Abstract
The tumour suppressor protein ARF provides a defence mechanism against hyperproliferative stresses that can result from the aberrant activation of oncogenes. Accordingly, ARF is silenced or deleted in many human cancers. Activation of ARF can arrest growth and cell cycle progression, or trigger apoptosis. A principle mediator of these effects is p53, which ARF stabilizes by binding and inhibiting MDM2. However, ARF has additional targets and remains able to block growth in the absence of p53, albeit less efficiently. For example, ARF can suppress rRNA production in a p53-independent manner. We have found that the synthesis of tRNA by RNA polymerase III is also inhibited in response to ARF. However, in contrast to its effects on rRNA synthesis, ARF is unable to inhibit tRNA gene transcription when p53 is ablated. These results add to the growing list of cellular changes that can be triggered by ARF induction.
INTRODUCTION
The tumour suppressor ARF is a small nucleolar protein, much of which is encoded by an alternative reading frame within exon 2 of the INK4a locus (1). It is induced by abnormal hyperproliferative stimuli and serves to provide protection against oncogenic stress (2–4). For example, ARF induction can be triggered by deregulation of cellular oncogenes, such as E2F-1, Myc and Ras (5–8). It has been described as a circuit breaker, which monitors the strength of mitogenic signals and allows growth and proliferation under normal circumstances, but not when cells receive inappropriate oncogenic stimuli (2–4). Loss of ARF expression, as occurs in many malignancies, can abrogate this surveillance mechanism and increase cancer susceptibility. Accordingly, mice lacking ARF are highly prone to tumour development (9). For example, Eμ-Myc transgenic mice develop much more rapid and aggressive lympholeukaemias if they lack both ARF alleles (10). Conversely, increased gene dosage of the Ink4a/ARF locus confers cancer resistance to transgenic mice (11).
A key way in which ARF protects against oncogenic stress is by raising p53 levels, which can lead to cell cycle arrest or apoptosis (2,4,12). ARF does this by binding to MDM2, a ubiquitin ligase that catalyses p53 degradation; this interaction sequesters MDM2 and also blocks its ligase activity, thereby stabilizing p53 (13,14). Recent data suggest that tumour suppression by p53 may hinge upon its ARF-dependent ability to respond to oncogenic stress (15,16). In addition, ARF can inhibit cell growth and proliferation in the absence of functional p53, albeit less efficiently (2,17,18). Thus, overexpressing ARF can prevent proliferation of p53-negative mouse embryonic fibroblasts (MEFs), although their arrest is slower than that of wild-type controls (17). Furthermore, mice engineered to lack both p53 and ARF develop cancers at a faster rate than animals lacking only one of these tumour suppressors, clear genetic evidence that ARF has p53-independent functions in vivo (18). Apart from MDM2, a variety of proteins have been reported to interact with ARF (4,19). Furthermore, there is strong evidence that ARF can influence ribosome biogenesis in a p53-indpendent manner (20,21). This may be of fundamental importance in ensuring that growth (accumulation of mass) is inhibited in parallel with cell cycle arrest.
ARF was found to reduce production of 5.8S, 18S and 28S rRNAs by inhibiting the processing of pre-rRNA precursors in a manner independent of p53 and MDM2 (21,22). This activity maps to a region of ARF that is especially well conserved through evolution and is required for it to suppress proliferation (21). Several proteins involved in ribosome biogenesis interact with ARF (20,22). One of these is nucleophosmin (B23), a nucleolar endoribonuclease that is required for rRNA processing and cell cycle progression (20,22,23). ARF can interfere with the function of nucleophosmin and inhibit rRNA maturation (20,22,23). ARF has also been reported to inhibit transcription of rRNA genes by RNA polymerase (pol) I (24). Since cell growth and proliferation depend on rRNA production (25,26), the ability of ARF to disrupt this process may contribute to its role as a tumour suppressor, even in the absence of p53. Indeed, it has been suggested that the primordial role of ARF was to slow ribosome biogenesis in response to hyperproliferative stresses caused by oncogene activation; its subsequent linkage to p53 via MDM2 may have evolved to provide a more efficient checkpoint for coupling ribosome production with p53-dependent inhibitors of the cell cycle (19,21).
In most situations, the production of large rRNAs by pol I is regulated co-ordinately with the production of 5S rRNA and tRNA by pol III (27,28). This is an economical arrangement, since the four rRNAs are required in equimolar amounts within the ribosome and the demand for tRNA must reflect the availability of ribosomes. Although pols I and III use very different sets of transcription factors, they are both targeted directly by several key growth regulators, including c-Myc, Erk, RB and p53 (28). For example, both the pol I-specific factor SL1 and the pol III-specific factor TFIIIB are bound and inactivated by p53 (29–32). ARF can therefore block rRNA accumulation by allowing p53 to inactivate SL1, as well as inhibiting pol I transcription and the processing of pre-rRNA independently of p53 (20,21,24,31). Precedent would suggest that a similar scenario may control the production of pol III products. This study confirms that pol III transcription is indeed inhibited in response to ARF. However, in contrast to the pol I system, we find no evidence for a p53-independent effect of ARF in this case.
MATERIALS AND METHODS
Cell lines and culture
F9 cells were cultured at 37°C in DMEM supplemented with 10% FCS. NARF2 cells, derived from the human osteosarcoma cell line U2OS and containing a stably integrated p14ARF gene under the control of IPTG (33), were cultured at 37°C in DMEM supplemented with 10% FCS, G418 at a concentration of 300 μg/ml and hygromycin at 150 μg/ml. E6-expressing NARF2 cells were cultured in the same way but with puromycin at a concentration of 1.5 μg/ml. Expression of p14ARF was induced by the addition of IPTG at a concentration of 1 mM.
Transient transfections
F9 cells were transiently transfected with pcDNA3HA-E2F1, an expression vector encoding full-length human E2F1, or pcDNA3 as a control. Transfection into F9 cells was achieved using superfect (Qiagen, Valencia, CA, USA) and cells were cultured for 24 h before harvesting. NARF2 cells were transfected with pSUPER-p53 or empty pSUPER control (34) using the Nucleofector system (Amaxa Biosystems, Gaithersburg, MD, USA), according to the manufacturer's instructions for U2OS cells. Cells were harvested after 48 h and RNA was extracted using TRI reagent (Sigma, St. Louis, MO, USA), according to the manufacturer's instructions.
Gene expression assays
Primer extension reactions were carried out as previously (35) using VA1-specific (5′-CACGCGGGCGGTAACCGCATG-3′) and GFP-specific (5′-CGTCGCCGTCCAGCTCGACCAG-3′) labelled primers.
RT-PCR analysis was performed as described previously (36). For p21 mRNA, primers were 5′-CACCTCCTCATGTACATACC-3′ and 5′-CAGGTCTGAGTGTCCAGGAA-3′ and the cycling parameters were: 95°C for 2 min, then 25 cycles of 95°C for 1 min, 58°C for 30 s and 72°C for 1 min, followed by 72°C for 3 min.
Antibodies and western blotting
Whole cell extracts were prepared as previously by a freeze–thaw protocol (37) and protein concentration determined by the Bradford assay. Western immunoblot analysis was performed as previously described (37) using anti-p53 antibodies DO-1 (Pharmingen, San Diego, CA, USA) and 1C12 (Cell Signaling Technology, Beverley, MA, USA), anti-p19 ARF antibody ab80 (Abcam, Cambridge, UK), anti-p14 ARF antibodies FL-132 and C-18, anti-p21 antibody C19, anti-E2F1 antibody KH95, anti-actin antibody C11 (all Santa Cruz Biotechnology, Santa Cruz, CA, USA) and anti-Brf1 antibody 330 (38).
RESULTS
Oncogenic stress caused by ectopic E2F1 overexpression results in repression of pol III transcription
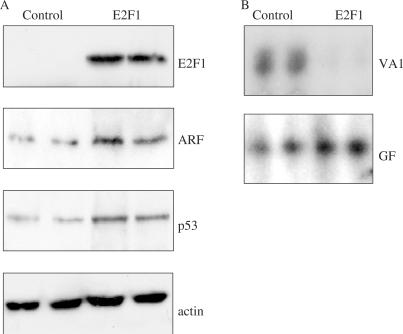
E2F1 has been shown to induce expression of ARF and thereby stabilize p53 (7,8,39). To test whether pol III transcription is regulated under these conditions, F9 cells were transfected with a plasmid encoding human E2F1, along with the VA1 gene as a pol III reporter and GFP as an internal control. Cells were harvested after 24 h and production of RNA from both VA1 and GFP reporters was assessed by quantitative primer extension. Western immunoblotting demonstrated that exogenous expression of E2F1 resulted in the induction of ARF and stabilization of p53 (Figure 1A). It also caused dramatic inhibition of the VA1 pol III reporter, whereas little or no effect was observed on pol II-dependent expression of the GFP control (Figure 1B). These data demonstrate that an oncogenic stress which induces ARF can trigger repression of a pol III reporter.
Figure 1.
Overexpression of E2F1 can induce ARF and p53 and inhibit pol III transcription. F9 cells were co-transfected in duplicate with pVA1 (1 μg), pGST (1 μg) and 15 μg of empty vector (control) or vector encoding full-length E2F1, as indicated. RNA and protein were isolated 48 h after transfection. (A) Western blot of exogenous E2F1 using anti-E2F1 (human) antibody KH95 and of endogenous p19ARF using anti-p19ARF antibody ab80, p53 using anti-p53 antibody 1C12 and actin, to confirm equal loading of protein, using anti-actin antibody C11. (B) Primer extension analysis of extracted RNA using primers specific for VA1 (upper panel), and GFP (lower panel).
Induction of ARF results in repression of pol III transcription
To test directly if pol III transcription responds to ARF, we used the NARF2 cell line, a derivative of U2OS osteosarcoma cells carrying an IPTG-inducible ARF gene (33). Immunoblotting revealed a substantial increase in expression of ARF after 24 h of IPTG treatment (Figure 2A). As expected, this was accompanied by an elevated level of p53, reflecting its stabilization by ARF (33). Functional activation of p53 was suggested by the induction of p21, a well-characterized p53 target (Figure 2A). This was supported by cell cycle analysis, which showed G1 arrest when ARF was induced (data not shown).
Figure 2.
Induction of ARF in NARF2 cells leads to stabilization of p53 and repression of pol III transcription. (A) NARF2 cells were treated with IPTG (1 mM) to induce ARF expression 24 h prior to harvesting. Western blot of extracted protein using anti-p14ARF antibody FL-132, anti-p53 antibody DO-1, anti-p21 antibody C19 and anti-actin antibody C11, as indicated. (B) RT-PCR analysis was performed on cDNAs generated by reverse transcription of total extracted RNA using primers specific for tRNA and ARPP P0 mRNA, as indicated. (C) Parental U2OS cells were treated with 1mM IPTG for 24 h prior to harvesting and RT-PCR analysis was performed using primers specific for tRNA and ARPP P0 mRNA, as indicated.
RT-PCR analysis of RNA extracted from these NARF2 cells revealed a marked decrease in tRNA expression following ARF induction (Figure 2B). These assays used intron-specific primers to detect primary tRNA transcripts and are considered to provide a measure of ongoing pol III transcription, because the primary products are processed very rapidly into mature tRNA (36). However, the possibility that ARF could affect tRNA processing has not been excluded. The observed effect of ARF induction on levels of pre-tRNA was specific, since expression of a pol II-dependent control transcript encoding acidic ribosomal phosphoprotein P0 (ARPP P0) remained undiminished. Because treatment of parental U2OS cells with IPTG did not influence pre-tRNA expression (Figure 2C), we believe that the response seen in NARF2 cells reflects the presence of the IPTG-inducible ARF gene.
A time course was conducted to compare the repression of pol III output with the induction of ARF and p53. Immunoblotting confirmed that ARF levels become elevated within 9 h of IPTG addition and remain high at all subsequent time points (Figure 3A). Induction of p53 was detected at the earliest time points, before maximal levels were reached by 9 h. One can infer from the p53 response that ARF is also induced at the earlier time points, but ARF is less readily detected than p53 due to inferior antibodies. The data suggest that induction of ARF stabilizes p53, as predicted. Analysis of tRNA expression revealed clear repression 3 h after IPTG addition, which was coincident with the accumulation of p53 in these cells (Figure 3B). Expression of the p53-responsive p21 gene increased following ARF induction, consistent with the stabilization of p53. However, activation of p21 was less dramatic than the pol III response and also slower to appear, only becoming evident at the 6-h time point. Thus, ARF can trigger a pol III response, which is faster and stronger than the well-documented induction of a paradigm p53 target gene. The rapidity with which pol III transcription is repressed indicates that regulation is likely to be direct, whereas an indirect response to cell cycle changes would follow p21 induction.
Figure 3.
Time course of ARF induction and downstream events. NARF2 cells were treated with IPTG (1 mM) and RNA and protein harvested at the time points indicated. (A) Western blot using anti-p14 antibody FL-132 (upper panel), anti-p53 antibody DO-1 (middle panel) and anti-actin antibody C11 (lower panel). (B) RT-PCR analysis using primers specific for tRNA (upper panel), p21 mRNA (middle panel) and ARPP P0 mRNA (bottom panel).
Repression of pol III transcription by ARF is dependent on p53
We used RNAi to test if the repression of tRNA gene expression in response to ARF is dependent on p53. NARF2 cells were transfected with pSUPER-p53, which encodes an shRNA against p53, or the empty pSUPER vector as control (34). Western blotting confirmed that p53 levels were reduced specifically by the p53 shRNA, and that this remained the case following ARF induction with IPTG (Figure 4A). This partial knock-down of p53 caused a substantial elevation in tRNA expression, consistent with the role of p53 in restraining pol III transcription (Figure 4B). Furthermore, it prevented the repression of tRNA expression that otherwise accompanies ARF induction in these cells. Thus, the level of tRNA is no longer diminished in response to ARF when p53 has been depleted (compare lanes 2 and 4).
Figure 4.
ARF-mediated repression of pol III-transcript expression is blocked by RNAi of p53. NARF2 cells were transfected with empty vector (lanes 1 and 3) or vector encoding p53-specific shRNA (lanes 2 and 4). IPTG was added 24 h later (lanes 3 and 4) and cells were harvested after a further 24 h. (A) Western blot of extracted protein using anti-p53 antibody DO-1 (upper panel), anti-p14 antibody FL-132 (middle panel) and anti-actin antibody C11 (lower panel). (B) RT-PCR analysis performed on cDNAs prepared from total extracted RNA using primers specific for tRNA (top panel), and ARPP P0 mRNA (bottom panel).
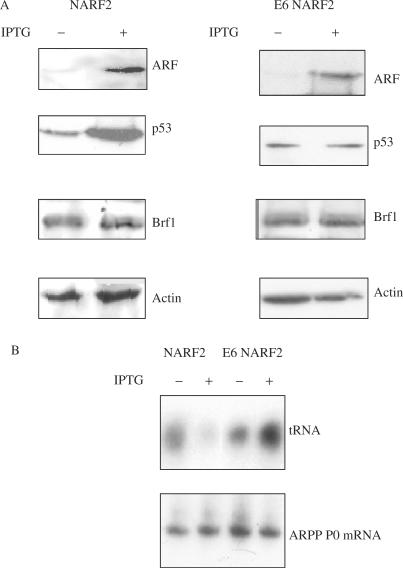
E6 NARF2 is a derivative of the NARF2 cell line that produces the E6 product of HPV16, an oncoprotein, which targets p53 for degradation (33). When ARF is induced in E6 NARF2 cells, p53 levels do not increase (Figure 5A) and tRNA expression is not repressed (Figure 5B). These data support the inference from RNAi (Figure 4) that ARF-mediated repression of pol III transcription requires induction of p53.
Figure 5.
ARF-mediated repression of pol III-transcript expression is blocked by HPV16 E6. Matched NARF2 and E6 NARF2 cells were cultured for 24 h with or without IPTG, as indicated. (A) Western blot analysis using anti-ARF antibody FL-132, anti-p53 antibody DO-1, anti-Brf1 antibody 330 and anti-actin antibody C11, as indicated. (B) RT-PCR analysis using primers specific for tRNA and ARPP P0 mRNA, as indicated.
DISCUSSION
This study establishes that pol III transcription is repressed in response to oncogenic stress caused by overexpression of E2F1. The ARF gene promoter is bound and activated by E2F1 (7,39) and we have shown directly that induction of ARF can trigger a potent repression of pol III transcription. This response is lost if p53 is neutralized using RNAi or the E6 viral oncoprotein. The data suggest that the ARF/p53 pathway can suppress pol III activity as part of the protective response to hyperproliferative stresses induced by aberrant oncogene activation. This may help limit the growth capacity of cells.
Even in the absence of p53, ARF is able to inhibit production of mature rRNA, an effect, which will undoubtedly have a major impact on ribosome accumulation and hence cell growth and proliferation (20,21,24). Since pol III output is frequently coordinated with the production of large rRNA (27,28), we wondered if ARF might also have a direct effect on the pol III system. However, our data argue that this is not the case and that the impact of ARF on pol III transcription is mediated through p53, at least in U2OS cells. This conclusion is consistent with the fact that p53-knockout fibroblasts display highly elevated levels of ARF (40) and yet also transcribe tRNA and 5S rRNA genes much more actively than matched wild-type cells (30). Although the possibility remains of a more direct effect under other circumstances, the available evidence indicates that ARF needs p53 in order to restrain pol III transcription.
The abundance and activity of p53 can be raised by a variety of different stresses, including exposure to DNA-damaging agents, hypoxia, ribonucleotide depletion and inappropriate growth signals resulting from aberrant oncogene activation (41–46). Under unstressed conditions, the level of p53 is kept low by proteolytic degradation, but stress causes p53 to accumulate rapidly, primarily due to its stabilization (41–46). It also triggers a complex series of modifications, which can determine the ability of p53 to bind and regulate its many targets (41–46). The nature of the response pathways and the resultant modifications can vary dramatically between different cell types and distinct kinds of stress (45,46). A good example is provided by serine 15 of p53, which is phosphorylated in response to DNA damage, but not in response to oncogenic stress (47). Because of these variations in the mode of activation, the sets of genes controlled by p53 depend upon the particular stress conditions. For example, DNA damage causes p53 to induce the p21 gene, but the same gene shows little or no response to p53 activation under hypoxic conditions (48). Although previous studies had established that pol III transcription can respond strongly to p53 in vitro or in vivo, they did not determine the physiological circumstances under which this control becomes most apparent. Indeed, artificial induction of p53 in TR9-7 cells from a tetracycline-dependent promoter had minimal effect on pol III activity (49). Furthermore, UV treatment of MCF-7 cells was found to induce endogenous p53, but again caused little or no repression of pol III transcription (50). UV-induced acetylation of p53 in its C-terminal domain encouraged its binding to the p21 gene but excluded it from pol III-transcribed genes, when tested in parallel (50). This striking result emphasizes the complexity of the p53 response and how it can vary between different genes and stimuli. A more robust effect on pol III transcription may occur in other cell types or with other types of DNA damage. Alternatively, the repression of pol III may not be a significant part of the protective repertoire of p53 under these conditions. Our data suggest that the ability of p53 to control pol III transcription is instead more important when a cell induces ARF. Indeed, under these circumstances we find a robust pol III response that is faster and more potent than the induction of p21, a paradigm direct target of p53. Repression of pol III transcription may therefore form part of the protective response to hyperproliferative signals triggered by aberrant oncogene activation. It is noteworthy that recent evidence suggests that the tumour suppression function of p53 may be mediated primarily by its ability to protect against oncogenic stress, whereas its responsiveness to DNA damage may be much less important (15,16).
It has been reported that p53 induction in TR9-7 cells causes selective degradation of the TFIIIB subunit Brf1 (49). We looked for a similar effect in NARF2 cells, but found no change in Brf1 levels following induction of ARF and the consequent stabilization of p53 (Figure 5A). Brf1 levels were also unchanged in response to p53 induction in H1299 cells (32). We therefore do not believe that destabilization of Brf1 is a general feature of the pol III response to p53.
In addition to E2F1, other proto-oncogenes including Ras and c-Myc can induce ARF expression and might therefore be expected to down-regulate pol III output (5). This seems contrary to the published evidence that Ras and c-Myc can stimulate pol III transcription robustly in a variety of cell lines (51,52). There is also evidence that ARF can directly antagonize the ability of c-Myc to activate a subset of its transcriptional targets (19,40,53) An explanation is suggested by the fact that most cell lines are compromised in the function of ARF and/or p53 and have therefore lost their protection against pol III activation by c-Myc or Ras (33). For example, the U2OS cells used in this study are ARF-negative, providing a null background to study the effects of artificial ARF induction (33). Indeed, loss of either ARF or p53 is essential for cell immortalization by c-Myc (5,10). When intact, the ARF-p53 pathway provides a protective safeguard against deregulated oncogenes, inhibiting a variety of processes that impact on growth and cell cycle progression, including pol III transcription. When either ARF or p53 are lost during cell immortalization, a restraint on pol III output is removed and Ras or c-Myc become free to activate the system fully.
Inappropriate hyperproliferative signals cause ARF to initiate mechanisms that inhibit both cell cycle progression and the accumulation of mass (i.e. growth). Coordination of these processes is necessary to maintain cell size. ARF's ability to restrain the production of rRNA and hence ribosomes provides a potent means of growth control. This pathway is bolstered by its ability to trigger p53-dependent repression of pol III transcription. Such an arrangement has clear benefits in terms of metabolic economy, as it will prevent the production of excess tRNA and 5S rRNA that is surplus to requirements when ribosome production is limited. ARF may be added to the growing list of oncogenes and tumour suppressors that, either directly or indirectly, can impinge upon the pol III-transcription machinery.
ACKNOWLEDGEMENTS
The authors are grateful to Gordon Peters for NARF2 and E6-NARF2 cell lines. This work was funded by the Wellcome Trust and Cancer Research UK. Funding to pay the Open Access publication charges for this article was provided by the Wellcome Trust.
Conflict of interest statement. None declared.
REFERENCES
- 1.Quelle D.E., Zindy F., Ashmun R.A., Sherr C.J. Alternative reading frames of the INK4a tumour suppressor gene encode two unrelated proteins capable of inducing cell cycle arrest. Cell. 1995;83:993–100. doi: 10.1016/0092-8674(95)90214-7. [DOI] [PubMed] [Google Scholar]
- 2.Lowe S.W., Sherr C.J. Tumour suppression by Ink4a-Arf: progress and puzzles. Curr. Opin. Gen. Dev. 2003;13:77–83. doi: 10.1016/s0959-437x(02)00013-8. [DOI] [PubMed] [Google Scholar]
- 3.Kim W.Y., Sharpless N.E. The regulation of INK4/ARF in cancer and aging. Cell. 2006;127:265–275. doi: 10.1016/j.cell.2006.10.003. [DOI] [PubMed] [Google Scholar]
- 4.Sherr C.J. Divorcing ARF and p53: an unsettled case. Nat. Rev. Cancer. 2006;6:663–672. doi: 10.1038/nrc1954. [DOI] [PubMed] [Google Scholar]
- 5.Zindy F., Eischen C.M., Randle D.H., Kamijo T., Cleveland J.L., Sherr C.J., Roussel M.F. Myc signaling via the ARF tumour suppressor regulates p53-dependent apoptosis and immortalization. Genes Dev. 1998;12:2424–2433. doi: 10.1101/gad.12.15.2424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Palmero I., Pantoja C., Serrano M. p19ARF links the tumour suppressor p53 to ras. Nature. 1998;395:125–126. doi: 10.1038/25870. [DOI] [PubMed] [Google Scholar]
- 7.Bates S., Phillips A.C., Clark P.A., Stott F., Peters G., Ludwig R.L., Vousden K.H. p14ARF links the tumour suppressors RB and p53. Nature. 1998;395:124–125. doi: 10.1038/25867. [DOI] [PubMed] [Google Scholar]
- 8.Lomazzi M., Moroni M.C., Jensen M.R., Frittoli E., Helin K. Suppression of the p53- or pRB-mediated G1 checkpoint is required for E2F-induced S-phase entry. Nat. Genet. 2002;31:190–194. doi: 10.1038/ng891. [DOI] [PubMed] [Google Scholar]
- 9.Kamijo T., Zindy F., Roussel M.F., Quelle D.E., Downing J.R., Ashmun R.A., Grosveld G., Sherr C.J. Tumour suppression at the mouse INK4a locus mediated by the alternative reading frame product p19ARF. Cell. 1997;91:649–659. doi: 10.1016/s0092-8674(00)80452-3. [DOI] [PubMed] [Google Scholar]
- 10.Eischen C.M., Weber J.D., Roussel M.F., Sherr C.J., Cleveland J.L. Disruption of the ARF-Mdm2-p53 tumor suppressor pathway in Myc-induced lymphomagenesis. Genes Dev. 1999;13:2658–2669. doi: 10.1101/gad.13.20.2658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Matheu A., Pantoja C., Efeyan A., Criado L.M., Martin-Caballero J., Flores J.M., Klatt P., Serrano M. Increased gene dosage of Ink4a/Arf results in cancer resistance and normal aging. Genes Dev. 2004;18:2736–2746. doi: 10.1101/gad.310304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Sherr C.J. The INK4a/ARF network in tumour suppression. Nature Revs. Mol. Cell Biol. 2001;2:731–737. doi: 10.1038/35096061. [DOI] [PubMed] [Google Scholar]
- 13.Weber J.D., Taylor L.J., Roussel M.F., Sherr C.J., Bar-Sagi D. Nucleolar Arf sequesters Mdm2 and activates p53. Nature Cell Biol. 1999;1:20–26. doi: 10.1038/8991. [DOI] [PubMed] [Google Scholar]
- 14.Honda R., Yasuda H. Association of p19ARF with Mdm2 inhibits ubiquitin ligase activity of Mdm2 for tumour suppressor p53. EMBO J. 1999;18:22–27. doi: 10.1093/emboj/18.1.22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Christophorou M.A., Ringshausen I., Finch A.J., Swigart L.B., Evan G.I. The pathological response to DNA damage does not contribute to p53-mediated tumour suppression. Nature. 2006;443:214–217. doi: 10.1038/nature05077. [DOI] [PubMed] [Google Scholar]
- 16.Efeyan A., Garcia-Cao I., Herranz D., Velasco-Miguel S., Serrano M. Policing of oncogene activity by p53. Nature. 2006;443:159. doi: 10.1038/443159a. [DOI] [PubMed] [Google Scholar]
- 17.Carnero A., Hudson J.D., Price C.M., Beach D.H. p16INK4A and p19ARF act in overlapping pathways in cellular immortalization. Nat. Cell Biol. 2000;2:148–155. doi: 10.1038/35004020. [DOI] [PubMed] [Google Scholar]
- 18.Weber J.D., Jeffers J.R., Rehg J.E., Randle D.H., Lozano G., Roussel M.F., Sherr C.J., Zambetti G.P. p53-independent functions of the p19ARF tumor suppressor. Genes Dev. 2000;14:2358–2365. doi: 10.1101/gad.827300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Cleveland J.L., Sherr C.J. Antagonism of Myc functions by Arf. Cancer Cell. 2004;6:309–311. doi: 10.1016/j.ccr.2004.09.020. [DOI] [PubMed] [Google Scholar]
- 20.Itahana K., Bhat K.P., Jin A., Itahana Y., Hawke D., Kobayashi R., Zhang Y. Tumor suppressor ARF degrades B23, a nucleolar protein involved in ribosome biogenesis and cell proliferation. Mol. Cell. 2003;12:1151–1164. doi: 10.1016/s1097-2765(03)00431-3. [DOI] [PubMed] [Google Scholar]
- 21.Sugimoto M., Kuo M.-L., Roussel M.F., Sherr C.J. Nucleolar Arf tumour suppressor inhibits ribosomal RNA processing. Mol. Cell. 2003;11:415–424. doi: 10.1016/s1097-2765(03)00057-1. [DOI] [PubMed] [Google Scholar]
- 22.Bertwistle D., Sugimoto M., Sherr C.J. Physical and functional interactions of the Arf tumor suppressor protein with nucleophosmin/B23. Mol. Cell Biol. 2004;24: 985–996. doi: 10.1128/MCB.24.3.985-996.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Brady S.N., Yu Y., Maggi L.B., Weber J.D. ARF impedes NPM/B23 shuttling in an Mdm2-sensitive tumor suppressor pathway. Mol. Cell Biol. 2004;24:9327–9338. doi: 10.1128/MCB.24.21.9327-9338.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ayrault O., Andrique L., Fauvin D., Eymin B., Gazzeri S., Seite P. Human tumor suppressor p14ARF negatively regulates rRNA transcription and inhibits UBF1 transcription factor phosphorylation. Oncogene. 2006;25:7577–7586. doi: 10.1038/sj.onc.1209743. [DOI] [PubMed] [Google Scholar]
- 25.Brandenburger Y., Jenkins A., Autelitano D.J., Hannan R.D. Increased expression of UBF is a critical determinant for rRNA synthesis and hypertrophic growth of cardiac myocytes. FASEB J. 2001;15:2051–2053. doi: 10.1096/fj.01-0853fje. [DOI] [PubMed] [Google Scholar]
- 26.Zhao J., Yuan X., Frodin M., Grummt I. ERK-dependent phosphorylation of the transcription initiation factor TIF-IA is required for RNA polymerase I transcription and cell growth. Mol. Cell. 2003;11:405–413. doi: 10.1016/s1097-2765(03)00036-4. [DOI] [PubMed] [Google Scholar]
- 27.White R.J. rRNA and tRNA production is closely tied to cell growth. In: Hall M., Raff M., Thomas G., editors. 2003. Cell Growth Control. Cold Spring Harbor Laboratory Press, Cold Spring Harbor. [Google Scholar]
- 28.White R.J. RNA polymerases I and III, growth control and cancer. Nat. Rev. Mol. Cell Biol. 2005;6:69–78. doi: 10.1038/nrm1551. [DOI] [PubMed] [Google Scholar]
- 29.Chesnokov I., Chu W.-M., Botchan M.R., Schmid C.W. p53 inhibits RNA polymerase III-directed transcription in a promoter-dependent manner. Mol. Cell Biol. 1996;16:7084–7088. doi: 10.1128/mcb.16.12.7084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Cairns C.A., White R.J. p53 is a general repressor of RNA polymerase III transcription. EMBO J. 1998;17:3112–3123. doi: 10.1093/emboj/17.11.3112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zhai W., Comai L. Repression of RNA polymerase I transcription by the tumour suppressor p53. Mol. Cell Biol. 2000;20:5930–5938. doi: 10.1128/mcb.20.16.5930-5938.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Crighton D., Woiwode A., Zhang C., Mandavia N., Morton J.P., Warnock L.J., Milner J., White R.J., Johnson D.L. p53 represses RNA polymerase III transcription by targeting TBP and inhibiting promoter occupancy by TFIIIB. EMBO J. 2003;22:2810–2820. doi: 10.1093/emboj/cdg265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Stott F.J., Bates S., James M.C., McConnell B.B., Starborg M., Brookes S., Palmero I., Ryan K., Hara E., et al. The alternative product from the human CDKN2A locus, p14Arf, participates in a regulatory feedback loop with p53 and MDM2. EMBO J. 1998;17:5001–5014. doi: 10.1093/emboj/17.17.5001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Brummelkamp T.R., Bernards R., Agami R. A system for stable expression of short interfering RNAs in mammalian cells. Science. 2002;296:550–553. doi: 10.1126/science.1068999. [DOI] [PubMed] [Google Scholar]
- 35.White R.J., Trouche D., Martin K., Jackson S.P., Kouzarides T. Repression of RNA polymerase III transcription by the retinoblastoma protein. Nature. 1996;382:88–90. doi: 10.1038/382088a0. [DOI] [PubMed] [Google Scholar]
- 36.Winter A.G., Sourvinos G., Allison S.J., Tosh K., Scott P.H., Spandidos D.A., White R.J. RNA polymerase III transcription factor TFIIIC2 is overexpressed in ovarian tumours. Proc. Natl Acad. Sci. USA. 2000;97:12619–12624. doi: 10.1073/pnas.230224097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.White R.J., Gottlieb T.M., Downes C.S., Jackson S.P. Mitotic regulation of a TATA-binding-protein-containing complex. Mol. Cell Biol. 1995;15:1983–1992. doi: 10.1128/mcb.15.4.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Sutcliffe J.E., Brown T.R.P., Allison S.J., Scott P.H., White R.J. Retinoblastoma protein disrupts interactions required for RNA polymerase III transcription. Mol. Cell Biol. 2000;20:9192–9202. doi: 10.1128/mcb.20.24.9192-9202.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Aslanian A., Iaquinta P.J., Verona R., Lees J.A. Repression of the Arf tumour suppressor by E2F3 is required for normal cell cycle kinetics. Genes Dev. 2004;18: 1413–1422. doi: 10.1101/gad.1196704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Qi Y., Gregory M.A., Li Z., Brousal J.P., West K., Hann S.R. p19ARF directly and differentially controls the functions of c-Myc independently of p53. Nature. 2004;431:712–717. doi: 10.1038/nature02958. [DOI] [PubMed] [Google Scholar]
- 41.Giaccia A.J., Kastan M.B. The complexity of p53 modulation: emerging patterns from divergent signals. Genes Dev. 1998;12:2973–2983. doi: 10.1101/gad.12.19.2973. [DOI] [PubMed] [Google Scholar]
- 42.Meek D.W. Multisite phosphorylation and the integration of stress signals at p53. Cell Signal. 1998;10:159–166. doi: 10.1016/s0898-6568(97)00119-8. [DOI] [PubMed] [Google Scholar]
- 43.Prives C. Signaling to p53: breaking the MDM2-p53 circuit. Cell. 1998;95:5–8. doi: 10.1016/s0092-8674(00)81774-2. [DOI] [PubMed] [Google Scholar]
- 44.Oren M. Regulation of the p53 tumor suppressor protein. J. Biol. Chem. 1999;274:36031–36034. doi: 10.1074/jbc.274.51.36031. [DOI] [PubMed] [Google Scholar]
- 45.Vousden K.H., Lu X. Live or let die: the cell's response to p53. Nature Rev. Cancer. 2002;2:594–604. doi: 10.1038/nrc864. [DOI] [PubMed] [Google Scholar]
- 46.Lu X. p53: a heavily dictated dictator of life and death. Curr. Opin. Gen. Dev. 2005;15:27–33. doi: 10.1016/j.gde.2004.12.008. [DOI] [PubMed] [Google Scholar]
- 47.de Stanchina E., McCurrach M.E., Zindy F., Shieh S.-Y., Ferbeyte G., Samulsen A.V., Prives C., Roussel M.F., Sherr C.J., et al. E1A signaling to p53 involves the p19ARF tumor suppressor. Genes Dev. 1998;12:2434–2442. doi: 10.1101/gad.12.15.2434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Ashcroft M., Taya Y., Vousden K.H. Stress signals utilize multiple pathways to stabilize p53. Mol. Cell Biol. 2000;20:3224–3233. doi: 10.1128/mcb.20.9.3224-3233.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Eichhorn K., Jackson S.P. A role for TAF3B2 in the repression of human RNA polymerase III transcription in nonproliferating cells. J. Biol. Chem. 2001;276:21158–21165. doi: 10.1074/jbc.M102295200. [DOI] [PubMed] [Google Scholar]
- 50.Gridasova A.A., Henry R.W. The p53 tumour suppressor protein represses human snRNA gene transcription by RNA polymerases II and III independently of sequence-specific DNA binding. Mol. Cell. Biol. 2005;25:3247–3260. doi: 10.1128/MCB.25.8.3247-3260.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Felton-Edkins Z.A., Fairley J.A., Graham E.L., Johnston I.M., White R.J., Scott P.H. The mitogen-activated protein (MAP) kinase ERK induces tRNA synthesis by phosphorylating TFIIIB. EMBO J. 2003;22:2422–2432. doi: 10.1093/emboj/cdg240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Gomez-Roman N., Grandori C., Eisenman R.N., White R.J. Direct activation of RNA polymerase III transcription by c-Myc. Nature. 2003;421:290–294. doi: 10.1038/nature01327. [DOI] [PubMed] [Google Scholar]
- 53.Datta A., Nag A., Pan W., Hay N., Gartel A.L., Colamonici O., Mori Y., Raychaudhuri P. Myc-ARF (alternative reading frame) interaction inhibits the functions of Myc. J. Biol. Chem. 2004;279:36698–36707. doi: 10.1074/jbc.M312305200. [DOI] [PubMed] [Google Scholar]