Abstract
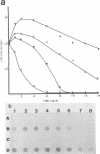
A method has been devised for directly detecting and monitoring genetically engineered microorganisms (GEMs) by using in vitro amplification of the target DNAs by a polymerase chain reaction and then hybridizing the DNAs with a specific oligonucleotide or DNA probe. A cloned 0.3-kilobase napier grass (Pennisetum purpureum) genomic DNA that did not hybridize to DNAs isolated from various microorganisms, soil sediments, and aquatic environments was inserted into a derivative of a 2,4-dichlorophenoxyacetic acid-degradative plasmid, pRC10, and transferred into Escherichia coli. This genetically altered microorganism, seeded into filter-sterilized lake and sewage water samples (10(4)/ml), was detected by a plate count method in decreasing numbers for 6 and 10 days of sample incubation, respectively. The new method detected the amplified unique marker (0.3-kilobase DNA) of the GEM even after 10 to 14 days of incubation. This method is highly sensitive (it requires only picogram amounts of DNA) and has an advantage over the plate count technique, which can detect only culturable microorganisms. The method may be useful for monitoring GEMs in complex environments, where discrimination between GEMs and indigenous microorganisms is either difficult or requires time-consuming tests.
Full text
PDF


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Barkay T., Fouts D. L., Olson B. H. Preparation of a DNA gene probe for detection of mercury resistance genes in gram-negative bacterial communities. Appl Environ Microbiol. 1985 Mar;49(3):686–692. doi: 10.1128/aem.49.3.686-692.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chaudhry G. R., Huang G. H. Isolation and characterization of a new plasmid from a Flavobacterium sp. which carries the genes for degradation of 2,4-dichlorophenoxyacetate. J Bacteriol. 1988 Sep;170(9):3897–3902. doi: 10.1128/jb.170.9.3897-3902.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Francisco D. E., Mah R. A., Rabin A. C. Acridine orange-epifluorescence technique for counting bacteria in natural waters. Trans Am Microsc Soc. 1973 Jul;92(3):416–421. [PubMed] [Google Scholar]
- Fuhrman J. A., Comeau D. E., Hagström A., Chan A. M. Extraction from natural planktonic microorganisms of DNA suitable for molecular biological studies. Appl Environ Microbiol. 1988 Jun;54(6):1426–1429. doi: 10.1128/aem.54.6.1426-1429.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holben William E., Jansson Janet K., Chelm Barry K., Tiedje James M. DNA Probe Method for the Detection of Specific Microorganisms in the Soil Bacterial Community. Appl Environ Microbiol. 1988 Mar;54(3):703–711. doi: 10.1128/aem.54.3.703-711.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kwok S., Mack D. H., Mullis K. B., Poiesz B., Ehrlich G., Blair D., Friedman-Kien A., Sninsky J. J. Identification of human immunodeficiency virus sequences by using in vitro enzymatic amplification and oligomer cleavage detection. J Virol. 1987 May;61(5):1690–1694. doi: 10.1128/jvi.61.5.1690-1694.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mancini P., Fertels S., Nave D., Gealt M. A. Mobilization of plasmid pHSV106 from Escherichia coli HB101 in a laboratory-scale waste treatment facility. Appl Environ Microbiol. 1987 Apr;53(4):665–671. doi: 10.1128/aem.53.4.665-671.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mullis K. B., Faloona F. A. Specific synthesis of DNA in vitro via a polymerase-catalyzed chain reaction. Methods Enzymol. 1987;155:335–350. doi: 10.1016/0076-6879(87)55023-6. [DOI] [PubMed] [Google Scholar]
- Ou C. Y., Kwok S., Mitchell S. W., Mack D. H., Sninsky J. J., Krebs J. W., Feorino P., Warfield D., Schochetman G. DNA amplification for direct detection of HIV-1 in DNA of peripheral blood mononuclear cells. Science. 1988 Jan 15;239(4837):295–297. doi: 10.1126/science.3336784. [DOI] [PubMed] [Google Scholar]
- Roszak D. B., Colwell R. R. Survival strategies of bacteria in the natural environment. Microbiol Rev. 1987 Sep;51(3):365–379. doi: 10.1128/mr.51.3.365-379.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saiki R. K., Bugawan T. L., Horn G. T., Mullis K. B., Erlich H. A. Analysis of enzymatically amplified beta-globin and HLA-DQ alpha DNA with allele-specific oligonucleotide probes. Nature. 1986 Nov 13;324(6093):163–166. doi: 10.1038/324163a0. [DOI] [PubMed] [Google Scholar]
- Saiki R. K., Gelfand D. H., Stoffel S., Scharf S. J., Higuchi R., Horn G. T., Mullis K. B., Erlich H. A. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science. 1988 Jan 29;239(4839):487–491. doi: 10.1126/science.2448875. [DOI] [PubMed] [Google Scholar]
- Saiki R. K., Scharf S., Faloona F., Mullis K. B., Horn G. T., Erlich H. A., Arnheim N. Enzymatic amplification of beta-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science. 1985 Dec 20;230(4732):1350–1354. doi: 10.1126/science.2999980. [DOI] [PubMed] [Google Scholar]
- Steffan R. J., Atlas R. M. DNA amplification to enhance detection of genetically engineered bacteria in environmental samples. Appl Environ Microbiol. 1988 Sep;54(9):2185–2191. doi: 10.1128/aem.54.9.2185-2191.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wong C., Dowling C. E., Saiki R. K., Higuchi R. G., Erlich H. A., Kazazian H. H., Jr Characterization of beta-thalassaemia mutations using direct genomic sequencing of amplified single copy DNA. 1987 Nov 26-Dec 2Nature. 330(6146):384–386. doi: 10.1038/330384a0. [DOI] [PubMed] [Google Scholar]