, we show that any statistics arising from linear transformation will not change the values of pretransformation statistics. To illustrate this point, let PA and P be the allele (haplotype) frequencies in cases and controls, respectively,  be a vector of differences in allele or haplotype frequencies between cases and controls, Σ be variance–covariance matrix of the vector of differences
be a vector of differences in allele or haplotype frequencies between cases and controls, Σ be variance–covariance matrix of the vector of differences  , and A be a linear transformation matrix, where linear transformation of the allele or haplotype frequencies is expressed as
, and A be a linear transformation matrix, where linear transformation of the allele or haplotype frequencies is expressed as  and AP, respectively. The popularly used
and AP, respectively. The popularly used  -test statistic can be derived from the statistic
-test statistic can be derived from the statistic
 |
where  is a generalized inverse of the matrix Σ.
is a generalized inverse of the matrix Σ.
The difference in linear transformation of allele or haplotype frequencies between cases and controls can be written as
 |
where A is assumed a nonsingular matrix. Then, the variance–covariance matrix is given by
 |
The new statistic resulting from transformation is
 |
This shows that linear transformation of allele or haplotype frequencies will not change test statistics.
APPENDIX B
Below we show that a similarity measure of the genome region is a quadratic function of allele or haplotype frequencies. Therefore, similarity measure-based statistics are nonlinear test statistics. For simplicity of presentation, we consider only haplotype similarity. However, the conclusions, in general, hold for other types of similarity of the genome region. Suppose that the numbers of  haplotypes in the affected and unaffected individuals are
haplotypes in the affected and unaffected individuals are  and
and  , respectively.
, respectively.  , and
, and  are defined as before. Then, we have
are defined as before. Then, we have  . Let
. Let  and
and  be the similarity measure of the haplotype
be the similarity measure of the haplotype  in the unaffected and affected individuals. Let
in the unaffected and affected individuals. Let  be a measure of the similarity between the haplotype
be a measure of the similarity between the haplotype  and the haplotype
and the haplotype  . Then, the similarity measure of the haplotype
. Then, the similarity measure of the haplotype  in the unaffected individuals is given by
in the unaffected individuals is given by
 |
Let  . Then the above equation can be further reduced to
. Then the above equation can be further reduced to
 |
The similarity measure of all the haplotypes in the unaffected individuals, which is referred to as the overall similarity measure and denoted by  , is defined as the summation of the similarity measure of the individual haplotype, i.e.,
, is defined as the summation of the similarity measure of the individual haplotype, i.e.,  . Let
. Let  be a similarity matrix. We have
be a similarity matrix. We have  . Then,
. Then,  can be written as
can be written as
 |
Similarly, for the affected individuals, we have
 |
where  ,
,  , and
, and  are similarly defined as those for the unaffected individuals. Clearly, similarities measures
are similarly defined as those for the unaffected individuals. Clearly, similarities measures  and
and  are quadratic functions of the haplotype frequencies and hence are nonlinear transformations of the haplotype frequencies. Both the overall similarity measure
are quadratic functions of the haplotype frequencies and hence are nonlinear transformations of the haplotype frequencies. Both the overall similarity measure  and the similarity measure
and the similarity measure  of the haplotype
of the haplotype  can be used to construct association tests.
can be used to construct association tests.
We first consider the overall similarity measure-based test statistic. Let  and
and  . The Jacobian matrix B of the overall similarity measure
. The Jacobian matrix B of the overall similarity measure  with respect to P is given by
with respect to P is given by
 |
Similarly, we have  . Let
. Let  ,
,  ,
,  , and
, and  be the corresponding estimators of
be the corresponding estimators of  ,
,  ,
,  , and
, and  , respectively. Then the variance of
, respectively. Then the variance of  and
and  can be approximated by
can be approximated by
 |
(Lehmann 1983). We define the overall haplotype similarity measure-based statistic as
 |
This is similar to the similarity measure-based test statistic D in Tzeng et al. (2003), where the variances of  and
and  are accurately calculated.
are accurately calculated.
Now consider the haplotype similarity measure-based test statistic. Let  ,
,  and
and  ,
,  .
.  and the Jacobian matrix C for the affected individuals are similarly defined. Let
and the Jacobian matrix C for the affected individuals are similarly defined. Let  . We define the haplotype similarity measure-based test statistic as
. We define the haplotype similarity measure-based test statistic as
 |
where  , and
, and  are the estimators of
are the estimators of  , and
, and  , respectively, and
, respectively, and  is the generalized inverse of the matrix
is the generalized inverse of the matrix  . Let
. Let  ; then, under the null hypothesis of no association between the haplotypes and the disease, the test statistic
; then, under the null hypothesis of no association between the haplotypes and the disease, the test statistic  is asymptotically distributed as a central
is asymptotically distributed as a central  . It is clear that both test statistics
. It is clear that both test statistics  and
and  are nonlinear test statistics. Therefore, the similarity measure-based statistics are special cases of the nonlinear test statistics.
are nonlinear test statistics. Therefore, the similarity measure-based statistics are special cases of the nonlinear test statistics.
APPENDIX C
Let  be a vector-valued nonlinear function of random vector P. Assume that the nonlinear function
be a vector-valued nonlinear function of random vector P. Assume that the nonlinear function  satisfies regularity conditions that ensure that Theorem 3.3A in Serfling (1980) holds. Then,
satisfies regularity conditions that ensure that Theorem 3.3A in Serfling (1980) holds. Then,  is asymptotically distributed as a multivariate normal distribution
is asymptotically distributed as a multivariate normal distribution  , where
, where
 |
Similarly,  is asymptotically distributed as
is asymptotically distributed as  , where
, where
 |
Therefore, under the null hypothesis  , which implies
, which implies  ,
,  is asymptotically distributed as
is asymptotically distributed as  , where
, where
 |
Let  and
and  . Then, under the null hypothesis, TN = ZTΛ−Z is asymptotically distributed as a central
. Then, under the null hypothesis, TN = ZTΛ−Z is asymptotically distributed as a central  -distribution (Greenwood and Nikulin 1996). The alternative hypothesis is
-distribution (Greenwood and Nikulin 1996). The alternative hypothesis is  . Under the alternative hypothesis,
. Under the alternative hypothesis,  is asymptotically distributed as a noncentral
is asymptotically distributed as a noncentral  -distribution with the following noncentrality parameter:
-distribution with the following noncentrality parameter:
 |
(C1) |
By Taylor expansion, we have
 |
(C2) |
where  ,
,  ,
,
 |
Equation C2 can be rewritten as
 |
(C3) |
Let  then
then
 |
(C4) |
Substituting  in Equation C4 into Equation C1 yields
in Equation C4 into Equation C1 yields
 |
(C5) |
Recall that
 |
(C6) |
where
 |
Thus,
 |
(C7) |
Substituting Equations C6 and C7 into Equation C5, we obtain
 |
(C8) |
Next we study geometric interpretation of the matrix S. Let  , where
, where  . Define the following parameter equations:
. Define the following parameter equations:
 |
As t varies,  defines a curve C in the space. The tangent vector of the curve C at the point P is given by
defines a curve C in the space. The tangent vector of the curve C at the point P is given by
 |
where
 |
Taking  as a new coordinate system, we obtain the change rates of the tangent vector of the curve over new coordinates,
as a new coordinate system, we obtain the change rates of the tangent vector of the curve over new coordinates,
 |
where
 |
The change rate of the tangent vector of the curve characterizes the strength of the nonlinearity of the nonlinear function (Bates and Watts 1980). The vector S has the following form:
 |
If the product terms of the haplotype frequencies are ignored, we obtain , where
, where
 |
 . Then, Equation C8 can be simplified to
. Then, Equation C8 can be simplified to
 |
For the standard  -test statistic, we have
-test statistic, we have  . Thus, its noncentrality parameter is given by
. Thus, its noncentrality parameter is given by
 |
and
 |
References
- Ahmadi, K. R., M. E. Weale, Z. Y. Xue, N. Soranzo, D. P. Yarnall et al., 2005. A single-nucleotide polymorphism tagging set for human drug metabolism and transport. Nat. Genet. 37: 84–89. [DOI] [PubMed] [Google Scholar]
- Akey, J., L. Jin and M. Xiong, 2001. Haplotypes vs single marker linkage disequilibrium tests: What do we gain? Eur. J. Hum. Genet. 9: 291–300. [DOI] [PubMed] [Google Scholar]
- Altshuler, D., and A. G. Clark, 2005. Genetics. Harvesting medical information from the human family tree. Science 307: 1052–1053. [DOI] [PubMed] [Google Scholar]
- Bates, D. M., and D. G. Watts, 1980. Relative curvature measures of nonlinearity. J. R. Stat. Soc. Ser. B 42: 1–25. [Google Scholar]
- Borsting, C., J. J. Sanchez and N. Morling, 2005. SNP typing on the NanoChip electronic microarray. Methods Mol. Biol. 297: 155–168. [DOI] [PubMed] [Google Scholar]
- Bourgain, C., E. Genin, P. Margaritte-Jeannin and F. Clerget-Darpoux, 2001. Maximum identity length contrast: a powerful method for susceptibility gene detection in isolated populations. Genet. Epidemiol. 21(Suppl. 1): S560–S564. [DOI] [PubMed] [Google Scholar]
- Chapman, N. H., and E. M. Wijsman, 1998. Genome screens using linkage disequilibrium tests: optimal marker characteristics and feasibility. Am. J. Hum. Genet. 63: 1872–1885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan, R., and K. Lange, 1998. Models for haplotype evolution in a nonstationary population. Theor. Popul. Biol. 53: 184–198. [DOI] [PubMed] [Google Scholar]
- Freimer, N., and C. Sabatti, 2004. The use of pedigree, sib-pair and association studies of common diseases for genetic mapping and epidemiology. Nat. Genet. 36: 1045–1051. [DOI] [PubMed] [Google Scholar]
- Greenwood, P. E., and M. S. Nikulin, 1996. A Guide to Chi-Square Testing. John Wiley & Sons, New York.
- Halldorsson, B. V., V. Bafna, R. Lippert, R. Schwartz, F. M. De La Vega et al., 2004. Optimal haplotype block-free selection of tagging SNPs for genome-wide association studies. Genome Res. 14: 1633–1640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jorde, L. B., 2000. Linkage disequilibrium and the search for complex disease genes. Genome Res. 10: 1435–1444. [DOI] [PubMed] [Google Scholar]
- Lehmann, E. L., 1983. Theory of Point Estimation. John Wiley & Sons, New York.
- Neale, B. M., and P. C. Sham, 2004. The future of association studies: gene-based analysis and replication. Am. J. Hum. Genet. 75: 353–362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nothnagel, M., 2002. Simulation of LD block-structured SNP haplotype data and its use for the analysis of case-control data by supervised learning methods. Am. J. Hum. Genet. 71(Suppl.): A2363. [Google Scholar]
- Pritchard, J. K., and P. Donnelly, 2001. Case-control studies of association in structured or admixed populations. Theor. Popul. Biol. 60: 227–237. [DOI] [PubMed] [Google Scholar]
- Risch, N., and K. Merikangas, 1996. The future of genetic studies of complex human diseases. Science 273: 1516–1517. [DOI] [PubMed] [Google Scholar]
- Saito, S., M. Ikeda, N. Iwata, T. Suzuki, T. Kitajima et al., 2005. No association was found between a functional SNP in ZDHHC8 and schizophrenia in a Japanese case-control population. Neurosci. Lett. 374: 21–24. [DOI] [PubMed] [Google Scholar]
- Serfling, R. J., 1980. Approximating Theorems of Mathematical Statistics. John Wiley & Sons, New York.
- Shifman, S., M. Bronstein, M. Sternfeld, A. Pisante-Shalom, E. Lev-Lehman et al., 2002. A highly significant association between a COMT haplotype and schizophrenia. Am. J. Hum. Genet. 71: 1296–1302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tzeng, J. Y., B. Devlin, L. Wasserman and K. Roeder, 2003. On the identification of disease mutations by the analysis of haplotype similarity and goodness of fit. Am. J. Hum. Genet. 72: 891–902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang, W. Y., B. J. Barratt, D. G. Clayton and J. A. Todd, 2005. Genome-wide association studies: theoretical and practical concerns. Nat. Rev. Genet. 6: 109–118. [DOI] [PubMed] [Google Scholar]
- Xiong, M., J. Zhao and E. Boerwinkle, 2003. Haplotype block linkage disequilibrium mapping. Front. Biosci. 8: a85–a93. [DOI] [PubMed] [Google Scholar]
- Yu, C., Y. Zhou, X. Miao, P. Xiong, W. Tan et al., 2004. Functional haplotypes in the promoter of matrix metalloproteinase-2 predict risk of the occurrence and metastasis of esophageal cancer. Cancer Res. 64: 7622–7628. [DOI] [PubMed] [Google Scholar]
- Zhang, S., Q. Sha, H. S. Chen, J. Dong and R. Jiang, 2003. Transmission/disequilibrium test based on haplotype sharing for tightly linked markers. Am. J. Hum. Genet. 73: 566–579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao, J., E. Boerwinkle and M. Xiong, 2005. An entropy-based statistic for genomewide association studies. Am. J. Hum. Genet. 77: 27–40. [DOI] [PMC free article] [PubMed] [Google Scholar]