Abstract
Glioma constitutes the most frequent brain tumour in man with glioblastoma as the most prevalent and malignant type. The average survival time of less than 16 months underlines the need for improvements in diagnosis and therapy. Here, we report the identification of a novel antigen termed glioma-expressed antigen 2 (GLEA2) causing a frequent immune response in glioma patients. Screening of 450 000 clones from a glioblastoma lambda zap expression library with autologous patient serum revealed a group of five serum-positive clones sharing a high sequence homology. Further sequence analysis showed a sequence homology to a hepatocellular carcinoma associated antigen 58 (HCA58). We localized the novel HCA homologous gene termed glioma-expressed antigen 2 (GLEA2) on chromosome 20 by somatic cell hybrid panel mapping. Using allogenic sera from 39 glioblastoma patients, we found an immune response against GLEA2 in 17 patients (43%). In addition, screening with allogenic sera from other glioma patients revealed GLEA2 directed antibodies in two out of five pilocytic astrocytomas and in one out of two astrocytomas. Unrelated tumour sera revealed no immune response and sera from healthy persons showed an immune response in two out of 14 cases (14%). Northern blot hybridization and RT-PCR showed ubiquitous GLEA2 gene expression in glioma and normal tissues. The novel HCA homologous gene, GLEA2, appears to induce a frequent immune response in glioma. In the light of the lack of useful glioma markers, it appears reasonable to consider GLEA2 as a potential future diagnostic marker.
Keywords: antigen, glioma, immune response
Introduction
Astrocytomas are the most common neoplasms of the central nervous system comprising more than 60% of primary brain tumours. Glioblastoma multiforme (GBM) is the most malignant of these tumours with an average survival time of less than 16 months and a five-year survival time of approximately 5% [1]. Cytogenetic and molecular genetic studies have permitted first insights into the complex genetic changes that are involved in the initiation and progression of human glioma [2]. The genetic alterations include gains and losses of partial or entire chromosomes indicating several tumour suppressor genes and oncogenes interacting in glioma development [2,3]. The p53 gene is the most frequently mutated tumour suppressor gene, and the EGFR gene is the most frequently amplified gene in astrocytic tumours [4,5]. However, as yet cytogenetic and molecular genetic studies have not unambiguously identified specific genetic aberrations as markers for the clinical behaviour of glioma.
There are numerous glioma-associated antigens, which are characterized by a differential expression in glioma versus normal cells. The majority of the glioma-associated antigens, however, have been found to be of minor therapeutic and diagnostic utility (for review see [6]). Until recently, several studies focused on the identification of tumour antigens that induce an immune response in the patient [7]. For several oncogenes, antibody responses have been demonstrated in tumour patients including antibodies against the c-myc protein in colorectal carcinoma patients, c-erbB2 in breast carcinoma patients and c-myb in patients with lymphoma [8–10]. Recent studies revealed antibodies against proteins other than oncogene proteins, including the eukaryotic translation initiation factor 4gamma in lung carcinoma and a novel hyaluronidase in meningioma [11].
Here, we report the cDNA cloning and the characterization of a novel antigen which is expressed in glioma and which induces an immune response in more than 43% of all glioma patients.
Materials and methods
CDNA library construction
cDNA was synthesized from poly(A) mRNA of a primary glioblastoma multiforme using oligo dT primers. Second strand synthesis was performed according to the manufacturer's instructions (Stratagene). Following EcoRI adapter ligation and digest with XhoI, the cDNA fragments were size fractionated with Sepharose CL-2B columns and cDNAs > 0·6 kb in length were ligated into ZAP Express™ vector. Packaging was done using Gigapack III Gold packaging extract from Stratagene. The library was amplified once prior to cDNA library screening.
Immunoscreening with patient serum
Blood samples were taken prior to surgery. Serum was isolated from 10 ml blood samples by using serum gel monovettes and stored by − 75°C. Preabsorption of serum with sonicated E. coli XL1 blue MRF′ cells and bacterial cells lysed by a nonrecombinant Zap Express phage was done as described by Heckel et al. 1997 [12]. The final concentration of the serum was 1 : 100 in 1 × TBS, 0·5% (w/v) dry milk and 0·01% thimerosal.
Phages were transfected into E. coli XL1 blue MRF′ cells at a density of 8000 phages per agar plate (Ø14·5 cm) and incubated at 42°C for 4–5 h until phages become visible. Fusion protein expression was induced by applying Duralose UVTM membranes (Stratagene) soaked in 10 mm IPTG on the agar surface. After an additional incubation for 4 h at 37°C, the plates were stored overnight at 4°C. Membranes were removed, washed twice for 15 min in 1 × TBST and blocked with 5% (w/v) dry milk in 1 × TBS, for 1 h. Following three wash steps for 10 min in 1 × TBS the membranes were incubated with the preabsorbed and diluted patient serum for 3·5–4 h. Unbound serum antibodies were removed by three washing steps in 1 × TBS for 10 min each and bound serum antibodies were detected with goat antihuman IgG antibodies conjugated to alkaline phosphatase (Dianova) diluted 1 : 5000 in 1 × TBS containing 5% dry milk followed by incubation in NBT/BCIP colour-substrate solution.
Sequencing and sequence analysis
Plasmid DNA was isolated for sequencing using the Qiagen-Mini kit. Sequencing was performed using vector specific primers close to the cloning site (T7-vector site: 5′ ACC CGG GTG GAA AAT CGA TGG 3′, T3-vector site: 5′ ACA AAA GCT GGA GCT CGC GCG 3′). Sequencing reactions were run on an ABI automated sequencer 373 A for 14 h. Sequence comparison was performed using BLASTN and BLASTX algorithms.
PCR localization of GLEA2 using somatic hybrid panel DNA
PCR was performed on human genomic DNA isolated from peripheral blood, hamster DNA, mouse DNA, two monochromosomal somatic hybrids (NA10791 containing human chromosome 7 and NA10478 containing human chromosome 20) and three polychromosomal somatic hybrids (NA09927 containing DNA from chromosomes 1, 2, 3, 4, 6, 7, 8, 10, 13, 4, 15, 17, 18, 19 and 20; NA09928 containing DNA from chromosomes 2, 3, 5, 6, 8, 10, 13, 14, 15, 17, 19, 21, 22 and Y; NA09931 containing DNA from chromosomes 5, 7, 10, 12, 14, 17, 20, 21 and Y). Primer sequences were as follows: for-5′GCT GGG CTT GTC TGA AGC T3′ and rev-5′ATG GAA TGG AGA AGT CAC TGG 3′. PCR was done for 26 cycles with 1 min denaturation at 94°C, 455 annealing at 58°C, 455 extension at 72°C using 100 ng of genomic DNA from somatic hybrids, genomic DNA from human peripheral blood and genomic DNA from mouse or hamster cell lines, respectively.
Northern blot analysis
Total RNA from frozen tissue samples were isolated using the RNA isolation kit from Stratagene. Fifteen micrograms of RNA from normal brain and glioblastoma samples were run to a 1% formaldehyde gel and blotted on nylon membrane (GeneScreen). Hybridization was carried out in 500 mm phosphate buffer (pH 7·2) as described previously [13]. Insert DNA (50 ng) was labelled with 32P-dATP by the random primer method of Feinberg and Vogelstein, 1984 [14].
Results
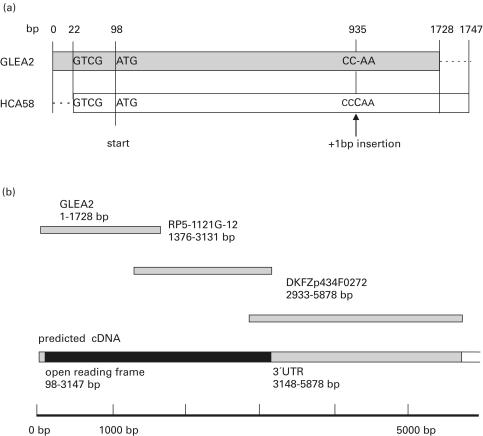
A cDNA expression library was established from a glioblastoma multiforme WHO grade IV. In brief, enriched mRNA was reverse transcribed into cDNA, inserted into the Zap ExpressTM expression vector and expressed in Escherichia coli cells. Recombinant proteins were screened with autologous patient serum to identify antigen–antibody complexes, which are indicative of an immune response against tumour-expressed antigens. Positive clones were isolated and subjected to a second round of screening with the autologous serum. In total, 450 000 clones were analysed in the first screening with autologous patient serum. The second screening and subsequent sequencing of the positive clones revealed a group of five clones that shared high sequence homology. The clones ranging in size from 1·9 kb to 2·4 kb represent a gene that was termed glioma-expressed antigen 2 (GLEA2/AccessNo AF258787). Sequence comparison performed by BLASTN revealed high homology to the hepatocellular carcinoma associated antigen 58 (HCA58). We found sequence identity to HCA58 except for a single nucleotide change disrupting the open reading frame (Fig. 1a). Further BLASTN analysis revealed an overlap to a predicted cDNA from the genomic clone RP5-1121G-12 and the latter revealed an overlap to a cDNA clone designated DKFZp434F0272. Combining those three sequences results in in the partial cDNA sequence for the GLEA2 gene as shown in Fig. 1b. As determined by BLASTX, the predicted amino acid sequence revealed a 36% similarity to a zinc finger motif. The N-terminal region of the protein has a high content (12%) of the basic amino acid lysine (Fig. 2).
Fig. 1.
(a) Sequence comparison between GLEA2 and the hepatocellular carcinoma associated antigen 58 (HCA58). The single nucleotide insertion is indicated. (b) Predicted partial cDNA sequence of the GLEA2 gene combining sequences GLEA2, RP5-1121G-12 and DKFZp434F0272. The resulting open reading frame is indicated.
Fig. 2.
Predicted amino acid sequence of GLEA2 protein. Basic amino acids are shown in bold letters.
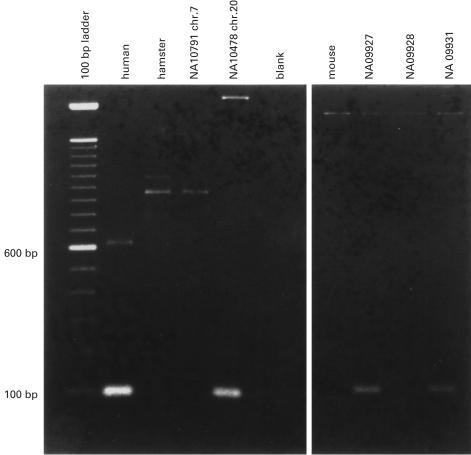
Since numerous chromosomal deviations have been described in glioblastoma, some of which have been associated with tumour progression, we determined the chromosomal location of GLEA2 gene. Somatic hybrid mapping with a panel of monochromosomal and polychromosomal cell hybrids localized the GLEA2 gene on chromosome 20 (Fig. 3). These results have been confirmed by recent results from the Sanger Center Chromosome 20 Mapping Group. GLEA2 shows a 100% homology to the exons of the clones RP-90108 and RP5-1121G12 localized on chromosome 20q11.1–11·23. There is circumstantial evidence that chromosome 20 bears specific genomic alterations associated with glioma development [15].
Fig. 3.
Chromosomal mapping of GLEA2 gene on human chromosome 20 using somatic hybrid DNA. 100 ng of human genomic DNA, mouse DNA, hamster DNA and DNA from the somatic cell hybrids were amplified by PCR for 26 cycles at 58°C. The expected PCR product of 100 basepairs was detected in human genomic DNA, DNA from hybrid NA10478 containing human chromosome 20 and DNA from polychromosomal hybrids NA09927 and NA09931 both of which containing chromosome 20.
To further evaluate the involvement of GLEA2 in the immune response of glioma patients, the clone GLEA2 was screened with allogenic sera from 39 glioblastoma patients. We found an immune response against GLEA2 protein in 17 of these patients (43%), making GLEA2 one of the few antigens known to elicit a frequent immune response in glioblastoma patients. Examples of the serum screening experiments are shown in Fig. 4. There was no obvious correlation between histopathological or clinical grade and the immune response to GLEA2. Survival time was only known for five of the tested glioma patients. For these cases we found no correlation to the GLEA2 immune response. In addition we did not find a correlation of the humoral GLEA2 response between patients with de novo and secondary glioblastoma.
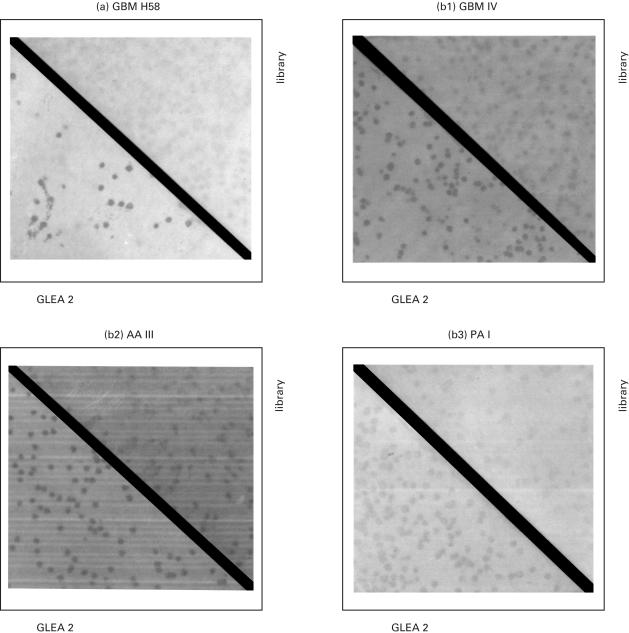
Fig. 4.
Results of immunoscreening using autologous patient serum (a) and heterologous sera from patients with glioma of different grades (b1–3).Phage recombinants are plated at a density of 8000 phages per agar plate (Ø14·5 cm) and hybridized with preabsorbed patient serum. A secondary antibody detected complexes of antigens with antibodies in the serum. Positive clones were isolated and subjected to a second round of screening. The immune response against enriched GLEA2 clones is demonstrated in the lower left part of each figure. The upper right part of each figure shows the hybridization of patient serum against the total library. GBM: Glioblastoma multiforme; AA: astrocytomas; PA: pilocytic astrocytomas.
Using allogenic sera from other glioma patients we found antibodies against GLEA2 protein in two out of five sera from patients with pilocytic astrocytomas and in one out of two patients with astrocytomas grade III (Fig. 4b2 and b3). Results from the screening with allogenic sera from glioma patients were summarized in Table 1. As controls, we used 22 sera from patients with unrelated tumours and from persons with no obvious disease. These results were summarized in Table 2. The frequency of GLEA2 antibodies in sera from glioblastoma patients is 43% and significantly higher than the frequency of the immune response against GLEA2 protein in the control sera (14%) from healthy persons.
Table 1.
Immune response against GLEA2 protein in glioma grade I–IV and oligoastrocytoma
| Tumorsera | Positive reactions |
|---|---|
| Glioblastoma multiforme grade IV | 17/39 (43%) |
| Anaplastic astrocytoma grade III | 1/2 (50%) |
| Astrocytoma grade II | 0/3 (0%) |
| Oligoastrocytoma grade II | 0/1 (0%) |
| Pilocytic astrocytoma | 2/5 (40%) |
Table 2.
Immune response against GLEA2 protein in glioma versus unrelated tumours and healthy persons
| Sera | Positive reactions |
|---|---|
| Glioma | 20/50 (40%) |
| Unrelated tumours and healthy persons | 2/22 (9%) |
| Meningioma | 0/5 (0%) |
| Squamous cell lung carcinoma | 0/3 (0%) |
| Healthy persons | 2/14 (14%) |
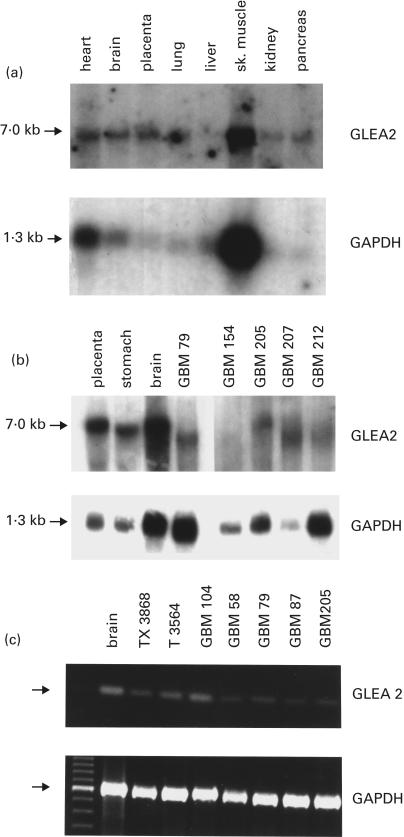
To analyse the expression of the GLEA2 gene, we performed Northern blot hybridization using RNA from glioblastoma and anaplastic astrocytoma. As a control, we included RNA from different normal human tissues in the same Northern blot experiments. GAPDH was used for normalization. Except for glioblastoma 154, we found GLEA2 gene expression in all tissues tested (Fig. 5b).
Fig. 5.
GLEA2 expression in RNA from normal tissues and gliomas. (a) Human multiple tissue Northern blot for GLEA2 (upper panel) and GAPDH (lower panel). (b) Northern hybridization using a GLEA2 specific probe (upper panel) and a GAPDH probe (lower panel). (c) RT-PCR using GLEA2 gene specific primers (upper panel) and GAPDH specific primers (lower panel).
The transcript size was 7·0 kilobases in normal tissues as well as in glioma tissues. Variations in the molecular size of the GLEA2 transcripts detected in stomach, placenta, brain and glioblastoma samples were caused by loading variations and do not reflect GLEA2 isoforms. These data were consistent with the results of a multiple tissue Northern blot experiment also showing ubiquitous GLEA2 gene expression, with muscle tissue having the strongest expression (Fig. 5a). Unfortunately, there was not sufficient RNA for Northern blot hybridizations of those tumours that were derived from patients recruited for the serum analysis including the original glioblastoma that was used to establish the expression library. The expression data were corroborated by RT-PCR analysis showing GLEA2 gene expression in glioma and normal tissues (Fig. 5c). The expression data demonstrate that the immune response against GLEA2 is not associated with an overexpression of GLEA2 transcripts and is likely to result from a different mechanism as discussed below.
Discussion
In our study, we report a novel glioma-expressed antigen that induces an immune response in more than 43% of all patients. The novel antigen showed a homology to the hepatocellular carcinoma associated antigen, which was recently reported. Unfortunately, for the time being no further information is available about the nature of this gene and its possible role in hepatocellular carcinoma. As for the potential function of the GLEA2 protein, the predicted amino acid sequence contains a high percentage of basic amino acids, which has previously been associated with DNA binding properties of the CTF transcription replication factor [16].
There are only very few antigens that cause an immune response with comparable frequency in glioma patients, specifically in patients with low grade glioma [6,17]. A well-known example for an immunogenic antigen in glioma is a deletion variant of the epidermal growth factor receptor (EGFR) which is frequently overexpressed in glioma. EGFR has been suggested as a target for therapeutic intervention [18,19]. Likewise, recombinant antibodies with high tumour specificity could be designed against GLEA2 protein. The high frequency of immune responses in glioma patients may lend this gene for monitoring disease and indicating prognosis. However, the immune response in individuals without obvious tumour formation has to be understood prior to using GLEA2 as a marker or even a target for therapy. The results from these normal control sera experiments have to be interpreted cautiously without having any information about a possible disease history of the donors. In addition, screening tumour-expressed antigens with a high number of normal sera frequently reveals immune responses at low frequencies in normal sera as indicated by our preliminary studies (data not shown).
A variety of mechanisms have been discussed that may cause an immune response against tumour antigens, including overexpression, nucleotide changes of the gene and post-translational modifications of the protein. Our expression data identified a GLEA2 transcript of 7·0 kb, which was found in the majority of glioma, and in all human control tissues. These data suggest that a GLEA2 gene overexpression does not cause the immune response. However, our study does not address the protein expression pattern that may be different from the RNA expression pattern. Without having analysed the protein expression of GLEA2 using a specific antibody, a possible link between GLEA2 expression and immune response cannot be completely ruled out. As for the nucleotide changes, we did not find differences between the hepatocellular carcinoma associated antigen and GLEA2 in the regions that were homologous between both genes. Another possibility that has to be discussed as the cause for the immunogenicity of GLEA2 is a change in the post-translational modification of the GLEA2 protein. However, we demonstrated the immune response against GLEA2 using recombinantly expressed GLEA2 in bacterial cells. Since bacteria do not modify proteins the same way as higher eukaryotic cells, and since the patient sera autoantibodies are able to detect the bacterial expressed proteins, our results do not support this hypothesis. A very different concept, which explains the induction of the immune response under a ‘geographical’ view, has been put forward by Zinkernagel and coworkers [20]. According to this hypothesis induction of immune reactivity occurs in a dose and time specific manner. To fulfil these conditions, GLEA2 should be presented in the draining lymph nodes of the brain (the cervical lymph nodes) after having been in contact with blood vessels and the perivascular space in the brain. As astrocytic tumours show an affected blood brain barrier, presentation of GLEA2 in the cervical lymph nodes is conceivable.
Since glioblastoma patients have anomalies in the immune response due to tumour-derived immunosuppressive molecules like TGF-β, IL-10 and prostaglandins, it would be interesting whether the humoral response is different in patients prior to and shortly after tumour removal. In our study blood samples were only taken prior to surgery and unfortunately we were not able to analyse blood samples taken shortly after tumour removal. In fact this question remains speculative since an incomplete resurrection of glioblastoma multiforme in the prevailing number of cases further complicates this analysis.
The gene encoding the GLEA2 antigen was mapped on chromosome 20. Molecular studies indicate at least three genetic pathways leading to the formation of glioblastoma multiforme. For none of these pathways, however, do alterations of chromosome 20 play a pivotal role as concluded from cytogenetic analysis. Recent comparative genomic hybridization revealed amplifications on chromosome 20 in glioblastoma (data not shown). It remains to be seen whether the chromosomal localization of the GLEA2 gene coincides with the amplification unit.
In summary the implications of our study are three-fold. First, we identified a novel immunoreactive antigen expressed in glioma. Second, the antigen induced an immune response in more than 43% of glioma lending this gene for diagnosis and therapy of glioma. Third, the frequent immune response against GLEA2 protein is likely not to be caused by elevated expression of GLEA2 gene.
Acknowledgments
This work was supported by a Grant from the Deutsche Forschungsgemeinschaft (SFB399 A1). A.-K. Struss and C. P. Pallasch were recipients of a grant from the Graduiertenkolleg ‘Zellulaere Regulation und Wachstum’.
References
- 1.Prados MD, Gutin PH, Philips TL, et al. Highly anaplastic astrocytoma: a review of 357 patients treated between 1977 and 1989. Int J Radiat Oncol Biol Phys. 1992;23:3–8. doi: 10.1016/0360-3016(92)90537-r. [DOI] [PubMed] [Google Scholar]
- 2.Lang FF, Miller DC, Koslow M, Newcomb EW. Pathways leading to glioblastoma multiforme: a molecular analysis of genetic alterations in 65 astrocytic tumors. J Neurosurg. 1994;81:427–36. doi: 10.3171/jns.1994.81.3.0427. [DOI] [PubMed] [Google Scholar]
- 3.Frankel RH, Bayona W, Koslow M, Newcomb EW. p53 mutations in human malignant gliomas. Comparison of loss of heterozygosity with mutation frequency. Cancer Res. 1992;52:1427–33. [PubMed] [Google Scholar]
- 4.Fults D, Brockmeyer D, Tullous MW, Podone CA, Cawthon RM. p53 mutation and loss of heterozygosity on chromosomes 17 and 10 during human astrocytoma progression. Cancer Res. 1992;52:674–79. [PubMed] [Google Scholar]
- 5.Wong AJ, Bigner SH, Bigner DD, Kinzler KW, Hamilton SR, Vogelstein B. Increased expression of the epidermal growth factor receptor gene in malignant gliomas is invariably associated with gene amplification. Proc Natl Acad Sci USA. 1987;84:6899–903. doi: 10.1073/pnas.84.19.6899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kurpad SN, Zhao XG, Wikstrand CJ, et al. Tumor antigens in astrocytic gliomas. GLIA. 1995;15:244–56. doi: 10.1002/glia.440150306. [DOI] [PubMed] [Google Scholar]
- 7.Sahin U, Tureci O, Schmitt H, et al. Human neoplasms elicit multiple specific immune responses in the autologous host. Proc Natl Acad Sci USA. 1996;92:11810–3. doi: 10.1073/pnas.92.25.11810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ben-Mahrez K, Sorokine I, Thierry D, et al. Circulating antibodies against c-myc oncogene product in sera of colorectal cancer patients. Int J Cancer. 1990;46:35–8. doi: 10.1002/ijc.2910460109. [DOI] [PubMed] [Google Scholar]
- 9.Disis ML, Calenoff E, McLaughlin G, et al. Existent T-cell and antibody immunity to HER-2/neu protein in patients with breast cancer. Cancer Res. 1994;54:16–20. [PubMed] [Google Scholar]
- 10.Soronkine I, Ben-Mahrez K, Bracone A, et al. Presence of circulating anti-c-myb oncogene product antibodies in human sera. Int J Cancer. 1991;47:665–9. doi: 10.1002/ijc.2910470506. [DOI] [PubMed] [Google Scholar]
- 11.Brass N, Heckel D, Sahin U, Pfreundschuh M, Sybrecht G, Meese E. Translation initiation factor eIF-4gamma is encoded by an amplified gene and induces an immune response in squamous cell lung carcinoma. Hum Mol Genet. 1997;6:33–9. doi: 10.1093/hmg/6.1.33. [DOI] [PubMed] [Google Scholar]
- 12.Heckel D, Brass N, Fischer U, et al. cDNA cloning and chromosomal mapping of a predicted coiled-coil proline-rich protein immunogenic in meningioma patients. Hum Mol Genet. 1997;6:2031–41. doi: 10.1093/hmg/6.12.2031. [DOI] [PubMed] [Google Scholar]
- 13.Fischer U, Meltzer P, Meese E. Twelve amplified and expressed genes localized in a single domain in glioma. Hum Genet. 1996;98:625–8. doi: 10.1007/s004390050271. 10.1007/s004390050271. [DOI] [PubMed] [Google Scholar]
- 14.Feinberg A, Vogelstein B. Addendum: a technique for radiolabeling DNA restriction fragments to a high specific activity. Anal Biochem. 1984;137:66–7. doi: 10.1016/0003-2697(84)90381-6. [DOI] [PubMed] [Google Scholar]
- 15.Schrock E, Thiel G, Lozanova T, et al. Comparative genomic hybridization of human malignant gliomas reveals multiple amplification sites and nonrandom chromosomal gains and losses. Am J Pathol. 1994;144:1203–18. [PMC free article] [PubMed] [Google Scholar]
- 16.Mitchell PJ, Tjian R. Transcriptional regulation in mammalian cells by sequence-specific DNA binding proteins. Science. 1989;245:371–8. doi: 10.1126/science.2667136. [DOI] [PubMed] [Google Scholar]
- 17.Bigner DD, Brown M, Coleman RE, et al. Phase I studies of treatment of malignant gliomas and neoplastic meningitis with 131I-radiolabeled monoclonal antibodies anti-tenascin 81C6 and anti-chondroitin proteoglycan sulfate Me1-14 F (ab′) 2 – a preliminary report. J Neurooncol. 1995;24:109–22. doi: 10.1007/BF01052668. [DOI] [PubMed] [Google Scholar]
- 18.Nishikawa R, Ji XD, Harmon RC, et al. A mutant epidermal growth factor receptor common in human glioma confers enhanced tumorigenicity. PNAS. 1994;91:7727–31. doi: 10.1073/pnas.91.16.7727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wikstrand CJ, McLendon RE, Friedman AH, Bigner DD. Cell surface localization and density of the tumor-associated variant of the epidermal growth factor receptor, EGFRvIII. Cancer Res. 1997;57:4130–45. [PubMed] [Google Scholar]
- 20.Zinkernagel RM, Ehl S, Oehen S, et al. Antigen localization regulates immune response in a dose- and time dependent fashion: geographical view of immune reactivity. Immunol Rev. 1997;196:199–209. doi: 10.1111/j.1600-065x.1997.tb00969.x. [DOI] [PubMed] [Google Scholar]