Abstract
Background
Diabetic nephropathy is the leading cause of end stage renal failure in the western world. There is substantial epidemiological evidence supporting a genetic predisposition to diabetic nephropathy, however the exact molecular mechanisms remain unknown. Transforming growth factor (TGFβ1) is a crucial mediator in the pathogenesis of diabetic nephropathy.
Methods
We investigated the role of five known single nucleotide polymorphisms (SNPs) in the TGFB1 gene for their association with diabetic nephropathy in an Irish, type 1 diabetic case (n = 272) control (n = 367) collection. The activity of TGFβ1 is facilitated by the action of type 1 and type 2 receptors, with both receptor genes (TGFBR1 and TGFBR2) shown to be upregulated in diabetic kidney disease. We therefore screened TGFBR1 and TGFBR2 genes for genomic variants using WAVE™ (dHPLC) technology and confirmed variants by direct capillary sequencing. Allele frequencies were determined in forty-eight healthy individuals. Data for all SNPs was assessed for Hardy Weinberg equilibrium, with genotypes and allele frequencies compared using the χ2 test for contingency tables. Patterns of linkage disequilibrium were established and common haplotypes estimated.
Results
Fifteen variants were identified in these genes, seven of which are novel, and putatively functional SNPs were subsequently genotyped using TaqMan™, Invader™ or Pyrosequencing® technology. No significant differences (p > 0.1) were found in genotype or allele distributions between cases and controls for any of the SNPs assessed.
Conclusion
Our results suggest common variants in TGFB1, TGFBR1 and TGFBR2 genes do not strongly influence genetic susceptibility to diabetic nephropathy in an Irish Caucasian population.
Background
Diabetic nephropathy is a major clinical complication of diabetes mellitus. Epidemiological evidence supporting a genetic contribution to this disease includes ethnicity differences [1], familial clustering [2,3] and the fact that only a subset of individuals with type 1 diabetes develops diabetic nephropathy regardless of metabolic control [4]. In addition, simulation studies have shown that environmental effects are insufficient to account for the familial aggregation of this disease [3,5].
Transforming growth factor beta (TGFβ1) is a multifunctional cytokine implicated in the pathogenesis of many forms of progressive renal disease, including diabetic nephropathy, by promoting renal hypertrophy and the accumulation of extracellular matrix [6]. Protein and mRNA levels of TGFβ1 are significantly increased in the renal glomeruli and tubulointerstitium of animal models of diabetes and in humans with diabetes [7]. TGFβ1 is transcriptionally activated by high extracellular glucose in murine glomerular mesangial cells [8]. Transgenic mice over-expressing TGFβ1 develop progressive renal failure, suggesting that chronically elevated levels of circulating TGFβ1 are integral to the pathogenesis of kidney disease [9,10]. Direct blockade of TGFβ protein by chronic administration of anti-TGFβ1 antibodies has been shown to decrease renal insufficiency [11]. In addition, antisense TGFβ1 oligonucleotides reduce the cellular hypertrophy and stimulation of matrix synthesis normally seen in renal cells exposed to high extracellular glucose [12].
The activity of TGFβ1 in regulating cell proliferation, differentiation and extracellular matrix production are mediated by a heterodimeric complex of type 1 and type 2 receptors. Upregulation of TGFβ1 receptors have been reported in animal models of glomerulosclerosis [13,14]. It has been proposed that upregulation of TGFβR2 induced by high extracellular glucose may contribute to distal tubular hypertrophy in diabetic nephropathy [15]. Isono and colleagues demonstrated that increased expression of TGFβR2 in the diabetic kidney is primarily due to stimulation of gene transcription rather than increased mRNA stability [16].
TGFβ1 is encoded by the TGFB1 gene located at chromosome 19q13.1 [17]. We have investigated the role of five known single nucleotide polymorphisms, which may influence TGFB1 gene expression (TGFB1: -800G>A, -509C>T, +72InsC, +869T>C, +915G>C) for their association with diabetic nephropathy. TGFβ receptors type 1 and type 2 are encoded by TGFBR1 and TGFBR2 genes respectively. At present there are over six hundred variants recorded in dbSNP for these genes, with little information available on the role of these variants in relation to renal complications of diabetes. We have screened the genomic draft sequence for the TGFBR1 and TGFBR2 genes in an Irish population to identify genomic variants. Allele frequencies were subsequently determined in a healthy control population and selected SNPs genotyped in a case-control collection. In summary, we investigated if putatively functional variants in three genes, TGFB1, TGFBR1 and TGFBR2, contribute to genetic susceptibility to diabetic nephropathy in type 1 diabetes.
Methods
Subjects
Ethical approval was obtained from the appropriate Research Ethics Committees in each country and written, informed consent obtained from individuals prior to conducting this study. The case and control groups used for this study (Table 1) have been described previously [18]. All patients were at least third generation Irish Caucasians diagnosed with type 1 diabetes mellitus before 31 years of age, and required insulin from diagnosis. Patients with nephropathy (cases, n = 272) had diabetes for at least 10 years before the onset of proteinuria (>0.5 g/24 h). Patients without nephropathy (controls, n = 367) had diabetes for at least 15 years, were not in receipt of antihypertensive medication, and had no evidence of non-diabetic renal disease. Patients with microalbuminuria were excluded from both groups.
Table 1.
Clinical characteristics of cases (n = 272) and controls (n = 367) (Data are n, mean ± SD)
| Criteria | Cases | Controls |
| N | 272 | 367 |
| Gender | ||
| Male | 61.4 % | 40.3 % |
| Female | 38.6 % | 59.7 % |
| Age at Diagnosis (years) | 17.1 ± 8.2 | 16.8 ± 8.1 |
| *Duration of Diabetes (years) | 26.9 ± 8.3 | 27.7 ± 9.0 |
| HbA1C (%) | 8.5 ± 1.7 | 8.4 ± 1.6 |
| BMI (kg/m2) | 26.0 ± 3.8 | 26.2 ± 3.5 |
| Systolic Blood Pressure (mm Hg) | 150.1 ± 22.6 | 126.9 ± 16.6 |
| Diastolic Blood Pressure (mm Hg) | 86.5 ± 11.4 | 76.1 ± 7.3 |
* Duration of diabetes was calculated from the date of diagnosis of diabetes to date of recruitment
* BMI = body mass index
In silico analysis
For TGFBR1, the nucleotide sequence of draft clone RP11-96L7 for human chromosome 9 was downloaded from the National Centre for Biotechnology Information [19]. Similarly, the sequence for TGFBR2 was obtained for draft clone RP11-1024P17 on human chromosome 3. Reference mRNA (NM_004612; NM_003242) and protein (NP_004603; NP_003233) sequences were also downloaded from NCBI for TGFBR1 and TGFBR2 respectively. These were used to determine intron-exon boundaries for genomic DNA using Vector NTI Advance (suite 2, version 8, Informax Inc (Europe), Oxford, UK). The nomenclature for all identified variants follows the Human Genome Variation Society recommendations for coding sequences, updated 21st May 2005 [20]. In addition, we have provided rs numbers for all previously identified SNPs and ss numbers for novel SNPs to facilitate ease of comparison between research groups.
Amplification and mutation screening
6464 bases of TGFBR1 and 5204 bases of TGFBR2 genomic sequences were divided into fragments with an average size of approximately 500 base pairs, for PCR and screening purposes in 15 case and 15 control individuals. As the TGFBR1 and TGFBR2 gene sequences cover approximately 45 kb and 84 kb respectively from start to stop codon, only the coding regions of these genes (including all exons, exon-intron boundaries and untranslated regions) were screened to prioritise the identification of potentially functional gene variants. Each PCR product was then evaluated using WaveMaker v3.4 software (Transgenomic Ltd, Crewe, UK) and analysed on the WAVE™ (dHPLC) DNA Fragment Analysis System (Transgenomic Ltd) following the manufacturer's recommendations. Differentially separating fragments (representing DNA variants) were bidirectionally sequenced to identify variants using an ABI PRISM® 3100 Genetic Analyser (Applied Biosystems, Warrington, UK). Forty-eight healthy controls (n = 96 chromosomes) from the Young Hearts collection [21] (a healthy Irish Caucasian population) were genotyped by direct capillary sequencing (Applied Biosystems) to establish allele frequencies for all gene variants.
Genotyping
Five SNPs were selected for genotyping in the TGFB1 gene as they have been previously suggested to influence the expression of TGFβ1, in addition to demonstrating a minor allele frequency greater than 5%. TaqMan assays were successfully designed for TGFB1: -800G>A (rs1800468), TGFB1: -509C>T (rs1800469), TGFB1: +869T>C (rs1982073) and TGFB1: +915G>C (rs1800471) SNPs, but proved problematic for TGFB1: +72InsC (rs1800999) due to the presence of a long C homopolymer. TGFB1: +72InsC was successfully genotyped using a biplex Invader™ assay (Third WAVE Technologies Inc, Madison, MI, USA). Genotyping was performed for receptor variants using Pyrosequencing® technology according to the manufacturer's instructions (Biotage, Uppsala, Sweden). Details of the primer sequences used for resequencing purposes, together with the WAVE conditions and the oligos used for the genotyping assays are listed (Tables 2, 3, 4, 5) with further details readily available from the authors on request. 272 case and 367 control samples were available for genotyping TGFB1 SNPs, however fewer samples were available (241 cases and 322 controls) for genotyping TGFBR2 gene variants. Genotype frequencies were assessed for Hardy-Weinberg equilibrium using a χ2 goodness-of-fit test. The χ2 test for contingency tables was used to compare genotype and allele frequencies between case and control subjects with the level of significance set to p < 0.05. Haploview [22] was used to visualise linkage disequilibrium (LD) and haplotype blocks within each gene.
Table 2.
Primer sequences used for screening TGFBR1 and TGFBR2 genes
| Primer Set | Primer 1 | Primer 2 |
| TGFBR1 i | tcctccttaaaaggttctgc | agaaagtcctcagatcccag |
| TGFBR1 ii | ggaggctatttgggggtgt | gcgagcgccggtttctg |
| TGFBR1 iii | actcacacagacacaccca | aagagcaggagcgagccag |
| TGFBR1 iv | ctaagagcaacaaacagtgc | gtcacttcttgcctctaaacg |
| TGFBR1 v | tgcaggaattgtgtaggattg | tggagctgacttattgattcg |
| TGFBR1 vi | ctccccagtgagataaattc | aatcttgaagaagttcctag |
| TGFBR1 vii | gcttactctgaggaactaaag | agatgcggttttgtcatgttg |
| TGFBR1 viii | aagtattgtaggtcatgtgg | gatattttctggaagggcaac |
| TGFBR1 ix | gtctgaaaggaggttcatc | caggaagagaatacactagg |
| TGFBR1 x | gtgatcttttaatgccttgg | aacattggtttgactgcta |
| TGFBR1 xi | caccagtaccctattgatgg | aaggagagttcaggcaaagc |
| TGFBR1 xii | gcaactcagtcaacaggaag | gaatcaaggaaactctagtgg |
| TGFBR1 xiii | agaaagtgatttactcct g | attcaaacatgaccatgc |
| TGFBR1 xiv | ctttctcctaccaaaatgtgc | ctgaattaaaagctgccttcc |
| TGFBR2 i | cctcctggctggcgagcg | ggaccaaacgtgccccgc |
| TGFBR2 ii | aagcaaatggctactcaacc | acacatacatgcagagaacacc |
| TGFBR2 iii | tgcgaatgctggagaacagg | ggaggacaccacctaacgtatg |
| TGFBR2 iv | agctgaagtttgaaggaagagc | gcacacggttgttgtagttggt |
| TGFBR2 v | catcatcttctactgctaccgc | ggttcccgttggatgtcctcat |
| TGFBR2 vi | ggagttggggaaacaatactgg | gggtcaagtcgtgtaaaaaagg |
| TGFBR2 vii | ctatctgtacctttctgtgc | ccaatacgatttgtcggatc |
| TGFBR2 viii | gttacttagtgcttcatgctcc | ccttccagggtaacacaagata |
| TGFBR2 ix | gtgttgggagtgttagtgtacc | ccgtaggtctaccacacactct |
| TGFBR2 x | accaactcatggtgccctttgg | cggtatggaacttttctctg |
| TGFBR2 xi | gctgtgttagcacttcctcagg | ggtttagaccccccgatcaaat |
| TGFBR2 xii | tgtttgaggaccagtgttcccg | ccgaggactaacgagttcgtgt |
Table 3.
WAVE conditions used for screening TGFBR1 and TGFBR2 genes.
| Primer Set | Tm°C | Melt Range°C | Run Temperature°C | WAVE Buffer B (%) |
| TGFBR1 i | 59.4 | 54 – 72 | 57 | 69–79 |
| 59 | 67–77 | |||
| 64 | 62–64 | |||
| TGFBR1 ii | 63.3 | 60 – 72 | 63 | 68–78 |
| 69 | 62–72 | |||
| TGFBR1 iii | 69.2 | 58 – 75 | 69 | 68–78 |
| 72 | 65–75 | |||
| TGFBR1 iv | 55.9 | 54 – 65 | 55 | 68–78 |
| 59 | 64–74 | |||
| TGFBR1 v | 57.2 | 54 – 65 | 55 | 62–72 |
| 59 | 58–68 | |||
| 61 | 56–58 | |||
| TGFBR1 vi | 56.1 | 54 – 65 | 56 | 68–78 |
| 59 | 65–75 | |||
| TGFBR1 vii | 56.5 | 54 – 68 | 55 | 69–79 |
| 58 | 66–76 | |||
| TGFBR1 viii | 57.0 | 54 – 65 | 57 | 67–77 |
| TGFBR1 ix | 57.4 | 55 – 67 | 56 | 67–77 |
| 58 | 64–75 | |||
| 62 | 61–71 | |||
| TGFBR1 x | 54.8 | 51 – 66 | 54 | 65–75 |
| 57 | 62–72 | |||
| TGFBR1 xi | 54.6 | 54 – 67 | 54 | 65–75 |
| 58 | 60–70 | |||
| TGFBR1 xii | 56.9 | 55 – 65 | 56 | 64–77 |
| 58 | 62–72 | |||
| TGFBR1 xiii | 55.8 | 52 – 62 | 56 | 67–77 |
| TGFBR1 xiv | 55.2 | 54 – 63 | 55 | 67–77 |
| 59 | 63–73 | |||
| TGFBR2 i | 66.1 | 59–74 | 60 | 63–73 |
| 65 | 58–68 | |||
| 67 | 56–66 | |||
| 70 | 53–63 | |||
| TGFBR2 ii | 66.3 | 61 – 72 | 64 | 58–68 |
| 66 | 56–66 | |||
| 69 | 52–62 | |||
| TGFBR2 iii | 58.5 | 57 – 61 | 59 | 61–71 |
| 61 | 59–69 | |||
| TGFBR2 iv | 59.8 | 55 – 65 | 56 | 67–77 |
| 61 | 60–70 | |||
| 63 | 58–68 | |||
| TGFBR2 v | 60.5 | 57–64 | 62 | 58–68 |
| TGFBR2 vi | 63.5 | 62–65 | 64 | 55–65 |
| TGFBR2 vii | 59.7 | 53–71 | 59 | 57–67 |
| 61 | 52–62 | |||
| TGFBR2 viii | 59.7 | 53 – 71 | 59 | 66–76 |
| 61 | 64–74 | |||
| TGFBR2 ix | 61.0 | 60–61 | 61 | 57–67 |
| TGFBR2 x | 57.0 | 51–66 | 52 | 63–73 |
| 58 | 57–67 | |||
| 62 | 53–63 | |||
| 64 | 50–60 | |||
| TGFBR2 xi | 54.4 | 53 – 66 | 54 | 67–77 |
| 56 | 65–75 | |||
| 60 | 61–71 | |||
| TGFBR2 xii | 57.0 | 56–60 | 59 | 63–73 |
Table 4.
TaqMan primers, probes, quencher and annealing temperature for relevant assays.
| Primer set | Primer 1 | Primer 2 | FAM™ Labelled Probe | Vic™ Labelled Probe | Fluorescent Quencher | Anneal (1 minute) |
| TGFB1 -800G>A | gctatcgcctgcacacagc | aggacagaagcggtcccat | tgcctccaacgtcaccaccatc | tctgcctccaacatcaccaccatc | TAMRA | 62°C |
| TGFB1 -509C>T | ttagccacatgggaggtgct | ccaggcggagaaggcttaa | acccttccatccctcaggtgtcct | ccctcccatccttcaggtgtcctg | TAMRA | 62°C |
| TGFB1 +869T>C | caccacaccagccctgttc | ccaggcgtcagcaccagta | agcagcggcagca | cagcagcagcagc | None | 60°C |
| TGFB1 +915G>C | Developed by Applied Biosystems (USA) as a Research and Development Kit | None | 56°C | |||
TGFBR1+72InsC was genotyped by a commercial Invader kit designed and manufactured by Third WAVE technologies (Madison, MI, USA)
Table 5.
Pyrosequencing primers, dispensation order and sequence to analyse for relevant assays
| Primer Set | Forward Primer | Reverse Primer | Sequence Primer | Dispensation order | Sequence to Analyse |
| TGFBR2 c.*747C>G | tcctgtgtgcccttatttctc | tgaaggtaaaaagtggggttc | agtttctaaactaggttgag | tcgagagtctac | c/ggagagtttctaaac |
| TGFBR2 c.1149G>A | gatcacactccatgtggg | ccagacgcagggaaagc | agagctccaatatcctc | tgatgagacgac tac | g/atgaagaacgacctaacc |
Results
We have submitted our annotated sequencing data for TGFBR1 and TGFBR2 genes as GenBank accession numbers DQ383416 – DQ383424 and DQ377553 – DQ377559 respectively. A total of fifteen variants were identified in these genes (TGFBR1, n = 5; TGFBR2, n = 10) of which eight were previously recorded in dbSNP; we have obtained unique NCBI identifiers for all novel SNPs (n = 7; Table 6).
Table 6.
Minor allele frequencies of identified variants in TGFBR1 and TGFBR2 genes, based on genotyping of 48 healthy control individuals. Accepted GenBank accession numbers for the reference sequences describing TGFBR1 and TGFBR2 gene variants are DQ383416 – DQ383424 and DQ377553 – DQ37759 respectively.
| Variant | Unique NCBI Identifier | Minor Allele Frequency (%) |
| aTGFBR1: c.694A>C | ss50394789 | 4.3 |
| aTGFBR1: c.*899T>C | ss50394790 | 2.2 |
| aTGFBR1: c.*921T>G | ss50394793 | 4.3 |
| aTGFBR1: c.*978G>A | ss50394791 | 1.1 |
| aTGFBR1: c.*1004A>T | ss50394792 | 2.2 |
| TGFBR2: c.263+7A>G | rs1155705 | 10.8 |
| aTGFBR2: c.263+17A>C | ss50394787 | 1.1 |
| TGFBR2: c.445-111A>G | rs17026161 | 2.2 |
| TGFBR2: c.445-4T>A | rs11466512 | 21.7 |
| aTGFBR2: c.1149G>A | ss50394788 | 1.1 |
| TGFBR2: c.1157C>T | rs2228048 | 2.2 |
| TGFBR2: c.1515-91C>A | rs2276767 | 4.3 |
| TGFBR2: c.*327-329delAT | rs4016180 | 21.7 |
| TGFBR2: c.*747C>G | rs11466531 | 7.6 |
| TGFBR2: c.*835C>A | rs17026332 | 2.2 |
aNovel SNPs accepted by dbSNP
The distribution of genotypes was found to be in Hardy-Weinberg equilibrium for all SNPs in both case and control groups. No significant differences were observed in genotype and allele frequencies between case and control groups for any of the SNPs assessed (Table 7). Logistic regression analysis for the clinical characteristics described in Table 1 did not reveal a significant association with any variant and diabetic nephropathy. Adjusted p values for these potential covariates are shown in Table 8.
Table 7.
Genotype and allele frequencies of selected SNPs in case and control groups. The five TGFB1 SNPs were selected on the basis of previous publications [23-27]. The remaining two TGFBR2 SNPs were selected from potentially functional SNPs identified through screening the gene as summarised in Table 1. Data are n (%)
| SNP | Genotype | Case | Control | p value | Allele | Case | Control | p value |
| TGFB1 -800G>A | GG | 188 (69.1) | 268 (73.0) | 0.56 | G | 454 (83.5) | 628 (85.6) | 0.30 |
| GA | 78 (28.7) | 92 (25.1) | ||||||
| AA | 6 (2.2) | 7 (1.9) | A | 90 (16.5) | 106 (14.4) | |||
| TGFB1 -509C>T | CC | 179 (65.8) | 245 (66.8) | 0.62 | C | 442 (81.3) | 595 (81.1) | 0.93 |
| CT | 84 (30.9) | 105 (28.6) | ||||||
| TT | 9 (3.3) | 17 (4.6) | T | 102 (18.7) | 139 (18.9) | |||
| TGFB1 +72InsC | - C | 221 (81.3) | 301 (82.0) | 0.88 | - C | 488 (89.7) | 663 (90.3) | 0.71 |
| +/- C | 46 (16.9) | 61 (16.6) | ||||||
| + C | 5 (1.8) | 5 (1.4) | +C | 56 (10.3) | 71 (9.7) | |||
| TGFB1 +869T>C | TT | 151 (55.5) | 204 (55.6) | 0.86 | T | 403 (74.1) | 540 (73.6) | 0.84 |
| TC | 101 (37.1) | 132 (36.0) | ||||||
| CC | 20 (7.4) | 31 (8.4) | C | 141 (25.9) | 194 (26.4) | |||
| TGFB1 +915G>C | GG | 219 (80.5) | 298 (81.2) | 0.98 | G | 488 (89.7) | 661 (90.1) | 0.84 |
| GC | 50 (18.4) | 65 (17.7) | ||||||
| CC | 3 (1.1) | 4 (1.1) | C | 56 (10.3) | 73 (9.9) | |||
| aTGFBR2 c.*747C>G | CC | 218 (90.5) | 287 (89.1) | 0.88 | C | 457 (94.8) | 606 (94.1) | 0.61 |
| CG | 21 (8.7) | 32 (9.9) | ||||||
| GG | 2 (0.8) | 3 (0.9) | G | 25 (5.2) | 38 (5.9) | |||
| aTGFBR2 c.1149G>A | GG | 232 (96.3) | 317 (98.4) | 0.10 | G | 473 (98.1) | 639 (99.2) | 0.10 |
| GA | 9 (3.7) | 5 (1.6) | ||||||
| AA | 0 | 0 | A | 9 (1.9) | 5 (0.8) |
a 272 cases and 367 controls were genotyped for TGFB1 SNPs with 241 cases and 322 controls genotyped for TGFBR2 variants.
Table 8.
Significance values following logistic regression analyses adjusting the association between diabetic nephropathy status and genotype for the listed potential confounders.
| p value adjusted for: | TGFB1 -800G>A | TGFB1 -509C>T | TGFB1 +72InsC | TGFB1 +869T>C | TGFB1 +915G>C | TGFBR2 c.*747C>G | TGFBR2 c.1149G>A |
| - | 0.56 | 0.62 | 0.89 | 0.86 | 0.98 | 0.88 | 0.11 |
| Gender | 0.56 | 0.45 | 0.98 | 0.92 | 0.94 | 0.88 | 0.09 |
| Age at Diagnosis | 0.56 | 0.61 | 0.88 | 0.86 | 0.98 | 0.88 | 0.11 |
| Duration of Diabetes | 0.62 | 0.69 | 0.94 | 0.94 | 0.98 | 0.56 | 0.26 |
| HbA1c | 0.44 | 0.63 | 0.96 | 0.80 | 0.97 | 0.86 | 0.16 |
| BMI | 0.79 | 0.72 | 0.63 | 0.79 | 0.97 | 0.22 | 0.72 |
| Mean BP | 0.94 | 0.93 | 0.68 | 0.99 | 0.77 | 0.31 | 0.85 |
| All of the above | 0.36 | 0.94 | 0.93 | 0.70 | 0.96 | 0.49 | 0.95 |
The level of observed LD between all genotyped variants within each gene, together with the raw |D'| and R2 scores are shown in the Figures (Figures 1, 2). The most common combinations of alleles observed 5' to 3' were GC-TG (49.4%); AC-TG (12.6%) and GT-CG (10.5%). In addition, we have identified a novel, rare variant at TGFB1: +728T>C (ss50394786).
Figure 1.
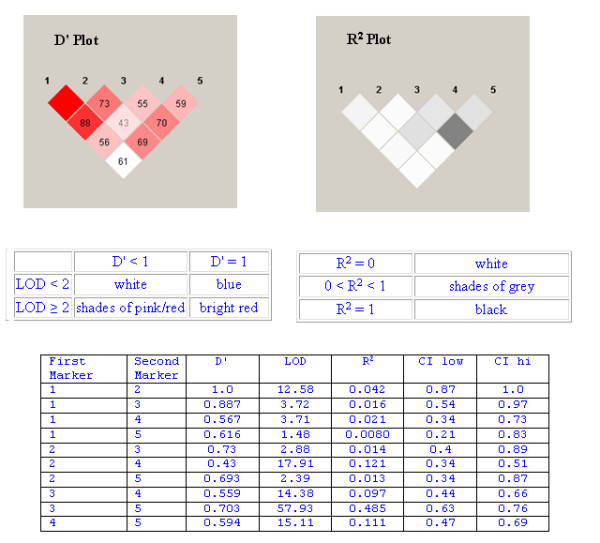
Although |D'| values were not particularly large for TGFB1 markers (D' Plot), they were statistically significant. R2 measure, there was little correlation observed between the genotyped markers (R2 Plot). Further details are displayed in the descriptive shown tables below the LD Plots. D' is the value of D primer between the two loci; LOD is the log of the likelihood odds ration (a measure of confidence in the value of D'); R2 is the correlation coefficient between the two loci and CI low/CI high represent 95% confidence limits for D' where the minor allele frequency is greater than 5%.
Figure 2.
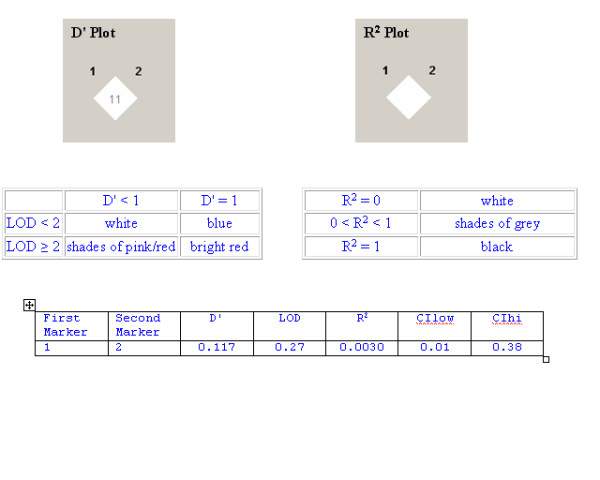
Strong linkage disequilibrium was not observed between the two genotyped markers for TGFBR2. Further details are displayed in the descriptive tables shown below the LD Plots.
Discussion
There is considerable evidence supporting the role of TGFB1, TGFBR1 and TGFBR2 genes in the development of diabetic nephropathy. The functional importance of the TGFβ1 protein is illustrated by the high degree of conservation observed across mammalian species. The TGFβ1 gene comprises seven exons separated by relatively large introns [23]. Transcriptional regulation is complex with two major start sites identified at positions +1 and +271 [24]; polymorphic positions are typically reported relative to the first major transcription start site for this gene. We have assessed five SNPs, suggested to play a role in TGFβ1 expression levels through their impact on regulatory regions of TGFB1 [25-27], for association with diabetic nephropathy. Genotype and allele frequencies were similar to those reported in other European populations, with a decreased frequency observed for the minor alleles of TGFB1: -509C>T and TGFB1: +869T>C compared to the ECTIM results [25,28,29]. As illustrated by the LD Plots (Figures 1, 2), we observed low correlation between the genotyped SNPs. The |D'| values suggest that these SNPs have been separated by recombination, however as the magnitude of |D'| is biased for smaller sample sizes |D'|, lower values may be difficult to interpret. To further clarify the patterns of LD between genetic markers, we also examined R2 values as this measure of LD is not as susceptible to small sample sizes. It is important to note that |D'| is less than one only when all haplotypes are observed and that markers with disparate alleles frequencies will also affect these measures of LD. The patterns of LD indicate that all genotyped loci within each gene are not contained within a single haplotype block, thereby suggesting limited value in performing haplotype analysis – i.e. the LD values suggest that these markers are not transmitted on a single haplotype, but are separated by recombination events. Our analysis suggests that all SNPs genotyped in this study are informative and the most common combinations of alleles observed are similar to those previously published [29]. Comparison of our data with Phase II, release 21 of the HapMap resource [30] supports our findings that the TGFB1 gene is located in a genomic region affected by a recombination hotspot. All recorded polymorphic sites in the HapMap CEPH (Utah residents with ancestry from northern and western Europe) population demonstrate poor correlation, suggesting that all should therefore be genotyped as 'tag' SNPs. None of the common variants genotyped for the TGFB1 gene in this study are recorded as polymorphic in the Phase II, release 21 HapMap Caucasian data set and the nearest polymorphic SNP is approximately 2 kb from our SNPs of interest.
We observed no significant differences in genotype or allele frequencies between case and control groups for any of the SNPs assessed. The TGFB1: +869T>C SNP has been associated with diabetic nephropathy in a Chinese population with type 2 diabetes [31], however our results do not support this finding for nephropathy in type 1 diabetes. The results from Wong and colleagues' smaller Chinese study (cases, n = 58; controls, n = 65), may be explained by a difference in genetic factors between type 1 and type 2 diabetes or differences between the Chinese and Irish populations.
Our study employed rigorous phenotypic criteria for inclusion of cases and controls. The annual incidence of diabetic nephropathy increases over the first fifteen to twenty years duration of type 1 diabetes, but after twenty-five years the absence of overt proteinuria makes the subsequent development of nephropathy unlikely [32,33]. The present report utilised cases and controls that were well matched for prolonged duration of diabetes (cases mean duration = 26.9 ± SD 8.3 years, control mean duration = 27.7 ± SD 9.0years). Our results for TGFB1 are in accord with Ng and colleagues' study which also failed to find an association between the TGFB1: -800G>A, -509C>T, +869T>C or +915G>C polymorphisms and diabetic nephropathy in US Caucasians with type 1 diabetes [34]. This is in contrast to a larger study in UK Caucasians where a significant association (p = 0.027) was identified between TGFB1: +869T>C and diabetic nephropathy [35]. This UK study utilised the Golden Years cohort of type 1 diabetic individuals as a control population [31] with all the recruited subjects (n = 410) having a very long duration of type 1 diabetes (> 50 years). Although these individuals did not have renal failure due to diabetic nephropathy 29% were taking antihypertensive medication and 35.7% had evidence of micro- or macroalbuminuria [36]. These clinical features (antihypertensive medication and micro- or macroalbuminura) form distinct exclusion criteria from our own diabetic control group. Although our sample size has ~90% power to detect a doubling in the minor allele frequency in cases relative to controls (e.g. 10% vs. 5%), there is a need for a collaborative genotyping effort in larger sample collections to definitively determine the role of these SNPs in predisposition to diabetic nephropathy.
TGFβ type I receptors form a heterodimeric complex with TGFβ type II receptors and bind to TGFβ to mediate many TGFβ activities including regulation of cell proliferation, differentiation and extracellular matrix production. It has been recently reported that TGFβ1-mediated epithelial-to-mesenchymal transition requires functional TGFBR2 [37]. Variants have been recorded for both TGFBR1 and TGFBR2 genes, however there is limited genomic information regarding their influence on diabetic nephropathy. TGFBR1 is composed of nine exons and maps to chromosome 9q33-q34 [38]. TGFBR2 is composed of seven exons and maps to 3p22 [39]. There are presently 409 validated SNPs recorded in dbSNP for these two genes (TGFBR1, n = 115; TGFBR2, n = 294; dbSNP, accessed 12/01/06). Due to the large number of reported SNPs and potential ethnic variation in SNP occurrence and frequency [40], we resequenced these genes in our population.
We prioritised screening of the protein coding regions of these genes to aid identification of potentially functional gene variants. We screened for variants directly affecting lariat regions, splice sites, exonic/intronic splice enhancers, signal sequences, protein coding sequence, polyadenylation signals and untranslated regions. It is possible that other variants may affect regulatory mechanisms, (promoter or enhancer elements, microRNA etc.) or that features such as post-translation modifications may affect these candidate genes and their subsequent protein activity. It is also possible that rare variants may play a role in susceptibility to diabetic nephropathy; however this study lacks sufficient sample numbers to definitively assess the role of rare variants in this disease. Five novel variants were identified in TGFBR1, of which none were at sufficient frequency to assess in this case-control collection. We have identified nine SNPs in TGFBR2. Two SNPs are located in exons (TGFBR2: c.1157C>T, TGFBR2: c.1149G>A) and one SNP in the 3' UTR which was found to be putatively functional (TGFBR2: c.*747C>G). Analysis of TGFBR2: c.*747C>G genotyping did not reveal a significant association with diabetic nephropathy. A microsatellite [AT]del was also identified in the 3' UTR of TGFBR2. TGFBR2: c.1157C>T in exon four was found in only one sample in the heterozygous state (MAF: 1.1%), and does not lead to a change in amino acid (aac → aat = N389N). TGFBR2: c.1149G>A was found in only two samples (MAF: 2.2%), but leads to a non-synonymous change in amino acid (gtg → atg = V387M) in the serine-threonine protein kinase active domain of the mature chain for TGFβR2 (PROSITE: PS00108; Pfam: PF00069, accessed 03/02/06). Genotyping TGFBR2: c.1149G>A did not reveal a significant association with diabetic nephropathy, however we did identify a doubling of the minor allele in cases (MAF: 1.9% in cases vs. 0.8% in controls). This finding may be due to the low frequency of minor allele, however our available sample numbers do not provide sufficient power to appropriately assess the association of this SNP with diabetic nephropathy. The search for causative variants for susceptibility to diabetic nephropathy is constrained by limited numbers of well-characterised, precisely phenotyped cases and controls, which represents a major challenge in the study of complex disease genetics. While the power to identify disease gene loci is influenced by many factors, the requirement for adequate samples sizes of stringently phenotyped individuals is critical to the success and validity of complex disease association studies. Our results warrant further investigations of rare variants, particularly the TGFBR2 exonic SNPs, provided the sample population is sufficiently powered to assess the association.
Conclusion
Although experimental evidence suggests TGFβ1 blockade may be an important therapeutic target we were unable to identify any association between TGFB1 gene variants and diabetic nephropathy. In resequencing the genes we identified eight novel variants for TGFB1, TGFBR1 and TGFBR2 genes but did not detect significant association between any of the common SNPs and nephropathy in this Caucasian population with type 1 diabetes.
Abbreviations
MAF: Minor allele frequency, SNP: Single nucleotide polymorphism, UTR: Untranslated region, TGF: Transforming growth factor
Competing interests
The author(s) declare that they have no competing interests.
Authors' contributions
AJM: Contributed to the conception and design of this study, carried out all resequencing and genotyping work, submitted all data to online repositories, drafted the manuscript and approved final manuscript.
DAS: Participated in study conception and design, critically reviewed the manuscript and approved final manuscript.
CCP: Performed all statistical analyses, contributed to data interpretation, critically reviewed the manuscript and approved final manuscript.
DS: Provided clinical supervision of patient recruitment in the Republic of Ireland, critically reviewed the manuscript and approved final manuscript.
APM: Participated in study conception and design, supervised recruitment of patients in Northern Ireland, contributed to data interpretation, co-wrote the manuscript and approved final manuscript.
Pre-publication history
The pre-publication history for this paper can be accessed here:
Acknowledgments
Acknowledgements
This work was funded by the Research and Development Office of Northern Ireland and the Health Research Board of Ireland. A. J. McK is supported by a Northern Ireland Kidney Research Fund postdoctoral fellowship.
Contributor Information
Amy Jayne McKnight, Email: a.j.mcknight@lineone.net.
David A Savage, Email: d.savage@qub.ac.uk.
Chris C Patterson, Email: c.patterson@qub.ac.uk.
Denise Sadlier, Email: DSADLIER@partners.org.
A Peter Maxwell, Email: a.p.maxwell@qub.ac.uk.
References
- USRDS 2005 United States Renal Data System, Annual Data Report (ADR) Atlas 2005 http://www.usrds.org/adr.htm
- Seaquist ER, Goetz FC, Rich S, Barbosa J. Familial clustering of diabetic kidney disease. Evidence for genetic susceptibility to diabetic nephropathy. N Engl J Med. 1989;320:1161–1165. doi: 10.1056/NEJM198905043201801. [DOI] [PubMed] [Google Scholar]
- Quinn M, Angelico MC, Warram JH, Krolewski A. Familial factors determine the development of diabetic nephropathy in patients with IDDM. Diabetologia. 1996;39:940–945. doi: 10.1007/BF00403913. [DOI] [PubMed] [Google Scholar]
- Krolewski M, Eggers PW, Warram JH. Magnitude of end-stage renal disease in IDDM: a 35 year follow-up study. Kidney Int. 1996;50:2041–2046. doi: 10.1038/ki.1996.527. [DOI] [PubMed] [Google Scholar]
- Khoury MJ, Beaty TH, Liang KY. Can familial aggregation of disease be explained by familial aggregation of environmental risk factors? Am J Epidemiol. 1988;127:674–683. doi: 10.1093/oxfordjournals.aje.a114842. [DOI] [PubMed] [Google Scholar]
- Reeves WB, Andreoli TE. Transforming growth factor beta contributes to progressive diabetic nephropathy. Proc Natl Acad Sci USA. 2000;97:7667–7669. doi: 10.1073/pnas.97.14.7667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto T, Nakamura T, Noble NA, Ruoslahti E, Border WA. Expression of transforming growth factor beta is elevated in human and experimental diabetic nephropathy. Proc Natl Acad Sci USA. 1993;90:1814–1818. doi: 10.1073/pnas.90.5.1814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffman BB, Sharma K, Zhu Y, Ziyadeh FN. Transcriptional activation of transforming growth factor-beta1 in mesangial cell culture by high glucose concentration. Kidney Int. 1998;54:1107–1116. doi: 10.1046/j.1523-1755.1998.00119.x. [DOI] [PubMed] [Google Scholar]
- Isaka Y, Fujiwara Y, Ueda N, Kaneda Y, Kamada T, Imai E. Glomerulosclerosis induced by in vivo transfection of transforming growth factor-beta or platelet-derived growth factor gene into the rat kidney. J Clin Invest. 1993;92:2597–2601. doi: 10.1172/JCI116874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kopp JB, Factor VM, Mozes M, Nagy P, Sanderson N, Bottinger EP, Klotman PE, Thorgeirsson SS. Transgenic mice with increased plasma levels of TGF-beta 1 develop progressive renal disease. Lab Invest. 1996;74:991–1003. [PubMed] [Google Scholar]
- Ziyadeh FN, Hoffman BB, Han DC, Iglesias-De La Cruz MC, Hong SW, Isono M, Chen S, McGowan TA, Sharma K. Long-term prevention of renal insufficiency, excess matrix gene expression, and glomerular mesangial matrix expansion by treatment with monoclonal antitransforming growth factor-beta antibody in db/db diabetic mice. Proc Natl Acad Sci USA. 2000;97:8015–8020. doi: 10.1073/pnas.120055097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han DC, Hoffman BB, Hong SW, Guo J, Ziyadeh FN. Therapy with antisense TGF-beta1 oligodeoxynucleotides reduces kidney weight and matrix mRNAs in diabetic mice. Am J Physiol Renal Physiol. 2000;278:F628–F634. doi: 10.1152/ajprenal.2000.278.4.F628. [DOI] [PubMed] [Google Scholar]
- Shankland SJ, Pippin J, Pichler RH, Gordon KL, Friedman S, Gold LI, Johnson RJ, Couser WG. Differential expression of transforming growth factor-beta isoforms and receptors in experimental membranous nephropathy. Kidney Int. 1996;50:116–124. doi: 10.1038/ki.1996.294. [DOI] [PubMed] [Google Scholar]
- Yamamoto T, Watanabe T, Ikegaya N, Fujigaki Y, Matsui K, Masaoka H, Nagase M, Hishida A. Expression of types I, II, and III TGF-beta receptors in human glomerulonephritis. J Am Soc Nephrol. 1998;9:2253–2261. doi: 10.1681/ASN.V9122253. [DOI] [PubMed] [Google Scholar]
- Yang YL, Guh JY, Yang ML, Lai YH, Tsai JH, Hung WC, Chang CC, Chuang LY. Interaction between high glucose and TGF-beta in cell cycle protein regulations in MDCK cells. J Am Soc Nephrol. 1998;9:182–193. doi: 10.1681/ASN.V92182. [DOI] [PubMed] [Google Scholar]
- Isono M, Mogyorosi A, Han DC, Hoffman BB, Ziyadeh FN. Stimulation of TGF-beta type II receptor by high glucose in mouse mesangial cells and in diabetic kidney. Am J Physiol Renal Physiol. 2000;278:F830–F838. doi: 10.1152/ajprenal.2000.278.5.F830. [DOI] [PubMed] [Google Scholar]
- Fujii D, Brissenden JE, Derynck R, Francke U. Transforming growth factor beta gene maps to human chromosome 19 long arm and to mouse chromosome 7. Somat Cell Mol Genet. 1986;12:281–288. doi: 10.1007/BF01570787. [DOI] [PubMed] [Google Scholar]
- McKnight AJ, Savage DA, Patterson CC, Brady HR, Maxwell AP. Resequencing of the characterised CTGF gene to identify novel or known variants and analysis of their association with diabetic nephropathy. J Hum Genet. 2006;51:383–386. doi: 10.1007/s10038-006-0368-7. [DOI] [PubMed] [Google Scholar]
- National Centre for Biotechnology Information http://www.ncbi.nlm.nih.gov
- Genome Variation Society recommendations for coding sequences http://www.genomic.unimelb.edu.au/mdi/mutnomen/recs.html#DNA
- Boreham C, Savage JM, Primrose D, Cran G, Strain J. Coronary risk factors in school-children. Arch Dis Child. 1993;68:182–186. doi: 10.1136/adc.68.2.182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21:263–265. doi: 10.1093/bioinformatics/bth457. [DOI] [PubMed] [Google Scholar]
- Derynck R, Rhee L, Chen EY, van Tilburg A. Intron-exon structure of the human transforming growth factor-beta precursor gene. Nucleic Acids Res. 1987;15:3188–3189. doi: 10.1093/nar/15.7.3188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim SJ, Glick A, Sporn MB, Roberts AB. Characterization of the promoter region of the human transforming growth factor-beta 1 gene. J Biol Chem. 1989;264:402–408. [PubMed] [Google Scholar]
- Cambien F, Ricard S, Troesch A, Mallet C, Generenaz L, Evans A, Arveiler D, Luc G, Ruidavets JB, Poirier O. Polymorphisms of the transforming growth factor-beta 1 gene in relation to myocardial infarction and blood pressure. The Etude Cas-Temoin de l'Infarctus du Myocarde (ECTIM) Study. Hypertension. 1996;28:881–887. doi: 10.1161/01.hyp.28.5.881. [DOI] [PubMed] [Google Scholar]
- Grainger DJ, Heathcote K, Chiano M, Snieder H, Kemp PR, Metcalfe JC, Carter ND, Spector TD. Genetic control of the circulating concentration of transforming growth factor type beta1. Hum Mol Genet. 1999;8:93–97. doi: 10.1093/hmg/8.1.93. [DOI] [PubMed] [Google Scholar]
- Awad MR, El Gamel A, Hasleton P, Turner DM, Sinnott PJ, Hutchinson IV. Genotypic variation in the transforming growth factor-beta1 gene: association with transforming growth factor-beta1 production, fibrotic lung disease, and graft fibrosis after lung transplantation. Transplantation. 1998;66:1014–1020. doi: 10.1097/00007890-199810270-00009. [DOI] [PubMed] [Google Scholar]
- Lympany PA, Avila JJ, Mullighan C, Marshall S, Welsh KI, du Bois RM. Rapid genotyping of transforming growth factor beta1 gene polymorphisms in a UK Caucasoid control population using the polymerase chain reaction and sequence-specific primers. Tissue Antigens. 1998;52:573–578. doi: 10.1111/j.1399-0039.1998.tb03089.x. [DOI] [PubMed] [Google Scholar]
- Syrris P, Carter ND, Metcalfe JC, Kemp PR, Grainger DJ, Kaski JC, Crossman DC, Francis SE, Gunn J, Jeffery S, Heathcote K. Transforming growth factor-beta1 gene polymorphisms and coronary artery disease. Clin Sci (Lond) 1998;95:659–667. doi: 10.1042/cs0950659. [DOI] [PubMed] [Google Scholar]
- HapMap http://www.HapMap.org accessed 12/10/06.
- Wong TY, Poon P, Chow KM, Szeto CC, Cheung MK, Li PK. Association of transforming growth factor-beta (TGF-beta) T869C (Leu10Pro) gene polymorphisms with type 2 diabetic nephropathy in Chinese. Kidney Int. 2003;63:1831–1835. doi: 10.1046/j.1523-1755.2003.00919.x. [DOI] [PubMed] [Google Scholar]
- Doria A, Warram JH, Krolewski AS. Genetic susceptibility to nephropathy in insulin-dependent diabetes: from epidemiology to molecular genetics. Diabetes Metab Rev. 1995;11:287–314. doi: 10.1002/dmr.5610110402. [DOI] [PubMed] [Google Scholar]
- Mogensen CE. How to protect the kidney in diabetic patients: with special reference to IDDM. Diabetes. 1997;46:S104–S111. doi: 10.2337/diab.46.2.s104. [DOI] [PubMed] [Google Scholar]
- Ng DP, Warram JH, Krolewski AS. TGF-beta 1 as a genetic susceptibility locus for advanced diabetic nephropathy in type 1 diabetes mellitus: an investigation of multiple known DNA sequence variants. Am J Kidney Dis. 2003;41:22–28. doi: 10.1053/ajkd.2003.50011. [DOI] [PubMed] [Google Scholar]
- Patel A, Scott WR, Lympany PA, Rippin JD, Gill GV, Barnett AH, Bain SC, Warren 3/UK GoKind Study Group The TGF-beta 1 gene codon 10 polymorphism contributes to the genetic predisposition to nephropathy in Type 1 diabetes. Diabet Med. 2005;22:69–73. doi: 10.1111/j.1464-5491.2005.01376.x. [DOI] [PubMed] [Google Scholar]
- Bain SC, Gill GV, Dyer PH, Jones AF, Murphy M, Jones KE, Smyth C, Barnett AH. Characteristics of type 1 diabetes of over 50 years duration (the Golden Years Cohort) Diabet Med. 2003;20:808–811. doi: 10.1046/j.1464-5491.2003.01029.x. [DOI] [PubMed] [Google Scholar]
- Han G, Lu SL, Li AG, He W, Corless CL, Kulesz-Martin M, Wang XJ. Distinct mechanisms of TGF-beta1-mediated epithelial-to-mesenchymal transition and metastasis during skin carcinogenesis. J Clin Invest. 2005;115:1714–1723. doi: 10.1172/JCI24399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson DW, Qumsiyeh M, Benkhalifa M, Marchuk DA. Assignment of human transforming growth factor-beta type I and type III receptor genes (TGFBR1 and TGFBR3) to 9q33-q34 and 1p32-p33, respectively. Genomics. 1995;28:356–357. doi: 10.1006/geno.1995.1157. [DOI] [PubMed] [Google Scholar]
- Matthew S, Murty VV, Cheifetz S, George D, Massague J, Chaganti RS. Transforming growth factor receptor gene TGFBR2 maps to human chromosome band 3p22. Genomics. 1994;20:114–115. doi: 10.1006/geno.1994.1134. [DOI] [PubMed] [Google Scholar]
- Mori M, Yamada R, Kobayashi K, Kawaida R, Yamamoto K. Ethnic differences in allele frequency of autoimmune-disease-associated SNPs. J Hum Genet. 2005;50:264–266. doi: 10.1007/s10038-005-0246-8. [DOI] [PubMed] [Google Scholar]


