Abstract
Regulation of the cell-specific transcription factor σF in the spore-forming bacterium Bacillus subtilis involves the antisigma factor SpoIIAB. Contributing to the activation of σF is the degradation of SpoIIAB in a manner that depends on the protease ClpCP. Here we show that the three residues (LCN) located at the extreme C terminus of SpoIIAB are both necessary and sufficient for this degradation. We also report that the use of the LCN extension as a degradation signal for ClpCP is unique to SpoIIAB.
Proteolysis is a common feature of the regulation of a wide range of processes in bacteria, including cell division in Escherichia coli (2, 14, 18), cell differentiation in Caulobacter crescentus (5, 24), and entry into the state of genetic competence in Bacillus subtilis (26). A crucial aspect of the involvement of proteolysis in regulation is the correct recognition of the proper substrates for degradation. Here we are concerned with the involvement of proteolysis in the activation of the transcription factor σF during the process of sporulation in B. subtilis. Sporulation involves the formation of an asymmetrically positioned (polar) septum that divides the developing cell (or sporangium) into unequally sized compartments called the forespore and the mother cell. The σF factor is a cell-specific regulatory protein that is produced in the predivisional cell but is activated only after the polar septum is formed when σF-directed gene transcription is confined to the forespore. The cell-specific activation of σF is governed by a multicomponent pathway, central to which is an antagonist of σF called SpoIIAB (for details, see reference 22). Briefly, SpoIIAB is an antisigma factor that binds to σF, holding it in an inactive complex (7, 17). Escape from the complex is mediated by an anti-antisigma factor called SpoIIAA (1, 4). The SpoIIAA protein reacts with SpoIIAB · σF in the forespore, causing it to dissociate into free (and active) σF and free SpoIIAB (10, 11, 13). We recently reported that free SpoIIAB (but not SpoIIAB in a complex with σF) is subject to proteolysis in a manner that depends on the ATP-dependent, intracellular protease ClpCP. We proposed that ClpCP-mediated degradation could set up a self-reinforcing cycle that would help to lock on σF activity in the forespore (21). Also, degradation of SpoIIAB could help to establish forespore-specific activation of σF by a previously described transient genetic asymmetry mechanism (8, 9).
The ClpCP protease is composed of ClpP, which contains the proteolytic active site, and ClpC, an ATPase that recognizes and unfolds substrates and translocates them into the catalytic chamber of ClpP (23, 25). In B. subtilis, transcription of the structural genes for ClpP and ClpC is repressed by CtsR. This repression is relieved when cells enter stationary phase and under conditions of stress (3, 15). Cells that are mutant for ClpCP are impaired in sporulation: cells with clpP deleted cannot initiate sporulation, and cells lacking clpC are reduced in their capacity to form polar septa (12, 19, 21). Importantly, SpoIIAB proteolysis is blocked in cells lacking either of the components of the ClpCP protease both during sporulation and when SpoIIAB is artificially produced in growing cells (21). This proteolysis depends on three residues, Leu-144, Cys-145, and Asn-146 (LCN), located at the extreme C terminus of the 146-residue-long polypeptide. A truncated form of SpoIIAB lacking these terminal residues is not subject to degradation, and mutant forms of the protein harboring glutamate substitutions at each of the three positions are substantially more stable than the wild-type protein. We concluded that the C-terminal residues of SpoIIAB were necessary for ClpCP-dependent proteolysis (21). Here we show that the LCN extension on the C terminus of SpoIIAB is both necessary and sufficient for proteolysis and that, as predicted by previous models, this proteolysis takes place in the forespore chamber of the sporangium.
Strains.
The strains used in this study are described in Table 1.
TABLE 1.
B. subtilis strains
| Strain | Relevant genotype |
|---|---|
| QPB656 | amyE::PspoIIA-gfpLCN |
| QPB657 | amyE::PspoIIA-gfp |
| QPB661 | clpCΔ amyE::PspoIIA-gfpLCN |
| QPB662 | clpCΔ amyE::PspoIIA-gfp |
| QPB671 | spoIIAΔ thrC::spoIIAB amyE::PspoIIA-gfpLCN |
| QPB672 | spoIIAΔ thrC::spoIIAB amyE::PspoIIA-gfp |
| QPB673 | clpCΔ spoIIAΔ thrC::spoIIAB amyE::PspoIIA-gfpLCN |
| QPB674 | clpCΔ spoIIAΔ thrC::spoIIAB amyE::PspoIIA-gfp |
| QPB687 | amyE::PspoIIA-gfpEEN |
| QPB725 | amyE::PspoIIQ-gfp |
| QPB726 | amyE::PspoIIQ-gfpLCN |
| QPB729 | amyE::PspoIID-gfp |
| QPB730 | amyE::PspoIID-gfpLCN |
| QPB734 | amyE::PspoIIA-gfpLCNE |
| QPB735 | amyE::PspoIIA-gfpLCNEEE |
The LCN extension is both necessary and sufficient for proteolysis.
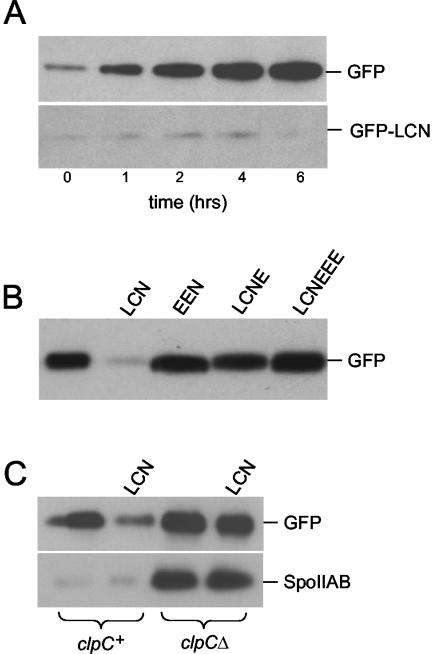
Our strategy to investigate the function of the LCN extension was to append codons for leucine, cysteine, and asparagine at the 3′ terminus of the coding sequence for the green fluorescent protein (GFP). The gene for this GFP-LCN fusion protein was initially placed under the control of the promoter (PspoIIA) for the operon (spoIIA) that contains the gene for SpoIIAB. Immunoblot analysis with anti-GFP antibodies showed that GFP accumulated to high levels during sporulation, whereas the fusion protein GFP-LCN showed little accumulation and was largely if not entirely absent at 6 h after the start of sporulation (Fig. 1A).
FIG. 1.
Accumulation of GFP and GFP fusion proteins during sporulation. Cells were induced to sporulate by exhaustion in DS medium. (A) Samples were collected at the indicated hours after the end of the exponential phase of growth. Cell pellets were lysed, and lysates were normalized according to protein concentration as described previously (21). Accumulation of GFP (QPB657) and GFP-LCN (QPB656) was monitored by immunoblot analysis. (B) Cells containing GFP (QPB657) and its variants (GFP-LCN [QPB656], GFP-EEN [QPB687], GFP-LCNE [QPB734], and GFP-LCNEEE [QPB735]) were collected 2.5 h after they entered stationary phase (T2.5) and used for immunoblot analysis. (C) Effect of clpCΔ on GFP-LCN and SpoIIAB accumulation during sporulation. The strains used were (from left to right) QPB672, -671, -674, and -673. Samples were collected at T2.5.
Next, to investigate the sequence specificity of the LCN degradation signal, we created a fusion protein harboring glutamates instead of leucine and cysteine in the extension. Immunoblot analysis showed that the GFP-EEN fusion protein accumulated to levels comparable to that of GFP (Fig. 1B). We also created a fusion protein in which a glutamate residue was appended at the C-terminal end of the LCN extension. Like GFP itself, the resulting GFP-LCNE fusion protein accumulated to high levels during sporulation (Fig. 1B).
These findings are consistent with the idea that LCN is both necessary and sufficient to prevent the accumulation of GFP during sporulation and that the three-residue extension must be displayed at the extreme C terminus of the protein to be effective. Pulse-chase experiments had previously demonstrated that the failure of free SpoIIAB to accumulate during sporulation is due to proteolysis. It seems likely that the effect of LCN in impeding the accumulation of GFP is also exerted at the level of protein degradation, and, for simplicity, we will assume this to be the case in what follows. With this assumption in mind, we conclude that LCN is a degradation signal and that its presence at the extreme C terminus of SpoIIAB is necessary and sufficient to account for the proteolytic instability of the antisigma factor.
LCN confers instability on GFP in a ClpCP-dependent manner.
Proteolysis of SpoIIAB is dependent on ClpCP. To investigate whether the effect of the LCN extension on GFP was also ClpCP dependent, we introduced a deletion mutation of clpC into cells producing the GFP-LCN fusion protein. Immunoblot analysis showed that the fusion protein accumulated to high levels in the absence of clpC but not in its presence (Fig. 1C). As an internal control and in agreement with previous results, immunoblot analysis of the same extracts used in the GFP-LCN analysis showed that accumulation of SpoIIAB exhibited a similar pattern of clpC-dependent accumulation (Fig. 1C).
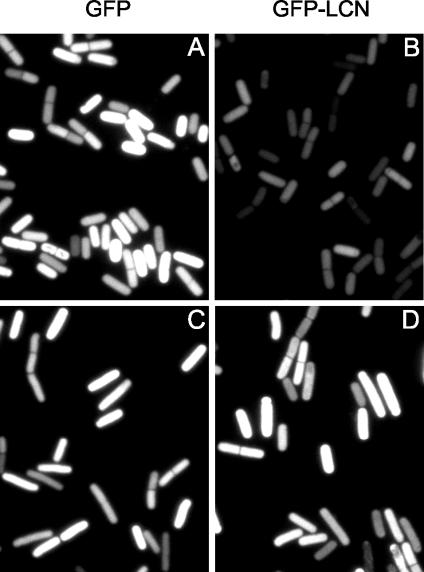
We also took advantage of the intrinsic fluorescence of GFP to visualize its accumulation in living cells by fluorescence microscopy. The results show that whereas GFP produced a strong fluorescence signal during sporulation (Fig. 2A), the GFP-LCN fusion protein did not (Fig. 2B). However, in a strain lacking clpC, many cells that exhibited a high level of fluorescence from GFP-LCN were observed (Fig. 2D). Notice that as observed before (21), sporulating cells with a deletion in clpC were slightly elongated and were impaired in asymmetric division (Fig. 2C and D and data not shown).
FIG. 2.
Visualization of GFP and GFP-LCN by fluorescence microscopy. Cells were induced to sporulate by exhaustion in DS medium. clpC+ cells (QPB657 [A] and QPB656 [B]) and clpCΔ cells (QPB662 [C] and QPB661 [D]) were collected for microscopy at 2.5 h after they entered stationary phase. The exposure time was 1 s.
ClpCP is active in both the forespore and the mother cell.
A previous model for the role of proteolysis in the activation of σF is based on the idea that ClpCP-dependent degradation of SpoIIAB takes place preferentially in the forespore (21). In this substrate availability model, it was imagined that ClpCP is present and active in both the mother cell and the forespore but that SpoIIAB only becomes available as a substrate for the protease as a result of the SpoIIAA-mediated dissociation of the SpoIIAB·σF complex into free σF and free SpoIIAB (recall that only free SpoIIAB is subject to ClpCP-dependent degradation). Therefore, preferential dissociation of SpoIIAB·σF in the forespore could lead to preferential degradation of the antisigma factor in the forespore.
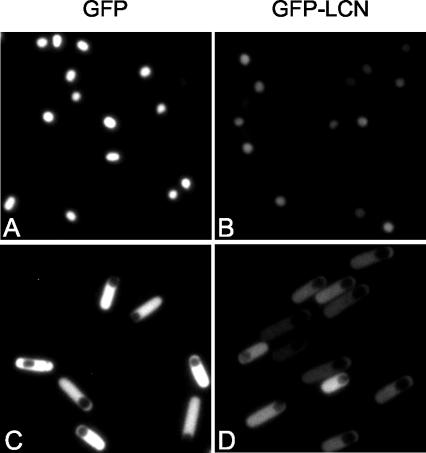
The discovery that the LCN degradation signal can confer ClpCP-dependent instability on GFP provided a convenient opportunity to test directly whether ClpCP is indeed active in both the forespore and mother-cell compartments of the postdivisional sporangium. To do so, we placed the coding sequences for GFP and for GFP-LCN under the control of a promoter (PspoIIQ) for a gene (spoIIQ) that is activated in the forespore under the control of σF and a promoter (PspoIID) for a gene (spoIID) that is switched on in the mother cell under the control of the early-acting, mother-cell-specific transcription factor σE (22). GFP and GFP-LCN were then visualized by fluorescence microscopy. Strong fluorescence was observed in the forespore of sporangia harboring the PspoIIQ-gfp fusion (Fig. 3A) but not in the forespore of sporangia harboring the PspoIIQ-gfpLCN fusion (Fig. 3B). Likewise, strong fluorescence was observed in the mother cell of sporangia harboring the PspoIID-gfp fusion (Fig. 3C) but not in the mother cell of sporangia harboring the PspoIID-gfpLCN fusion (Fig. 3D).
FIG. 3.
Visualization of GFP and GFP-LCN in postdivisional sporangia. Cells were induced to sporulate by exhaustion in DS medium. Cells were collected for microscopy at 3.5 h after they entered stationary phase. Exposure times were 0.3 s (A and B) and 0.5 s (C and D). The strains used were QPB725 (PspoIIQ-gfp) (A), QPB726 (PspoIIQ-gfpLCN) (B), QPB729 (PspoIID-gfp) (C), and QPB730 (PspoIID-gfpLCN) (D).
We also took advantage of the LCN-dependent instability of GFP to investigate the activity of ClpCP at a later stage of development in both the forespore and the mother cell. For this purpose, we created fusions of the coding sequences for GFP and GFP-LCN to a promoter (PsspB) under the control of the late-acting, forespore-specific transcription factor σG and a promoter (PgerE) under the control of the late-acting, mother- cell-specific transcription factor σK. In both cases, much lower accumulation was observed for GFP-LCN than for GFP (data not shown). We conclude that the ClpCP protease is active throughout sporulation, both before (Fig. 2B) and after asymmetric division and in both compartments of the postdivisional sporangium (Fig. 3B and D).
Use of LCN as a degradation signal is unique to SpoIIAB.
Strikingly, SpoIIAB is the only protein from the annotations of the B. subtilis genome project that contains LCN at its extreme C terminus (16). Evidently, the use of LCN as a degradation determinant is unique to SpoIIAB. On the other hand, the LCN C-terminal extension is conserved identically (in B. anthracis, B. cereus, B. coagulans, B. licheniformis, B. megaterium, and B. stearothermophilus) or approximately so (ICN in B. halodurans) in SpoIIAB orthologs of other Bacillus species, with the exception of B. sphaericus (AVM). Conceivably, σF regulation in B. sphaericus does not involve ClpCP-dependent proteolysis. Alternatively, ClpCP of B. sphaericus is capable of recognizing a different degradation signal at the C terminus of SpoIIAB. Interestingly, a known substrate (Spx) for ClpXP in B. subtilis (20) has a similar extension (LAN) at its C terminus. ClpXP is an ATP-dependent protease that has the same catalytic core (ClpP) as ClpCP but uses a different ATPase component (ClpX) for substrate recognition. To investigate whether the LAN extension would trigger degradation in a ClpCP-dependent manner, we created a GFP-LAN fusion protein. The presence of the LAN extension did indeed impair the accumulation of the fusion protein, but the accumulation of GFP-LAN was not measurably increased in mutant cells lacking ClpC (data not shown). Conceivably, a single residue difference (C→A) at the penultimate position is responsible for rendering Spx a substrate for ClpXP.
Finally, we come to the question of the role of the LCN extension in the ClpCP-dependent proteolysis of SpoIIAB. The simplest interpretation of our results would be that the LCN degradation signal is directly recognized by ClpCP. However, several lines of evidence argue against this. First, we have failed to detect proteolysis of SpoIIAB with purified ClpCP under conditions in which efficient degradation of a bona fide ClpCP substrate, MecA, was observed (data not shown). Second, we have failed to detect an interaction between SpoIIAB (or GFP-LCN) and ClpC, the substrate recognition and denaturing component of the protease, by chemical cross-linking or use of the yeast two-hybrid system (data not shown). The most likely explanation for these negative results is that recognition of SpoIIAB by ClpCP depends on an unknown adaptor protein that recruits the sporulation protein to the proteolytic enzyme. Indeed, many of the ATPase-associated proteases in bacteria, including ClpCP itself, depend on adaptor proteins for recognition of certain substrates (6). It will be interesting to learn the identity of such an adaptor, if it exists, and to determine whether it is dedicated uniquely to the recognition of SpoIIAB.
Acknowledgments
We thank K. Carniol, M. Fujita, E. Gonzalez-Pastor, D. Kearns, D. Rudner, and K. Turgay for strains and helpful advice.
This work was supported by National Institutes of Health grant GM18568 to R.L.
REFERENCES
- 1.Alper, S., L. Duncan, and R. Losick. 1994. An adenosine nucleotide switch controlling the activity of a cell type-specific transcription factor in B. subtilis. Cell 77:195-205. [DOI] [PubMed] [Google Scholar]
- 2.Canceill, D., E. Dervyn, and O. Huisman. 1990. Proteolysis and modulation of the activity of the cell division inhibitor SulA in Escherichia coli lon mutants. J. Bacteriol. 172:7297-7300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Derre, I., G. Rapoport, and T. Msadek. 1999. CtsR, a novel regulator of stress and heat shock response, controls clp and molecular chaperone gene expression in gram-positive bacteria. Mol. Microbiol. 31:117-131. [DOI] [PubMed] [Google Scholar]
- 4.Diederich, B., J. F. Wilkinson, T. Magnin, M. Najafi, J. Erringston, and M. D. Yudkin. 1994. Role of interactions between SpoIIAA and SpoIIAB in regulating cell-specific transcription factor σF of Bacillus subtilis. Genes Dev. 8:2653-2663. [DOI] [PubMed] [Google Scholar]
- 5.Domian, I. J., K. C. Quon, and L. Shapiro. 1997. Cell type-specific phosphorylation and proteolysis of a transcriptional regulator controls the G1-to-S transition in a bacterial cell cycle. Cell 90:415-424. [DOI] [PubMed] [Google Scholar]
- 6.Dougan, D. A., A. Mogk, K. Zeth, K. Turgay, and B. Bukau. 2002. AAA+ proteins and substrate recognition, it all depends on their partner in crime. FEBS Lett. 529:6-10. [DOI] [PubMed] [Google Scholar]
- 7.Duncan, L., and R. Losick. 1993. SpoIIAB is an anti-sigma factor that binds to and inhibits transcription by regulatory protein σF from Bacillus subtilis. Proc. Natl. Acad. Sci. USA 90:2325-2329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Dworkin, J., and R. Losick. 2001. Differential gene expression governed by chromosomal spatial asymmetry. Cell 107:339-346. [DOI] [PubMed] [Google Scholar]
- 9.Frandsen, N., I. Barak, C. Karmazyn-Campelli, and P. Stragier. 1999. Transient gene asymmetry during sporulation and establishment of cell specificity in Bacillus subtilis. Genes Dev. 13:394-399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Garsin, D. A., L. Duncan, D. M. Paskowitz, and R. Losick. 1998. The kinase activity of the antisigma factor SpoIIAB is required for activation as well as inhibition of transcription factor σF during sporulation in Bacillus subtilis. J. Mol. Biol. 284:569-578. [DOI] [PubMed] [Google Scholar]
- 11.Garsin, D. A., D. M. Paskowitz, L. Duncan, and R. Losick. 1998. Evidence for common sites of contact between the antisigma factor SpoIIAB and its partners SpoIIAA and the developmental transcription factor σF in Bacillus subtilis. J. Mol. Biol. 284:557-568. [DOI] [PubMed] [Google Scholar]
- 12.Gerth, U., E. Kruger, I. Derre, T. Msadek, and M. Hecker. 1998. Stress induction of the Bacillus subtilis clpP gene encoding a homologue of the proteolytic component of the Clp protease and the involvement of ClpP and ClpX in stress tolerance. Mol. Microbiol. 28:787-802. [DOI] [PubMed] [Google Scholar]
- 13.Ho, M. S., K. Carniol, and R. Losick. Evidence in support of a docking model for the release of the transcription factor σF from the antisigma factor SpoIIAB in Bacillus subtilis. J. Biol. Chem. 278:20898-20905. [DOI] [PubMed]
- 14.Kanemori, M., H. Yanagi, and T. Yura. 1999. The ATP-dependent HslVU/ClpQY protease participates in turnover of cell division inhibitor SulA in Escherichia coli. J. Bacteriol. 181:3674-3680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kruger, E., and M. Hecker. 1998. The first gene of the Bacillus subtilis clpC operon, ctsR, encodes a negative regulator of its own operon and other class III heat shock genes. J. Bacteriol. 180:6681-6688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kunst, F., N. Ogasawara, I. Moszer, A. M. Albertini, G. Alloni, V. Azevedo, M. G. Bertero, P. Bessieres, A. Bolotin, S. Borchert, R. Borriss, L. Boursier, A. Brans, M. Braun, S. C. Brignell, S. Bron, S. Brouillet, C. V. Bruschi, B. Caldwell, V. Capuano, N. M. Carter, S. K. Choi, J. J. Codani, I. F. Connerton, A. Danchin, et al. 1997. The complete genome sequence of the gram-positive bacterium Bacillus subtilis. Nature 390:249-256. [DOI] [PubMed] [Google Scholar]
- 17.Min, K. T., C. M. Hilditch, B. Diederich, J. Errington, and M. D. Yudkin. 1993. σF, the first compartment-specific transcription factor of B. subtilis, is regulated by an anti-sigma factor that is also a protein kinase. Cell 74:735-742. [DOI] [PubMed] [Google Scholar]
- 18.Mizusawa, S., and S. Gottesman. 1983. Protein degradation in Escherichia coli: the lon gene controls the stability of SulA protein. Proc. Natl. Acad. Sci. USA 80:358-362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Msadek, T., V. Dartois, F. Kunst, M. L. Herbaud, F. Denizot, and G. Rapoport. 1998. ClpP of Bacillus subtilis is required for competence development, motility, degradative enzyme synthesis, growth at high temperature and sporulation. Mol. Microbiol. 27:899-914. [DOI] [PubMed] [Google Scholar]
- 20.Nakano, S., G. Zheng, M. M. Nakano, and P. Zuber. 2002. Multiple pathways of Spx (YjbD) proteolysis in Bacillus subtilis. J. Bacteriol. 184:3664-3670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Pan, Q., D. A. Garsin, and R. Losick. 2001. Self-reinforcing activation of a cell-specific transcription factor by proteolysis of an anti-sigma factor in B. subtilis. Mol. Cell 8:873-883. [DOI] [PubMed] [Google Scholar]
- 22.Piggot, P. J., and R. Losick. 2002. Sporulation genes and intercompartmental regulation, p. 483-518. In A. L. Sonenshein, J. A. Hoch, and R. Losick (ed.), Bacillus subtilis and its closest relatives: from genes to cells. American Society for Microbiology, Washington, D.C.
- 23.Porankiewicz, J., J. Wang, and A. K. Clarke. 1999. New insights into the ATP-dependent Clp protease: Escherichia coli and beyond. Mol. Microbiol. 32:449-458. [DOI] [PubMed] [Google Scholar]
- 24.Ryan, K. R., E. M. Judd, and L. Shapiro. 2002. The CtrA response regulator essential for Caulobacter crescentus cell-cycle progression requires a bipartite degradation signal for temporally controlled proteolysis. J. Mol. Biol. 324:443-455. [DOI] [PubMed] [Google Scholar]
- 25.Schirmer, E. C., J. R. Glover, M. A. Singer, and S. Lindquist. 1996. HSP100/Clp proteins: a common mechanism explains diverse functions. Trends Biochem. Sci. 21:289-296. [PubMed] [Google Scholar]
- 26.Turgay, K., J. Hahn, J. Burghoorn, and D. Dubnau. 1998. Competence in Bacillus subtilis is controlled by regulated proteolysis of a transcription factor. EMBO J. 17:6730-6738. [DOI] [PMC free article] [PubMed] [Google Scholar]