Abstract
Background
More than half of the approximately 500,000 women diagnosed with cervical cancer worldwide each year will die from this disease. Investigation of genes expressed in precancer lesions compared to those expressed in normal cervical epithelium will yield insight into the early stages of disease. As such, establishing a baseline from which to compare to, is critical in elucidating the abnormal biology of disease. In this study we examine the normal cervical tissue transcriptome and investigate the similarities and differences in relation to CIN III by Long-SAGE (L-SAGE).
Results
We have sequenced 691,390 tags from four L-SAGE libraries increasing the existing gene expression data on cervical tissue by 20 fold. One-hundred and eighteen unique tags were highly expressed in normal cervical tissue and 107 of them mapped to unique genes, most belong to the ribosomal, calcium-binding and keratinizing gene families. We assessed these genes for aberrant expression in CIN III and five genes showed altered expression. In addition, we have identified twelve unique HPV 16 SAGE tags in the CIN III libraries absent in the normal libraries.
Conclusion
Establishing a baseline of gene expression in normal cervical tissue is key for identifying changes in cancer. We demonstrate the utility of this baseline data by identifying genes with aberrant expression in CIN III when compared to normal tissue.
Background
Approximately 500,000 women are diagnosed with cervical cancer worldwide each year and more than half of them will die from this disease [1]. The highest incidence rates are observed in developing countries where it is the second most prevalent cancer in women and remains a leading cause of cancer related death [1]. Widely implemented screening programs have been responsible for the much lower incidence and mortality rates seen in the developed world. Present day screening methods primarily identify precancer lesions termed cervical intraepithelial neoplasia (CIN). CIN lesions are classified into three subgroups, CIN I, CIN II and CIN III, corresponding to mild, moderate and severe dysplasia/carcinoma in situ (CIS), respectively. CIN III lesions have a high likelihood of progression to invasive disease if left untreated [2]. Human Papillomavirus (HPV) has long been established as a necessary but not sufficient cause for cervical carcinoma development. HPV is detected in 99% of invasive disease, 94% of CIN lesions and 46% of normal cervical epithelium [2]. The high risk strains HPV 16 and HPV 18 are most prevalent in invasive disease.
A comprehensive characterization of gene expression of the normal cervical tissue is critical to establish a baseline for comparison against transcriptomes of precancer and cancer. A recent report described the global expression of genes in cervical epithelium using a serial analysis of gene expression (SAGE) based method, enumerating 30,418 sequence tags generated from one normal uterine ectocervical tissue [3]. Another study compared cDNA microarray profiles of cervical tissue to exfoliated cervical cells used in cytology-based cancer screening [4].
In this study, we increased the depth of our understanding of the normal cervical transcriptome and identified gene expression changes in CINIII. We achieved this (i) by using an unbiased Long SAGE (L-SAGE) approach to improve the accuracy of tag-to-gene mapping [5-7], and (ii) by examining 691,390 L-SAGE tags thus increasing publicly available cervical SAGE data by greater than 20 fold.
Results
In this study, we sequenced 691,390 SAGE tags from four libraries. Cervical L-SAGE libraries N1, N2, C1, and C2 were sequenced to 165,624, 181,224, 173,534, and 171,008 tags, respectively. Duplicate ditags were eliminated from analysis resulting in 136,276, 139,656, 154,828 and 136,386 useful tags respectively and a total of 24, 058 unique tags (Figure 1A). 15,438 of the unique tags mapped to annotated UniGene identifiers. The raw data of the sequence tags have been made publicly available (Gene Expression Omnibus, series accession number GSE6252). We characterized the transcriptome of normal cervical tissue and evaluated the highly expressed genes in terms of tissue specificity, concordant expression among the normal libraries and their altered expression in CIN III lesions (Figure 1B).
Figure 1.
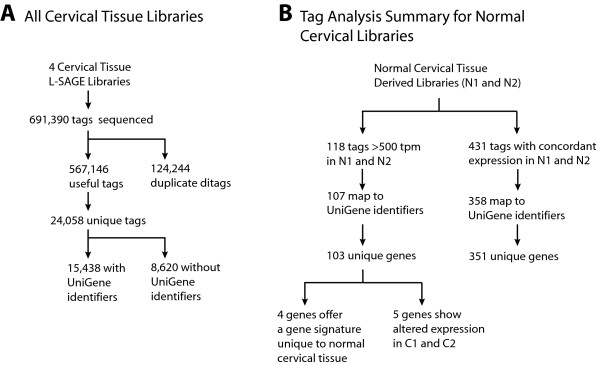
Flow diagram of SAGE analysis and tag-to-gene mapping. A. Sequence tags yielded from the four SAGE libraries were catagorized. Useful tags indicate all sequenced tags less duplicate ditags. B. The abundance and classification of unique tags in the SAGE libraries of normal cervix tissue (N1, N2) are summarized.
Genes Highly Expressed in Normal Cervical Epithelium
118 unique tags were found to be highly expressed in the normal cervical epithelium (at >500 tpm in both normal libraries). 103 of these tags mapped to UniGene clusters and represent 100 unique genes and hypothetical proteins (Figure 1). Manual examination of tags not mapped by SAGE Genie yielded three additional tags. This results in a total of 107 unique tag-to-gene mappings and 103 unique genes. The abundance of the 118 tags and the genes they represent are summarized in Table 1.
Table 1.
Tags expressed in normal cervical libraries at ≥500 tags per million.
| Gene Symbol | N1 tpm | N2 tpm | C1 tpm | C2 tpm | SAGE Tag |
| unknown | 2510 | 3623 | 2655 | 873 | TGATTTCACTTCCACTC |
| unknown | 2517 | 3430 | 3307 | 1672 | CTAAGACTTCACCAGTC |
| unknown | 594 | 559 | 497 | 594 | GTAGGGGTAAAAGGAGG |
| unknown | 2033 | 1683 | 2280 | 1239 | CACCTAATTGGAAGCGC |
| unknown | 807 | 852 | 1298 | 0 | CAAGCATCCCCGTTCCA |
| unknown | 822 | 680 | 814 | 631 | AGGTGGCAAGAAATGGG |
| unknown | 3757 | 2714 | 4134 | 2757 | ACTTTTTCAAAAAAAAA |
| unknown | 616 | 687 | 782 | 374 | ACTAACACCCTTAATTC |
| unknown | 1519 | 1525 | 2222 | 2126 | AAAAAAAAAAAAAAAAA |
| unknown | 565 | 780 | 588 | 308 | TTTAACGGCCGCGGTAC |
| unknown | 6289 | 6602 | 6510 | 4326 | TTCATACACCTATCCCC |
| ACTB | 2099 | 1031 | 1292 | 1459 | GCTTTATTTGTTTTTTT |
| ACTG1 | 1424 | 1955 | 1195 | 1598 | CTAGCCTCACGAAACTG |
| ANXA1 | 3713 | 6480 | 3701 | 10162 | AGAAAGATGTCTATGTA |
| ANXA2 | 1644 | 1754 | 1298 | 887 | CTTCCAGCTAACAGGTC |
| AQP3 | 609 | 573 | 930 | 1136 | TTTGCTTTTGTTTTGTT |
| ATP5G3 | 543 | 773 | 743 | 851 | GGAATGTACGTTATTTC |
| B2M | 8718 | 2821 | 9443 | 5712 | GTTGTGGTTAATCTGGT |
| BZRP | 550 | 960 | 794 | 675 | GAATTTTATAAGCTGAA |
| CCNB1IP1 | 580 | 924 | 988 | 565 | CCACTGCACTCCAGCCT |
| CD24 | 851 | 559 | 775 | 697 | GGAACAAACAGATCGAA |
| CD74 | 2348 | 573 | 1789 | 1444 | GTTCACATTAGAATAAA |
| CEACAM7 | 793 | 866 | 891 | 594 | AATCACAAATAAAAGCC |
| CFL1 | 1937 | 1031 | 1744 | 1679 | GAAGCAGGACCAGTAAG |
| COX4I1 | 646 | 523 | 969 | 616 | CCTATTTACTGGAAACC |
| CSTB | 712 | 3881 | 1227 | 2676 | ATGAGCTGACCTATTTC |
| CTSD | 2502 | 931 | 2894 | 1987 | GAAATACAGTTGTTGGC |
| EEF1A1 | 2245 | 1353 | 1737 | 1496 | TGTGTTGAGAGCTTCTC |
| EEF1B2 | 961 | 1210 | 1072 | 1298 | GCATTTAAATAAAAGAT |
| EEF1G | 726 | 501 | 807 | 418 | TGGGCAAAGCCTTCAAT |
| FLJ20701 | 550 | 501 | 685 | 264 | ATTTGAGAAGCCTTCGC |
| FTH1 | 2311 | 1275 | 2325 | 5176 | TTGGGGTTTCCTTTACC |
| GAPD | 1666 | 2363 | 2312 | 2816 | TACCATCAATAAAGTAC |
| GJA1 | 741 | 1310 | 504 | 513 | TGTTCTGGAGAGTGTTC |
| GLUL | 1658 | 1275 | 1905 | 2845 | TACAGTATGTTCAAAGT |
| IGHG1 | 14118 | 788 | 349 | 4715 | GAAATAAAGCACCCACC |
| IGHG1 | 18286 | 773 | 601 | 6548 | GAAATAAAGCACCCAGC |
| ITM2B | 1203 | 1031 | 1369 | 865 | TAAGTAGCAAACAGGGC |
| K-ALPHA-1 | 1181 | 795 | 1783 | 1239 | TGTACCTGTAATATTTT |
| KRT13 | 1380 | 8206 | 1815 | 2046 | AAAGCGGGGCTGGAGAA |
| KRT15 | 1152 | 3494 | 2713 | 2427 | TAATAAAGAATTACTTT |
| KRT5 | 2216 | 1539 | 1892 | 1371 | GCCCCTGCTGACACGAG |
| KRT6A | 1181 | 2657 | 975 | 1166 | AAAGCACAAGTGACTAG |
| LAMR1 | 2987 | 2370 | 3268 | 2911 | GAAAAATGGTTGATGGA |
| LGALS7 | 682 | 2220 | 446 | 425 | TAAACCTGCTGTGCGGG |
| LOC44051 | 719 | 702 | 588 | 719 | GGGTTGGCTTGAAACCA |
| LY6D | 1424 | 2248 | 1796 | 3072 | GAGATAAATGATTTAAA |
| MGC71999 | 998 | 1353 | 924 | 565 | TCACAAGCAAATGTGTC |
| MYL6 | 1115 | 559 | 995 | 1063 | GTGCTGAATGGCTGAGG |
| NET-5 | 1783 | 780 | 1298 | 997 | CACAAACGGTAGTTTTG |
| PERP | 1306 | 2828 | 1828 | 2236 | CCACAGGAGAATTCGGG |
| PLCB4 | 866 | 752 | 1072 | 733 | AAAACATTCTCCTCCGC |
| PPIA | 1504 | 1697 | 1951 | 1452 | CCTAGCTGGATTGCAGA |
| PTMA | 1049 | 1017 | 1518 | 697 | TTCATTATAATCTCAAA |
| RPL10 | 2223 | 1797 | 1944 | 1283 | AGGGCTTCCAATGTGCT |
| RPL10A | 998 | 652 | 1414 | 821 | GGCAAGCCCCAGCGCCT |
| RPL11 | 1130 | 773 | 1124 | 829 | CGCTGGTTCCAGCAGAA |
| RPL12 | 932 | 580 | 865 | 726 | ACATCATCGATGACATC |
| RPL13A | 2818 | 1998 | 2855 | 2383 | AGGCTACGGAAACAGGC |
| RPL13A | 1490 | 716 | 1434 | 1195 | CTCCTCACCTGTATTTT |
| RPL21 | 2091 | 1353 | 1899 | 1554 | GCATAATAGGTGTTAAA |
| RPL23 | 1196 | 1031 | 1072 | 1114 | ATTCTCCAGTATATTTG |
| RPL27A | 1923 | 1310 | 1989 | 1899 | GAGGGAGTTTCATTAAA |
| RPL28 | 1996 | 1504 | 1983 | 1833 | GCAGCCATCCGCAGGGC |
| RPL29 | 1049 | 773 | 1104 | 814 | GGGCTGGGGTCCTCCTG |
| RPL3 | 917 | 702 | 840 | 983 | GGACCACTGAAGAAAGA |
| RPL30 | 1541 | 1017 | 1259 | 961 | CCAGAACAGACTGGTGA |
| RPL31 | 609 | 573 | 568 | 491 | AAGGAGATGGGAACTCC |
| RPL32 | 1512 | 1024 | 1550 | 1371 | TGCACGTTTTCTGTTTA |
| RPL35 | 638 | 551 | 665 | 572 | CGCCGCCGGCTCAACAA |
| RPL36 | 1570 | 1275 | 1628 | 1173 | AGGAAAGCTGCTGCCAA |
| RPL37 | 5063 | 3272 | 6362 | 6064 | CAATAAATGTTCTGGTT |
| RPL37A | 1350 | 938 | 1259 | 953 | AAGACAGTGGCTGGCGG |
| RPL4 | 521 | 537 | 465 | 359 | CGCCGGAACACCATTCT |
| RPL41 | 2649 | 1812 | 2609 | 1906 | TTGGTCCTCTGCCCTGG |
| RPL7 | 1702 | 1160 | 1427 | 1180 | ATTATTTTTCTAAGCTG |
| RPLP0 | 815 | 1239 | 846 | 601 | CTCAACATCTCCCCCTT |
| RPLP1 | 3574 | 2735 | 3507 | 2398 | TTCAATAAAAAGCTGAA |
| RPLP2 | 2422 | 1461 | 2196 | 2376 | GGATTTGGCCTTTTTGA |
| RPS11 | 1365 | 802 | 917 | 770 | TCTGTACACCTGTCCCC |
| RPS12 | 690 | 695 | 601 | 396 | GCCGAGGAAGGCATTGC |
| RPS14 | 1159 | 859 | 1240 | 843 | TAAAAAAAAAAAAAAAA |
| RPS18 | 1071 | 1053 | 1085 | 858 | TGGTGTTGAGGAAAGCA |
| RPS19 | 2598 | 1604 | 2357 | 2618 | CTGGGTTAATAAATTGC |
| RPS20 | 903 | 695 | 1020 | 667 | GCTTTTAAGGATACCGG |
| RPS23 | 961 | 637 | 833 | 704 | CTGTTGGTGATATTCCT |
| RPS25 | 1042 | 988 | 924 | 660 | AATAGGTCCAACCAGCT |
| RPS26 | 2576 | 1002 | 1938 | 873 | TAAGGAGCTGAGTTCTT |
| RPS27A | 815 | 623 | 988 | 887 | AACTAAAAAAAAAAAAA |
| RPS28 | 998 | 902 | 1053 | 1034 | GACGACACGAGCCGATC |
| RPS29 | 2245 | 1940 | 2325 | 2317 | ATAATTCTTTGTATATA |
| RPS3A | 1218 | 823 | 1427 | 1232 | GTGAAGGCAGTAGTTCT |
| RPS4X | 1541 | 952 | 1615 | 924 | TCAGATCTTTGTACGTA |
| RPS8 | 4021 | 3587 | 4282 | 3996 | TAATAAAGGTGTTTATT |
| RPS9 | 587 | 530 | 407 | 521 | CCAGTGGCCCGGAGCTG |
| S100A10 | 514 | 501 | 543 | 506 | AGCAGATCAGGACACTT |
| S100A2 | 2950 | 723 | 2235 | 3505 | GATCTCTTGGGCCCAGG |
| S100A7 | 2899 | 752 | 388 | 5990 | GAGCAGCGCCCTGTTCC |
| S100A8 | 690 | 1031 | 827 | 1466 | GCTTTTTTTGTGGGCTG |
| S100A8 | 11880 | 25205 | 8855 | 24629 | TACCTGCAGAATAATAA |
| S100A9 | 13561 | 17078 | 10089 | 18660 | GTGGCCACGGCCACAGC |
| SERPINB3 | 1137 | 1454 | 704 | 917 | CCTTTCTCTCTTTCTCT |
| SFN | 1255 | 1976 | 1505 | 1569 | TTTCCTCTCAATAAAGT |
| SMAD2 | 2935 | 2341 | 3378 | 1987 | CCCATCGTCCTAGAATT |
| SPINK5 | 712 | 4189 | 0 | 1305 | TCCACCAAGTCTGAGCC |
| SPRR3 | 1600 | 11757 | 3888 | 5968 | TTTCCTGCTCTGCCCTC |
| SPRR3 | 1636 | 13068 | 4121 | 8329 | TTTCCTGCTCTTCCCTC |
| STX3A | 646 | 1905 | 1318 | 1312 | TAAAATGTTTATGATAA |
| SUI1 | 1446 | 1239 | 1259 | 2009 | CAATAAACTGAAAAGAG |
| TAGLN2 | 660 | 616 | 769 | 543 | GTCTGGGGCTTGAGGAA |
| TMSB4X | 2245 | 616 | 1175 | 917 | TTGGTGAAGGAAGAAGT |
| TPI1 | 947 | 1382 | 1589 | 1584 | TGAGGGAATAAACCTGG |
| TPT1 | 4726 | 3659 | 4915 | 4568 | TAGGTTGTCTAAAAATA |
| TXNL5 | 660 | 838 | 258 | 924 | TTAGCAATAAATGATGT |
| UQCRH | 646 | 508 | 930 | 726 | GGTTTGGCTTAGGCTGG |
| VAMP8 | 521 | 795 | 459 | 491 | TGGCTGGGAAACTGTTG |
| VPS13B | 763 | 594 | 911 | 0 | CACTACTCACCAGACGC |
| YWHAZ | 587 | 702 | 517 | 814 | TAAGTGGAATAAAAGTT |
To determine cervical tissue specific expression, we first investigated the expression of the 107 genes using expression data available at the National Center for Biotechnology Information (NCBI) Unigene database and the National Cancer Institute (NCI) Cancer Genome Anatomy Project (CGAP) SAGE Anatomical Viewer. Based on CGAP information, only four of the 107 genes were unique to cervical tissue: carcinoembryonic antigen-related cell adhesion molecule 7 (CEACAM7), keratin 6A (KRT6A), small proline-rich protein 3 (SPRR3) and S100 calcium binding protein A7 (S100A7). These genes were further investigated for expression by RT-PCR in 20 different tissue types and three normal cervical specimens (Figure 2). CEACAM7 was found to be expressed in colon, larynx, pancreas and two of the three normal cervical specimens. KRT6A expression was detected in placenta, thymus, tongue, prostate, larynx, colon, skin and in all three of the normal cervical specimens. SPRR3 was found strongly expressed in placenta, thymus, colon, tongue, larynx and all three of the normal cervical cases. S100A7 showed expression in placenta, thymus, and tongue and in all three of the normal cervical specimens. All four genes were prominently expressed in the cervical epithelium but this combination of genes was not expressed in the tissues examined (Figure 2).
Figure 2.
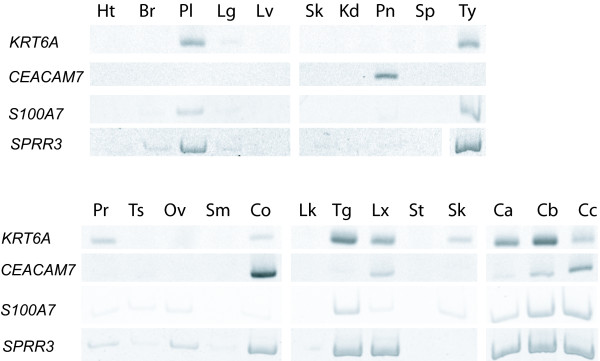
Validation of tissue specificity of gene expression. Reverse transcriptase PCR of four genes in 20 tissue types and three normal cervical specimens. Heart (Ht), breast (Br), placenta (Pl), lung (Lg), liver (Lv), skeletal muscle (Sk), kidney (Kd), pancreas (Pn), spleen (Sp), thymus (Ty), prostate (Pr), testis (Ts), ovary (Ov), small intestine (Sm), colon (Co), peripheral leukocytes (Lk), tongue (Tg) larynx (Lx), stomach (St), skin (Sn). Ca, Cb and Cc are three individual normal cervical tissue specimens.
Disrupted Gene Expression in CIN III
All tags were assessed for altered expression in CIN III. Four hundred and seventy-six tag show greater than two fold increase in CIN III and are expressed at greater than 15 tpm (see Additional file 1) while 315 tags were decreased in CIN III (see Additional file 2).
We determined if the expression of the 107 unique tags, that were highly expressed in normal cervical libraries (> 500 TPM), were disrupted in CIN III. Comparison of expression levels in N1, N2 to the CIN III libraries using the Z-test revealed five differentially expressed genes (Table 2). Annexin 2 (ANXA2), galectin 7 (LGALS7) and connexin 43 (GJA1) exhibited decreased expression in CIN III (Z < -1.96) while aquaporin 3 (AQP3) and ribosomal-like protein 37 (RPL37) increased in expression (Z > 1.96). Real-time PCR was performed on a panel of 6 new cervical specimens, three each of normal and CIN III for all five of these genes (Figure 3). Expression results were normalized to housekeeping gene ACTB and 18S (Figure 3A and 3B, respectively). Decrease in expression of ANXA2, LGALS7 and GJA1 in CIN III was confirmed while increase in expression of AQP3 and RPL37 were not.
Table 2.
Highly expressed genes with altered expression in CIN III.
| Z_Score a | |||||||||
| Tag | N1 tpm | N2 tpm | C1 tpm | C2 tpm | N1 vs C2 | N1 vs C1 | N2 vs C2 | N2 vs C1 | Gene Symbol |
| TTTGCTTTTGTTTTGTT | -4.7 | -3.1 | -5.1 | -3.5 | AQP3 | ||||
| CAATAAATGTTCTGGTT | 5063 | 3272 | 6362 | 6064 | -3.5 | -4.6 | -10.8 | -12 | RPL37 |
| TGTTCTGGAGAGTGTTC | 741 | 1310 | 504 | 513 | 2.4 | 2.6 | 6.9 | 7.3 | GJA1 |
| TAAACCTGCTGTGCGGG | 682 | 2220 | 446 | 425 | 2.9 | 2.7 | 12.9 | 13.4 | LGALS7 |
| CTTCCAGCTAACAGGTC | 1644 | 1754 | 1298 | 887 | 5.6 | 2.4 | 6.3 | 3.2 | ANXA2 |
a Z > 1.96; reduced expression in CIN III, Z < -1.96; increased expression in CIN III.
Figure 3.
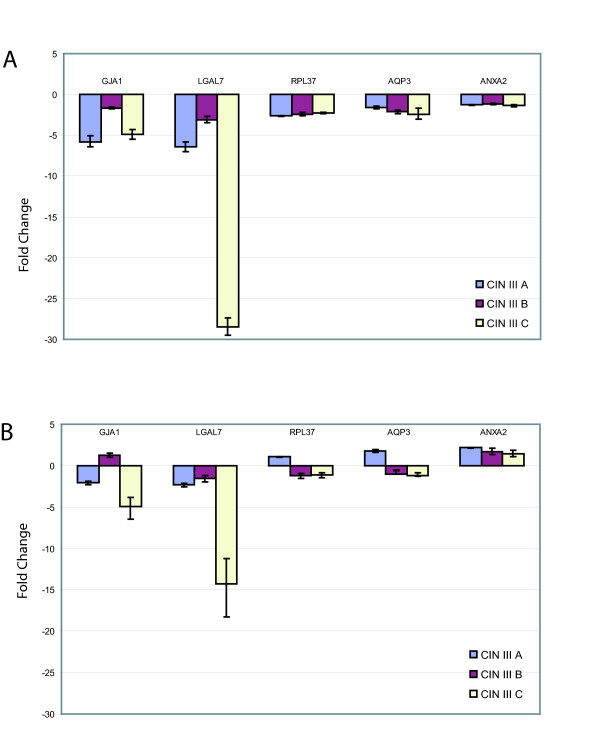
Summary of test panel quantitative PCR results of genes with altered expression in CIN III L-SAGE libraries. A panel of three new CIN III cases (CIN III A, CIN III B, CIN III C) were investigated for expression and compared to three new normal specimens. Gene expression was normalized to ACTB and 18S (Figure 3A and 3B, respectively). Zero on the Y-axis denotes mean expression levels of the respective genes in normal cervical tissue. All five genes investigated showed decreased expression.
Viral (HPV 16) tags in L-SAGE libraries
HPV transcripts were also detected by L-SAGE. Tags from all four libraries were mapped against the genomes of HPV 16 and HPV 18. While no tags mapped to HPV 18, twelve tags from the CIN III libraries mapped to the more prevalent HPV 16 genome (Table 3). The highest transcript counts of known genes belonged to E5 at 1,180 and 290 tpm and E2 at 240 and 20 tpm, in libraries C2 and C1, respectively. Compared by BLAST [8] against the RefSeq Genome collection, none of the twelve tags matched 100% to the human genome. All twelve tags were also mapped against human transcript sets (mitochondrial genome, RefSeq, UCSC gene set, Unigene, Ensembl, UCSC mRNA, UCSC EST, SAGEmap and SAGEgenie SAGE tag sets). No tags matched to any of the described transcript sets with the exception of CATGCACGCTTTTTAATTACA and CATGTGTATGTATTAAAAATA which mapped to human EST BF909200. The full length EST sequence is 97% identical with the HPV 16 E5 gene and was likely amplified from HPV sequences in the originating uterine tumour lesion.
Table 3.
Tags mapping to HPV16 genome.
| Tag | Strand | Gene | N1 tpm | N2 tpm | C1 tpm | C2 tpm |
| CATGCACGCTTTTTAATTACA | + | E5 | 0 | 0 | 290 | 1,180 |
| CATGCAACATAAATAAACTTA | + | - | 0 | 0 | 6 | 770 |
| CATGGCATTGGACAGGACATA | + | E2 | 0 | 0 | 6 | 240 |
| CATGTGTATGTATTAAAAATA | - | E5 | 0 | 0 | 6 | 155 |
| CATGACACAATAGTTACACAA | + | - | 0 | 0 | 6 | 125 |
| CATGGGGAGGAATATGATTTA | + | L1/L2 | 0 | 0 | 0 | 30 |
| CATGTAGACGACACTGCAGTA | - | E2 | 0 | 0 | 0 | 20 |
| CATGGTAGATTATGGTTTCTG | - | E7 | 0 | 0 | 6 | 7 |
| CATGTCGTAGGTACTCCTTAA | - | L1/L2 | 0 | 0 | 0 | 7 |
| CATGGGGATCCTTTGCCCCAG | - | L1/L2 | 0 | 0 | 0 | 7 |
| CATGATAATATATGTTTGTGC | - | L1/L2 | 0 | 0 | 0 | 7 |
| CATGCGCCTAGAATGTGCTAT | + | E2 | 0 | 0 | 0 | 0 |
Discussion
This study represents the most comprehensive gene expression analysis of cervical tissue reported to date. In total, 691,390 L-SAGE tags were sequenced (Figure 1A). The length of the L-SAGE tags (21 bp as compared to 14 bp in conventional SAGE) greatly reduces tag-to-gene mapping ambiguity [6]. 107 of the 118 (88%) highly expressed tags (i.e. >500 tpm) were mapped to known genes or hypothetical proteins encompassing 103 unique genes (Figure 1B).
Assessing Highly Expressed Tags by Functional Group
Of the 107 highly expressed tags (>500 tpm), 47 were expressed at extremely high levels (>103 tpm) including genes frequently used as controls in expression analysis, GAPDH and ACTB. High expression of 20 genes in normal cervical tissue was reported in a previous study [3]. Fifteen of these genes are encompassed by our list of 107 high expressers. The most highly expressed tags expressed at >104 tpm (GTGGCCACGGCCACAGC and TACCTGCAGAATAATAA) mapped to the genes S100A9 and S100A8, respectively. Both genes belong to the calcium binding protein family. These findings are in agreement with a previous report of high S100A8 and S100A9 levels in cervical tissue [3]. Although the function of these genes is not well understood, genes within this family have been proposed to participate in a variety of cellular process including cell cycle, wound healing and cell differentiation [9].
Assigning the 103 highly expressed genes to one of eleven broad functional groups allowed for an assessment of those cellular processes represented by the most abundant transcripts. These cellular processes include calcium binding proteins, cell cycle or cell death, cytoskeleton, immune functioning, keratinization, membrane proteins, mitochondrial, protein processing, translation (ribosomal proteins), translation (non ribosomal proteins) with a small fraction of tags mapping to other functional groups or to genes with no known function (Figure 4A). The 41 ribosomal genes account for the greatest proportion of highly expressed genes at 28% and 31% (normal and CIN III, respectively). In contrast, only five calcium binding genes account for the second largest functional subgroup of highly expressed tags, 18% and 19% (normal and CIN III, respectively).
Figure 4.
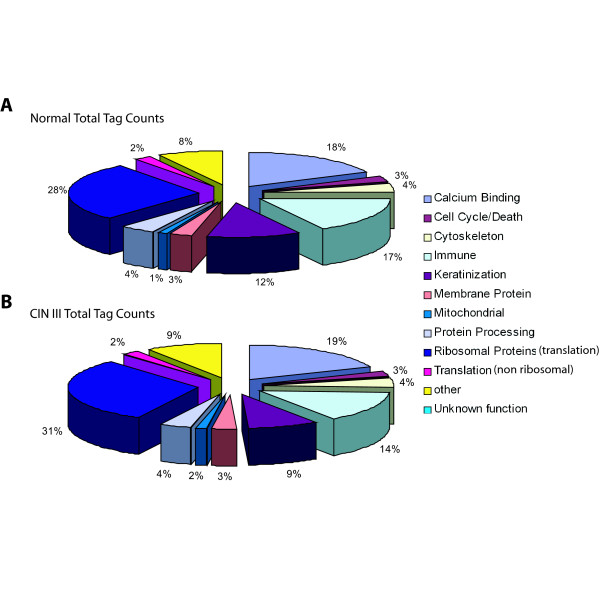
Functional groupings of tags highly expressed (>500 tpm) in normal libraries. Categories are as described. The Other group consists of tags which map to known genes but are not encompassed by any of the ten categories. The Unknown function group consists of tags mapping to no known genes. A) Tag counts in the normal libraries are categorized by functional group. B) Tags found in A were quantified in both CIN III libraries and categorized according to the same functional grouping scheme. In both groups Ribosomal genes accounted for the greatest number of tags while only the keratins changed in expression or were decreased in the CIN III libraries.
The relative expression levels of the functional groups do not change greatly between the normal and CIN III libraries however the keratin and immune related functional groups show slight decrease from 12% and 17% in CIN III to 9% and 14% in normal tissue (keratin and immune groups, respectively).
All tags expressed at or greater than 15 tpm in Normal and CIN III libraries (2,814 and 3,279 respectively) were also evaluated according functional group using Onto-Express (see Additional File 3) [10]. The most represented groups included DNA dependent transcription regulators and transcription in both the Normal and CIN III libraries.
Cervical Tissue Gene Signature
Four of the 103 unique genes we found to be abundantly expressed in normal cervical tissue were documented to have limited or no expression in other tissues according to the web resources NCBI UniGene [11] and NCI CGAP SAGE Anatomical Viewer [12,13] (Figure 1B). These genes (CEACAM7, SPRR3, S100A7 and KRT6A) are our candidates for an expression signature unique to normal cervical tissue and were further investigated in a panel of 20 different tissue types and three new normal cervical specimens. We found that all four of these genes were not abundantly expressed simultaneously in any of the 20 tissues examined (Figure 2). Placenta, thymus and tongue were found to express a combination of three genes (S100A7, SPRR3 and KRT6A), while colon expressed another combination (CEACAM7, SPRR3 and KRT6A). KRT6A and SPRR3 expression was observed in larynx tissue with only minimal expression detected for S100A7 and CEACAM7. In contrast, two of the cervical cases strongly expressed all four genes investigated while only the third showed very low CEACAM7 expression. Significantly, our study is the first to document CEACAM7 expression in cervix. The data suggest that abundant expression of CEACAM7 and S100A7 collectively, are unique to cervical tissue and have the potential to serve as useful biomarkers in identifying origins of metastatic disease.
It is interesting to note that decreased expression of three of the genes is linked to abnormal growth and organization of epithelium. For example, CEACAM7 is a member of the carcinoembryonic antigen family of genes and expression has been documented in highly differentiated normal colon epithelium and the apical surface of normal ductal pancreas epithelium, while loss of expression has been reported in colon hyperplastic polyps [14]. Another member of this gene family, CEACAM1, is shown to have no or very low expression in cervical carcinoma [15]. Decreased expression of KRT6A and S100A7 have been associated with breast, lung and ovarian cancer [16-20]. SPRR3 belongs to the class of small proline rich genes which are expressed in differentiated keratinocytes and has previously been shown to be highly expressed in normal cervical tissue [21].
Genes Altered in Expression in CIN III
The 107 highly expressed unique tags in the normal libraries were assessed for expression changes in CIN III. Two genes showed increased expression (AQP3 and RPL37) while three genes declined in transcript counts in the CIN III libraries (ANXA2, GJA1 and LGALS7). All five were evaluated by real-time PCR in a new cervical tissue panel. Results for ANXA2, LGALS7 and GJA1 confirmed L-SAGE findings.
Panel results for GJA1 are in agreement with those reported by King et al. in that GJA1 expression was detected in normal cervical epithelium while reduced expression was observed in CIN III lesions investigated [22]. It has been suggested that this pattern may be a consequence of epithelial disorganization and not causative in dysplasia development [23,24].
The high expression of LGALS7 in normal cervical epithelium contrasted by the low expression seen in the CIN III lesions in the tissue panel we report here is similar to those expression patterns seen in studies of other normal tissue types compared to their respective carcinomas including cornea and larynx [25]. LGALS7 expression has been hypothesized in all stratified epithelium tissue types and has been experimentally detected in human cornea, heart, larynx, tongue, skin, thymus and thyroid [26-28]. Though, LGALS7 has been investigated in the context of cell line models, it is interesting to note that expression of this gene in cervical epithelium has not been previously reported [26-28]. LGALS7 is one of fifteen members of the B-galactoside-binding lectin family, some of which have been shown to influence cell growth, cell cycle, apoptosis and cell migration via their predicted role in homeostasis, however, the role of LGALS7 in cancer is unclear [27,29]. Support for the pro-apoptotic function of LGALS7 was reported by Kuwabara et al. and Bernard et al. who identified cells more sensitive to apoptosis when LGALS7 expression was high in the epithelium derived cells [28,30,31]. In contrast, Demers et al. showed an increase in LGALS7 expression in lymphoma cells and suggested a positive role in cell growth and dispersal through induced matrix metalloproteinase 9 (MMP9) expression [32]. We did not observe a statistically significant change in MMP9 in the four libraries investigated. This variance in expression suggests multiple roles for LGALS7 that may be tissue-type dependent.
The third gene investigated, ANXA2, is known to be highly expressed in epithelial cells and is localized to the plasma membrane and endosome. It has been suggested to function in linking membrane to membrane and membrane to cytoskeleton [33]. The decrease in expression we observe suggests that the loss of ANXA2 may be a causative factor in disorganisation of the epithelial architecture, which is characteristic of cervical neoplastic lesions. ANXA2 is also known to bind with S100A10 and participates in transport channel function across the plasma membrane [33]. Interestingly, the ANXA2 binding site on S100A10 also binds NS3, a viral protein from the bluetongue virus, and therefore directly competes with ANXA2. The S100A10 protein has also been shown inhibit Hepatitis B virus polymerase (HBV pol) activity [34]. It is plausible that a S100A10-ANXA2 complex may have a role in HPV infection or viral lifecycle. S100A10 expression was high and consistent in Normal and CIN III libraries whereas ANXA2 decreased in the CIN III libraries.
The above real-time PCR results were normalized to the widely accepted housekeeping gene ACTB (Figure 3A). For comparison we also normalized the genes to a second housekeeping gene 18S (Figure 3B). Briefly, results for GJA1 and LGALS7 were in agreement to those when normalizing to ACTB, however ANXA2 was shown to be increased in CIN III as expected by the SAGE data. but this does not concur with the QPCR results when normalized to ACTB. One possible explanation is that on average, the ACTB cycle threshold was >1.3 Ct lower in the CIN III cases indicating an increase in ACTB expression in CIN III lesions. Any Ct decrease less than this in the genes investigated would appear as a decrease in gene expression in CIN III when normalized to ACTB but and increase when normalized to 18S.
The real-time PCR results for AQP3 and RPL37 did not concur with L-SAGE data and may be due to interindividual differences rather than a representation of changes present in CIN lesions or cancer. It is interesting to note that RPL37 overexpression in prostate cancer, colon cancer cell lines and clinical specimens have been reported [35,36]. Our L-SAGE results also suggest a similar pattern in cervical neoplasia. Results for these genes when normalized to 18S showed an increase in expression in only CIN III A (see Additional file 3).
Wong et al investigated gene expression in invasive cervical carcinoma by DNA microarray [37]. We investigated this publicly available data through NCBI GEO [38]. Briefly, in this data, GJA1 showed a small decrease in expression in invasive disease while LGALS7 was detected in only four of the 26 specimens, three of which were normal tissue. Moderate AQP3 expression was detected in the majority of cases including the control group. Expression of ANXA2 and RPL37 was not assessed in the microarray study.
Human L-SAGE Tags Map to the HPV Genome
HPV is an established etiological factor in cervical cancer [2]. There are over 100 known strains of HPV, however HPV 16 and HPV 18 are considered to be the frequent high risk types owing to higher rates of persistent infection, higher rates of progression to cervical neoplasia, and shorter median progression times than other HPV strains [2]. HPV 16 is the most common strain and can be detected in approximately 60% of cervical cancers, while HPV 18 infection occurs in approximately 10–20%. Uncontrolled expression of E6 and E7 genes from strains 16 or 18 are considered to be essential for oncogenic transformation and function through inhibition of host cell tumour suppressors p53 and the retinoblastoma protein (Rb) [2].
This is the first study to mine human SAGE libraries for viral transcripts. Overall, CIN III library C2 (2,548 tpm) possessed a greater number of tag counts from the more prevalent HPV 16 strain when compared to C1 (320 tpm) (Table 3). HPV 18 tags were not found in any library and no viral tags were detected in the normal libraries. With the exception of two tags, the viral tags expressed in cervical SAGE did not map to known human genes or expressed sequences. The exceptions, CATGCACGCTTTTTAATTACA and CATGTGTATGTATTAAAAATA, mapped to a single human EST isolated from a uterine tumour (EST BF909200). The full length EST sequence is 97% identical with the HPV 16 genome, more specifically the E5 gene, and therefore was likely amplified from HPV sequences in the original lesion.
Tags mapping to the E5 gene accounted for the greatest proportion of HPV tags mapping to known transcripts in both C1 and C2 (93% and 52%, respectively). E5 is considered to be one of three HPV 16 oncogenes (E5, E6 and E7) and is highly expressed in basal cells of premalignant cervical lesions [39]. This expression declines as cells differentiate and move toward the apical face of the epithelium whereas E6 and E7 expression increases [39]. E5 is detected throughout all epithelium layers in high grade lesions such as CIN III. In contrast, expression is restricted to layers closest to the basal cells in low grade lesions, implying that E5 expression may be limited to undifferentiated basal cells [40,41]. The high expression of E5 we observe in the CIN III libraries and the absence of HPV 16 genes in the normal libraries is in concordance with such studies.
An increase in sample size and inclusion of mild and moderate stages of cervical intraepithelial neoplasia will aid in quantifying the relationship between viral gene expression and disease. This will also assist in further elucidating genes important in early lesion transcriptome events. A comparison of such events with those seen in later stages will help to identify genes important in the molecular pathogenesis of the disease.
Conclusion
In this study we have described the transcriptome of normal cervical tissues and compared against that of CIN III lesions. This was achieved by construction of four L-SAGE libraries and sequencing to the depth of 172,848 tags per library on average. We highlighted that the Long-SAGE technique provides a comprehensive profile of the transcriptome without focusing on only known genes. Potent tumour suppressors (e.g. PTEN), cell cycle mediators (e.g. CCND1), and cellular respiration genes (e.g. NDUFA1) were found to be tightly regulated in the normal libraries. An expression signature of four highly expressed genes (KRT6A, CEACAM7, S100A7 and SPRR3) in normal cervical epithelium was identified and confirmed, and three abundantly expressed genes (ANXA2, GJA1 and LGALS7) were found to have altered expression in CIN III. Furthermore, this is the first study to have identified viral tags in human SAGE libraries demonstrating the versatile nature of SAGE data, which allows for mining and re-mining according to newly posed questions. HPV 16 E5 transcripts were found most highly expressed while few E7 and no E6 transcripts were enumerated.
The identification of expression changes associated with stages of disease progression will help further our understanding of cervical cancer development and potentially elucidate novel targets for diagnosis and treatment. Establishing a baseline from which to compare is essential to the identification of such aberrations and the 20 fold increase in cervical gene expression data presented here is a significant contribution to this effort.
Methods
Sample selection
The specimens were collected immediately prior to the LEEP (Loop electrosurgical excision procedure) cone biopsy targeting a small portion of the affected epithelium. These specimens were collected with patient consent at the Vancouver General Hospital Women's Clinic at Vancouver Hospital & Health Science Centre. Cases were assessed by cervical cancer pathologists at Vancouver Hospital and Health Science Centre and were selected without prior knowledge of HPV status. Specimens N1 and N2 in this study were observed to be normal squamous epithelia whereas C1 and C2 were identified as high grade dysplasia or CIN III. Detailed information on specimen pathology based on the LEEP cone specimens can be found in Additional file 4. All samples were stored immediately in RNA later and stored at -80°C. Three cases each of CIN III (CIN III A, CIN III B and CIN III C) and normal cervical tissue (NA, NB and NC) which were used for target validation through real-time PCR were also collected, assessed and stored in the same manner.
L-SAGE Library Construction and Sequence Tag Analysis
The biopsies were individually homogenised in Lysis Binding buffer (100 mM Tris-HCl, pH7.5, 500 mM LiCl, 10 mM EDTA, pH 8.0, 1% LiDS, 5 mM dithiothreitol). Long SAGE libraries were constructed according to the L-SAGE kit manual (Invitrogen, Ontario, Canada). Sequencing was performed at the BC Cancer Agency Michael Smith Genome Sciences Centre. L-SAGE employs 21 basepair sequence tags, reducing the ambiguity in tag-to-gene mapping that is sometimes encountered in classic SAGE libraries which use 14 basepair sequence tags.
Data Analysis
Tags were mapped using the February 12, 2006 version of SAGE Genie [13], and raw tag counts excluding duplicate ditags were normalized to tags per million (tpm). A Z-test analysis, standard for SAGE data analysis, was performed as previously established by Kal et al. for comparison of one SAGE library to another using an established cut-off of 1.96 on the absolute Z-score to determine statistically significant differences in expression levels between normal and CIN III [42].
Reverse Transcriptase PCR
For validation of cervical tissue gene signature, human Multiple Tissue cDNA Panel I and II (Clontech, Mississauga, Ontario) and total RNA for human larynx, skin, stomach and tongue (Stratagene, Cedar Creek, Texas) were used. Five-hundred nanograms from larynx, skin, stomach and tongue was used to generate cDNA. Final concentrations of PCR reagents for cervical tissue signature were 0.5 μM primer, 2 mM MgCl2, 0.2 mM dNTP, 1× PCR buffer (Invitrogen), 0.5 U Taq polymerase and 1 μL of cDNA (annealing temperatures: 55°C (ACTB, CEACAM7), 60°C (KRT6A), 65°C (SPRR3 and S100A7). Total RNA for the panel of six cervical specimens used for validation of genes was isolated using Trizol (Invitrogen) and cDNA was generated using the High Capacity TaqMan Reverse Transcription Reagents, according to the manufacturer's instructions (Applied Biosystems, Foster City, CA).
Expression of genes selected by data analysis were analyzed by real-time PCR using TaqMan® Gene Expression Assays on the ABI 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA), according to manufacturer's instructions. Samples were run in duplicate and normalized against an beta-actin (ACTB) endogenous control (HuACTB, Applied Biosytems). Assay IDs include AQP3, Hs00185020_m1; GAL7, Hs00170104_m1; RPL37, Hs02340038_g1; and GJA-1, Hs00748445_s1. The relative quantification of these target genes in CIN III (CIN III A, CIN III B and CIN III C) samples compared to normal tissue (NA, NB and NC) samples was performed using the established 2-ΔΔCt method, (Applied Biosystems, Relative Quantitation Of Gene Expression, ABI PRISM 7700 Sequence Detection System: User Bulletin #2). The cycle threshold (Ct) value of the target gene was normalized to ACTB by subtracting the Ct value for ACTB from that of the target gene (ΔCt gene = Ct gene - Ct ACTB). This number was then averaged from the three normal cases (ave ΔCt normal). The mean fold change of each gene between normal and CIN III was calculated using the following equation: 2 -(ΔCt gene - ave ΔCt normal). The relative quantification values were then plotted, one indicating no change with respect to normal cervical tissue.
HPV Tag-to-Gene Mapping
Viral genomic sequence files (viral1.genomic.fna and HPV11.txt) were downloaded from public repositories and processed into 21 bp SAGE tags at every Nla III site in both orientations using custom Perl scripts [43,44].
Abbreviations
Cervical Intraepithelial Neoplasia (CIN), Expressed Sequence Tag (EST), Human Papillomavirus (HPV), Serial Analysis of Gene Expression (SAGE), Tags per million (tpm)
Authors' contributions
AS designed, organized and performed this study and wrote the manuscript. RC and GV performed data analysis. JC performed gene specific expression assays. KL contributed to the project design, directed library construction and edited the manuscript. JM, DN, TE and DM were responsible for clinical diagnosis, sample acquisition and pathology assessment. MF, WL and CA are principle investigators of this study. All authors read and approved the final draft of this manuscript.
Supplementary Material
Supplemental Table 1. Increased Expression in CIN III (>15 tpm and >2 fold change)
Supplemental Table 2. Decreased expression in CIN III (≥15 tpm and ≥2 fold change)
Supplemental Figure 1. Functional annotation of all genes expressed in Normal and CIN III lesions (Supplemental Figure A and B, respectively). Tags expressed >15 TPM included.
Supplemental Table 3. Cervical Specimen Description based on LEEP cone biopsy Pathology Report
Acknowledgments
Acknowledgements
We are grateful to Anita Carraro for sample coordination and retrieval and Blair Gervan, Lana Galac and Ariane Williams for their technical assistance. This work was made possible by support from the National Institute of Health (5 P0l CA 82710-07 and 7P0l CA 103830).
Contributor Information
Ashleen Shadeo, Email: ashadeo@bccrc.ca.
Raj Chari, Email: rchari@bccrc.ca.
Greg Vatcher, Email: gvatcher@bccrc.ca.
Jennifer Campbell, Email: jcampbel@bccrc.ca.
Kim M Lonergan, Email: klonergan@bccrc.ca.
Jasenka Matisic, Email: jmatisic@bccancer.bc.ca.
Dirk van Niekerk, Email: dvanniek@bccancer.bc.ca.
Thomas Ehlen, Email: tehlen@vanhosp.bc.ca.
Dianne Miller, Email: dmmiller@vanhosp.bc.ca.
Michele Follen, Email: mfollen@mdanderson.org.
Wan L Lam, Email: wanlam@bccrc.ca.
Calum MacAulay, Email: cmacaula@bccrc.ca.
References
- Parkin DM, Bray F, Ferlay J, Pisani P. Global cancer statistics, 2002. CA Cancer J Clin. 2005;55:74–108. doi: 10.3322/canjclin.55.2.74. [DOI] [PubMed] [Google Scholar]
- Scheurer ME, Tortolero-Luna G, Adler-Storthz K. Human papillomavirus infection: biology, epidemiology, and prevention. Int J Gynecol Cancer. 2005;15:727–746. doi: 10.1111/j.1525-1438.2005.00246.x. [DOI] [PubMed] [Google Scholar]
- Perez-Plasencia C, Riggins G, Vazquez-Ortiz G, Moreno J, Arreola H, Hidalgo A, Pina-Sanchez P, Salcedo M. Characterization of the global profile of genes expressed in cervical epithelium by Serial Analysis of Gene Expression (SAGE) BMC Genomics. 2005;6:130. doi: 10.1186/1471-2164-6-130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steinau M, Lee D, Rajeevan M, Vernon S, Ruffin M, Unger E. Gene expression profile of cervical tissue compared to exfoliated cells: Impact on biomarker discovery. BMC Genomics. 2005;6:64. doi: 10.1186/1471-2164-6-64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siddiqui AS, Delaney AD, Schnerch A, Griffith OL, Jones SJ, Marra MA. Sequence biases in large scale gene expression profiling data. Nucleic Acids Res. 2006;34:e83. doi: 10.1093/nar/gkl404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saha S, Sparks AB, Rago C, Akmaev V, Wang CJ, Vogelstein B, Kinzler KW, Velculescu VE. Using the transcriptome to annotate the genome. Nat Biotechnol. 2002;20:508–512. doi: 10.1038/nbt0502-508. [DOI] [PubMed] [Google Scholar]
- Velculescu VE, Zhang L, Vogelstein B, Kinzler KW. Serial analysis of gene expression. Science. 1995;270:484–487. doi: 10.1126/science.270.5235.484. [DOI] [PubMed] [Google Scholar]
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Eckert RL, Broome AM, Ruse M, Robinson N, Ryan D, Lee K. S100 proteins in the epidermis. J Invest Dermatol. 2004;123:23–33. doi: 10.1111/j.0022-202X.2004.22719.x. [DOI] [PubMed] [Google Scholar]
- Draghici S, Khatri P, Martins RP, Ostermeier GC, Krawetz SA. Global functional profiling of gene expression. Genomics. 2003;81:98–104. doi: 10.1016/S0888-7543(02)00021-6. [DOI] [PubMed] [Google Scholar]
- Wheeler DL, Church DM, Federhen S, Lash AE, Madden TL, Pontius JU, Schuler GD, Schriml LM, Sequeira E, Tatusova TA, Wagner L. Database resources of the National Center for Biotechnology. Nucleic Acids Res. 2003;31:28–33. doi: 10.1093/nar/gkg033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strausberg RL, Buetow KH, Emmert-Buck MR, Klausner RD. The cancer genome anatomy project: building an annotated gene index. Trends in Genetics. 2000;16:103–106. doi: 10.1016/S0168-9525(99)01937-X. [DOI] [PubMed] [Google Scholar]
- Boon K, Osorio EC, Greenhut SF, Schaefer CF, Shoemaker J, Polyak K, Morin PJ, Buetow KH, Strausberg RL, De Souza SJ, Riggins GJ. An anatomy of normal and malignant gene expression. Proc Natl Acad Sci U S A. 2002;99:11287–11292. doi: 10.1073/pnas.152324199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scholzel S, Zimmermann W, Schwarzkopf G, Grunert F, Rogaczewski B, Thompson J. Carcinoembryonic antigen family members CEACAM6 and CEACAM7 are differentially expressed in normal tissues and oppositely deregulated in hyperplastic colorectal polyps and early adenomas. Am J Pathol. 2000;156:595–605. doi: 10.1016/S0002-9440(10)64764-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Albarran-Somoza B, Franco-Topete R, Delgado-Rizo V, Cerda-Camacho F, Acosta-Jimenez L, Lopez-Botet M, Daneri-Navarro A. CEACAM1 in cervical cancer and precursor lesions: association with human papillomavirus infection. J Histochem Cytochem. 2006;54:1393–1399. doi: 10.1369/jhc.6A6921.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cury PM, Butcher DN, Fisher C, Corrin B, Nicholson AG. Value of the mesothelium-associated antibodies thrombomodulin, cytokeratin 5/6, calretinin, and CD44H in distinguishing epithelioid pleural mesothelioma from adenocarcinoma metastatic to the pleura. Mod Pathol. 2000;13:107–112. doi: 10.1038/modpathol.3880018. [DOI] [PubMed] [Google Scholar]
- Emberley ED, Alowami S, Snell L, Murphy LC, Watson PH. S100A7 (psoriasin) expression is associated with aggressive features and alteration of Jab1 in ductal carcinoma in situ of the breast. Breast Cancer Res. 2004;6:R308–15. doi: 10.1186/bcr791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emberley ED, Niu Y, Njue C, Kliewer EV, Murphy LC, Watson PH. Psoriasin (S100A7) expression is associated with poor outcome in estrogen receptor-negative invasive breast cancer. Clin Cancer Res. 2003;9:2627–2631. [PubMed] [Google Scholar]
- Ordonez NG. Value of cytokeratin 5/6 immunostaining in distinguishing epithelial mesothelioma of the pleura from lung adenocarcinoma. Am J Surg Pathol. 1998;22:1215–1221. doi: 10.1097/00000478-199810000-00006. [DOI] [PubMed] [Google Scholar]
- Tsuda H, Birrer MJ, Ito YM, Ohashi Y, Lin M, Lee C, Wong WH, Rao PH, Lau CC, Berkowitz RS, Wong KK, Mok SC. Identification of DNA copy number changes in microdissected serous ovarian cancer tissue using a cDNA microarray platform. Cancer Genet Cytogenet. 2004;155:97–107. doi: 10.1016/j.cancergencyto.2004.03.002. [DOI] [PubMed] [Google Scholar]
- Gibbs S, Fijneman R, Wiegant J, van Kessel AG, van De Putte P, Backendorf C. Molecular characterization and evolution of the SPRR family of keratinocyte differentiation markers encoding small proline-rich proteins. Genomics. 1993;16:630–637. doi: 10.1006/geno.1993.1240. [DOI] [PubMed] [Google Scholar]
- King TJ, Fukushima LH, Hieber AD, Shimabukuro KA, Sakr WA, Bertram JS. Reduced levels of connexin43 in cervical dysplasia: inducible expression in a cervical carcinoma cell line decreases neoplastic potential with implications for tumor progression. Carcinogenesis. 2000;21:1097–1109. doi: 10.1093/carcin/21.6.1097. [DOI] [PubMed] [Google Scholar]
- Aasen T, Hodgins MB, Edward M, Graham SV. The relationship between connexins, gap junctions, tissue architecture and tumour invasion, as studied in a novel in vitro model of HPV-16-associated cervical cancer progression. Oncogene. 2003;22:7969–7980. doi: 10.1038/sj.onc.1206709. [DOI] [PubMed] [Google Scholar]
- Steinhoff I, Leykauf K, Bleyl U, Durst M, Alonso A. Phosphorylation of the gap junction protein Connexin43 in CIN III lesions and cervical carcinomas. Cancer Lett. 2006;235:291–297. doi: 10.1016/j.canlet.2005.04.031. [DOI] [PubMed] [Google Scholar]
- Chovanec M, Smetana K, Jr., Plzak J, Betka J, Plzakova Z, Stork J, Hrdlickova E, Kuwabara I, Dvorankova B, Liu FT, Kaltner H, Andre S, Gabius HJ. Detection of new diagnostic markers in pathology by focus on growth-regulatory endogenous lectins. The case study of galectin-7 in squamous epithelia. Prague Med Rep. 2005;106:209–216. [PubMed] [Google Scholar]
- Magnaldo T, Fowlis D, Darmon M. Galectin-7, a marker of all types of stratified epithelia. Differentiation. 1998;63:159–168. doi: 10.1046/j.1432-0436.1998.6330159.x. [DOI] [PubMed] [Google Scholar]
- Saussez S, Kiss R. Galectin-7. Cell Mol Life Sci. 2006;63:686–697. doi: 10.1007/s00018-005-5458-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ueda S, Kuwabara I, Liu FT. Suppression of tumor growth by galectin-7 gene transfer. Cancer Res. 2004;64:5672–5676. doi: 10.1158/0008-5472.CAN-04-0985. [DOI] [PubMed] [Google Scholar]
- Dumic J, Dabelic S, Flogel M. Galectin-3: An open-ended story. Biochimica et Biophysica Acta (BBA) - General Subjects. 2006;1760:616–635. doi: 10.1016/j.bbagen.2005.12.020. [DOI] [PubMed] [Google Scholar]
- Bernerd F, Sarasin A, Magnaldo T. Galectin-7 overexpression is associated with the apoptotic process in UVB-induced sunburn keratinocytes. Proc Natl Acad Sci U S A. 1999;96:11329–11334. doi: 10.1073/pnas.96.20.11329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuwabara I, Kuwabara Y, Yang RY, Schuler M, Green DR, Zuraw BL, Hsu DK, Liu FT. Galectin-7 (PIG1) exhibits pro-apoptotic function through JNK activation and mitochondrial cytochrome c release. J Biol Chem. 2002;277:3487–3497. doi: 10.1074/jbc.M109360200. [DOI] [PubMed] [Google Scholar]
- Demers M, Magnaldo T, St-Pierre Y. A novel function for galectin-7: promoting tumorigenesis by up-regulating MMP-9 gene expression. Cancer Res. 2005;65:5205–5210. doi: 10.1158/0008-5472.CAN-05-0134. [DOI] [PubMed] [Google Scholar]
- Rescher U, Gerke V. Annexins--unique membrane binding proteins with diverse functions. J Cell Sci. 2004;117:2631–2639. doi: 10.1242/jcs.01245. [DOI] [PubMed] [Google Scholar]
- Choi J, Chang JS, Song MS, Ahn BY, Park Y, Lim DS, Han YS. Association of hepatitis B virus polymerase with promyelocytic leukemia nuclear bodies mediated by the S100 family protein p11. Biochem Biophys Res Commun. 2003;305:1049–1056. doi: 10.1016/S0006-291X(03)00881-7. [DOI] [PubMed] [Google Scholar]
- Vaarala MH, Porvari KS, Kyllonen AP, Mustonen MV, Lukkarinen O, Vihko PT. Several genes encoding ribosomal proteins are over-expressed in prostate-cancer cell lines: confirmation of L7a and L37 over-expression in prostate-cancer tissue samples. Int J Cancer. 1998;78:27–32. doi: 10.1002/(SICI)1097-0215(19980925)78:1<27::AID-IJC6>3.0.CO;2-Z. [DOI] [PubMed] [Google Scholar]
- Zhang L, Zhou W, Velculescu VE, Kern SE, Hruban RH, Hamilton SR, Vogelstein B, Kinzler KW. Gene expression profiles in normal and cancer cells. Science. 1997;276:1268–1272. doi: 10.1126/science.276.5316.1268. [DOI] [PubMed] [Google Scholar]
- Wong YF, Selvanayagam ZE, Wei N, Porter J, Vittal R, Hu R, Lin Y, Liao J, Shih JW, Cheung TH, Lo KW, Yim SF, Yip SK, Ngong DT, Siu N, Chan LK, Chan CS, Kong T, Kutlina E, McKinnon RD, Denhardt DT, Chin KV, Chung TK. Expression genomics of cervical cancer: molecular classification and prediction of radiotherapy response by DNA microarray. Clin Cancer Res. 2003;9:5486–5492. [PubMed] [Google Scholar]
- Barrett T, Troup DB, Wilhite SE, Ledoux P, Rudnev D, Evangelista C, Kim IF, Soboleva A, Tomashevsky M, Edgar R. NCBI GEO: mining tens of millions of expression profiles--database and tools update. Nucleic Acids Res. 2007;35:D760–5. doi: 10.1093/nar/gkl887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Disbrow GL, Hanover JA, Schlegel R. Endoplasmic reticulum-localized human papillomavirus type 16 E5 protein alters endosomal pH but not trans-Golgi pH. J Virol. 2005;79:5839–5846. doi: 10.1128/JVI.79.9.5839-5846.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang CH, Tsai LC, Chen ST, Yuan CC, Hung MW, Hsieh BT, Chao PL, Tsai TH, Lee TW. Radioimmunotherapy and apoptotic induction on CK19-overexpressing human cervical carcinoma cells with Re-188-mAbCx-99. Anticancer Res. 2005;25:2719–2728. [PubMed] [Google Scholar]
- Kell B, Jewers RJ, Cason J, Best JM. Cellular proteins associated with the E5 oncoprotein of human papillomavirus type 16. Biochem Soc Trans. 1994;22:333S. doi: 10.1042/bst022333s. [DOI] [PubMed] [Google Scholar]
- Kal AJ, van Zonneveld AJ, Benes V, van den Berg M, Koerkamp MG, Albermann K, Strack N, Ruijter JM, Richter A, Dujon B, Ansorge W, Tabak HF. Dynamics of gene expression revealed by comparison of serial analysis of gene expression transcript profiles from yeast grown on two different carbon sources. Mol Biol Cell. 1999;10:1859–1872. doi: 10.1091/mbc.10.6.1859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ftp://ftp.ncbi.nih.gov/refseq/release/viral/
- http://http:www.ncbi.nlm.nih.gov/entrez/viewer.fcgi?db=nucleotide&val=333026
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental Table 1. Increased Expression in CIN III (>15 tpm and >2 fold change)
Supplemental Table 2. Decreased expression in CIN III (≥15 tpm and ≥2 fold change)
Supplemental Figure 1. Functional annotation of all genes expressed in Normal and CIN III lesions (Supplemental Figure A and B, respectively). Tags expressed >15 TPM included.
Supplemental Table 3. Cervical Specimen Description based on LEEP cone biopsy Pathology Report


