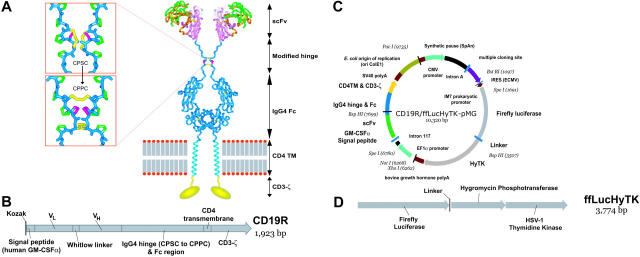
Figure 1.
Schematics of CD19-specific chimeric immunoreceptor and DNA plasmid coexpressing CD19R and trifunctional reporter gene. (A) Ribbon-model of the CD19-specific chimeric immunoreceptor (CD19R) shown dimerized on the cell surface. CD19R is composed of murine scFv (coupled using the Whitlow linker71), a modified serine → proline (CPSC changed to CPPC,72 single-letter amino acid code) human Fc region, human CD4 transmembrane (TM) region, and human CD3-ζ domain. Coloring: pink = VH; green = VL; orange = CDR loops; white = Whitlow linker (GSTSGSGKPGSGEGSTKG); dark blue = hinge/Fc; light blue = transmembrane helix; yellow = CD3-ζ chain (schematic); red/gray = lipid bilayer (schematic); yellow sticks = disulfides; magenta = second proline in CPPC sequence. Boxes show stick models of CPSC to CPPC amino acid change in the modified IgG4 hinge region. Computer modeling demonstrates that substitution of serine with proline at position 241 (Kabat numbering system)73 in the IgG4 hinge region creates a molecule predisposed to interchain disulfide bridging, whereas the native hinge is predisposed to intrachain disulfide bond formation. In the top panel the flexible serine in the sequence CPSC allows intrachain disulfides to form, thus preventing covalent bonding of heavy chain pairs. In the bottom panel the rigid proline in CPPC prevents the cysteines in the mutant IgG4 hinge from forming intrachain bonds, thus favoring covalent bonding of heavy chain pairs. Coloring: dark blue = backbone atoms; green = proline side chains; magenta = serine wild-type (top panel) or proline mutation (bottom panel); yellow = disulfide-bonded cysteines. (B) Schematic of fusion gene of CD19R showing component parts. (C) Schematic of DNA plasmid CD19R/ffLucHyTK-pMG to expresses CD19R gene under control of human elongation factor (EF) 1α promoter and ffLucHyTK gene under control of the human cytomegalovirus (CMV) major immediate early promoter. This plasmid is similar to the clinical vector CD19R/HyTK-pMG, with the exception in the clinical vector the IRES element was deleted, intron A was reduced, and a 20-base pair sequence was deleted 5′ to the IM7 prokaryotic promoter. Selected restriction enzyme sites are shown. (D) Schematic of fusion gene composed of firefly luciferase linked via amino acid (linker = QLISGANGV) to hygromycin phosphotransferase and thymidine kinase (predicted molecular weight of ∼137 kDa).