Abstract
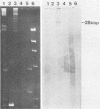
The replication region of a 28-kilobase-pair (kbp) cryptic plasmid from Lactococcus lactis subsp. lactis biovar diacetylactis SSD207 was cloned in L. lactis subsp. lactis MG1614 by using the chloramphenicol resistance gene from the streptococcal plasmid pGB301 as a selectable marker. The resulting 8.1-kbp plasmid, designated pVS34, was characterized further with respect to host range, potential cloning sites, and location of replication gene(s). In addition to lactococci, pVS34 transformed Lactobacillus plantarum and, at a very low frequency, Staphylococcus aureus but not Escherichia coli or Bacillus subtilis. The 4.1-kbp ClaI fragment representing lactococcal DNA in pVS34 contained unique restriction sites for HindIII, EcoRI, XhoII, and HpaII, of which the last three could be used for molecular cloning. A region necessary for replication was located within a 2.5-kbp fragment flanked by the EcoRI and ClaI restriction sites. A 3.8-kbp EcoRI fragment derived from a nisin resistance plasmid, pSF01, was cloned into the EcoRI site of pVS34 to obtain a nisin-chloramphenicol double-resistance plasmid, pVS39. From this plasmid, the streptococcal chloramphenicol resistance region was subsequently eliminated. The resulting plasmid, pVS40, contains only lactococcal DNA. Potential uses for this type of a nisin resistance plasmid are discussed.
Full text
PDF

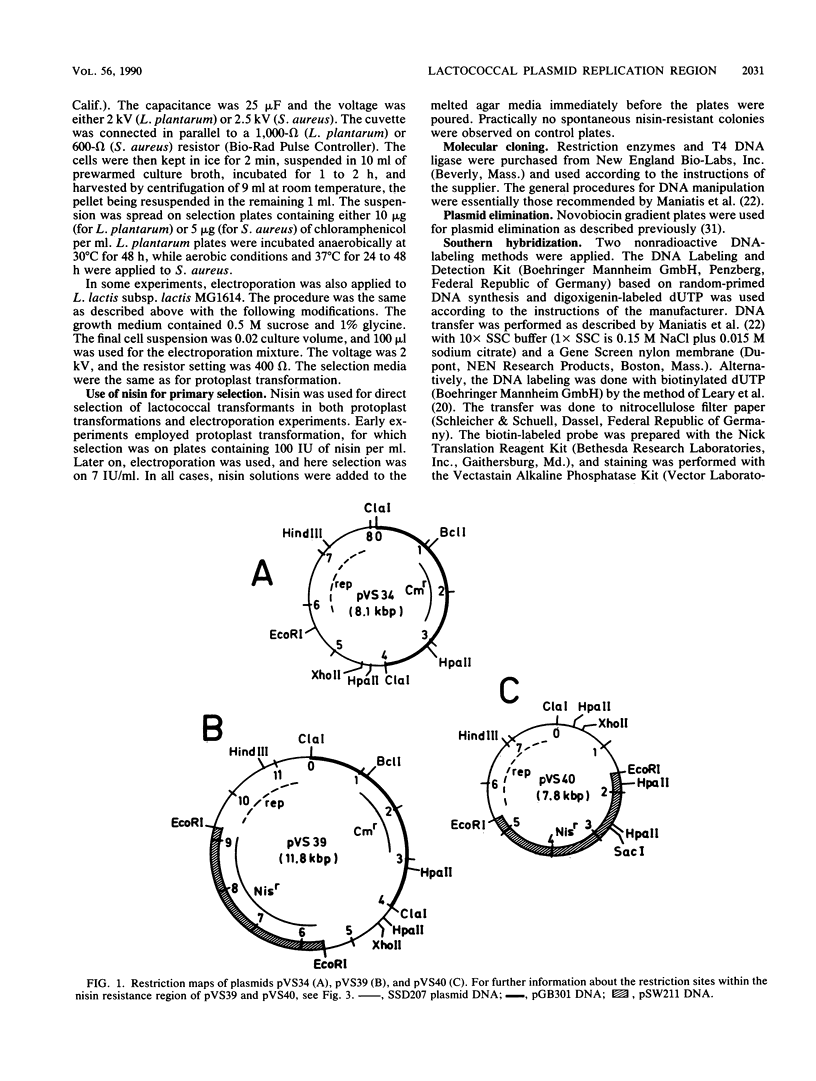
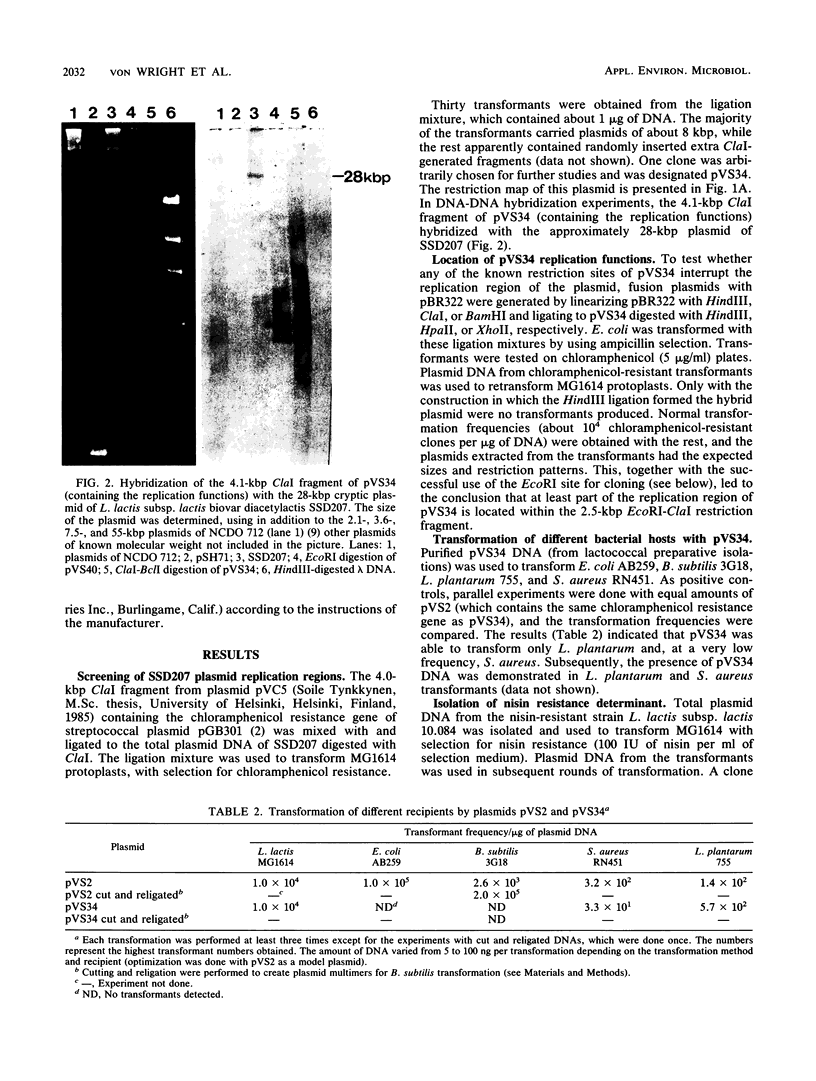
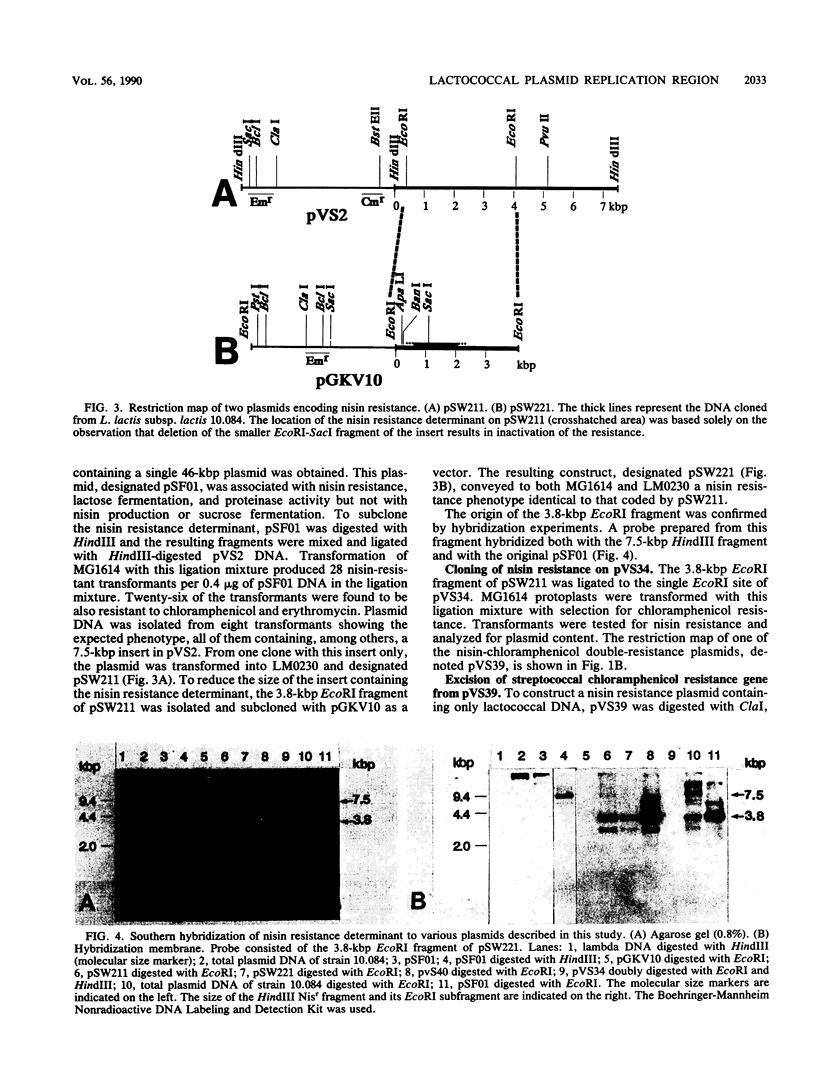


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anderson D. G., McKay L. L. Simple and rapid method for isolating large plasmid DNA from lactic streptococci. Appl Environ Microbiol. 1983 Sep;46(3):549–552. doi: 10.1128/aem.46.3.549-552.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behnke D., Gilmore M. S. Location of antibiotic resistance determinants, copy control, and replication functions on the double-selective streptococcal cloning vector pGB301. Mol Gen Genet. 1981;184(1):115–120. doi: 10.1007/BF00271206. [DOI] [PubMed] [Google Scholar]
- Birnboim H. C. A rapid alkaline extraction method for the isolation of plasmid DNA. Methods Enzymol. 1983;100:243–255. doi: 10.1016/0076-6879(83)00059-2. [DOI] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Boylan R. J., Mendelson N. H., Brooks D., Young F. E. Regulation of the bacterial cell wall: analysis of a mutant of Bacillus subtilis defective in biosynthesis of teichoic acid. J Bacteriol. 1972 Apr;110(1):281–290. doi: 10.1128/jb.110.1.281-290.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Froseth B. R., Herman R. E., McKay L. L. Cloning of nisin resistance determinant and replication origin on 7.6-kilobase EcoRI fragment of pNP40 from Streptococcus lactis subsp. diacetylactis DRC3. Appl Environ Microbiol. 1988 Aug;54(8):2136–2139. doi: 10.1128/aem.54.8.2136-2139.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gasson M. J. Plasmid complements of Streptococcus lactis NCDO 712 and other lactic streptococci after protoplast-induced curing. J Bacteriol. 1983 Apr;154(1):1–9. doi: 10.1128/jb.154.1.1-9.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzalez C. F., Kunka B. S. Transfer of Sucrose-Fermenting Ability and Nisin Production Phenotype among Lactic Streptococci. Appl Environ Microbiol. 1985 Mar;49(3):627–633. doi: 10.1128/aem.49.3.627-633.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gruss A., Ehrlich S. D. The family of highly interrelated single-stranded deoxyribonucleic acid plasmids. Microbiol Rev. 1989 Jun;53(2):231–241. doi: 10.1128/mr.53.2.231-241.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes F., Daly C., Fitzgerald G. F. Identification of the Minimal Replicon of Lactococcus lactis subsp. lactis UC317 Plasmid pCI305. Appl Environ Microbiol. 1990 Jan;56(1):202–209. doi: 10.1128/aem.56.1.202-209.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaletta C., Entian K. D. Nisin, a peptide antibiotic: cloning and sequencing of the nisA gene and posttranslational processing of its peptide product. J Bacteriol. 1989 Mar;171(3):1597–1601. doi: 10.1128/jb.171.3.1597-1601.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klaenhammer T. R., Sanozky R. B. Conjugal transfer from Streptococcus lactis ME2 of plasmids encoding phage resistance, nisin resistance and lactose-fermenting ability: evidence for a high-frequency conjugative plasmid responsible for abortive infection of virulent bacteriophage. J Gen Microbiol. 1985 Jun;131(6):1531–1541. doi: 10.1099/00221287-131-6-1531. [DOI] [PubMed] [Google Scholar]
- Kok J., van der Vossen J. M., Venema G. Construction of plasmid cloning vectors for lactic streptococci which also replicate in Bacillus subtilis and Escherichia coli. Appl Environ Microbiol. 1984 Oct;48(4):726–731. doi: 10.1128/aem.48.4.726-731.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leary J. J., Brigati D. J., Ward D. C. Rapid and sensitive colorimetric method for visualizing biotin-labeled DNA probes hybridized to DNA or RNA immobilized on nitrocellulose: Bio-blots. Proc Natl Acad Sci U S A. 1983 Jul;80(13):4045–4049. doi: 10.1073/pnas.80.13.4045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mandel M., Higa A. Calcium-dependent bacteriophage DNA infection. J Mol Biol. 1970 Oct 14;53(1):159–162. doi: 10.1016/0022-2836(70)90051-3. [DOI] [PubMed] [Google Scholar]
- Marinus M. G. Location of DNA methylation genes on the Escherichia coli K-12 genetic map. Mol Gen Genet. 1973 Dec 14;127(1):47–55. doi: 10.1007/BF00267782. [DOI] [PubMed] [Google Scholar]
- McKay L. L., Baldwin K. A. Conjugative 40-megadalton plasmid in Streptococcus lactis subsp. diacetylactis DRC3 is associated with resistance to nisin and bacteriophage. Appl Environ Microbiol. 1984 Jan;47(1):68–74. doi: 10.1128/aem.47.1.68-74.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKay L. L. Functional properties of plasmids in lactic streptococci. Antonie Van Leeuwenhoek. 1983 Sep;49(3):259–274. doi: 10.1007/BF00399502. [DOI] [PubMed] [Google Scholar]
- Terzaghi B. E., Sandine W. E. Improved medium for lactic streptococci and their bacteriophages. Appl Microbiol. 1975 Jun;29(6):807–813. doi: 10.1128/am.29.6.807-813.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Vossen J. M., Kok J., Venema G. Construction of cloning, promoter-screening, and terminator-screening shuttle vectors for Bacillus subtilis and Streptococcus lactis. Appl Environ Microbiol. 1985 Aug;50(2):540–542. doi: 10.1128/aem.50.2.540-542.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- von Wright A., Taimisto A. M., Sivelä S. Effect of Ca2+ ions on plasmid transformation of Streptococcus lactis protoplasts. Appl Environ Microbiol. 1985 Oct;50(4):1100–1102. doi: 10.1128/aem.50.4.1100-1102.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- von Wright A., Tynkkynen S. Construction of Streptococcus lactis subsp. lactis Strains with a Single Plasmid Associated with Mucoid Phenotype. Appl Environ Microbiol. 1987 Jun;53(6):1385–1386. doi: 10.1128/aem.53.6.1385-1386.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- von Wright A., Tynkkynen S., Suominen M. Cloning of a Streptococcus lactis subsp. lactis Chromosomal Fragment Associated with the Ability To Grow in Milk. Appl Environ Microbiol. 1987 Jul;53(7):1584–1588. doi: 10.1128/aem.53.7.1584-1588.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]