Abstract
We describe a computer-based method that selects representative clones for full-length sequencing in a full-length cDNA project. Our method classifies end sequences using two kinds of criteria, grouping, and clustering. Grouping places together variant cDNAs, family genes, and cDNAs with sequencing errors. Clustering separates those cDNA clones into distinct clusters. The full-length sequences of the clones selected by grouping are determined preferentially, and then the sequences selected by clustering are determined. Grouping reduced the number of rice cDNA clones for full-length sequencing to 21% and mouse cDNA clones to 25%. Rice full-length sequences selected by grouping showed a 1.07-fold redundancy. Mouse full-length sequences showed a 1.04-fold redundancy, which can be reduced by ∼30% from the selection using our previous method. To estimate the coverage of unique genes, we used FANTOM (Functional Annotation of RIKEN Mouse cDNA Clones) clusters (the RIKEN Genome Exploration Research Group 2001). Grouping covered almost all unique genes (93% of FANTOM clusters), and clustering covered all genes. Therefore, our method is useful for the selection of appropriate representative clones for full-length sequencing, thereby greatly reducing the cost, labor, and time necessary for this process.
[The programs used in this paper are available online at http://genome.gsc.riken.go.jp/software/2C.]
Full-length cDNA projects attempt to collect all mRNAs transcribed from the genome and determine the full-length cDNA sequences in their entirety. Considerable effort has been expended to improve the techniques involved. At the completion of mouse and human full-length cDNA projects, 30,000 to 100,000 full-length cDNA sequences will have been determined. The determination of full-length sequences requires large amounts of reagents, manpower, and time, which can be reduced by removing redundant full-length cDNA clones that potentially number in the thousands. As an experimental approach, mRNAs transcribed at high levels can be removed by normalization and subtraction during the construction of cDNA libraries (Carninci et al. 1996, 1997, 1998, 2000; Carninci and Hayashizaki 1999). However, because experimental normalization and subtraction are very difficult to apply to similar clones belonging to the same gene family, redundancies remain in the normalized cDNA library. Another possibility is, after end sequencing of the cDNAs, to identify negligibly redundant clones by a computational approach to classify the end sequences.
In general, when several full-length cDNA sequences are determined, redundancy can be reduced using homology search software by aligning the full-length cDNA sequences with their end sequences. Genomic sequences are also useful for reducing the redundancy. However, early in a project or when genome sequences are unavailable, the end sequences need to be classified. Several programs for clustering single-read expressed sequence tags (EST) have been reported (Adams et al. 1995, Boguski and Schuler 1995; Sutton et al. 1995; Schuler et al. 1996, 1997; Burke et al. 1998, 1999; Miller et al. 1999; Parsons and Rodriguez-Tome 2000; Haas et al. 2000; Christoffels et al. 2001; Quackenbush et al. 2001). However, the results of classification vary slightly depending on the program used (Bouck et al. 1999). Table 1 shows the characteristics of several gene indexing databases and the databases used for our full-length cDNA projects.
Table 1.
Characteristics of Gene Indexing Databases and Classification Methods
| Feature | Databases and clustering methods | |||||
|---|---|---|---|---|---|---|
| TGIa | UniGeneb | STACKc | GeneNestd | Mouse cDNA projecte | Rice cDNA project (our method) | |
| Clustering program | MegaBlastf and CAP3g | MegaBlast | d2_clusterh | BLAST | BLAST | BLAST |
| Representative clones | No | Yes | No | Yes | Yes | Yes |
| Consensus sequences | Yes | No | Yes | Yes | no | No |
| Alignments | Yes | No | Yes | Yes | Yes | No |
| Alternative splices | Different clusters | Same clusters | Different clusters | Same clusters | Same clusters | Same and different clusters |
| Redundancy of groups | High | Low | High | Low | Low | Low |
| Visualization tool | Yes | No | Yes | Yes | Yes | No |
| Other information | Coding potential | Annotation tissue | Tissue | Sequence quality | Tissue | Tissue |
Burke et al. 1999. Four gene indexing databases and two classification methods used for our full-length cDNA projects are compared based on eight distinct points of view.
In our mouse full-length cDNA project (http://genome.gsc.riken.go.jp), we classified 1,100,000 end sequences using BLAST homology search software (Pearson and Lipman 1988). We focused on the 100-bp 3′ end sequences of each cDNA sequence and placed into the same cluster end sequences those ≥90% identical over a ≥90-bp region (Konno et al. 2001). The accuracy and average read-length of the sequences were not consistent at the beginning of the project but have improved as the project has progressed. However, the criteria of classification were determined in light of the accuracy and average read-length of the early stages of the project and have been applied without further modification.
After determining the full-length sequences of the mouse cDNAs, we analyzed the redundancy of 21,076 mouse full-length sequences selected by the previous classification method. The analysis showed a 1.35-fold redundancy in the sequences. The method selects a single representative clone from each cluster, rearrays it from the master plate onto another plate, and does the sequencing in full. When we determine the full-length sequences of additional clones from the same cluster to analyze variant mRNAs and various members of the same gene family, we have to return to the master plates. However, this step requires much time and the enormous number of master plates increases the possibility of error.
To address these problems, we developed a two-step classification method, which consists of two distinct criteria for classifying end sequences, grouping, and clustering. Grouping places cDNAs with similar sequences together because they are derived from the same gene family and those cDNAs whose differences are a result of sequencing errors. Clustering segregates variant clones into different clusters. We designed the methods for classifying each process and adjusted the parameters of the homology searches by using the 213,404 3′ end sequences of mouse cDNAs determined in our laboratory. We selected the representative clones for full-length sequencing from each group and cluster in light of the results of classifying the end sequences. To ensure the effectiveness of the classification, we calculated the redundancy of the representative clones selected by grouping and estimated the coverage of unique genes. Although grouping does not cover a family gene, clustering can cover family genes derived from several loci on the mouse genome (ftp://ftp.sanger.ac.uk/pub/image/tmp/ssahaAssemble/mouse).
In 2000, we applied our method to the rice full-length cDNA project (S. Kikuchi, K. Satoh, T. Nagata, N. Kawagashira, K. Doi, N. Kishimoto, J. Yazaki, M. Ishikawa, K. Kojima, T. Namiki et al. in prep.) and classified both end sequences of cDNAs. During the project, we plotted a graph showing the increase in the number of novel groups and clusters in proportion to the increasing number of end sequences. We describe these efforts here.
RESULTS
Two-Step Classification
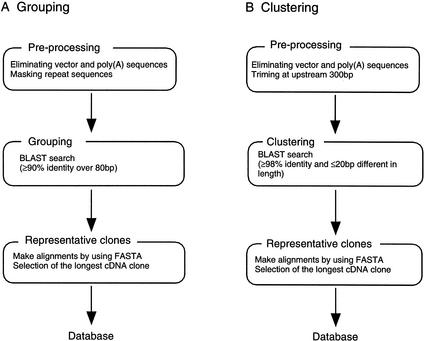
We established grouping and clustering criteria for the classification of end sequences to collect minimally redundant and variant clones in parallel (Fig. 1). Overlap length, percentage identity, direction of strand, and aligned positions extracted from the results of BLAST searches were used, but not other information such as annotations.
Figure 1.
Flow chart of two-step classification of end sequences. After the determination of the 5′ and/or 3′ end sequences, we classify them on the basis of two distinct criteria: (A) grouping and (B) clustering. Each classification is performed separately and consists of three steps: preprocessing, grouping or clustering, and selection of the representative clones. The criteria of each step are determined as stated in the Methods section.
For the grouping, the entire set of end sequences was searched, and transcripts of various lengths but containing a similar region were placed together. To further consolidate similar sequences into a single group, those with a common clone were merged, resulting in each clone belonging to only one group. The grouping procedure is as follows:
- 1.
Vector and poly(A) sequences are eliminated by computational analysis (Konno et al. 2001).
- 2.
Repeat sequences in end sequences are masked using RepeatMasker software (A. Smit, unpubl.; http://www.genome.washington.edu/uwgc/analysistools/repeatmask.htm).
- 3.
An entire end sequence is searched against all other end sequences using BLAST homology search software.
- 4.
Clones containing cDNAs that are ≥90% identical over ≥80 bp are placed into the same group in light of the pairwise alignments constructed in Step 3.
- 5.
Steps 3 and 4 are repeated for all remaining end sequences.
- 6.
Groups with a clone in common are merged so that each clone appears in only one group.
- 7.
End sequences in each group are aligned using FASTA homology search software (Pearson and Lipman 1988). In light of the alignment results of 5′ and 3′ end sequences, the clone containing the longest cDNA sequence in each group is typically selected as representative of that group. If either the 5′ or 3′ end sequence of the same clone is available, the longest cDNA sequence is selected using either end.
- 8.
The group information is stored in a relational database.
In clustering, repeat sequences were not masked, because masked regions are not regarded as matched regions and cannot satisfy stringent clustering criteria. In clustering, previously unclassified sequences that satisfied the clustering criteria were selected and allocated to a new cluster to not consolidate similar sequences as groups to prevent the same clone from appearing in more than one cluster. In summary, the clustering procedure is as follows:
- 1.
Vector and poly(A) sequences are eliminated by computational analysis (Konno et al. 2001).
- 2.
All end sequences are trimmed at upstream 300 bp.
- 3.
An entire end sequence is searched against all other end sequences using BLAST homology search software.
- 4.
Sequences ≥98% identical and ≤20 bp different in the lengths of their overlapping regions and their ends are placed together. However, a clone that is already assigned to a cluster is not placed into another; each clone belongs to a single cluster.
- 5.
Steps 3 and 4 are repeated for all end sequences.
- 6.
The end sequences in each cluster are aligned using FASTA homology search software. In light of the alignment results of 5′ and 3′ end sequences, the clone containing the longest cDNA insert in each cluster is selected as representative of that cluster. If either the 5′ or 3′ end sequence of the same clone is available, the longest cDNA sequence is selected using either end.
- 7.
The cluster information is stored in a relational database.
We processed the grouping and clustering of about 100,000 sequences in 2 d using a Compaq AlphaServer ES40 (four 500-MHz Alpha 21264 processors) with 6534 MB memory. Grouping and clustering can be performed faster in parallel, especially when the analyses involve more than 100,000 sequences.
Reduction of the Number of Clones for Full-Length Sequencing
We used our two-step classification method for the rice full-length cDNA project (S. Kikuchi, K. Satoh, T. Nagata, N. Kawagashira, K. Doi, N. Kishimoto, J. Yazaki, M. Ishikawa, K. Kojima, T. Namiki et al. in prep.) in 2000. We used the results of classification of 5′ as well as 3′ end sequences to determine the population of representative clones. When clones could be classified into several groups according to the 5′ end of the clones, we selected representative clones from all separated groups of the 5′ end. After the grouping and clustering of 97,808 cDNA clones, the number of representative clones for full-length sequencing decreased to 20,806 (21%) by grouping and to 62,888 (64%) by clustering. Here, we also classified 213,404 mouse 3′ end sequences by grouping and clustering and calculated the numbers of representative clones. The number for full-length sequencing decreased to 54,371 (25%) by grouping and to 194,977 (91%) by clustering. The clustering of rice end sequences eliminated 36% of the total cDNA clones, whereas the clustering of mouse end sequences eliminated only 9%. Because mouse end sequences used in this study were determined several years ago, they contain sequences with low accuracy (<98%), and it is difficult to judge whether these sequences were derived from sequencing errors or gene variants. Recently, sequencing accuracy has improved to ≥98%, so that in the case of the rice full-length cDNA project we could distinguish between sequencing errors and gene variants using clustering criteria.
The 7802 rice cDNA clones selected on the basis of the grouping results were sequenced completely, and there was a 1.07-fold redundancy in the full-length sequences according to the Smith-Waterman algorithm, with the restriction of ≥90% base identity over 80% overall length. When both end sequences were used, the cDNAs inserts of the clones selected as representatives were longer. On the other hand, the mouse full-length cDNA sequences of FANTOM clones (the RIKEN Genome Exploration Research Group 2001; Bono et al. 2002) selected by grouping showed a 1.04-fold redundancy according to the Smith-Waterman algorithm with the same restriction in the rice full-length sequences. When we aligned the 4% redundant full-length sequences with their end sequences, we found that none of the redundant clones could be removed from the collection of representative clones by using only the information in the end sequences because of differences in the identity, overall length, and overlap length between the end sequences and full-length sequences (data not shown). In comparison, the full-length sequences of the FANTOM clones selected by using the clustering method previously used in the mouse full-length cDNA project (Konno et al. 2001) showed a 1.35-fold redundancy according to the Smith-Waterman algorithm with the same restriction. Approximately 30% of these redundant clones could be removed from the collection of representative clones for full-length sequencing using our grouping process.
Coverage of Genes by the Two-Step Classification
Because the criteria for selection by grouping are looser than those for clustering, some unique genes may not be selected as representative clones after grouping. Therefore we used the FANTOM clusters to evaluate the coverage of genes by grouping. The representative clones selected by grouping accounted for 93% (13,359 clusters) of the FANTOM clusters, whereas those selected by clustering covered 98% (14,084 clusters). However, the functional annotations and cDNA sequences of the remaining 2% of the FANTOM clusters were the same as those of the already covered clusters (98%), so that clustering covered all the FANTOM clusters. Therefore, when the full-length sequences of the representative clones selected by grouping are determined predominantly in a full-length cDNA project, 93% of the unique genes can be collected with less redundancy. In addition, the remaining unique genes will already be rearrayed to allow easy selection and determination of their full-length sequences without the need to handle a large number of master plates. This combination of grouping and clustering enabled us to efficiently and thoroughly collect unique genes.
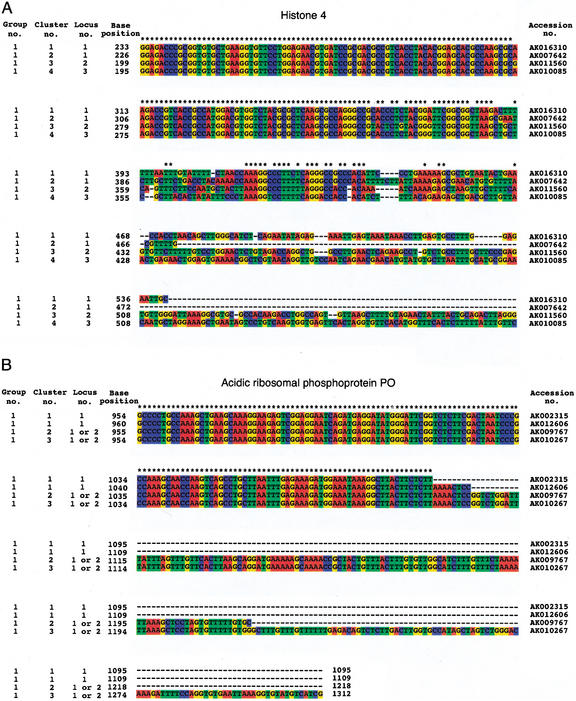
Clustering separated variant clones into distinct clusters. Figure 2 shows examples of family genes that were placed into a single group but separated into several clusters. Figure 2A presents the situation of family member genes containing similar and variable regions. This group includes four clones for histone 4 protein (AK016310, AK007642, AK011560, and AK010085) that were separated into four clusters. These full-length cDNA sequences matched other loci on the mouse draft genome (ftp://ftp.sanger.ac.uk/pub/image/tmp/ssahaAssemble/mouse). Grouping led to the selection of a clone as the representative for full-length sequencing from the family genes, but clustering covered the remaining genes as representative clones. Figure 2B presents the situation of family member genes that contain different polyadenylation sites. This group includes four clones for acidic ribosomal phosphoprotein PO (AK002315, AK009767, AK010267, and AK012606) that were separated into three clusters. These full-length cDNA sequences were similar to each other, but the lengths of their 5′ and 3′ ends differed; differences in the length of the 3′ end can be caused by differential poly(A) sites (Gautheret et al. 1998), internal priming, or artifacts incurred during the construction of the cDNA library. These clones matched one or two loci on the mouse draft genome.
Figure 2.
Multiple alignment of full-length cDNAs belonging to the same group and having the same functional annotation using the ClustalX program (Thompson et al. 1997). (A) This group contains four cDNAs of the histone 4 protein (AK016310, AK007642, AK011560, and AK010085). These sequences have a homologous region and a variable region, so that these sequences were placed together in grouping but separated into distinct clusters. We compared these sequences with mouse draft genome sequences. As a result, these sequences matched all distinct loci on the mouse draft genome, indicating that they were derived from distinct genes. With regard to grouping, one clone is selected as the representative, but all clones are selected as representatives after clustering. (B) This group contains four cDNAs of the acidic ribosomal phosphoprotein PO (AK002315, AK009767, AK010267, and AK012606). The 3′ ends of the upper two cDNA sequences differ in length by ≤20 bp; therefore, these sequences are regarded as the same cDNAs after clustering. These sequences matched the same locus on the mouse genome. However, the lengths of the 3′ ends of the other two cDNA sequences differ by >20 bp, so that these sequences separate into distinct clusters. These sequences may match another locus on the mouse genome. With regard to grouping, one clone is selected as the representative, but three clones are selected as representatives after clustering.
Number of Groups and Clusters Formed from a cDNA Library
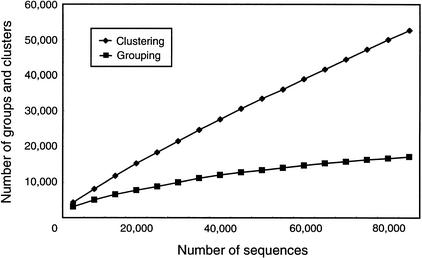
In a full-length cDNA project, it is necessary to estimate the number of clones with unique cDNA inserts that can be derived from a library to avoid redundant full-length sequencing. This can be effectively performed with a graph showing the increase in the number of novel groups and clusters in proportion to the increase in the number of end sequences (Fig. 3). As the number of sequences increases, the number of new groups and clusters increase. After 10,000 sequences had been determined, the addition of 10,000 new sequences yielded about 2699 novel groups. After 50,000 sequences had been determined, the addition of 10,000 new sequences yielded only about 1331 novel groups. However, the numbers of groups and clusters are not subject to saturation under these circumstances. From the graph in Figure 3, we can use the increase in the number of novel groups to determine whether we can collect almost all cDNA clones in a cDNA library.
Figure 3.
An increased number of groups and clusters of 3′ end sequences in the rice full-length cDNA project. As end sequences were determined, the number of novel groups and clusters increased, whereas the rate of the increase gradually decreased.
DISCUSSION
A full-length cDNA library includes similar sequences that have been duplicated on the genome during evolution and are derived from sequencing errors. Among them, distinct full-length cDNA sequences should be determined first, before the variant sequences, for efficient execution of the full-length cDNA project. In this study, we have shown that grouping can collect negligible redundant cDNAs and clustering can select variant cDNAs.
Among several gene index projects (Table 1), TGI, UniGene, and GeneNest adopted single clustering criteria, which allow splice variants to be incorporated into the same cluster or to be separated into distinct clusters. STACK categorizes splice variants according to the tissue from which they were derived. However, our method can collect and separate splice variants using two distinct criteria; at present, our method collects splice variants that occur on end sequences by computationally comparing their length and identity. However, we cannot select each splice variant by examining their alignments. A visualization tool should be added to our method as in the mouse full-length cDNA project. In the case of UniGene, a sequence of a single cluster, which may be derived from a low-quality sequence, is merged with a cluster at a lower level of stringency, and GeneNest removes low-quality sequences before clustering. Our method does not take sequence quality into consideration, so that low-quality sequences will be separated into distinct clusters according to clustering criteria. We also compared our method with the d2_cluster program in STACK and StackPack (Burke et al. 1999). The results of the classification of 213,404 mouse 3′ end cDNA sequences using a d2_cluster were nearly the same as for those from grouping; d2_cluster covered 93% of the FANTOM clusters. The results of the classification of variant clones selected by clustering were assessed with CRAW (Burke et al. 1999). However, neither of these programs can automatically select representative clones for full-length sequences. Because a full-length cDNA project must identify likely candidates for full-length sequencing, we designed our two-step classification method to automatically and simultaneously collect representative clones that were negligibly redundant and variant. Our method also allows the researcher to change the priority with which clones derived from gene variants and artifacts are sequenced in full.
Additional methods might facilitate the collection of rare genes or reduction of artifact products by changing the priority of clone sequencing. In a full-length cDNA project, the representative clones selected by grouping and clustering need to be rearrayed from the master plates before beginning full-length sequencing, and then the full-length sequences of the representative clones selected by grouping should be determined by collecting cDNAs that are as distinctive as possible. Therefore, by aligning the full-length cDNA and end sequences, we can identify variant sequences with alternative transcriptional start sites, alternative polyadenylation sites, and artifacts such as internal priming (Gautheret et al. 1998). We then can change the priority of the sequencing of these clones.
Here we propose another method for selecting the representative clones. Clones in single-member groups or clusters are likely to be derived from rare genes or to be the result of contamination by genomic sequences. Therefore, after grouping or clustering of the end sequences, we can rearray the representative clones derived from the single-member groups and clusters on the same plates and postpone determining the full-length sequences of these clones. Clones belonging to groups or clusters with two or more members are not likely to be derived from artifacts. Therefore, the full-length sequencing of these clones can be our first priority.
In the rice full-length cDNA project, our two-step classification method yielded many of the same results as for the mouse cDNA sequences. This indicates that our method may be useful for full-length cDNA projects of various species. Clones selected by grouping should also be useful for the construction of a DNA microarray and proteome analysis, because these clones include negligibly redundant and unique genes, and the full-length sequences of these clones are determined preferentially.
METHODS
Criteria for Classification of End Sequences
We established grouping and clustering criteria for the classification of end sequences to collect minimally redundant and variant clones in parallel. Because BLAST homology search software can process many sequences quickly, we used this software in the classification of end sequences. For the purpose of grouping, we determined the criteria of the BLAST homology search by examining the number of groups obtained when we varied the lower limits of identity and overlap length between the end sequences under comparison. We used our laboratory's 213,404 3′ end sequences, which include those from the full-length cDNA sequences of the 21,076 FANTOM clones (the RIKEN Genome Exploration Research Group 2001), to carry out this experiment.
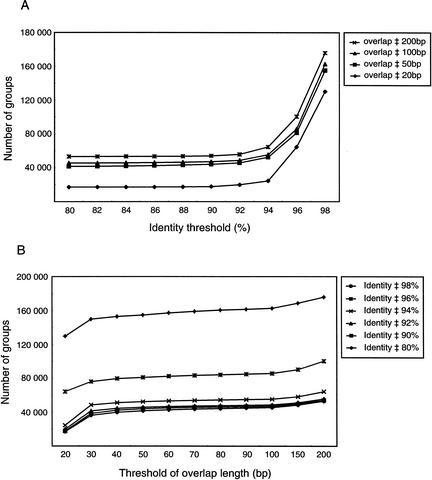
First, we set the lower limit for the length of the overlap between two end sequences at 50 to 200 bp and varied the identity threshold for these sequences from 10% to 98%; the expected (E) value threshold was set at 100 (Fig. 4A). The number of groups was relatively constant between 80% and 92% identity but dramatically increased at >94% identity. Gene variants and cDNAs with sequencing errors will increasingly be placed in the same group when the identity threshold increases from 80% to 92%; therefore we adopted a 90% identity threshold as a grouping criterion for this study.
Figure 4.
Determination of the criteria of identity value and overlapping length in grouping condition. We classified 213,404 mouse 3′ end sequences in light of the results of homology searches using BLAST software (Pearson and Lipman 1988) to determine the criteria of grouping. (A) Clones whose end sequences were more similar than the identity threshold were placed together. Here, the identity threshold varied from 80% to 98%. The number of resulting groups increased as the identity threshold increased from 90%; therefore, an identity threshold of 90% is appropriate for placing similar sequences together. (B) Clones whose end sequences exceeded the overlap threshold were placed in the same group. In this example, the overlap threshold varied from 20 to 200 bp. The number of resulting groups was almost constant between 30 and 150 bp; therefore, an overlap threshold of 30 to 150 bp is appropriate for grouping.
Next, we analyzed the change in the number of groups obtained when the threshold for the length of the overlap between two end sequences varied from 20 to 200 bp (Fig. 4B). The number of groups dramatically increased when the overlap threshold was 20 to 30 bp but only gradually increased when this limit ranged from 150 to 200 bp. Therefore, an overlap threshold of 30 to 150 bp was appropriate for the grouping criterion; here, we adopted a limit of 80 bp.
Because the goal of the clustering process is to separate variant sequences into distinct clusters, the criteria for the BLAST homology search needed to be sufficiently stringent to prevent the merging of collections of similar end sequences, as happens in grouping. When using the 213,404 mouse cDNAs, we classified them by evaluating only their 3′ end sequences. For the rice full-length cDNA project, we used 5′ and 3′ end sequences. The lower limit for the identity value was set at 98%, because the accuracy of sequencing typically is ≥98% and because sequences that were ≥98% identical were regarded as the same gene by human inspection. End sequences used in clustering are trimmed at upstream 300 bp, because the identity threshold of clustering is high (98%); therefore a high-quality part of the sequences should be used. At the 3′ end of the cDNAs, the polyadenylation site starts 10 to 30 bp downstream of the poly(A) signal (Wahle and Keller 1992; Wahle 1995; Edwalds-Gilbert et al. 1997). Therefore, we placed in the same cluster, clones whose 3′ end sequences differed by ≤20 bp in length. Because the transcriptional start site typically lies mainly 10 to 80 bp from the 5′ end of the sequence (Suzuki et al. 2001), we also clustered clones whose 5′ ends differed by ≤10 bp.
WEB SITE REFERENCES
http://genome.gsc.riken.go.jp; Mouse full-length cDNA project.
http://genome.gsc.riken.go.jp/software/2C; Programs and sequences used in this paper.
http://www.genome.washington.edu/uwgc/analysistools/repeatmask.htm; Web site for RepeatMasker software.
Acknowledgments
This work would not have been possible without the encouragement and support offered by Keiji Kainuma. We thank Hidemasa Bono, Shiro Fukuda, Jun Adachi, Rintaro Saito, Tetsuya Saito, Takeya Kasukawa, Masaaki Furuno, Shigeyasu Yoshida, and Yoshimi Ota for their discussion, encouragement, and technical assistance. We also thank Katsunori Aizawa, Norihito Hayatsu, Tomoko Hirozane, Yoshiyuki Ishii, Ayako Yasunishi, Ayako Hara, Daisuke Sasaki, and the members of the RIKEN Genome Exploration Research Group Science Center for the data preparation.
This study was supported by a research grant for the RIKEN Genome Exploration Research Project from the Ministry of Education, Culture, Sports, Science, and Technology of the Japanese Government to Y.H. and ACT-JST (Research and Development for Applying Advanced Computational Science and Technology) of the Japan Science and Technology Corporation (JST) to Y.H. This work was also supported by a research grant for the Rice Genome Full-Length cDNA Library Construction Project from BRAIN (Bio-oriented Technology Research Advancement Institution) to Y.H.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
Footnotes
E-MAIL rgscerg@gsc.riken.go.jp; FAX: 81 45 503 9222.
Article and publication are at http://www.genome.org/cgi/doi/10.1101/gr.75202. Article published online before print in June 2002.
REFERENCES
- Adams MD, Kerlavage AR, Fleischmann RD, Fuldner RA, Bult CJ, Lee NH, Kirkness EF, Weinstock KG, Gocayne JD, White O. Initial assessment of human gene diversity and expression patterns based upon 83 million nucleotides of cDNA sequence. Nature. 1995;377 (6547 Suppl):3–174. [PubMed] [Google Scholar]
- Boguski MS, Schuler GD. Establishing a human transcript map. Nat Genet. 1995;10:369–371. doi: 10.1038/ng0895-369. [DOI] [PubMed] [Google Scholar]
- Bono H, Kasukawa T, Furuno M, Hayashizaki Y, Okazaki Y. FANTOM DB: Database of Functional Annotation of RIKEN Mouse cDNA Clones. Nucleic Acids Res. 2002;30:116–118. doi: 10.1093/nar/30.1.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bouck J, Yu W, Gibbs R, Worley K. Comparison of gene indexing databases. Trends Genet. 1999;15:159–162. doi: 10.1016/s0168-9525(99)01709-6. [DOI] [PubMed] [Google Scholar]
- Burke J, Wang H, Hide W, Davison DB. Alternative gene form discovery and candidate gene selection from gene indexing projects. Genome Res. 1998;8:276–290. doi: 10.1101/gr.8.3.276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burke J, Davison D, Hide W. d2_Cluster: A validated method for clustering EST and full-length cDNA sequences. Genome Res. 1999;9:1135–1142. doi: 10.1101/gr.9.11.1135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carninci P, Hayashizaki Y. High-efficiency full-length cDNA cloning. Methods Enzymol. 1999;303:19–44. doi: 10.1016/s0076-6879(99)03004-9. [DOI] [PubMed] [Google Scholar]
- Carninci P, Kvam C, Kitamura A, Ohsumi T, Yasushi O, Itoh I, Kamiya K, Shibata K, Sasaki N, Izawa M. High-efficiency full-length cDNA cloning by biotinylated CAP trapper. Genomics. 1996;137:327–336. doi: 10.1006/geno.1996.0567. [DOI] [PubMed] [Google Scholar]
- Carninci P, Nishiyama Y, Westover A, Itoh M, Nagaoka S, Sasaki N, Okazaki Y, Muramatsu M, Hayashizaki Y. Thermostabilization and thermoactivation of thermolabile enzymes by trehalose and its application for the synthesis of full-length cDNA. Proc Natl Acad Sci. 1998;95:520–524. doi: 10.1073/pnas.95.2.520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carninci P, Shibata Y, Hayatsu N, Sugahara Y, Shibata K, Itoh M, Konno H, Okazaki Y, Muramatsu M, Hayashizaki Y. Normalization and subtraction of cap-trapper-selected cDNAs to prepare full-length cDNA libraries for rapid discovery of new genes. Genome Res. 2000;10:1617–1630. doi: 10.1101/gr.145100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carninci P, Westover A, Nishiyama Y, Ohsumi T, Itoh M, Nagaoka S, Sasaki N, Okazaki Y, Muramatsu M, Schneider C. High efficiency selection of full-length cDNA by improved biotinylated cap trapper. DNA Res. 1997;4:61–66. doi: 10.1093/dnares/4.1.61. [DOI] [PubMed] [Google Scholar]
- Christoffels A, van Gelder A, Greyling G, Miller R, Hide T, Hide W. STACK: Sequence tag alignment and consensus knowledge base. Nucleic Acids Res. 2001;29:234–238. doi: 10.1093/nar/29.1.234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edwalds-Gilbert G, Veraldi KL, Milcarek C. Alternative poly(A) site selection in complex transcription units: Means to an end? Nucleic Acids Res. 1997;25:2547–2561. doi: 10.1093/nar/25.13.2547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gautheret G, Poirot O, Lopez F, Audic S, Claverie JM. Alternate polyadenylation in human mRNAs: A large-scale analysis by EST clustering. Genome Res. 1998;8:524–530. doi: 10.1101/gr.8.5.524. [DOI] [PubMed] [Google Scholar]
- Haas SA, Beissbarth T, Rivals E, Krause A, Vingron M. GeneNest: Automated generation and visualization of gene indices. Trends Genet. 2000;16:521–523. doi: 10.1016/s0168-9525(00)02116-8. [DOI] [PubMed] [Google Scholar]
- Huang X, Madan A. CAP3: A DNA Sequence Assembly Program. Genome Res. 1999;9:868–877. doi: 10.1101/gr.9.9.868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konno H, Fukunishi Y, Shibata K, Itoh M, Carninci P, Sugahara Y, Hayashizaki Y. Computer-based methods for the mouse full-length cDNA encyclopedia: Real-time sequence clustering for construction of a nonredundant cDNA library. Genome Res. 2001;11:281–289. doi: 10.1101/gr.145701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller RT, Christoffels AG, Gopalakrishnan C, Burke J, Ptitsyn AA, Broveak TR, Hide WA. A comprehensive approach to clustering to expressed human gene sequence: The sequence tag alignment and consensus knowledge base. Genome Res. 1999;9:1143–1155. doi: 10.1101/gr.9.11.1143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parsons JD, Rodriguez-Tome P. JESAM: CORBA software components to create and publish EST alignments and clusters. Bioinformatics. 2000;16:313–325. doi: 10.1093/bioinformatics/16.4.313. [DOI] [PubMed] [Google Scholar]
- Pearson WR, Lipman DJ. Improved tools for biological sequence comparison. Proc Natl Acad Sci. 1988;85:2444–2448. doi: 10.1073/pnas.85.8.2444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quackenbush J, Cho J, Lee D, Liang F, Holt I, Karamycheva S, Parvizi B, Pertea G, Sultana R, White J. The TIGR Gene Indices: Analysis of gene transcript sequences in highly sampled eukaryotic species. Nucleic Acids Res. 2001;29:159–164. doi: 10.1093/nar/29.1.159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- The RIKEN Genome Exploration Research Group Phase II Team and the FANTOM Consortium. Functional annotation of a full-length mouse cDNA collection. Nature. 2001;409:685–690. doi: 10.1038/35055500. [DOI] [PubMed] [Google Scholar]
- Schuler GD, Boguski MS, Stewart EA, Stein LD, Gyapay G, Rice K, White RE, Rodriguez-Tome P, Aggarwal A, Bajorek E. A gene map of the human genome. Science. 1996;274:540–546. [PubMed] [Google Scholar]
- Schuler GD. Pieces of the puzzle: Expressed sequence tags and the catalogue of human genes. J Mol Med. 1997;75:694–698. doi: 10.1007/s001090050155. [DOI] [PubMed] [Google Scholar]
- Sutton G, White O, Adams M, Kerlavage A. TIGR assembler: A new tool for assembling large shotgun sequencing projects. Genome Sci Tech. 1995;1:9–19. [Google Scholar]
- Suzuki Y, Taira H, Tsunoda T, Mizushima-Sugano J, Sese J, Hata H, Ota T, Isogai T, Tanaka T, Morishita S, et al. Diverse transcriptional initiation revealed by fine, large-scale mapping of mRNA start sites. EMBO Rep. 2001;2:388–393. doi: 10.1093/embo-reports/kve085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG. The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997;25:4876–4882. doi: 10.1093/nar/25.24.4876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wahle E. 3′-End cleavage and polyadenylation of mRNA precursors. Biochim Biophys Acta. 1995;1261:183–194. doi: 10.1016/0167-4781(94)00248-2. [DOI] [PubMed] [Google Scholar]
- Wahle E, Keller W. The biochemistry of 3′-end cleavage and polyadenylation of messenger RNA precursors. Ann Rev Biochem. 1992;61:419–440. doi: 10.1146/annurev.bi.61.070192.002223. [DOI] [PubMed] [Google Scholar]