Abstract
DNA in living cells is generally processed via the generation and the protection of single-stranded DNA involving the binding of ssDNA-binding proteins (SSBs). The studies of SSB-binding mode transition and cooperativity are therefore critical to many cellular processes like DNA repair and replication. However, only a few atomic force microscopy (AFM) investigations of ssDNA nucleoprotein filaments have been conducted so far. The point is that adsorption of ssDN A–SSB complexes on mica, necessary for AFM imaging, is not an easy task. Here, we addressed this issue by using spermidine as a binding agent. This trivalent cation induces a stronger adsorption on mica than divalent cations, which are commonly used by AFM users but are ineffective in the adsorption of ssDNA–SSB complexes. At low spermidine concentration (<0.3 mM), we obtained AFM images of ssDNA–SSB complexes (E. coli SSB, gp32 and yRPA) on mica at both low and high ionic strengths. In addition, partially or fully saturated nucleoprotein filaments were studied at various monovalent salt concentrations thus allowing the observation of SSB-binding mode transition. In association with conventional biochemical techniques, this work should make it possible to study the dynamics of DNA processes involving DNA–SSB complexes as intermediates by AFM.
INTRODUCTION
Single-stranded binding (SSB) proteins bind with a high affinity to single-stranded DNA (ssDNA), which is a transient state in many DNA metabolic processes such as replication, repair and recombination (1–4). SSB proteins share a common conserved domain called oligonucleotides/oligosaccharides-binding (OB) fold to bind to ssDNA on which they stabilize intermediates, remove secondary structure (e.g. hairpins, cruciforms) and protect it from DNA-damaging agents. Most kinds of SSB proteins bind non-specifically along ssDNA according to different binding modes depending on their structure, their degree of cooperativity, their level of oligomerization and the environmental conditions (e.g. ionic strength, buffer composition). ssDNA–SSB complexes have been generally studied by ensemble-average biochemical approaches. Besides, imaging and analyses of ssDNA–SSB complexes at the single molecule level can reveal conformational changes which are essential in many processes. For this reason, transmission electron microscopy (TEM) has been extensively used to study the binding properties of various SSB proteins (5–11). In this context, the atomic force microscope (AFM) also appears as a unique instrument for studying DNA–protein interactions, owing to its high resolution and capability of working in liquid. Its potential has already been highlighted by the numerous investigations of double-stranded DNA (dsDNA) interacting with ligands (12–18). However, comparatively to EM, only a few AFM investigations of ssDNA or DNA–SSB complexes have been undertaken so far (16,19–23). The reason is that the spreading of ssDNA–SSB complexes on an atomically flat surface, generally mica, is not an easy task (16). In the few attempts to adsorb ssDNA nucleoprotein filaments on mica, glutaraldehyde, a strong cross-linking agent, was generally used (19,20) leading to a poor resolution. More importantly, future dynamical studies in liquid by AFM are then precluded under such strong fixation conditions.
Our purpose in this study is to develop an experimental method to adsorb ssDNA–SSB complexes on mica in order to extend the application domain of the AFM to ssDNA and its partner proteins. It is generally known that the addition of divalent cations to the deposition buffer overcomes the natural repulsion between the negatively charged DNA and the negatively charged mica surface, which leads to the adsorption of dsDNA (24–26) or dsDNA nucleoprotein filaments on mica (16,18). However ssDNA–SSB nucleoprotein filaments cannot be properly adsorbed by using the same method. The adsorption mediated by multivalent counterion correlations at the polyelectrolyte–mica interface (27) decreases sharply with the decrease of the polyelectrolyte surface charge density. Thus a loose binding of ssDNA–SSB complexes to mica, resulting from both the ssDNA neutralization and the enlargement of the diameter of the complex upon SSB binding, may explain the unsatisfactory result.
A solution to this problem is to use spermidine, a trivalent polyamine, as a powerful binding agent allowing the spreading of ssDNA nucleoprotein filaments on mica. As previously shown for dsDNA (28), spermidine can induce a strong DNA adsorption even at submillimolar concentration and can still be effective at high NaCl concentration (up to 300 mM). This is highly relevant for studying E. coli SSB since this homotetrameric protein (4 × 18.8 kDa) forms different types of complexes with ssDNA depending on the NaCl concentration. At low ionic strengths, the (E. coli SSB)35 binding mode for which the nucleic acid interacts with two monomers is preferred. However, the (E. coli SSB)65 binding mode for which the nucleic acid interacts with all four monomers is predominant at high ionic strengths. The transition between the two binding modes is reversible and is modulated by NaCl concentration (29).
In this article, we present high-resolution AFM images in air of ssDNA–E. coli SSB nucleoprotein filaments adsorbed on mica at both low and high ionic strengths. Partly and fully saturated ssDNA nucleoprotein filaments were observed by varying the E. coli SSB concentration, paving the way for the characterization of the E. coli SSB cooperative binding to ssDNA by AFM. Finally, we demonstrate that this method can be extended to many other ssDNA–SSB complexes as shown in this study for Bacteriophage T4 gene 32 protein (gp32) and yeast Replication Protein A (yRPA)–ssDNA complexes.
MATERIALS AND METHODS
DNA and SSB proteins
The chemical compounds (MgCl2, NaCl), spermidine (SpdCl3) and M13mp18 plasmid phage DNA (M13 ssDNA) were purchased from Sigma-Aldrich (Saint Quentin Fallavier, France). Bacteriophage T4 gene 32 protein (gp32) and the E. coli ssDNA-binding protein (E. coli SSB) were purchased from USB Corporation (Staufen, Germany) and were used without further purification. Yeast RPA (yRPA) was purified essentially as described by Kantake et al. (30).
Electrophoretic mobility shift assay experiments
M13 ssDNA–E. coli SSB complexes with variation of the ratio of E. coli SSB to ssDNA, were formed in two spermidine buffers: Tris 20 mM pH 7.5, NaCl 20 mM, SpdCl3 50 µM, and Tris 20 mM pH 7.5, NaCl 300 mM, SpdCl3 300 µM. Various concentrations of E. coli SSB were incubated for 10 min at 37°C with 100 ng of M13 ssDNA plasmid in both spermidine buffers. Samples were loaded onto 1% agarose gels in 1× TBE buffer (Tris 89 mM pH 8.3, boric acid 89 mM, EDTA 2 mM). The gels were run at 3 V/cm at 4°C for 3 h and ssDNA was stained in 1:10 000 equimolar mix of SYBR Green I and II (Molecular Probes) for 60 min.
AFM sample preparation
M13 ssDNA is diluted to a concentration of 2 µg/ml in a buffer solution containing Tris 20 mM pH 7.5, and different concentrations of NaCl, MgCl2 or spermidine and SSB proteins. Solutions were incubated at 37°C for 10 min. A 5-µl droplet of ssDNA–SSB solution was deposited onto the surface of freshly cleaved mica (muscovite) for 1 min. Then, the surface was rinsed with 0.02% diluted uranyl acetate solution in order to stabilize the ssDNA–SSB complexes in their 3D conformations for AFM imaging in air (31). The sample is then rapidly rinsed with pure water (Millipore) to obtain a clean surface after drying with filter paper.
The use of uranyl acetate discriminates between weak and firmly adsorbed molecules (28). Indeed, the addition of uranyl acetate triggers DNA aggregation in bulk solution (32). When the DNA molecules are loosely adsorbed on the surface, they adopt a nearly 3D conformation on the surface. Consequently, the addition of uranyl acetate leads to monomolecular DNA compaction on the surface. It is worth noting that the results presented here are not dependent on the uranyl acetate concentration for a large concentration range (0.2 and 0.02% uranyl acetate solutions have been tested).
AFM imaging
Imaging was performed in Tapping Mode™ with a Multimode™ AFM (Veeco, Santa Barbara, CA) operating with a Nanoscope IIIa™ controller. We used Olympus (Hamburg, Germany) silicon cantilevers AC160TS with nominal spring constants between 36 and 75 N/m. The scan frequency was typically 1.5 Hz per line and the modulation amplitude was a few nanometres. We only used a first or second order polynomial function to remove the background slope.
RESULTS AND DISCUSSION
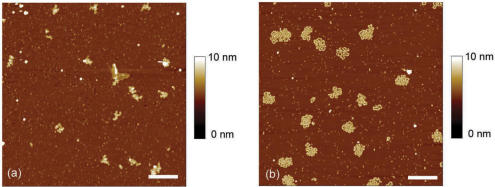
Mg2+ cations or other divalent cations are generally used to adsorb DNA–protein complexes on mica at low NaCl concentration. Figure 1a shows M13 ssDNA–E. coli SSB complexes adsorbed on mica in Mg buffer (Tris 20 mM pH 7.5, MgCl2 10 mM and NaCl 20 mM). It can be observed that the complexes are not properly spread on the surface even though the protein:nucleotide concentration ratio is larger than that necessary to saturate the ssDNA–E. coli SSB nucleoprotein filaments. The bright spots in the image may represent aggregated ssDNA–E. coli SSB complexes. It is worth noting that a loosely bound biopolymer can be transformed into a globule during the drying step, as it occurs for loosely bound dsDNA (28). Several experiments were performed at lower ionic strengths in order to increase the attraction force but failed to spread nucleoprotein filaments on mica.
Figure 1.
AFM images of M13 ssDNA–E. coli SSB on mica with a concentration ratio R [=(SSB tetramers)/(nucleotides)] equal to 1/30 in (a) Tris 20 mM pH 7.5, NaCl 20 mM, MgCl2 10 mM and, (b) Tris 20 mM pH 7.5, NaCl 20 mM, SpdCl3 50 µM (scale bars 500 nm).
In a previous model (28), we have shown that DNA binding to mica is expected to be very sensitive to the valence of the multivalent cations. The use of trivalent cations rather than divalent ones could significantly enhance the complex binding to mica. Indeed, the energy benefit of multivalent counterion correlations between the mica and ssDNA–SSB complex counterions, which trigger complex adsorption, is roughly proportional to z2, with z being the multivalent cation charge. In addition, higher valence counterions are better competitors for both mica and complex neutralizations and are not easily replaced by monovalent cations. Therefore, a submillimolar concentration of trivalent cation is sufficient to trigger the nucleoprotein filament adsorption at moderate or high ionic strengths. Spermidine, a naturally occurring trivalent polyamine (33), which interacts non-specifically with nucleic acids, is a good candidate to spread DNA on mica (28). According to constructed diagrams of DNA adsorption on mica mediated by spermidine (28), a spermidine concentration of 50 µM could be enough to trigger ssDNA–E. coli SSB filament adsorption on mica at moderate ionic strengths. Figure 1b shows M13 ssDNA–E. coli SSB complexes adsorbed on mica in spermidine buffer (Tris 20 mM pH 7.5, NaCl 20 mM, SpdCl3 50 µM) with the same protein:nucleotide concentration ratio as used in Figure 1a. It turns out that nucleoprotein filaments are then properly spread on the surface and their contours can be observed at high resolution on the mica surface. Besides the high-resolution imaging, AFM allows to study the cooperativity of the E. coli SSB binding to ssDNA. Indeed, E. coli SSB is known to bind cooperatively to single-stranded polynucleotides. The type and the magnitude of this cooperativity depend on the binding mode. The highly unlimited cooperative binding that results in a formation of long protein clusters on ssDNA occurs only when the tetramer adopts the (E. coli SSB)35 binding mode, i.e. at low ionic strengths (34). By contrast, (E. coli SSB)65 binding mode displays a limited type of cooperativity in which protein clusters are limited to the formation of dimers of tetramers (35).
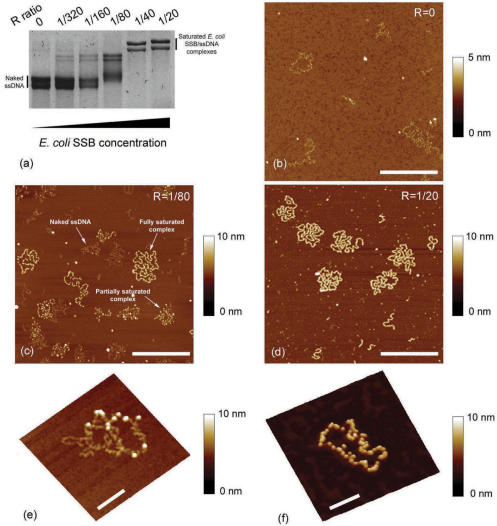
The AFM observation of E. coli SSB cooperativity requires to adsorb the nucleoprotein filament at low ionic strengths with a lower protein:nucleotide concentration ratio than that required for the saturation of the nucleotides. It is worth noting that a spermidine concentration of 50 µM may induce a transition from (E. coli SSB)35 to (E. coli SSB)65 and thus may prevent the unlimited cooperative binding observation (36). To address this issue, electrophoretic mobility shift assays (EMSA) were performed with increasing E. coli SSB concentration, in spermidine buffer (Tris 20 mM pH 7.5, NaCl 20 mM, SpdCl3 50 µM) in order to control the gradual formation of M13 ssDNA–E. coli SSB complexes. The results are presented in Figure 2a. We observe that saturation occurs at a site size of ∼40 (i.e. the number of nucleotides wrapped around one E. coli SSB tetramer). This is in excellent agreement with previous results reported for 20 mM NaCl (29) and allows the AFM observation of E. coli SSB unlimited cooperativity in the (E. coli SSB)35 binding mode. We then imaged naked M13 ssDNA (Figure 2b) and M13 ssDNA–E. coli SSB complexes for a ratio R of E. coli SSB tetramer:nucleotide concentrations below (R = 1/80) and above (R = 1/20) the saturation (Figure 2c and d). Fully saturated M13 ssDNA–E. coli SSB complexes were visualized for the first time with such high resolution. Figure 2c clearly shows the cooperative binding property of E. coli SSB. Indeed, naked ssDNA, partially and fully saturated M13 ssDNA–E. coli SSB complexes coexist on the mica surface.
Figure 2.
(a) Agarose gel electrophoresis of M13 ssDNA–E. coli SSB complexes formed in spermidine buffer Tris 20 mM pH 7.5, NaCl 20 mM, SpdCl3 50 µM with increasing E. coli SSB protein concentration: free ssDNA (lane 1), R = 1/320 (lane 2), R = 1/160 (lane 3), R = 1/80 (lane 4), R = 1/40 (lane 5) and R = 1/20 (lane 6). (b) AFM image of free M13ssDNA. (c) AFM image of M13 ssDNA–E. coli SSB complexes associated to lane 4. Free ssDNA, partly formed complex and nearly fully saturated complex coexist and are a typical feature of cooperative binding of E. coli SSB to ssDNA. (d) AFM image of fully saturated M13 ssDNA–E. coli SSB complexes associated to lane 6 (scale bars 500 nm). (e) Zoom on M13 ssDNA–E. coli SSB complexes (R = 1/320 and scale bar 100 nm) at which few SSB proteins can be distinguished. (f) Zoom on a nearly fully saturated M13 ssDNA–E. coli SSB complex (R = 1/40 and scale bar 100 nm). Even at high R value, single SSB protein can be resolved on the complex thread.
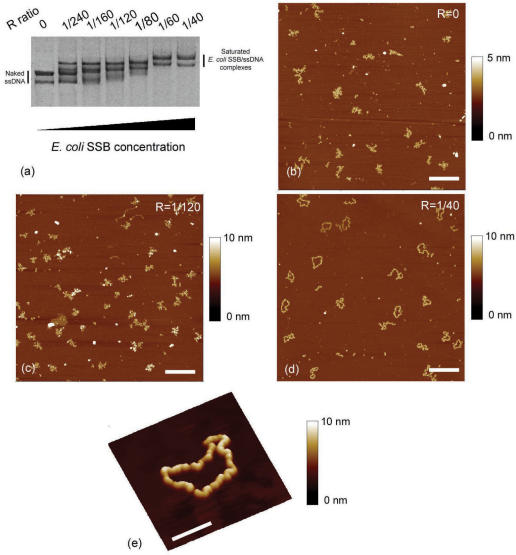
The (E. coli SSB)65 binding mode, which is associated with a limited cooperativity, requires a higher NaCl concentration (above 200 mM). As we discussed in a previous article (28), the adsorption of dsDNA mediated by spermidine allows to observe DNA by AFM even at high NaCl concentrations (up to 300 mM). The only requirement is to increase the spermidine concentration to compensate the effect of the spermidine counterion replacement by monovalent cations, which weakens the binding at high monovalent salt concentrations. Thus, with 300 mM NaCl, a spermidine concentration higher than 100 μM is necessary to trigger DNA adsorption, according to the diagram of dsDNA adsorption on mica in presence of spermidine (28). M13 ssDNA–E. coli SSB complexes, formed and adsorbed in a high monovalent salt concentration buffer (Tris 20 mM pH 7.5, NaCl 300 mM, SpdCl3 300 µM), were then studied by AFM (Figure 3b-3e). Prior to AFM imaging, we checked by gel shift assay (Figure 3a) that the formation of the nucleoprotein filaments took place under the conditions used for AFM. The measured concentration ratio at saturation indicates that the average number of nucleotides occluded by the E. coli SSB tetramer is ∼70 at such high ionic strengths which is close to the expected value in the (E. coli SSB)65 binding mode. At E. coli SSB tetramer: nucleotide concentration ratio equal to 1/120 (see Figure 3c), below the nucleoprotein filament saturation, different structures coexist in the AFM image, from naked ssDNA to more or less saturated nucleoprotein filaments. This is an illustration of the limited cooperativity which occurs at such high ionic strengths. At larger E. coli SSB tetramer concentration (E. coli SSB tetramer: nucleotide concentration ratio of 1/40), i.e. above the concentration ratio at saturation, we observed saturated filaments, the contours of which were well defined (see Figure 3d and e). Compared to Figure 2d, the M13 ssDNA–E. coli SSB complexes in Figure 3d have a lower contour length which indicates a different binding mode of E. coli SSB. This is in agreement with the model proposed by Lohman et al. (37) assuming that the E. coli SSB tetramer can be viewed as a ‘dimer of dimers’. The ssDNA is wrapped around only one dimer in (E. coli SSB)35 binding mode, whereas, for the (E. coli SSB)65 binding mode, the ssDNA is wrapped around both dimers, involving a lower contour length of the nucleoprotein complex. The AFM values of the adsorbed nucleoprotein contour length are 920 ± 30 nm and 560 ± 40 nm for the (E. coli SSB)35 and (E. coli SSB)65 binding mode, respectively.
Figure 3.
(a) Agarose gel electrophoresis of M13 ssDNA–E. coli SSB complexes formed in spermidine buffer Tris 20 mM pH 7.5, NaCl 300 mM, SpdCl3 300 μM with increasing E. coli SSB protein concentration: free ssDNA (lane 1), R = 1/240 (lane 2), R = 1/160 (lane 3), R = 1/120 (lane 4), R = 1/80 (lane 5), R = 1/60 (lane 6) and R = 1/40 (lane 7). (b) AFM image of free M13ssDNA. (c) AFM image of M13 ssDNA–E. coli SSB complexes associated to lane 4. Due to the higher ionic strength, the cooperative binding of E. coli SSB to ssDNA is less pronounced than in Figure 2c. (d) AFM image of fully saturated M13 ssDNA–E. coli SSB complexes associated to lane 7 (scale bars 500 nm). (e) Zoom on a fully saturated M13 ssDNA–E. coli SSB complex (scale bar 100 nm).
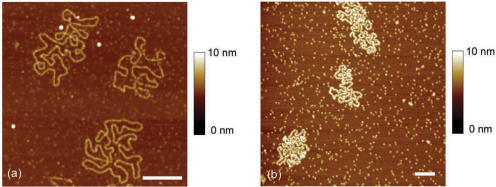
Finally, in order to test our protocol, we extended our investigations to two other SSB proteins : the Bacteriophage T4 gene 32 protein (gp32) and the yeast Replication Protein A (yRPA). For a gp32:ssDNA nucleotide concentrations ratio equal to 1/7, which corresponds to the average number of nucleotides occluded by gp32 (2), M13 ssDNA–gp32 complexes were adsorbed on mica surface by using spermidine. In these conditions, the nucleoprotein filaments are fully saturated as observed by gel shift assay (data not shown). Figure 4a shows properly adsorbed ssDNA–gp32 filaments. AFM images of M13 ssDNA–yRPA nucleofilaments were also obtained with the same success (see Figure 4b), which show that this method can be generalized to many ssDNA–SSB complexes.
Figure 4.
(a) AFM image of M13 ssDNA–gp32 complexes formed in spermidine buffer Tris 20 mM pH 7.5, NaCl 300 mM, SpdCl3 300 µM with R = [(gp32)/(nucleotides)] = 1/7. (b) AFM image of M13 ssDNA–yRPA complexes formed in spermidine buffer Tris 20 mM pH 7.5, NaCl 20 mM, SpdCl3 50 µM with R = [(yRPA)/(nucleotides)] = 1/10 (scale bars 500 nm).
The adsorption of E. coli SSB nucleoprotein filaments on mica over a large range of NaCl concentrations is now possible which makes this method unique to study the E. coli SSB binding to ssDNA at high resolution and its cooperativity. In addition, AFM provides information at the single molecule level whereas typical biochemical techniques indicate only average values obtained from a large number of molecules. The results obtained with other SSB seem to validate this method for many ssDNA–SSB complexes. Its additional benefit lies in its ability to image nucleoprotein filaments under physiological conditions by AFM since strong fixation agents are not required. Future AFM observations should then provide further insights into the structural and dynamical properties of ssDNA–SSB complexes.
ACKNOWLEDGEMENTS
Funding to pay the Open Access publication charge was provided by INSERM.
Conflict of interest statement. None declared.
REFERENCES
- 1.Chase JW, Williams KR. Single-stranded DNA binding proteins required for DNA replication. Annu. Rev. Biochem. 1986;55:103–136. doi: 10.1146/annurev.bi.55.070186.000535. [DOI] [PubMed] [Google Scholar]
- 2.Karpel RL. The Biology of Non-Specific DNA-Protein Interactions. Boca Raton, Florida: CRC Press; 1990. T4 bacteriophage gene 32 protein. In Revzin,A. (ed), pp. 103–130. [Google Scholar]
- 3.Lohman TM, Ferrari ME. Escherichia coli single-stranded DNA-binding protein: multiple DNA-binding modes and cooperativities. Annu. Rev. Biochem. 1994;63:527–570. doi: 10.1146/annurev.bi.63.070194.002523. [DOI] [PubMed] [Google Scholar]
- 4.Wold MS. REPLICATION PROTEIN A: a heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism. Annu. Rev. Biochem. 1997;66:61–92. doi: 10.1146/annurev.biochem.66.1.61. [DOI] [PubMed] [Google Scholar]
- 5.Chrysogelos S, Griffith J. Escherichia coli single-strand binding protein organizes single-stranded DNA in nucleosome-like units. Proc. Natl Acad. Sci. USA. 1982;79:5803–5807. doi: 10.1073/pnas.79.19.5803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Griffith JD, Harris LD, Register J., 3rd. Visualization of SSB-ssDNA complexes active in the assembly of stable RecA-DNA filaments. Cold Spring Harb. Symp. Quant. Biol. 1984;49:553–559. doi: 10.1101/sqb.1984.049.01.062. [DOI] [PubMed] [Google Scholar]
- 7.Syvanen A.-C, Alanen M, Soderlund H. A complex of single-strand binding protein and M13 DNA as hybridization probe. Nucleic Acids Res. 1985;13:2789–2802. doi: 10.1093/nar/13.8.2789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Soengas MS, Gutierrez C, Salas M. Helix-destabilizing activity of [phi]29 single-stranded DNA binding protein: effect on the elongation rate during strand displacement DNA replication. J. Mol. Biol. 1995;253:517–529. doi: 10.1006/jmbi.1995.0570. [DOI] [PubMed] [Google Scholar]
- 9.Lefebvre SD, Wong ML, Morrical SW. Simultaneous interactions of bacteriophage T4 DNA replication proteins gp59 and gp32 with single-stranded (ss) DNA. J. Biol. Chem. 1999;274:22830–22838. doi: 10.1074/jbc.274.32.22830. [DOI] [PubMed] [Google Scholar]
- 10.Hameau L, Jeusset J, Lafosse S, Coulaud D, Delain E, Unge T, Restle T, Le Cam E, Mirambeau G. Human immunodeficiency virus type 1 central DNA flap: dynamic terminal product of plus-strand displacement DNA synthesis catalyzed by reverse transcriptase assisted by nucleocapsid protein. J. Virol. 2001;75:3301–3313. doi: 10.1128/JVI.75.7.3301-3313.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Delius H, Mantell NJ, Alberts B. Characterization by electron microscopy of the complex formed between T4 bacteriophage gene 32-protein and DNA. J. Mol. Biol. 1972;67:341–350. doi: 10.1016/0022-2836(72)90454-8. [DOI] [PubMed] [Google Scholar]
- 12.Bustamante C, Rivetti C. Visualizing protein-nucleic acid interactions on a large scale with the scanning force microscope. Annu. Rev. Biophys. Biomol. Struct. 1996;25:395–429. doi: 10.1146/annurev.bb.25.060196.002143. [DOI] [PubMed] [Google Scholar]
- 13.Hansma HG. Surface biology of DNA by atomic force microscopy. Annu. Rev. Phys. Chem. 2001;52:71–92. doi: 10.1146/annurev.physchem.52.1.71. [DOI] [PubMed] [Google Scholar]
- 14.Sattin BD, Goh MC. Direct observation of the assembly of RecA/DNA complexes by atomic force microscopy, Biophys. J. 2004;87:3430–3436. doi: 10.1529/biophysj.104.045567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lyubchenko YL. DNA structure and dynamics: an atomic force microscopy study. Cell Biochem. Biophys. 2004;41:75–98. doi: 10.1385/CBB:41:1:075. [DOI] [PubMed] [Google Scholar]
- 16.Ristic D, Modesti M, van der Heijden T, van Noort J, Dekker C, Kanaar R, Wyman C. Human Rad51 filaments on double- and single-stranded DNA: correlating regular and irregular forms with recombination function. Nucleic Acids Res. 2005;33:3292–3302. doi: 10.1093/nar/gki640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Piétrement O, Pastré D, Landousy F, David MO, Fusil S, Hamon L, Zozime A, Le Cam E. Studying the effect of a charged surface on the interaction of bleomycin with DNA using an atomic force microscope. Eur. Biophys. J. 2005;34:200–207. doi: 10.1007/s00249-004-0443-y. [DOI] [PubMed] [Google Scholar]
- 18.Shi WX, Larson RG. Atomic force microscopic study of aggregation of RecA-DNA nucleoprotein filaments into left-handed supercoiled bundles. Nano Lett. 2005;5:2476–2481. doi: 10.1021/nl051783v. [DOI] [PubMed] [Google Scholar]
- 19.Hansma HG, Sinsheimer RL, Groppe J, Bruice TC, Elings V, Gurley G, Bezanilla M, Mastrangelo IA, Hough PV, et al. Recent advances in atomic force microscopy of DNA. Scanning. 1993;15:296–299. doi: 10.1002/sca.4950150509. [DOI] [PubMed] [Google Scholar]
- 20.Lyubchenko YL, Jacobs BL, Lindsay SM, Stasiak A. Atomic force microscopy of nucleoprotein complexes. Scanning Microsc. 1995;9:705–724. [PubMed] [Google Scholar]
- 21.Brown TA, Cecconi C, Tkachuk AN, Bustamante C, Clayton DA. Replication of mitochondrial DNA occurs by strand displacement with alternative light-strand origins, not via a strand-coupled mechanism. Genes Dev. 2005;19:2466–2476. doi: 10.1101/gad.1352105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Woolley AT, Kelly RT. Deposition and characterization of extended single-stranded DNA molecules on surfaces. Nano Lett. 2001;1:345–348. [Google Scholar]
- 23.Adamcik J, Klinov DV, Witz G, Sekatskii SK, Dietler G. Observation of single-stranded DNA on mica and highly oriented pyrolytic graphite by atomic force microscopy. FEBS Lett. 2006;580:5671–5675. doi: 10.1016/j.febslet.2006.09.017. [DOI] [PubMed] [Google Scholar]
- 24.Hansma HG, Laney DE. DNA binding to mica correlates with cationic radius: assay by atomic force microscopy. Biophys. J. 1996;70:1933–1939. doi: 10.1016/S0006-3495(96)79757-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rivetti C, Guthold M, Bustamante C. Scanning force microscopy of DNA deposited onto mica: equilibration versus kinetic trapping studied by statistical polymer chain analysis. J. Mol. Biol. 1996;264:919–932. doi: 10.1006/jmbi.1996.0687. [DOI] [PubMed] [Google Scholar]
- 26.Shao Z, Mou J, Czajkowksy DM, Yang J, Yuan JY. Biological atomic force microscopy: what is achieved & what is needed. Adv. Phys. 1996;45:1–86. [Google Scholar]
- 27.Pastré D, Piétrement O, Fusil S, Landousy F, Jeusset J, David MO, Hamon L, Le Cam E, Zozime A. Adsorption of DNA to mica mediated by divalent counterions: a theoretical and experimental study. Biophys. J. 2003;85:2507–2518. doi: 10.1016/s0006-3495(03)74673-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Pastré D, Hamon L, Landousy F, Sorel I, David MO, Zozime A, Le Cam E, Piétrement O. Anionic polyelectrolyte adsorption on mica mediated by multivalent cations: a solution to DNA imaging by atomic force microscopy under high ionic strengths. Langmuir. 2006;22:6651–6660. doi: 10.1021/la053387y. [DOI] [PubMed] [Google Scholar]
- 29.Lohman TM, Overman LB. Two binding modes in Escherichia coli single strand binding protein-single stranded DNA complexes. Modulation by NaCl concentration. J. Biol. Chem. 1985;260:3594–3603. [PubMed] [Google Scholar]
- 30.Kantake N, Sugiyama T, Kolodner RD, Kowalczykowski SC. The recombination-deficient mutant RPA (rfa1-t11) is displaced slowly from single-stranded DNA by Rad51 protein. J. Biol. Chem. 2003;278:23410–23417. doi: 10.1074/jbc.M302995200. [DOI] [PubMed] [Google Scholar]
- 31.Revet B, Fourcade A. Short unligated sticky ends enable the observation of circularised DNA by atomic force and electron microscopes. Nucleic Acids Res. 1998;26:2092–2097. doi: 10.1093/nar/26.9.2092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zobel CR, Beer M. Electron stains. I. chemical studies on the interaction of DNA with uranyl salts. J. Biophys. Biochem. Cytol. 1961;10:335–346. doi: 10.1083/jcb.10.3.335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Tabor CW, Tabor H. Polyamines. Annu. Rev. Biochem. 1984;53:749–790. doi: 10.1146/annurev.bi.53.070184.003533. [DOI] [PubMed] [Google Scholar]
- 34.Ferrari ME, Bujalowski W, Lohman TM. Co-operative binding of Escherichia coli SSB tetramers to single-stranded DNA in the (SSB)35 binding mode. J. Mol. Biol. 1994;236:106–123. doi: 10.1006/jmbi.1994.1122. [DOI] [PubMed] [Google Scholar]
- 35.Overman LB, Bujalowski W, Lohman TM. Equilibrium binding of Escherichia coli single-strand binding protein to single-stranded nucleic acids in the (SSB)65 binding mode. Cation and anion effects and polynucleotide specificity. Biochemistry. 1988;27:456–471. doi: 10.1021/bi00401a067. [DOI] [PubMed] [Google Scholar]
- 36.Wei TF, Bujalowski W, Lohman TM. Cooperative binding of polyamines induces the Escherichia coli single-strand binding protein-DNA binding mode transitions. Biochemistry. 1992;31:6166–6174. doi: 10.1021/bi00141a029. [DOI] [PubMed] [Google Scholar]
- 37.Bujalowski W, Lohman TM. Escherichia coli single-strand binding protein forms multiple, distinct complexes with single-stranded DNA. Biochemistry. 1986;25:7799–7802. doi: 10.1021/bi00372a003. [DOI] [PubMed] [Google Scholar]