Abstract
To gain insight into the mechanism by which Gli-similar 2 (Glis2) regulates transcription, we performed yeast-two hybrid cDNA library screening using Glis2 as bait. This screening identified β-catenin as a potential Glis2-interacting protein. Mammalian two-hybrid, co-immunoprecipitation, and GST-pulldown analyses supported the interaction between Glis2 and β-catenin. Pulldown analyses with several Glis2 deletion mutants indicated that the 1st zinc finger motif of Glis2 is critical for its interaction with β-catenin, while the armadillo repeats of β-catenin are important in its interaction with Glis2. Reporter analyses showed that Glis2 represses T-Cell Factor (TCF)-mediated transcriptional activation. In addition, Glis2 represses the expression of the TCF target gene cyclin D1. Our results indicate that Glis2 interacts with β-catenin and suggest that Glis2 functions as a negative modulator of β-catenin/TCF-mediated transcription.
Keywords: Gli-similar, Krüppel-like zinc finger protein, β-catenin, gene expression
1. Introduction
Gli-similar (Glis) 1–3 constitute a subfamily of Krüppel-like zinc finger proteins that are closely related to members of the Gli subfamily [1–5]. The highly conserved zinc finger domain mediates the binding of these proteins to Gli-binding sites (GBS), containing the consensus sequence GACCACCCAC, in the regulatory region of target genes [2,5,6].
The Glis2 gene encodes a 56 kD transcriptional regulator that [4]. It is highly expressed in the ureteric bud and induced during neuronal differentiation suggesting that Glis2 is involved in the regulation of gene transcription at specific stages of kidney development and neurogenesis [3,4]. Its transcriptional activity is likely mediated through interactions with other nuclear proteins that function as co-repressors or co-activators. This is supported by findings showing that the transcriptional repression by Glis2 is mediated through recruitment of the co-repressors carboxyl-terminal binding protein 1 (CtBP1) and histone deacetylase 3 (HDAC3) [7]. However, the precise molecular mechanisms by which Glis2 regulate transcription, are still poorly understood.
To identify additional mediators or modulators of Glis2 activity, we performed a yeast two-hybrid screening using the amino terminus of Glis2 as bait. This analysis identified β-catenin as a potential Glis2-interacting protein. Previous studies have shown that β-catenin is a multifunctional protein [8–10]. The cadherin-bound form functions as a regulator of cell adhesion and migration while the cytoplasmic form functions as a transducer in the canonical Wnt signaling pathway. Recent studies have indicated that β-catenin can also interact with proteins other than those associated with the canonical Wnt signaling pathway. These include several nuclear receptors, inhibitor of β-catenin and TCF-4 (ICAT), pontin, and reptin [11–13]. Moreover, several studies have provided evidence for a link between the hedgehog/Gli and Wnt/β-catenin signaling pathways [14,15].
In this study, we provide evidence for an interaction between Glis2 and β-catenin. The 1st zinc finger motif of Glis2 and the armadillo repeats of β-catenin are critical in the interaction between these two proteins. In addition we demonstrate that Glis2 represses β-catenin/TCF-mediated transcription. Our observations indicate an interaction between the Glis2 and β-catenin/TCF signaling pathways and suggest that Glis2 can function as a negative regulator of β-catenin/TCF-induced gene expression.
2. Materials and Methods
2.1. Plasmids
pEGFP-Glis2 was obtained by cloning full-length Glis2 into pEGFP/C1. The sequences of all plasmids were verified by restriction enzyme analyses and DNA sequencing analyses. p3xFlag-CMV-Glis2ZF1mut, containing a point mutation (C175A) in the first zinc finger (ZF1) of Glis2, was generated using a site-directed mutagenesis kit from Stratagene following the manufacturer’s instructions. For other plasmids see Supplements.
2.2. Immunoprecipitation (IP) assay and Western Blot Analysis
Cells were transiently transfected with p3xFlag-CMV, p3xFlag-CMV-Glis2, pEGFP-β-catenin, V5-β-catenin, and truncated V5-β-catenin mutants as indicated, using Fugene 6 transfection reagent (Roche, Indianapolis, IN). After 48 h incubation, cells were harvested and lysed in RIPA buffer [50 mM Tris-HCl (pH 7.4), 150 mM NaCl, 0.25% deoxycholate, 1% NP-40, 1 mM EDTA] containing protease inhibitor cocktails I and II (Sigma). Flag-Glis2 protein complexes were isolated using anti-Flag M2 affinity gel (Sigma). The bound protein complexes were solubilized in sample buffer and analyzed by Western blot analysis using mouse anti-Flag M2 (Sigma) or anti-GFP (Miltenyi Biotec, Auburn, CA) antibodies. In certain cases proteins were immunoprecipitated with anti-β-catenin agarose resin (Santa Cruz Biotechnology, Santa Cruz, CA) and bound proteins examined by Western blot analysis with an anti-β-catenin (BD Pharmingen, San Jose, CA) and anti-Flag M2 antibodies. To examine the regulation of cyclin D1 expression by Glis2, HCT116 cells were transfected with p3xFlag-CMV-Glis2 and V5-β-catenin expression vectors as indicated. Thirty-six h later, cells were lysed directly in 2X sample buffer (Sigma). Protein lysates were examined by Western blot analysis with an anti-Flag M2 antibody, and antibodies against V5 (Invitrogen), cyclin D1, and β-tubulin (Santa Cruz Biotechnology).
2.3. In vitro pulldown assay
Equal amounts of GST-β-catenin recombinant protein or GST were incubated with gluthathione-Sepharose 4B beads and washed in PBS. [35S]-methionine-labeled Glis2 was obtained using the TNT Quick Coupled Transcription/Translation system (Promega). The GST and GST-β-catenin bound beads were then incubated with the [35S]-methionine-labeled Glis2 in 0.2 ml binding buffer (20 mM Tris-HCl, pH 7.6, 100 mM KCl, 0.05% Nonidet P-40, 0.1 mM EDTA, 10% glycerol and 1 mM PMSF). After 1 h incubation at 4 °C, beads were washed five times in binding buffer and then boiled in 15 ml of 2X SDS-PAGE loading buffer. Solubilized proteins were then separated by 4–20% SDS-PAGE and the radiolabeled proteins visualized by autoradiography.
2.4. Reporter gene assay
Cells were transfected with the reporter plasmids pFR-LUC (Stratagene), TOPFlash, FOPFlash, Super8XTOPFlash, Super8XFOPFlash, PCD1, pCMVbeta or pRL-SV40, and various Glis2 and β-catenin constructs as indicated. Transfection was performed with Fugene 6 transfection reagent. phRL-SV40 served as internal control. Luciferase activity was measured 48 h later using the Dual-Luciferase® Reporter Assay System from Promega. All transfections were performed in triplicate. Experiments were carried out at least twice.
3. Results
3.1. Identification of β-catenin as a Glis2-interacting protein
Previous studies have shown that Glis2 contains a transcriptional activation as well as a repressor domain and that the transcriptional repression is mediated through interaction with other proteins [4,7]. To identify additional proteins that modulate or mediate Glis2 activity, we performed yeast two-hybrid cDNA screening using the amino-terminus (aa 1–196) and the carboxyl-terminus (aa 319–522) of Glis2 as baits and two different cDNA libraries as preys (see Supplements). Glis2(319–522) turned out to be self-activating in yeast and therefore could not be used in yeast-two hybrid analysis. However, yeast two-hybrid analysis with Glis2(1–196) as bait yielded 8 positive clones encoding proteins with widely different functions (Table I). These included β-catenin, lysine-deficient protein kinase 1 (WNK1), and two tetratricopeptide repeat domain containing proteins, the GTPase XPA binding protein 1 (XAB1) and G-protein signalling modulator 2 (GPSM2). Two independently-derived β-catenin clones were isolated; one clone using a mixed human breast/prostate cancer cDNA library as prey, the other from a mixed mouse embryo cDNA (E7–E19) library. Further studies were carried out to analyze the putative interaction of Glis2 with β-catenin.
Table I.
Putative Glis2-interacting proteins identified by yeast to hybrid analysis
| Genbank # | Symbol | Description | Function |
|---|---|---|---|
| AK035311 | CTNNB | beta-catenin | Cell adhesion and signal transduction |
| NM_014309 | RBM9 | RNA binding motif protein 9, isoform 2 | Negative regulator of transcription |
| NM_015330 | KIAA0376 | cytospin A | Unknown |
| NM_053193 | CPSF1 | cleavage and polyadenylation specific factor 1 | Nucleic acid binding |
| NM_029522 | GPSM2 | G-protein signaling modulator 2 (AGS3-like) | GTPase |
| NM_133671 | U2AF2 | U2 small nuclear ribonucleoprotein auxiliary factor 2 | RNA splicing |
| NM_018979 | WNK1 | WNK lysine deficient protein kinase 1 | Kinase, Ion transport |
| NM_020196 | XAB2 | XPA binding protein 2 | DNA repair, transcription |
The interaction between Glis2 and β-catenin was confirmed by mammalian two-hybrid analysis. HCT116 colorectal carcinoma cells were co-transfected with (UAS)5-LUC reporter, pM or pM-Glis2 in the presence of increasing amounts of pVP16-β-catenin. Co-expression of VP16-β-catenin significantly increased reporter activity in a dose-dependent manner due to the interaction of β-catenin with Glis2 (Fig. 1).
Fig. 1.
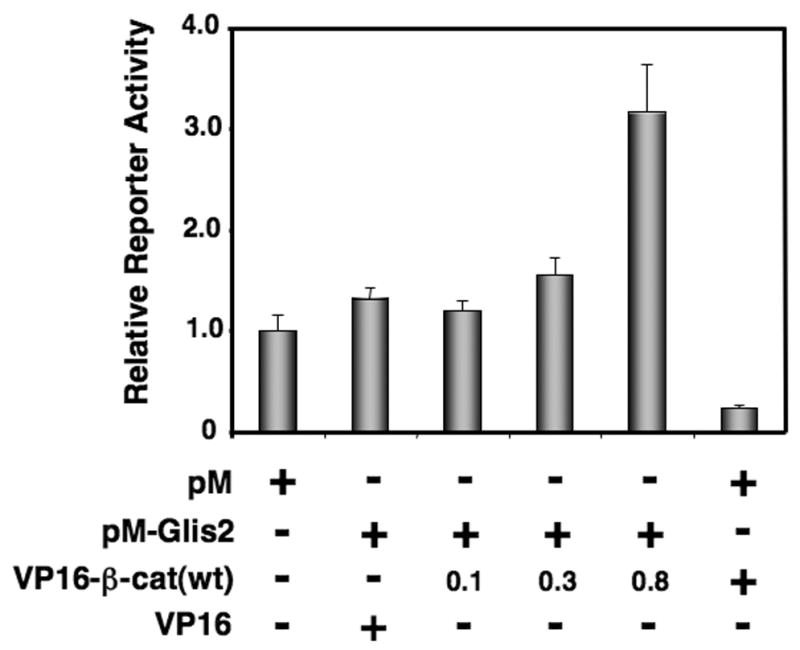
(A) Identification of β-catenin as a Glis2-interacting protein by mammalian two-hybrid analysis. HCT116 cells were co-transfected with (UAS)5-LUC, pRL-SV40, pM, pVP16, pM-Glis2, and increasing amounts of pVP16-β-catenin as indicated. After 48 h, cells were harvested and assayed for reporter activities. The relative LUC activity was calculated and plotted.
In order to be physiologically relevant the two proteins need to be co-expressed at least in some mammalian cells. Therefore, we examined the expression of Glis2 and β-catenin in normal colon and several colorectal carcinoma cell lines (Supplement Fig. 1). Glis2 and β-catenin are both expressed in normal colon and in Caco-2, HCT15, HCT116, Lovo, SW620, and SW480 cells. However, colorectal carcinoma Colo205 and KM12 cells expressed only β-catenin.
Next, we confirmed the interaction of Glis2 and β-catenin by co-immunoprecipitation analysis. Flag-Glis2 and EGFP-β-catenin were transiently expressed in HCT116 cells and Flag-Glis2 protein complexes immunoprecipitated using anti-Flag-M2 agarose resin. As shown in Fig. 2A, Western blot analysis using an anti-GFP antibody identified EGFP-β-catenin as part of the immunoprecipitated protein complexes. We next examined whether Glis2 was able to bind endogenous β-catenin in HCT116 cells expressing Flag-Glis2. As shown in Fig. 2B, endogenous β-catenin was co-immunoprecipitated with anti-Flag-M2 affinity resin. Inversely, immunoprecipitation of protein complexes containing endogenous β-catenin by anti-β-catenin affinity resin co-immunoprecipitated Glis2 (Supplement Fig. 2). These results are in agreement with the data obtained by mammalian two-hybrid analysis and support the conclusion that Glis2 and β-catenin interact with each other.
Fig. 2.
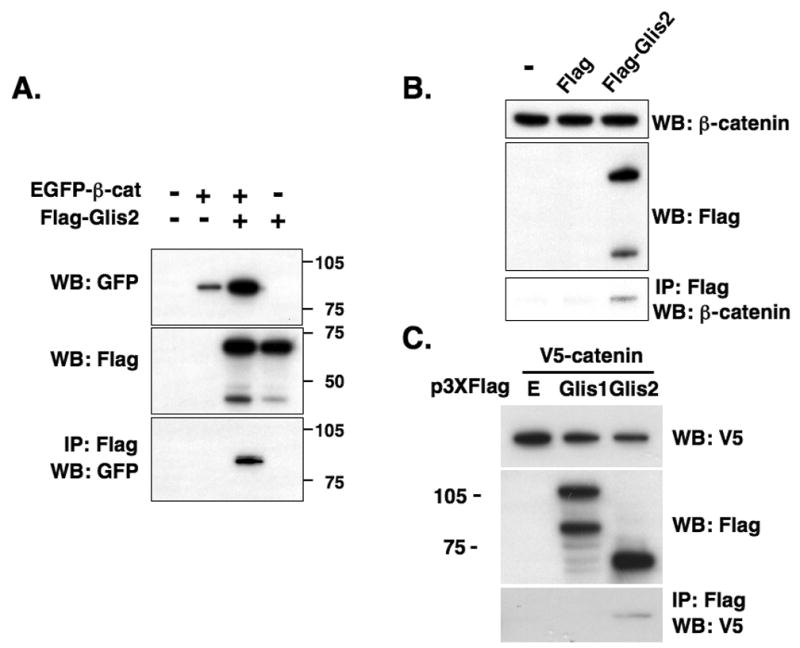
Glis2 and β-catenin are associated with the same protein complex. (A) HCT116 cells were transfected with p3xFlag-CMV-Glis2, p3xFlag-CMV, and pEGFP-wt-β-catenin as indicated. Forty-eight h after transfection, protein lysates were prepared and one part was used in Western blot (WB) analysis with an anti-GFP (upper panel) or anti-Flag M2 antibody (middle panel). The remaining was used for immunoprecipitation (IP) using anti-Flag M2 affinity resin. Bound proteins were then examined by Western blot analysis with anti-GFP antibody (lower panel). (B) Glis2 interacts with endogenous β-catenin. HCT116 cells were transfected with p3xFlag-CMV-Glis2 or p3xFlag-CMV. Forty-eight h after transfection, cell lysates were prepared and Flag-Glis2 protein complexes immunoprecipitated with anti-Flag affinity resin. Bound proteins were examined by Western blot analysis with anti-β-catenin or anti-Flag antibodies. (C) The interaction of β-catenin is specific for Glis2. HCT116 cells were transfected with V5-β-catenin, p3xFlag-CMV, p3xFlag-CMV-Glis1, or p3xFlag-CMV-Glis2. Proteins in the cellular lysates and immunoprecipitated proteins were examined by Western blot analysis using anti-Flag M2 or anti-V5 antibodies.
To determine whether this interaction was specific for Glis2, we examined the interaction between β-catenin and another member of the Glis subfamily. As shown in Fig. 2C, Glis2 but not Glis1 co-immunoprecipitated β-catenin. These data suggest that β-catenin specifically interacts with Glis2.
3.2. In vitro interaction of Glis2 with β-catenin
To determine whether β-catenin physically interacted with Glis2, we performed in vitro pulldown analysis using GST-β-catenin and in vitro translated, [35S]-labeled Glis2. GST-β-catenin was incubated with [35S]-labeled Glis2 and GST-β-catenin protein complexes isolated with glutathione-Sepharose beads. As shown in Fig. 3A, GST-β-catenin was able to pulldown radiolabeled Glis2 while GST (control) did not. Moreover, GST-β-catenin protein was unable to pulldown [35S]-labeled Glis1 in agreement with the results shown in Fig. 2C. These data support the conclusion that β-catenin and Glis2 physically interact with each other and demonstrate that this interaction is specific.
Fig. 3.
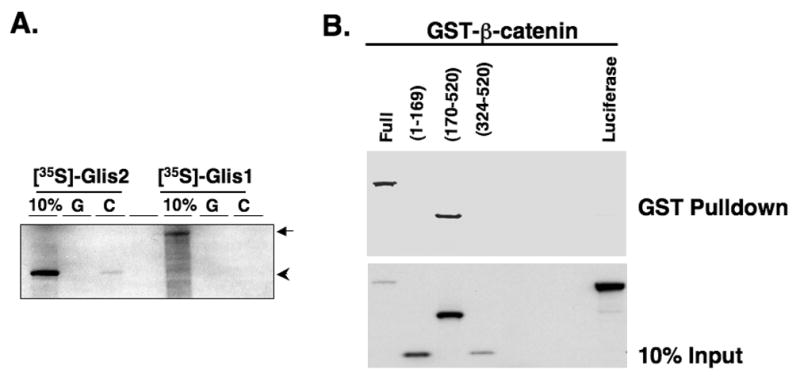

Glis2 interacts directly with β-catenin. (A) Interaction of GST and GST-β-catenin fusion proteins with [35S]-methionine-labeled Glis2 or Glis1 was examined as described in Materials and Methods. First lane, 10% input; G, GST; C; GST-β-catenin. Arrow indicated Glis1 and arrow head indicated Glis2. (B) Glis2 interacts with β-catenin through its ZFD. GST-pulldown analysis with [35S]-methionine-labeled Glis2, Glis2(1–169), Glis2(170–520), Glis2(324–520) or luciferase which served as negative control. Bound radiolabeled proteins were separated by PAGE and visualized by autoradiography. (C, D) Co-immunoprecipitation assay using Glis2 mutants. HCT116 cells were transfected with V5-β-catenin, p3xFlag-CMV empty vector, p3xFlag-CMV-Glis2, p3xFlag-CMV-Glis2(1–382), p3xFlag-CMV-Glis2(1–200), p3xFlag-CMV-Glis2(1–143), p3xFlag-CMV-Glis2(170–377), or p3xFlag-CMV-Glis2ZF1mut containing the C175A mutation in ZF1 (ZF1mut). Proteins in the cellular extracts and immunoprecipitated proteins were examined by Western blot analysis using anti-Flag M2 or anti-V5 antibodies.
3.3. The first zinc finger motif of Glis2 is essential for its interaction with β-catenin
To obtain greater insight into what region of Glis2 was involved in the interaction with β-catenin, we performed GST-pulldown analysis using GST-β-catenin and several [35S]-labeled Glis2 deletion mutants. Fig. 3B shows that GST-β-catenin was able to bind full-length Glis2 and Glis2(170–520), but not Glis2(1–169) or Glis2(324–520). GST-β-catenin also did not bind luciferase which was used as a negative control. These results suggest that the zinc finger domain of Glis2, localized between amino acids 170 and 324, is important in the interaction with β-catenin.
Co-immunoprecipitation analysis with V5-β-catenin and different Flag-Glis2 expression vectors supported the importance of the zinc finger domain. Fig. 3C shows that β-catenin was able to immunoprecipitate full-length Glis2 and the mutants Glis2(1–382), Glis2(1–200), and Glis2(170–377) but did not immunoprecipitate Glis2(1–143). These data suggest that the 1st zinc finger (ZF1) region of Glis2 (from aa 170 to 193) is critical for its interaction with β-catenin.
To obtain further support for this conclusion, we analyzed the ability of β-catenin to interact with the mutant Glis2ZF1mut, which contains a C175A mutation in the first zinc finger that destroys the tetrahedral configuration in the zinc finger. Fig. 3D shows that V5-β-catenin did not co-immunoprecipitate Glis2ZF1mut suggesting that the structural integrity of the first zinc finger of Glis2 is important for its interaction with β-catenin.
3.4. The armadillo repeats of β-catenin are important for its interaction with Glis2
To determine which region of β-catenin is involved in its interaction with Glis2, we performed GST-pulldown analysis using GST-β-catenin and different mutant GST-β-catenin fusion proteins and [35S]-labeled Glis2. As shown in Fig. 4, full-length β-catenin, and the β-catenin mutants (1–423), (132–423), and (423–781) were able to pulldown Glis2. In contrast, control GST and the β-catenin mutant (1–131) did not pulldown Glis2. These results indicate that the region of β-catenin containing the armadillo repeats are important in its interaction with Glis2 but that the amino terminus is not required.
Fig. 4.
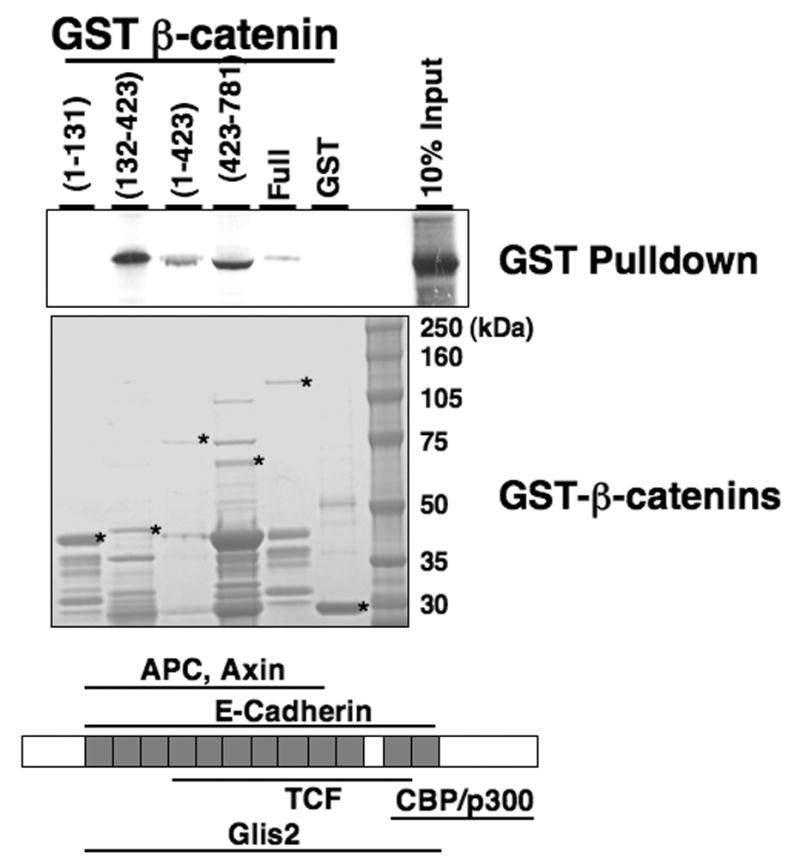
β-catenin interacts with Glis2 through its specific regions. GST and several GST-β-catenin fusion proteins were bound to glutathione-Sepharose 4B beads and then incubated with [35S]-methionine-labeled Glis2. Bound radiolabeled proteins were visualized by autoradiography. The input of the various GST-β-catenin proteins is shown in the lower panel. Asterisks indicate the respective GST-β-catenin proteins.
3.5. Glis2 enhances the nuclear localization of β-catenin
We next examined the subcellular localization of Glis2 and β-catenin in HEK293 cells by confocal microscopy. In 79% of the cells expressing pEGFP-β-catenin only, β-catenin was found equally distributed between cytoplasm and nucleus, while in 21% of the cells β-catenin was confined largely to the nucleus (Fig. 5). However, when pEGFP-β-catenin and Flag-Glis2 were co-expressed, the percentage of cells in which β-catenin was restricted mainly to the nucleus, was greatly increased to about 90% (Fig. 5). Similar results were obtained with HCT116 cells (data not shown). These observations suggest that Glis2 enhances the nuclear location of β-catenin. Under all conditions analyzed, Glis2 localized only to the nucleus. In contrast to Glis2, Glis2ZF1mut did not change the nuclear translocation of β-catenin significantly (Fig. 5). The mutation in the 1st zinc finger motif, however, did not have any effect on the nuclear localization of Glis2. These observations are in agreement with our conclusion that the integrity of the 1st zinc finger domain of Glis2 is essential in its interaction with β-catenin.
Fig. 5.
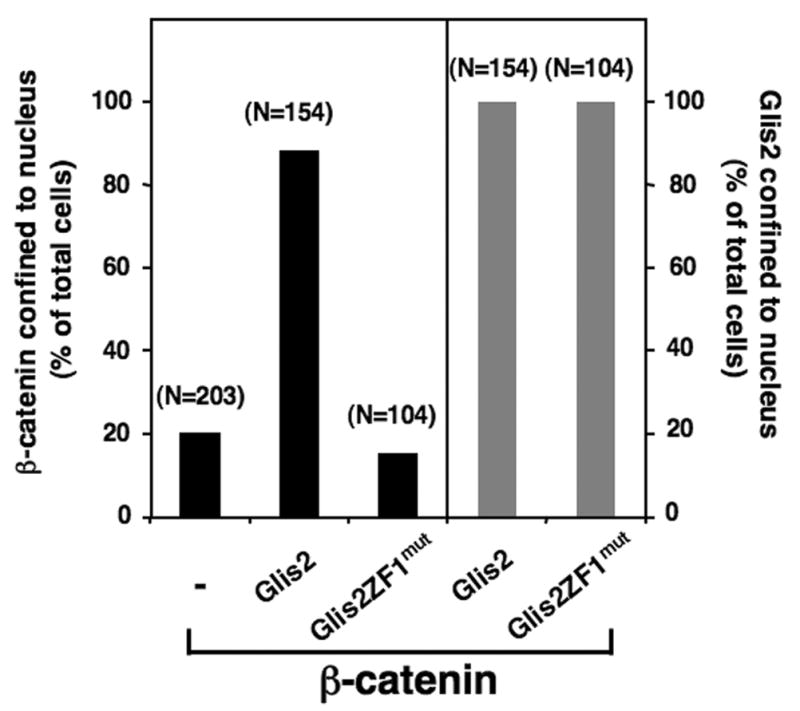
Glis2 enhances nuclear localization of β-catenin. NEH293 cells were transiently transfected with p3xFlag-CMV-Glis2, p3xFlag-CMV-Glis2ZF1mut, and/or pEGFP-β-catenin. The percentage of cells in which the localization of Glis2 or β-catenin was restricted to the nucleus, was calculated and plotted. The number of cells counted is indicated above each bar.
3.6. Glis2 represses β-catenin/TCF-mediated transcriptional activation
Previous studies have shown that Glis2 contains an activation and a repressor domain [4]. To determine whether Glis2 interfered with β-catenin-induced transcription, we analyzed the effect of Glis2 on TCF-mediated transcriptional activation using TOPFlash and FOPFlash TCF-reporters which contain multiple copies of an optimal TCF-binding element (TBE) and mutant TBE, respectively. As shown in Fig. 6A, pEGFP-ΔN90-β-catenin, a constitutively active form of β-catenin, induced reporter activity highly (6-fold) in HCT116 cells. This increase was greatly (67%) inhibited by co-expression of Glis2 suggesting that Glis2 inhibits TCF-mediated transcriptional activation (Fig. 6A). This was confirmed by reporter analysis using Super8XTOPFlash and Super8XFOPFlash reporters which contain 8 copies of TBE or mutant TBE, respectively [16]. The constitutively active β-catenin (S37C) caused a 7-fold increase in transcriptional activation, this activity was dramatically (70%) suppressed by Glis2 (Supplement Fig. 3). No such effects were observed with the TCF mutant reporter Super8XFOPFlash (Supplement Fig. 3). Glis2 repressed transcriptional activation by β-catenin in a dose dependent manner (Fig. 6B). At high levels of Glis2 the reporter activity was below that of control HCT116 cells suggesting that Glis2 also inhibited activation by the constitutively active form of β-catenin present in these cells (compare bar 1 with bar 5, Fig. 6B). These data suggest that Glis2 functions as a negative modulator of β-catenin/TCF-mediated transcriptional activation.
Fig. 6.
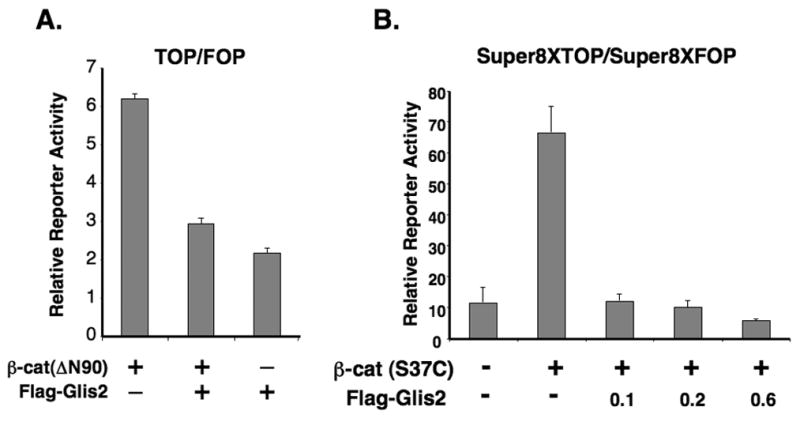
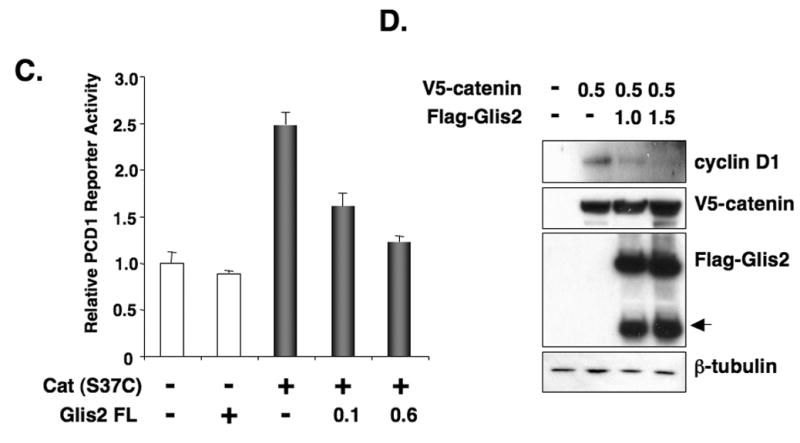
Glis2 represses TCF-mediated transcriptional activity and the induction of cyclin D1 by β-catenin. (A) HCT116 cells were cotransfected with TOPFlash and FOPFlash reporters, pCMVbeta, β-catenin(ΔN90), and p3xFlag-CMV-Glis2 as indicated. Forty-eight h after transfection, cells were assayed for reporter activities as described in Materials and Methods. (B) HCT116 cells were cotransfected with Super8XTOPFlash and Super8XFOPFlash reporters, pRL-SV40, pCMV-V5-β-catenin (β-cat(S37C)) and different concentrations of p3xFlag-CMV-Glis2 expression vector as indicated and then processed as described under A. (C) HCT116 cells were cotransfected with the cyclin D1 reporter PCD1, pRL-SV40, the constitutively-active pCMV-V5-β-catenin (S37C) expression vector, and p3xFlag-CMV-Glis2 as indicated and then processed as described under A. (D) HCT116 cells were transfected with pCMV-V5-β-catenin and p3xFlag-CMV-Glis2 (1.0 and 1.5 μg) as indicated. Thirty-six h later, cellular proteins were analyzed by Western blot analysis using antibodies against V5, Flag, β-tubulin, and cyclin D1. The lower molecular weight band (arrow head) in the Flag-Glis2 Western blot represents a proteolytically processed form of Glis2.
3.7. Glis2 represses cyclin D1 promoter activity
We next examined the repressor activity of Glis2 on the activation of the cyclin D1-promoter (PCD1), a well-studied target of the Wnt signaling pathway [17], in HCT116 cells. As shown in Fig. 6C, the constitutively active β-catenin (S37C) enhanced PCD1 reporter activity about 2.5 fold. Coexpression of Glis2 significantly reduced cyclin D1-reporter activity in a dose-dependent manner (Fig. 6C). We next examined the effect of β-catenin and Glis2 on the expression of endogenous cyclin D1 in HCT116 cells. The level of cyclin D1 protein was enhanced by β-catenin (Fig. 6D). Glis2 overexpression diminished the β-catenin-induced increase in cyclin D1. These results support the concept that Glis2 functions as a repressor of the activation of target genes in the Wnt/β-catenin signaling pathway.
4. Discussion
To obtain greater insights into the mechanisms by which Glis2 regulates transcription, we performed yeast two-hybrid screening using the amino-terminus of Glis2 as bait. This analysis identified several putative Glis2-interacting proteins, including WNK1, XAB1, GPSM2, and β-catenin. XAB1 and GPSM2 contain a tetratricopeptide repeat domain which are thought to mediate protein-protein interactions in a variety of biological systems [18,19]. β-catenin is a versatile protein with a function in cell adhesion and in the canonical Wnt signaling pathway [8–10]. Binding of Wnt to receptors of the Frizzled family results in the translocation of β-catenin to the nucleus where it interacts with TCF family members replacing co-repressors and promoting the recruitment of co-activators and activation of transcription of target genes, including cyclin D1.
Because Glis proteins are closely related to members of the Gli subfamily [1,2,4,5] and because of reported interactions between the hedgehog/Gli and Wnt/β-catenin signaling pathways [14,15] and the fact that two independent clones of β-catenin were isolated by yeast two-hybrid analysis, we decided to focus on the interaction between β-catenin and Glis2. Mammalian two-hybrid and co-immunoprecipitation analysis confirmed that β-catenin interacts with Glis2; this was further supported by GST-pulldown analysis. We demonstrated by deletion analysis that the region between aa 170 and 193 is important for the interaction of Glis2 with β-catenin. This region contains the first zinc finger motif of Glis2. The critical role of the first zinc finger motif was confirmed by the observation that a point mutation in the first zinc finger motif, that destroys its tetrahedral configuration, abolished the interaction. β-catenin contains several functional domains, the amino terminus interacts with GSK3β and casein kinase Iα (CK1) binding sites while its 12 armadillo repeats provides an interface for TCF/LEFs, the co-activator CBP, and several other proteins [8–10]. Our results show that the amino-terminus of β-catenin is not essential for the interaction with Glis2 but that its armadillo repeat domain is required. The interaction of β-catenin with TCF members, axin, cadherins, and the peroxisome proliferator-activated receptor γ (PPARγ) has been shown to involve the central region of the armadillo repeats while interaction with the androgen receptor involves the first 6 armadillo repeats [13]. Each armadillo repeat consists of three α-helices and all 12 repeats form a superhelical structure with a long positively charged groove [20]. Interestingly, the second loop in zinc finger 1 of Glis2 consists of a negatively charged α-helix containing 3 Glu/Asp residues that could facilitate the interaction of Glis2 with the armadillo repeats.
Previous studies have demonstrated that Glis2 can function as a repressor of gene transcription and recruit the co-repressors CtBP1 and HDAC3 [7]. Our results indicate that Glis2 can inhibit β-catenin/TCF-mediated transcriptional activation of TOPFlash. In addition, expression of Glis2 represses the promoter activity of the human cyclin D1 gene, a natural Wnt target, and inhibits the induction of cyclin D1 expression by β-catenin. These observations strongly suggest that Glis2 may be a repressor of Wnt/β-catenin signaling. A number of other transcriptional factors, including several nuclear receptors, have been shown to be potent repressors of β-catenin/TCF signaling [13]. This negative regulation can involve different mechanisms, including competition for a limited pool of β-catenin, enhanced degradation of β-catenin, competition for co-factors, or recruitment of co-repressors to TCF/β-catenin complexes. Since Glis2 acts as a repressor and associates with CtBP1 and HDAC3, it may be unlikely that it competes with β-catenin/TCF for the binding of co-activators. The nuclear proteins Pontin52 and Reptin52 have been reported to interact with the armadillo repeats of β-catenin and to form a multiprotein complex that includes TCF and TATA-box binding protein (TBP) [12]. Glis2 may be part of a similar multiprotein transcriptional complex. Elucidating the mechanisms by which Glis2 modulates transcription requires further study.
We show that Glis2 is co-expressed with β-catenin in normal colon and several colorectal carcinoma cell lines (Supplement Fig. 1). However, Glis2 expression was undetectable in two cell lines. Since Glis2 functions as a negative modulator of β-catenin signaling, lack of Glis2 may enhance transcriptional activation by β-catenin and therefore promote tumorigenesis. However, the role of Glis2 in β-catenin signaling and tumorigenesis is likely to be much more complex and dependent on the activity of Glis2. This may involve various posttranslational modifications and proteolytic processing of Glis2. Additional studies have to determine the significance of the interaction between Glis2 and Wnt4/β-catenin signaling pathways in tumorigenesis and embryonic development.
Supplementary Material
Acknowledgments
We would like to thank Drs. T. Eling and H.Y. Cho for their comments on the manuscript. The authors would like to thank Drs. R.T. Moon (University of Washington) Seattle, WA), Y. Sekido (Nagoya University School of Medicine), A. Barth (Stanford University), A. Raz (Wayne State University), and R.G. Pestell (Georgetown University) for the various β-catenin and promoter constructs. The authors thank Robert M. Petrovich and Lori Edward from the NIEHS protein core facility, and Dr. X-P. Yang for their technical assistance. This research was supported by the Intramural Research Program of the NIH, National Institute of Environmental Health Sciences.
Abbreviations
- Glis
Gli-similar
- TCF
T-cell factor
- TOP
TOPFlash
- FOP
FOPFlash
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Kim YS, Lewandoski M, Perantoni AO, Kurebayashi S, Nakanishi G, Jetten AM. Identification of Glis1, a novel Gli-related, Kruppel-like zinc finger protein containing transactivation and repressor functions. J. Biol. Chem. 2002;277:30901–30913. doi: 10.1074/jbc.M203563200. [DOI] [PubMed] [Google Scholar]
- 2.Kim YS, Nakanishi G, Lewandoski M, Jetten AM. GLIS3, a novel member of the GLIS subfamily of Kruppel-like zinc finger proteins with repressor and activation functions. Nucleic Acids Res. 2003;31:5513–5525. doi: 10.1093/nar/gkg776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Lamar E, Kintner C, Goulding M. Identification of NKL, a novel Gli-Kruppel zinc-finger protein that promotes neuronal differentiation. Development. 2001;128:1335–1346. doi: 10.1242/dev.128.8.1335. [DOI] [PubMed] [Google Scholar]
- 4.Zhang F, Nakanishi G, Kurebayashi S, Yoshino K, Perantoni A, Kim YS, Jetten AM. Characterization of Glis2, a novel gene encoding a Gli-related, Kruppel-like transcription factor with transactivation and repressor functions. Roles in kidney development and neurogenesis. J. Biol. Chem. 2002;277:10139–10149. doi: 10.1074/jbc.M108062200. [DOI] [PubMed] [Google Scholar]
- 5.Kasper M, Regl G, Frischauf AM, Aberger F. GLI transcription factors: mediators of oncogenic Hedgehog signalling. Eur J Cancer. 2006;42:437–445. doi: 10.1016/j.ejca.2005.08.039. [DOI] [PubMed] [Google Scholar]
- 6.Kinzler KW, Vogelstein B. The GLI gene encodes a nuclear protein which binds specific sequences in the human genome. Mol. Cell. Biol. 1990;10:634–642. doi: 10.1128/mcb.10.2.634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kim SC, Kim YS, Jetten AM. Kruppel-like zinc finger protein Gli-similar 2 (Glis2) represses transcription through interaction with C-terminal binding protein 1 (CtBP1) Nucleic Acids Res. 2005;33:6805–6815. doi: 10.1093/nar/gki985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bienz M, Clevers H. Armadillo/beta-catenin signals in the nucleus--proof beyond a reasonable doubt? Nat. Cell Biol. 2003;5:179–182. doi: 10.1038/ncb0303-179. [DOI] [PubMed] [Google Scholar]
- 9.Sharpe C, Lawrence N, Martinez Arias A. Wnt signalling: a theme with nuclear variations. Bioessays. 2001;23:311–318. doi: 10.1002/bies.1045. [DOI] [PubMed] [Google Scholar]
- 10.van Es JH, Barker N, Clevers H. You Wnt some, you lose some: oncogenes in the Wnt signaling pathway. Curr. Opin. Genet. Dev. 2003;13:28–33. doi: 10.1016/s0959-437x(02)00012-6. [DOI] [PubMed] [Google Scholar]
- 11.Tago K, Nakamura T, Nishita M, Hyodo J, Nagai S, Murata Y, Adachi S, Ohwada S, Morishita Y, Shibuya H, Akiyama T. Inhibition of Wnt signaling by ICAT, a novel beta-catenin-interacting protein. Genes Dev. 2000;14:1741–1749. [PMC free article] [PubMed] [Google Scholar]
- 12.Bauer A, Chauvet S, Huber O, Usseglio F, Rothbacher U, Aragnol D, Kemler R, Pradel J. Pontin52 and reptin52 function as antagonistic regulators of beta-catenin signalling activity. Embo J. 2000;19:6121–6130. doi: 10.1093/emboj/19.22.6121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Mulholland DJ, Dedhar S, Coetzee GA, Nelson CC. Interaction of nuclear receptors with the Wnt/beta-catenin/Tcf signaling axis: Wnt you like to know? Endocr. Rev. 2005;26:898–915. doi: 10.1210/er.2003-0034. [DOI] [PubMed] [Google Scholar]
- 14.Gill PS, Rosenblum ND. Control of murine kidney development by sonic hedgehog and its GLI effectors. Cell Cycle. 2006;5:1426–1430. doi: 10.4161/cc.5.13.2928. [DOI] [PubMed] [Google Scholar]
- 15.van den Brink GR, Bleuming SA, Hardwick JC, Schepman BL, Offerhaus GJ, Keller JJ, Nielsen C, Gaffield W, van Deventer SJ, Roberts DJ, Peppelenbosch MP. Indian Hedgehog is an antagonist of Wnt signaling in colonic epithelial cell differentiation. Nat. Genet. 2004;36:277–82. doi: 10.1038/ng1304. [DOI] [PubMed] [Google Scholar]
- 16.Veeman MT, Slusarski DC, Kaykas A, Louie SH, Moon RT. Zebrafish prickle, a modulator of noncanonical Wnt/Fz signaling, regulates gastrulation movements. Curr. Biol. 2003;13:680–685. doi: 10.1016/s0960-9822(03)00240-9. [DOI] [PubMed] [Google Scholar]
- 17.Shtutman M, Zhurinsky J, Simcha I, Albanese C, D'Amico M, Pestell R, Ben-Ze'ev A. The cyclin D1 gene is a target of the beta-catenin/LEF-1 pathway. Proc Natl Acad Sci USA. 1999;96:5522–5527. doi: 10.1073/pnas.96.10.5522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Marty C, Browning DD, Ye RD. Identification of tetratricopeptide repeat 1 as an adaptor protein that interacts with heterotrimeric G proteins and the small GTPase Ras. Mol. Cell. Biol. 2003;23:3847–3858. doi: 10.1128/MCB.23.11.3847-3858.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Nitta M, Saijo M, Kodo N, Matsuda T, Nakatsu Y, Tamai H, Tanaka K. A novel cytoplasmic GTPase XAB1 interacts with DNA repair protein XPA. Nucleic Acids Res. 2000;28:4212–4218. doi: 10.1093/nar/28.21.4212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Huber AH, Nelson WJ, Weis WI. Three-dimensional structure of the armadillo repeat region of beta-catenin. Cell. 1997;90:871–882. doi: 10.1016/s0092-8674(00)80352-9. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


