Abstract
Comparative analysis of complex developmental pathways depends on our ability to resolve the function of members of gene families across taxonomic groups. LIN5, which belongs to a small gene family of apoplastic invertases in tomato (Lycopersicon esculentum), is a quantitative trait locus that modifies fruit sugar composition. We have compared the genomic organization and expression of this gene family in the two distantly related species: tomato and Arabidopsis. Invertase family members reside on segmental duplications in the near-colinear genomes of tomato and potato (Solanum tuberosum). These chromosomal segments are syntenically duplicated in the model plant Arabidopsis. On the basis of phylogenetic analysis of genes in the microsyntenic region, we conclude that these segmental duplications arose independently after the separation of the tomato/potato clade from Arabidopsis. Rapid regulatory divergence is characteristic of the invertase family. Interestingly, although the processes of gene duplication and specialization of expression occurred separately in the two species, synteny-based orthologs from both clades acquired similar organ-specific expression. This similar expression pattern of the genes is evidence of comparable evolutionary constraints (parallel evolution) rather than of functional orthology. The observation that functional orthology cannot be identified through analysis of expression similarity highlights the caution that needs to be exercised in extrapolating developmental networks from a model organism.
The primary sources for evolutionary novelties are duplications of individual genes, chromosomal segments, or entire genomes (polyploidy; Lynch and Conery, 2000). Analysis of the Arabidopsis sequence indicates that 60% of the genome is segmentally duplicated, more than any of the sequenced eukaryotic genomes (Samonte and Eichler, 2002; Arabidopsis Genome Initiative, 2000). Segmental duplications within a genome can be followed by functional divergence of the paralogs either by the evolution of a novel role for one of the genes or by specialization in some aspects of their ancestral role (Sankoff, 2001). Diversification of gene function among distantly related species is difficult to assess because attribution of orthology is not trivial, particularly for members of gene families (Hofer and Ellis, 2002; TheiBen, 2002). One way to determine gene orthology is to analyze their syntenic relations, because a conserved linear order of loci is a strong anchor for inferring common ancestry.
While the tomato (Lycopersicon esculentum) and potato (Solanum tuberosum) genetic maps are nearly colinear (Tanksley et al., 1992), complex syntenic relationships have been described for tomato and Arabidopsis that belong to two different families (Solaneceae and Brassicaceae, respectively). Ku et al. (2000) have demonstrated that a 105-kb bacterial artificial chromosome (BAC) of tomato shows conservation of gene content and order with four segments of different Arabidopsis chromosomes. The degree of microcolinearity between these two species was found to be much higher for five open reading frames (ORFs) that lie in a 57-kb region of tomato, whereas all of these ORFs could be identified on a 30-kb region of Arabidopsis, with two inversion events distinguishing the arrangement of these genes in the two species (Rossberg et al., 2001). Overall, these comparative studies of the gene content in tomato and Arabidopsis showed a complex network of synteny encompassing relatively small stretches that are often interrupted by non-colinear genes.
In this study, we explored the evolution of an ancestral invertase gene that encodes for an enzyme that functions in the cleavage of Suc into Glc and Fru. The apoplastic invertases are a subfamily of genes that show highly differential tissue-specific expression, mainly in the phloem conductive tissues (Tymowska-Lalanne and Kreis, 1998a; Sturm, 1999). Invertases play an important role in supplying carbohydrates to sink tissues via the apoplastic pathway (Sturm, 1999), in influencing developmental processes (Tang et al., 1999), and in linking different intracellular and extracellular stimuli to the regulation of source/sink relations (Roitsch, 1999). One member of this gene family in tomato (LIN5) is a quantitative trait locus (QTL) that modifies sugar content in the fruits (Fridman et al., 2000). A wild tomato species allele of LIN5 increases total soluble solids (mainly sugars) of the fruit by 20%. We describe segmental duplications of an ancestral invertase-containing chromosomal segment and demonstrate a high degree of evolutionary plasticity in organ expression patterns for the descending gene family members.
RESULTS AND DISCUSSION
Syntenic Relationships of the Apoplastic Invertase in Tomato and Arabidopsis
LIN5 is part of a small family of four genes in tomato (Godt and Roitsch, 1997). Low-stringency Southern analysis with all four members revealed no additional closely related copies in the tomato genome. We determined the full genomic and cDNA sequence of the four genes (see “Materials and Methods”) and mapped them on the tomato genetic map using the Lycopersicon pennellii introgression lines (Eshed and Zamir, 1995). LIN5, residing on chromosome 9, is arranged in a direct tandem repeat with LIN7 (Fig. 1A). To the telomeric end of the invertases is a KINESIN gene and downstream of LIN7 resides a 40S RIBOSOMAL gene in opposite orientation. The remaining family members, LIN6 and LIN8, are tandemly arranged on chromosome 10, flanked by an additional KINESIN and 40S RIBOSOMAL genes and therefore represent a segmental duplication between the two tomato chromosomes. Similar linear order of genes is found on the segmental duplications of Arabidopsis chromosomes 2 and 3 (Arabidopsis Genome Initiative, 2000) but with a single invertase (ATβFRUCT5 [At2g36190] and ATβFRUCT2 [At3g52600], respectively; Fig. 1B). The Arabidopsis chromosome 3 locus contains the 40S ribosomal gene but not the KINESIN ORF, and both Arabidopsis loci have a UBIQUITIN EXTENSION ORF downstream of the invertase. A complete Arabidopsis genome BLAST screen with the above invertases revealed two additional closely related genes that are arranged in tandem on a separate locus on chromosome 3 (ATβFRUCT1 [At3g13790] and ATβFRUCT6 [At3g13784]) with no conserved gene order as described for the other invertases. The UBQ and 40S RIBOSOMAL genes are also found in the genomic region of these tandemly arranged genes; however, the four genes are found on different BACs, with gaps that are more than 1 Mbp and thus represent a more complex order of the synteny network (INVERTASES on BAC MMM17; RIBOSOMAL 40S on BAC F24K9; UBQ on BAC F8A24; http://www.arabidopsis.org/). Our results indicate that the tomato invertases LIN5, LIN7, LIN6, and LIN8 are synteny-based orthologs with the Arabidopsis genes ATβFRUCT5 and ATβFRUCT2 and therefore we focused our study on these orthologous genes.
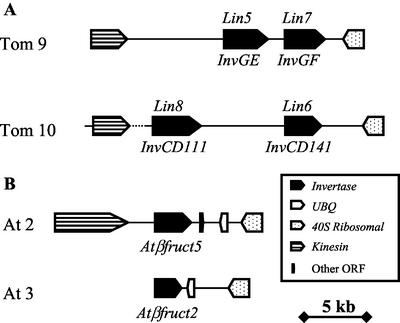
Figure 1.
Genomic organization of the invertase locus in tomato, potato, and Arabidopsis. Different patterns denote the ORFs (solid black, invertase; horizontal bars, kinesin; dotted, 40S ribosomal; and empty, UBIQUITIN EXTENSION). A, The tomato invertase loci on chromosome 9 (top) and chromosome 10 (bottom). The invertase gene names are indicated for tomato (above) and potato (below). Dotted line denotes a gap in the sequence. B, The Arabidopsis invertase loci on chromosome 2 (AC007135; top) and chromosome 3 (AL050300; bottom).
Evolution of the Syntenic Block
The primary question considering the timing of the block duplication in the both species is whether the segmental duplication occurred before or after the divergence of these two distantly related species. An immediate answer to this question lies in the phylogenetic relations between the invertase peptides from tomato (LIN5, LIN6, LIN7, and LIN8) and the two synteny-based orthologous proteins from Arabidopsis (ATβFRUCT5 and ATβFRUCT2; Fig. 2A). The 100% reliability of the separation between the clades of the two species (based on 500 bootstrapping runs) implies that these proteins are significantly clustered in a species-specific manner, i.e. they are significantly more similar within a species than between the species. This species-specific clustering of the peptides strongly suggests that the segmental duplication occurred independently in the two species. However, in case of gene conversion within each species, paralogs might look so much more alike than they otherwise should (Fitch, 2000). Gene conversion, in which copies of block of DNA from one gene replace the homologous residues in its paralog, can obscure the evidence for an early timing of a duplication that occurred before the species divergence (in the ancestor). The origin of the segmental duplications must be explored in reference to the surrounding sequences. In a similar manner to the invertases, for the 40S RIBOSOMAL proteins and the KINESINs, the tomato paralogs are significantly more similar within the species than they are to their Arabidopsis orthologs (Fig. 2, B and C). This triple protein phylogeny implies that the most parsimonious scenario is that the segmental duplications in tomato and Arabidopsis arose independently after their divergence from a common ancestor (Fig. 3). This conclusion is in agreement with the dating of species divergence to 150 million years ago (MYA; Ku et al., 2000) and the origin of the Arabidopsis segmental duplication of chromosomes 2 and 3, which occurred later (100 MYA; Vision et al., 2000). The main deduction from the above “molecular archeology” analysis is that the ancestral species carried a chromosome segment that harbored a single progenitor invertase locus. This segment duplicated independently in the two species. In tomato and potato, the phylogenetic clustering of the invertases according to their chromosomal locations suggests that the tandem duplications within each locus occurred independently in the two chromosomes.
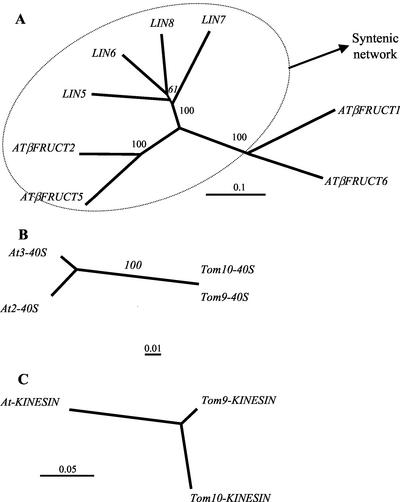
Figure 2.
Phylogeny of the ORFs in the syntenic region. Neighbor-joining phylogenetic trees for INVERTASE proteins (A), 40S RIBOSOMAL peptides (B), and KINESIN peptides (C). A total of 500 bootstrapping runs were performed for each tree, and the percent reliability is labeled next to each branch. The scales indicate the average substitutions per site for each cladogram. The four potato invertases (Fig. 1A) cluster with their tomato orthologs in a locus-specific manner (bootstrap value of 100; data not shown).
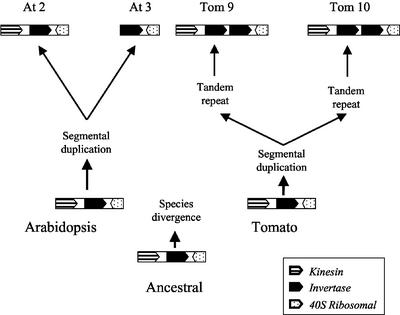
Figure 3.
Evolution of the invertase ancestral locus. The cascade starts from the bottom (ancestral) and proceeds upwards to the contemporary blocks in chromosome 2 and 3 of Arabidopsis and chromosomes 9 and 10 of tomato. Different patterns denote the ORFs within the segment (solid black, invertase; horizontal bars, kinesin; and dotted, 40S ribosomal).
Interestingly, the synteny network presented here for the INVERTASE and 40S RIBOSOMAL can be extended to two duplicated chromosomal regions in the partial genomic sequence of the monocot rice genome (http://www.tigr.org/tdb/e2k1/osa1/). Arabidopsis and rice diverged 240 million years ago (Gale and Devos, 1998), and recent comparative genomics of the two species showed scant collinearity in gene order between the two species (Liu et al., 2001). In cases where gene order was preserved, it encompassed relatively small stretches that were often interrupted by noncollinear genes. The rice sequence indicates that the INVERTASE/40S RIBOSOMAL-containing block is an ancient ancestral block that was preserved over long evolutionary times in different plant lineages.
Functional Divergence of the Invertase Genes
Expression of the orthologs invertase gene family in tomato and Arabidopsis was analyzed using gene-specific primers in a PCR with reverse transcription (RT-PCR; Fig. 4). The chromosome 9 LIN genes are specific to flowers and fruits: LIN5 was highly expressed in the ovary and in the developing fruits, showing lower expression levels in the petals and stamens; whereas LIN7 transcripts were detected only in stamens and pollen. The chromosome 10 LIN genes were primarily expressed in vegetative tissues: LIN8 in roots and leaves and LIN6 in all tested organs except the ovary and pollen. These findings are consistent with published data (Godt and Roitsch, 1997) and with the in silico analysis of expressed sequence tags (ESTs) generated from different organs of the tomato plant (http://www.tigr.org/tdb/lgi/; Table I). Whereas the invertases show organ-specific expression, both tomato 40S RIBOSOMAL genes were constitutively expressed (Table I). The Arabidopsis synteny-based orthologs ATβFRUCT5 and ATβFRUCT2 were expressed only in the flower and pod but not in the vegetative tissues (Fig. 4B). These results are consistent with published data and the EST representation of ATβFRUCT5 and ATβFRUCT2 in silique and flower libraries, respectively (Tymowska-Lalanne and Kreis, 1998b). Apoplastic invertase expression in vegetative tissues of Arabidopsis is mediated by the non-syntenic and tandemly arranged apoplastic invertases on chromosome 3 (ATβFRUCT1 and ATβFRUCT6; Fig. 4B; Tymowska-Lalanne and Kreis, 1998b).
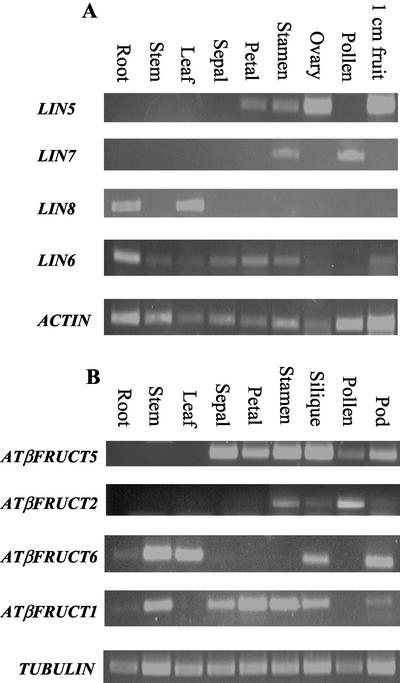
Figure 4.
RT-PCR analysis of the expression profiles of tomato (A) and Arabidopsis (B) apoplastic invertases using gene-specific primers. Ethidium bromide-stained RT-PCR products separated on a 1% (w/v) agarose gel. ACTIN and TUBULIN transcripts were used as a control for the cDNA quantity in the different samples of tomato and Arabidopsis, respectively. The LIN8 transcripts are the product of 37 cycles of PCR, 30 cycles for the other genes.
Table I.
In silico analysis of the tomato INVERTASE and 40S RIBOSOMAL genes expression
| Gene | TC No. | Tissue
|
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Germinating Seed (I) | Root
|
Shoot/ Meristem (TOF) | Shoot (B) | Leaf
|
|||||||
| Nutrient Deficient (W) | Fruit Set (X) | Pre-Anthesis (Y) | Etyiolated Radicle (Z) | pto Resistant (R) | pto Sensitive (S) | Mixed Elicitor (T) | |||||
| Library ESTs No. | |||||||||||
| 3,927 | 3,219 | 3,277 | 3,349 | 2,448 | 9,124 | 1,049 | 5,195 | 5,317 | 9,544 | ||
| No. of ESTs | |||||||||||
| LIN5 | 90043 | ||||||||||
| LIN6 | 86272 | 1 | 1 | ||||||||
| LIN7 | 84912 | ||||||||||
| LIN8 | 86570 | 3 | 1 | ||||||||
| Tom9-40S | 84990 | 1 | 5 | 1 | 1 | 13 | 1 | 2 | 5 | 5 | |
| Tom10-40S | 84989 | 1 | 2 | 1 | 2 | 3 | 3 | ||||
| Gene | Tissue
|
|||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Flower
|
Pollen (PP) | Ovary (D) | Fruit
|
Callus (C) | Suspension Culture (TOS) | Crown Gall (TOE) | ||||||
| 0–3 mm (TOA) | 3–8 mm (TOB) | 8 mm (TOC) | Anthesis (TOD) | Mixed Stages (L) | 20-d Old Pericarp | Breaker (G) | ||||||
| Library ESTs No. | ||||||||||||
| 2,586 | 2,474 | 2,629 | 2,660 | 7,010 | 5,426 | 9,956 | 6 | 8,659 | 9,576 | 8,026 | 5,204 | |
| No. of ESTs | ||||||||||||
| LIN5 | 1 | 10 | ||||||||||
| LIN6 | 1 | 2 | ||||||||||
| LIN7 | 6 | 4 | 145 | |||||||||
| LIN8 | ||||||||||||
| Tom9-40S | 4 | 2 | 2 | 3 | 1 | 3 | 1 | 6 | ||||
| Tom9-40S | 1 | 1 | 1 | |||||||||
BLAST analysis was performed with the full-length cDNAs of the four INVERTASE genes and the deduced coding sequence of the two 40S RIBOSOMAL genes against the Tomato Gene Index at The Institute for Genomic Research. Web site (http://www.tigr.org/tdb/lgi/). Each gene is represented by number of ESTs in the cDNA libraries. Tom9-40S and Tom10-40S are the deduced ORFs of the 40S RIBOSOMAL proteins from chromosome 9 and chromosome 10, respectively. TC, Tentative consensus sequences that are created by assembling ESTs into virtual transcripts. For each tissue's cDNA library, the number of ESTs is indicated below and the cod of this library is indicated in brackets.
The tomato paralogous pairs LIN5/LIN7 and LIN8/LIN6 reflect the rapid regulatory divergence within the family. Since their separation from the common ancestor, the chromosome 9 LINs specialized in flower expression, whereas the other pair shows expression in the vegetative tissue as well. Moreover, within the pairs that apparently represent the closest paralogs, there is significant differentiation in expression. This rapid regulatory divergence of the invertase gene family is also implied from invertase expression data in other Solanaceae species. The crucial functional role of invertase in pollen development was demonstrated recently with the induction of male sterility in tobacco (Nicotiana tabacum) plants by antisense suppression of the tobacco homolog of LIN7, NIN88 (Goetz et al., 2001). The potato ortholog of LIN7 (InvGF) is also expressed in the stamen and pollen, suggesting that the activity of this apoplastic invertase in the Solanaceae is functionally related and monophyletic (Maddison et al., 1999). Assuming that the single ancestral invertase had a repertoire of expression covering diverse organs and that it duplicated independently in tomato and Arabidopsis (after their divergence), we propose that the similar stamen and pollen specificity of LIN7 and AtβFRUCT2 resulted from comparable selection pressures acting on both of them (so-called parallel evolution). However, this similarity was not reflected in convergent evolution of the promoter region of these genes: Phylogenetic analysis of the promoter regions of the six genes from the two species revealed a similar species-specific relatedness of the genes' promoters (data not shown). Although tomato and potato diverged 12 MYA (deSa and Drouin, 1996), the potato LIN5 ortholog (InvGE) is characterized by a wider spectrum of expression: InvGE is expressed in leaves in addition to its expression in flowers (Maddison et al., 1999). Interestingly, the InvGE promoter fused to the β-glucuronidase reporter gene mediates expression also in the developing sugar-accumulating tuber (Viola et al., 2001). The expression of LIN5/InvGE in the sink tissues combined with the QTL effect of the tomato gene and the linkage of the potato InvGE with a QTL for tuber starch content (Schafer-Pregl et al., 1998) suggest that this invertase may be an early determinant of source-sink relationships. Taken together, our results indicate that control elements of the orthologous invertase genes have undergone rapid evolutionary change, resulting in divergence in organ expression. Such an analysis of the function of orthologous invertases can be extended to additional species, including monocots, that preserve the same syntenic block.
CONCLUSIONS
The above results highlight the caution that needs to be exercised in extrapolating developmental networks from a model organism (Arthur, 2002). Attempts to determine functional orthology of genes as part of mapping developmental processes onto phylogeny should take into account the plasticity of developmental programs, even when comparison is made between closely related species for genes that underline an essential and presumably conservative process, as was demonstrated for the Hox genes in nematodes (Eizinger et al., 1999). Moreover, determination of functional orthology across great genetic distances cannot easily be inferred, and sequence homology combined with similar expression between genes of different species do not necessarily imply common ancestry of the expression. Such similarity may rather reflect similar evolutionary constraints or similar developmental biases (Arthur, 2002) and may be the result of parallel evolution.
MATERIALS AND METHODS
Isolation and Sequencing of Tomato BACs and Invertase cDNAs
Screening of a tomato (Lycopersicon esculentum) BAC library (Budiman et al., 2000) with a 750-bp fragment originating from the third exon of the potato (Solanum tuberosum) apoplastic invertase (Hedley et al., 1994) yielded 45 clones. On the basis of Southern-blot and PCR analyses, three BACs were selected representing each of the four LINs: BAC 91A4 for LIN5 and LIN7, BAC 28O22 for LIN6, and BAC 95H1 for LIN8. BACs DNA was digested with EcoRI, PstI, HindIII, and BamHI and subcloned into pBS vectors. Subclones from each of the resulting sublibraries were selected based on different TaqI restriction patterns and sequenced using the two flanking T7 and T3 primers on an Abi Prism 3700 (Applied Biosystems, Weiterstadt, Germany). Sequences were assembled using the Sequencher Software package (Gene Codes Corporation, Ann Arbor, MI) to determine uni-sequences and contigs that ranged from 388 bp to 23,155 kb (GenBank accession nos. AF506004 and AJ272306). This approach yielded a total of 71,587 and 109,553 bp of unique sequence for chromosomes 9 and 10, respectively. cDNA clones were isolated from flower cDNA library (LIN5 and LIN7; accessions nos. AJ272304 and AF506006) and from root cDNA library (LIN6 and LIN8; accessions nos. AF506005 and AF506007). Exon/intron organization of the four genes was determined based on comparison of the genomic sequences with the corresponding cDNA sequences.
BLAST Searches and Construction of Phylogenetic Trees
The tomato sequences were searched against The Arabidopsis Information Resource Web site (http://www.Arabidopsis.org/) using the BlastX algorithm (Altschul et al., 1997). The threshold for reporting a match between a tomato ORF and a specific Arabidopsis BAC was an expected value of <E−20. The chromosome 9 and 10 tomato invertase-containing contigs (approximately 23 kb each) were annotated based on BlastN against invertase cDNA sequences and the EST database (http://www.tigr.org/tdb/lgi/). Construction of the phylogenetic trees using the amino acid sequences as performed using the GCG9 software package (University of Wisconsin-Genetics Computer Group, Madison) as described previously (Pan et al., 2000).
Plant Material
Tomato plants (var M82) were grown in pots in the greenhouse and used as the source for the harvested tissues, except for the roots that were collected from young seedlings. Arabidopsis (ecotype Columbia) was grown as described previously (Tymowska-Lalanne and Kreis, 1998). Pollen collection from Arabidopsis was performed as described previously (Preuss et al., 1993) using water instead of germination medium. Harvested tissues were immediately frozen in liquid nitrogen and stored at −80°C until RNA extraction.
RT-PCR Analysis
The four tomato/Arabidopsis apoplastic invertase cDNAs were aligned. On the basis of maximum nucleotide polymorphism, a pair of specific primers was designed for each of the genes. To determine the expression patterns of the apoplastic invertase genes, total RNA was isolated from various parts of the plant using TRIzol reagent (Invitrogen, Carlsbad, CA) according to the manufacturer's protocol. The RT steps were carried out according to the manufacturer's protocol using 1 μg of RNA. The following forward and reverse primers, respectively, were used: for LIN5, 5′-CTGGATATGTAGATGTAGAT-3′ and 5′-CACACTTTGCCTTCTAAATT-3′; for LIN6, 5′-GTAGATGTAGATTTAGCAGA-3′ and 5′-CTATGGTTTCTTTGTGACGT-3′; for LIN7, 5′-GTTGATGTTGATTTGGCTGA-3′ and 5′-TTGACTCGAGGGATATCTAA-3′; for LIN8, 5′-CTGGATATGTAGATGTAGAT-3′ and 5′-ACGTCTGCTCAAATATCATG-3′; for ACTIN, 5′-ATTCCCTGACTGTTT-GCTAGT-3′ and 5′-TCCAACACAATACCGGTGGT-3′; for ATβFRUCT1, 5′-TCTTCAAAGCCCGTCAAAAC-3′ and 5′-AATGCATGCTCTTC-CCTTTC-3′; for ATβFRUCT2, 5′-TGGTTTGATCACGTTGGCTA-3′ and 5′-GAGGCTTCTGCATGTTCCAT-3′; for ATβFRUCT5, 5′-GACATCAAGAT-GGGTCAACG-3′ and 5′-CATTCATTTGCAGAGGACGA-3′; for AtβFRUCT6, 5′-GAAACGGCTCCGAGATATGA-3′ and 5′-TGCAC-CAATCTCTTTCCTGA-3′; and for TUBULIN, 5′-CTCAAGAGGTTCT-CAGCAGTA-3′ and 5′-TCACCTTCTTCATCCGCAGTT-3′. PCR conditions were 30 s at 94°C, followed by 30 cycles of 20 s at 94°C, 20 s at 54°C (LIN5 and ACTIN)/56°C (Arabidopsis genes)/58.5°C (LIN6 and LIN8)/64°C (LIN7), 40 s at 68°C, and additional 2 min at 68°C. The PCR for LIN8 was carried out for an additional seven cycles because transcript level was non-detectable after 30 cycles of PCR. The four cDNA of the tomato genes and a genomic template for two species were used as controls to eliminate the possibility of a nonspecific amplification or the existence of DNA in the samples. Ten microliters of the PCR was separated on a 1% (w/v) gel, stained with ethidium bromide, and visualized and captured using NIH Imager. The PCR products of each gene were cloned into a pGEM vector (Promega, Madison, WI) and sequenced for verification.
ACKNOWLEDGMENTS
We thank Y. Eshed (Weizmann Institute, Rehovot, Israel), C. Gebhardt (Max-Planck-Institut fur Zuchtungsforschung, Koln, Germany, N. Ori (Hebrew University, Rehovot, Israel), S. Tanksley (Cornell University, Ithaca, NY), and E. Pichersky (University of Michigan, Ann Arbor, MI) for useful comments; T. Pleban and S. Dagan (Hebrew University, Rehovot, Israel) for valuable technical assistance; R. Wing, and D. Frisch from the Clemson University Genomics Institute for the tomato BACs.
Footnotes
This work was supported by the United States-Israel Binational Science Foundation.
Article, publication date, and citation information can be found at www.plantphysiol.org/cgi/doi/10.1104/pp.014431.
LITERATURE CITED
- Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature. 2000;408:796–815. doi: 10.1038/35048692. [DOI] [PubMed] [Google Scholar]
- Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arthur W. The emerging conceptual framework of evolutionary developmental biology. Nature. 2002;415:757–764. doi: 10.1038/415757a. [DOI] [PubMed] [Google Scholar]
- Budiman MA, Mao L, Wood TC, Wing RA. A deep-coverage tomato BAC library and prospects toward development of an STC framework for genome sequencing. Genome Res. 2000;10:129–136. [PMC free article] [PubMed] [Google Scholar]
- deSa MM, Drouin G. Phylogeny and substitution rates of angiosperm actin genes. Mol Biol Evol. 1996;13:1198–1212. doi: 10.1093/oxfordjournals.molbev.a025685. [DOI] [PubMed] [Google Scholar]
- Eizinger A, Jungblut B, Sommer RJ. Evolutionary change in the functional specificity of genes. Trends Genet. 1999;15:197–202. doi: 10.1016/s0168-9525(99)01728-x. [DOI] [PubMed] [Google Scholar]
- Eshed Y, Zamir D. An introgression lines population of Lycopersicon pennellii in the cultivated tomato enables the identification and fine mapping of yield-associated QTL. Genetics. 1995;141:1147–1162. doi: 10.1093/genetics/141.3.1147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fitch WM. Homology: a personal view on some of the problems. Trends Genet. 2000;16:227–231. doi: 10.1016/s0168-9525(00)02005-9. [DOI] [PubMed] [Google Scholar]
- Fridman E, Pleban T, Zamir D. A recombination hotspot delimits a wild-species quantitative trait locus for tomato sugar content to 484 bp within an invertase gene. Proc Natl Acad Sci USA. 2000;97:4718–4723. doi: 10.1073/pnas.97.9.4718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gale MD, Devos KM. Plant comparative genetics after 10 years. Science. 1998;282:656–659. doi: 10.1126/science.282.5389.656. [DOI] [PubMed] [Google Scholar]
- Godt DE, Roitsch T. Regulation and tissue-specific distribution of mRNAs for three extracellular invertase isoenzymes of tomato suggests an important function in establishing and maintaining sink metabolism. Plant Physiol. 1997;115:273–282. doi: 10.1104/pp.115.1.273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goetz M, Godt DE, Guivarc'h A, Kahmann U, Chriqui D, Roitsch T. Induction of male sterility in plants by metabolic engineering of the carbohydrate supply. Proc Natl Acad Sci USA. 2001;98:6522–6527. doi: 10.1073/pnas.091097998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hedley PE, Machray GC, Davies HV, Burch L, Waugh R. Potato (Solanum tuberosum) invertase-encoding cDNAs and their differential expression. Gene. 1994;145:211–214. doi: 10.1016/0378-1119(94)90007-8. [DOI] [PubMed] [Google Scholar]
- Hofer J, Ellis N. Conservation and diversification of gene function in plant development. Curr Opin Plant Biol. 2002;5:56–61. doi: 10.1016/s1369-5266(01)00228-x. [DOI] [PubMed] [Google Scholar]
- Ku HM, Vision T, Liu J, Tanksley SD. Comparing sequenced segments of the tomato and Arabidopsis genomes: large-scale duplication followed by selective gene loss creates a network of synteny. Proc Natl Acad Sci USA. 2000;97:9121–9126. doi: 10.1073/pnas.160271297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu H, Sachidanandam R, Stein L. Comparative genomics between rice and Arabidopsis shows scant collinearity in gene order. Genome Res. 2001;11:2020–2026. doi: 10.1101/gr.194501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lynch M, Conery JS. The evolutionary fate and consequences of duplicate genes. Science. 2000;290:1151–1155. doi: 10.1126/science.290.5494.1151. [DOI] [PubMed] [Google Scholar]
- Maddison AL, Hedley PE, Meyer RC, Aziz N, Davidson D, Machray GC. Expression of tandem invertase genes associated with sexual and vegetative growth cycles in potato. Plant Mol Biol. 1999;41:741–751. doi: 10.1023/a:1006389013179. [DOI] [PubMed] [Google Scholar]
- Pan Q, Liu YS, Budai-Hadrian O, Sela M, Carmel-Goren L, Zamir D, Fluhr R. Comparative genetics of nucleotide binding site-leucine rich repeat resistance gene homologues in the genomes of two dicotyledons: tomato and Arabidopsis. Genetics. 2000;155:309–322. doi: 10.1093/genetics/155.1.309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Preuss D, Lemieux B, Yen G, Davis RW. A conditional sterile mutation eliminates surface components from Arabidopsis pollen and disrupts cell signaling during fertilization. Genes Dev. 1993;7:974–985. doi: 10.1101/gad.7.6.974. [DOI] [PubMed] [Google Scholar]
- Roitsch T. Source-sink regulation by sugars and stress. Curr Opin Plant Biol. 1999;2:198–206. doi: 10.1016/S1369-5266(99)80036-3. [DOI] [PubMed] [Google Scholar]
- Rossberg M, Theres K, Acarkan A, Herrero R, Schmitt T, Schumacher K, Schmitz G, Schmidt R. Comparative sequence analysis reveals extensive microcolinearity in the lateral suppressor regions of the tomato, Arabidopsis, and capsella genomes. Plant Cell. 2001;13:979–988. doi: 10.1105/tpc.13.4.979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samonte RV, Eichler EE. Segmental duplications and the evolution of the primate genome. Nat Rev Genet. 2002;3:65–72. doi: 10.1038/nrg705. [DOI] [PubMed] [Google Scholar]
- Sankoff D. Gene and genome duplication. Curr Opin Genet Dev. 2001;11:681–684. doi: 10.1016/s0959-437x(00)00253-7. [DOI] [PubMed] [Google Scholar]
- Schäfer-Pregl R, Ritter E, Concilio L, Hesselbach J, Lovatti L, Walkemeier B, Thelen H, Salamini F, Gebhardt C. Analysis of quantitative trait loci (QTLs) and quantitative trait alleles (QTAs) for potato tuber yield and starch content. Theor Appl Genet. 1998;97:834–846. [Google Scholar]
- Sturm A. Invertases: primary structures, functions, and roles in plant development and sucrose partitioning. Plant Physiol. 1999;121:1–8. doi: 10.1104/pp.121.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang GQ, Luscher M, Sturm A. Antisense repression of vacuolae and cell wall invertase in transgenic carrot alters early plant development and sucrose partitioning. Plant Cell. 1999;11:177–189. doi: 10.1105/tpc.11.2.177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanksley SD, Ganal MW, Prince JP, de Vicente MC, Bonierbale MW, Broun P, Fulton TM, Giovannoni JJ, Grandillo S, Martin GB et al. High-density molecular linkage maps of the tomato and potato genomes. Genetics. 1992;132:1141–1160. doi: 10.1093/genetics/132.4.1141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- TheiBen G. Orthology: secret life of genes. Nature. 2002;415:741. doi: 10.1038/415741a. [DOI] [PubMed] [Google Scholar]
- Tymowska-Lalanne Z, Kreis M. The plant invertases: physiology, biochemistry and molecular biology. Adv Bot Res. 1998a;28:71–117. [Google Scholar]
- Tymowska-Lalanne Z, Kreis M. Expression of the Arabidopsis thaliana invertase gene family. Planta. 1998b;207:259–265. doi: 10.1007/s004250050481. [DOI] [PubMed] [Google Scholar]
- Viola R, Roberts AG, Haupt S, Gazzani S, Hancock RD, Marmiroli N, Machray GC, Oparka KJ. Tuberization in potato involves a switch from apoplastic to symplastic phloem unloading. Plant Cell. 2001;13:385–398. doi: 10.1105/tpc.13.2.385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vision TJ, Brown DG, Tanksley SD. The origins of genomic duplications in Arabidopsis. Science. 2000;290:2114–2117. doi: 10.1126/science.290.5499.2114. [DOI] [PubMed] [Google Scholar]