Abstract
Approximately 94% of spinal muscular atrophy (SMA) patients lack both copies of SMN1 exon 7. We report our SMA genetic testing experience (total 1281 cases), using SMA linkage analysis (32 families), SMA diagnostic testing by PCR-RFLP (restriction fragment length polymorphism) to detect the homozygous absence of SMN1 exon 7 (and exon 8) (533 cases), and an assay to determine copy number of SMN1 exon 7 (SMN1 gene dosage analysis) (716 cases). SMN1 gene dosage analysis is used for SMA carrier testing as well as for the confirmation of a heterozygous SMN1 deletion in symptomatic individuals who do not lack both copies of SMN1. We conclude that comprehensive SMA testing, including SMN1 deletion analysis, SMN1 gene dosage analysis, and linkage analysis, offers the most complete evaluation of SMA patients and their families.
Autosomal recessive spinal muscular atrophy (SMA) is a disorder that affects motor neurons in the anterior horn of the spinal cord. 1, 2 SMA is the second most common lethal autosomal recessive disease after cystic fibrosis in the western hemisphere, and is clinically classified into three types based on the severity of symptoms. Type I SMA (Werdnig-Hoffmann disease) is the most severe form, type III SMA (Kugelberg-Wellander disease) is the mildest form, and type II SMA is an intermediate form. Localization of the SMA critical region made linkage analysis possible as the first genetic test for SMA. 3, 4, 5
SMA of all three types is associated with deletions and point mutations in the Survival Motor Neuron 1 gene (SMN1 or SMNt). 6 SMN1 and its centromeric homologue, SMN2 (or SMNc), lie within the telomeric and centromeric halves, respectively, of a large inverted repeat on chromosome 5q13. 6 SMN1 and SMN2 coding regions differ in only one base in exon 7. 7 This single-base C-to-T substitution in SMN2 exon 7 affects the activity of an exonic splice enhancer and the splicing pattern of SMN2 mRNA, 8 resulting in a lower level of full-length SMN protein from SMN2 compared to SMN1. 9, 10
Takingadvantage of the fact that approximately 94% of clinically typical SMA patients lack both copies of SMN1 exon 7 (reviewed by Wirth 11 ), a relatively sensitive, cost-effective polymerase chain reaction (PCR) method of testing for SMA was developed. 12 This test, herein referred to as the SMA diagnostic test, is used widely in the United States, including in our laboratory. It detects homozygous deletions of SMN1 exon 7 (and exon 8) and can be used to confirm a diagnosis of SMA or for prenatal testing. 12 However, it has the following limitations: it cannot distinguish the types of SMA, since all three types of SMA typically show a homozygous absence of SMN1 exon 7; it cannot distinguish SMA carriers from non-carriers, since both carriers and non-carriers are expected to have negative results; and a minority of SMA patients who have a small intragenic mutation in one SMN1 allele instead of a homozygous SMN1 deletion will have negative results.
To allow identification of carriers, a method to determine the copy number of SMN1 was originally developed by McAndrew et al 13 and modified as a non-radioisotopic assay in our laboratory. 14 These methods are collectively referred to herein as SMN1 gene dosage analysis. A one-copy SMN1 result confirms carrier status in unaffected individuals or supports a diagnosis of SMA in an individual who has typical symptoms of SMA and does not lack both copies of SMN1 exon 7. In this article, we describe our SMA genetic testing experience, including SMA diagnostic testing, SMN1 gene dosage analysis, and SMA linkage analysis.
Materials and Methods
Subject Samples, DNA Isolation, SMA Diagnostic Test, and Linkage Analysis
Testing was performed in the Molecular Pathology Laboratory of the University of Pennsylvania during the period from April 1992 to June 2000. The subject population referred for SMA genetic testing was widely distributed in the U.S. with the majority being from the Northeast region. Rare samples were from outside the U.S., including Canada, Mexico, and the Middle East. Subjects referred for SMA genetic testing included patients clinically suspected to be affected by SMA, family members of SMA patients, individuals without a family history of SMA (typically those who marry someone with a family history of SMA), and fetuses of parents who had affected children or a family history of SMA. In addition, we tested five fetuses and two neonates with arthrogryposis. Patients were categorized into SMA type I, type II, and type III based on the clinical diagnosis provided by the ordering clinicians or genetic counselors according to standard criteria. 1 If a patient had clinical features that overlapped two types of SMA, then type I-II or type II-III was assigned to such a patient by the referring clinician. Adult-onset SMA, so-called type IV, is not a well-established category and many regard it as a variant of type III. Therefore, we included adult-onset SMA in the type III category.
Genomic DNA was extracted from peripheral blood, prenatal samples [chorionic villous sampling (CVS) tissues and amniocytes, before and/or after culture], or fresh frozen tissue using the Puregene DNA Isolation Kit (Gentra Systems, Inc., Minneapolis, MN) and from formalin-fixed, paraffin-embedded tissue sections using the QIAmp Tissue Kit (Qiagen, Inc., Valencia, CA). The SMA diagnostic test for homozygous deletions of SMN1 exon 7 (and exon 8 until March 1997) was performed as described previously. 12 Prenatal samples were analyzed in duplicate since July 1998. SMA linkage analysis was performed as described previously. 3, 4, 15
SMN1 Gene Dosage Analysis and Genetic Risk Assessment
SMN1 gene dosage analysis was originally developed by McAndrew et al 13 and modified as a non-radioisotopic assay in our laboratory. 14 All samples were analyzed in duplicate. Briefly, exon 7 of SMN1 and SMN2 was co-amplified with exon 4 of the cystic fibrosis transmembrane regulator gene (CFTR), an SMN1 internal standard, and a CFTR internal standard using fluorescently labeled primers. PCR products were then digested with DraI, which digests only SMN2 exon 7 PCR products, followed by analysis on the ABI 373a Sequencer (Applied Biosystems Inc., Foster City, CA) or the ABI 310 Genetic Analyzer (Applied Biosystems Inc.). 16
Genetic risk assessment was performed routinely as an integral part of SMA genetic testing in our laboratory. Remaining risk of being a carrier was analyzed and reported in cases with two or more copies of SMN1 by gene dosage analysis, using estimates of various SMN1 normal and disease allele frequencies (Ogino S, Leonard DGB, Rennert H, Ewens WJ, Wilson RB, submitted).
Sequence Analysis for SMN1 PCR Products
The PCR products from five samples showing the expected SMN1 peak and a one-base-larger peak on ABI 310 or 373a electropherograms by SMN1 gene dosage analysis (Figure 1) were sequenced. First, PCR was performed by the same method used for the SMA diagnostic test. Then, PCR products were purified from the SMN1 bands on agarose gels and sequenced with the SMN forward primer, R1116 and the SMN reverse primer, X-7 Dra 12 using the BigDye Terminator Reaction Kit (Applied Biosystems Inc.) and the ABI 310 Genetic Analyzer (Applied Biosystems Inc.).
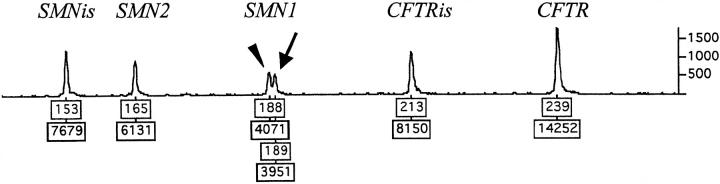
Figure 1.
An example of an ABI 373a electropherogram showing a one-base-larger SMN1 peak (actual size 188 bp, shown as 189 bp, arrow) in addition to the expected SMN1 peak (actual size 187 bp, shown as 188 bp, arrowhead), each corresponding to one SMN1 copy by gene dosage analysis. Each peak corresponds to PCR products of an SMN internal standard (SMNis), SMN2 genomic sequence (SMN2), SMN1 genomic sequence (SMN1, with a doublet), a CFTR internal standard (CFTRis), and CFTR genomic sequence (CFTR), from left to right, respectively. Numbers under each peak represent approximate product size in bp (top) and fluorescence intensity measured as an area under curve (bottom).
Results
SMA Diagnostic Test: Symptomatic and Prenatal Testing
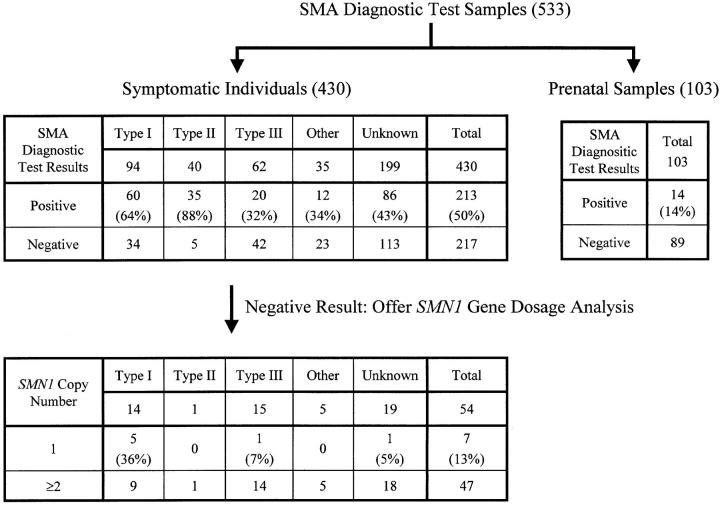
A total of 533 SMA diagnostic tests were performed: 430 for symptomatic individuals with a suspected diagnosis of SMA and 103 for prenatal diagnosis (Figure 2) . Of the 430 tests on symptomatic individuals, 213 were positive (a homozygous deletion of exon 7). Of the 213 positive patients, 16 were positive only for an exon 7 deletion but negative for exon 8, whereas 110 were positive for deletions of both exon 7 and exon 8. The other 87 patients were tested only for an exon 7 deletion with positive results. There were no symptomatic individuals with an exon 8 deletion without an exon 7 deletion. When the subjects were divided into SMA types, type II patients had the highest percentage of positive results (88%; 35 of 40), followed by type I (64%; 60 of 94), type I-II (44%; 7 of 16), type II-III (33%; 4 of 12) and type III (32%; 20 of 62). SMA type was unknown in 199 cases (46%). Reasons for unknown SMA type included a lack of description by the health care provider and atypical symptoms precluding classification by clinicians.
Figure 2.
A flow chart of the results of SMA diagnostic tests and SMN1 gene dosage analyses for symptomatic and prenatal cases.
The 103 samples for prenatal SMA diagnostic testing consisted of 31 CVS samples without prior culture, 10 cultured cell samples derived from CVS, 42 cultured cell samples derived from amniotic fluid, 12 amniotic fluid samples without prior culture, 3 cultured cell samples not specified as CVS or amniotic fluid derived, and 5 other prenatal samples. In all, 14 of 103 prenatal cases were positive and 89 were negative for a homozygous SMN1 deletion by the SMA diagnostic test (Figure 2) . To exclude significant maternal DNA contamination, polymorphic short tandem repeat (STR) analysis was performed (data not shown).
SMN1 Gene Dosage Analysis
The primary purpose of SMN1 gene dosage analysis was SMA carrier testing, which was performed on a total of 663 asymptomatic individuals (Table 1) . Of these individuals, 191 individuals had no family history of SMA and 472 had a family history of suspected or definitive chromosome 5q13-linked SMA. Among individuals without a family history, the percentage of individuals with a one-copy result was 1.6%, which is comparable to the ∼2% carrier frequency in the general population. Among parents whose affected children lacked both copies of SMN1 (0-copy SMN1) (n = 118), 110 had one copy of SMN1, confirming a carrier status. Of the remaining eight parents who had two copies of SMN1, five had two copies of SMN1 on one chromosome 5 and a deletion mutation of SMN1 on the other chromosome 5, also referred to as the 2 + 0 genotype. 14 Two other parents with two copies of SMN1 had one copy of SMN1 on each chromosome 5 (1 + 1 genotype) and had passed a de novo deletion mutation associated with a crossover to their affected children. The 2 + 0 or 1 + 1 genotype was confirmed by a combination of SMN1 gene dosage analysis and SMA linkage analysis. 14
Table 1.
SMN1 Gene Dosage Analyses for SMA Carrier Testing
| SMN1 copy number | Number of cases | |||||||
|---|---|---|---|---|---|---|---|---|
| No family history | Family history of SMA | |||||||
| Parent of a child with SMA and the child has | Other family history | Subtotal | Total | |||||
| 0-copy SMN1 | 1-copy SMN1 | Negative diagnostic test | Not been tested | |||||
| 1 | 3 | 110 | 0 | 2 | 39 | 117 | 268 | 271 |
| 2 | 174 | 8* | 3 | 6 | 8 | 157 | 182 | 356 |
| 3 | 12 | 0 | 1 | 0 | 1 | 20 | 22 | 34 |
| 4 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 2 |
| % 1-copy results | 1.6% | 93% | 0% | 25% | 81% | 40% | 57% | 48% |
| Total | 191 | 118 | 4 | 8 | 48 | 294 | 472 | 663 |
Results are listed by type of family history.
Five individuals showed the “2 + 0” genotype and two showed the “1 + 1” genotype. The “2 + 0” genotype refers to two SMN1 exon 7 copies on one chromosome 5 and an exon 7 deletion on the other chromosome 5. The “1 + 1” genotype refers to one SMN1 exon 7 copy on each chromosome 5. The other individual’s genotype could not be determined because additional family members were not available for testing.
Three asymptomatic individuals from independent families were shown to have one SMN1 copy and no SMN2 copies. Therefore, only one copy of SMN1, without any SMN2 copies is apparently sufficient for a normal phenotype. All of them had an affected child with a homozygous SMN1 deletion, but with SMN2 copies present.
The second purpose of SMN1 gene dosage analysis was for the detection of heterozygous SMN1 deletions to support a diagnosis of SMA in symptomatic individuals with a negative SMA diagnostic test result (Figure 2) . For this purpose, we tested a total of 54 samples that initially were negative for homozygous SMN1 deletions. The percentage of heterozygous SMN1 deletions (one copy of SMN1) was highest for type I SMA patients (36%; 5 of 14) but was low for all other SMA types.
We also performed SMN1 gene dosage analysis on four prenatal samples from three different families, where the affected child was negative by the SMA diagnostic test and subsequently was found to have one copy of SMN1 by gene dosage analysis. These prenatal SMN1 gene dosage analyses were combined with linkage analyses (see SMA Linkage Analysis). Two prenatal samples showed one copy of SMN1 and the other two showed two copies of SMN1.
SMA Linkage Analysis
Linkage analysis was performed for a total of 32 families, with 2 to 12 members per family. For 22 families, linkage analysis was performed before the SMA diagnostic test was developed. However, linkage analysis continues to be useful even with the availability of the SMA diagnostic testing and SMN1 gene dosage analysis. Linkage analysis is particularly important for families with an affected child who has a homozygous deletion of SMN1 exon 7 and a parent with two copies of SMN1. 14 We used linkage analysis in combination with SMN1 gene dosage analysis in six such families. Four of the six parents with two copies of SMN1 were carriers with a deletion allele plus two copies of SMN1 on the other chromosome 5, and the remaining two parents had two copies of SMN1 and their children had a de novo deletion associated with a cross-over event. Linkage analysis was also used in five cases in four independent families with an affected child having a non-deletion mutation in one SMN1 allele, to determine whether a new fetus would be likely to inherit the same disease alleles as the affected child. In three cases both the maternal and paternal haplotypes of the fetus were shared with the affected child, in one case only the paternal haplotype was shared with the affected child, and in the remaining one case neither haplotype was shared with the affected child.
SMA Diagnostic Testing in Unusual Cases
One unusual case was a two-month-old white male who was diagnosed with sudden infant death syndrome (SIDS) at autopsy. However, the parents reported decreased movements of extremities and choking. After completion of the autopsy, SMA diagnostic testing showed a homozygous deletion of SMN1 exon 7.
We tested seven samples from five fetuses and two neonates with clinical arthrogryposis. Only one case (a 6-week-old neonate) had a homozygous SMN1 exon 7 deletion. One of the six negative cases, subsequently tested by SMN1 gene dosage analysis, had two copies of SMN1.
Sequence Analysis of SMN1 PCR Products
Five samples showed the expected size SMN1 peak (actual size 187 bp) and a one-base-larger peak (actual size 188 bp) on ABI 310 or 373a electropherograms by SMN1 gene dosage analysis (Figure 1) . The clinical features of these five cases included two symptomatic individuals (a 29-year-old pregnant woman with type III SMA and a 2-year-old boy with type II SMA) and three without symptoms. The two symptomatic individuals were initially tested for SMN1 homozygous deletions with negative results and subsequently tested by SMN1 gene dosage analysis. The three asymptomatic individuals included two with family histories of SMA and one without a family history of SMA. These samples were analyzed by sequencing of the SMN1 exon 7 PCR product. All five cases showed the presence of two alleles, which differed in length by one base due to a single thymidine insertion in the pre-existing polythymidine tract (8T) in SMN1 intron 6.
Discussion
Our laboratory has been using homozygous deletion analysis and SMN1 gene dosage analysis for SMN1 exon 7 only and not for exon 8. Our rationale is based on the critical functional importance of exon 7 in splicing. 17 A gene conversion event, in which a single nucleotide C→T change in the exon 7 sequence (from SMN1 to SMN2), but without a change in the SMN1 exon 8 sequence, has been shown to be associated with SMA 18, 19 (reviewed by Burghes 20 ). The significance of a homozygous deletion of only exon 8 without an exon 7 homozygous deletion is unknown. Two patients (type II and type III) with an SMN1 exon 8 homozygous deletion but exon 7 present have been reported. 21 However, the possibility of missing a deletion located between the distal half of exon 7 and exon 8 cannot be excluded, because the method used was that of van der Steege et al, 12 which cannot detect a deletion limited to the distal half of SMN1 exon 7. Therefore, an SMN1 allele with a deletion limited to exon 8 but with an intact exon 7 does not necessarily constitute a disease allele.
In our laboratory, 88% of clinically-defined type II SMA patients lacked both copies of SMN1, compared to 64% of type I patients and 32% of type III patients. The reported percentages of SMA patients who lack both copies of SMN1 are 96% (400 of 418) for type I SMA, 94% (241 of 257) for type II, and 86% (185 of 216) for type III (reviewed by Wirth 11 ). These studies included only patients selected by the standard criteria with clinical manifestations typical of each type of SMA. By contrast, our results were derived from patients tested in a clinical molecular pathology laboratory. Many of the patients we tested had atypical clinical features, suggesting that their diseases might be similar to SMA but unrelated to SMN1 mutations. Our results suggest that hypotonia with intermediate onset and severity is more likely due to 5q13-linked SMA than early or late onset hypotonia. On the other hand, among patients with negative SMA diagnostic test results whom we tested by SMN1 gene dosage analysis, suspected type I patients had the highest percentage (36%) of one-copy results. Our results suggest that negative results in SMA diagnostic testing on possible type III SMA patients significantly decreases the likelihood of 5q13-linked SMA, as only 1 of 14 suspected type III SMA patients without a homozygous SMN1 deletion who were subsequently tested by SMN1 gene dosage analysis had one copy of SMN1. The positive predictive value of a one-copy SMN1 result in a symptomatic individual depends largely on the degree of clinical diagnostic certainty, due to the relatively high prevalence (∼2%) of carriers with one SMN1 copy in the general population. Therefore, we have not recommended SMN1 gene dosage analysis unless clinical suspicion for SMA remains high even after a negative SMA diagnostic test.
We obtained a positive result in 14% (14 of 103) of prenatal samples by the SMA diagnostic test. This is significantly lower than the 25% positivity we would have expected by simple Mendelian genetics if we had tested only prenatal samples from carrier couples with heterozygous SMN1 deletions. However, we also tested prenatal samples from carrier parents with new partners whose genetic status was unknown, as well as samples from parents whose previous affected children had not been tested for SMN1 deletions. These prenatal samples are expected to have a lower positive rate than those from carrier parents with confirmed SMN1 deletion mutations. Negative selection by early miscarriage of fetuses with lethal mutations, such as contiguous gene deletions that comprise deletions of both SMN1 and SMN2 in the 5q13 region, may also have contributed to the lower-than-expected percentage of positive results. There have been no documented cases of SMA with homozygous deletions of both SMN1 and SMN2. Homozygous deletion of the mouse SMN homolog (Smn −/−) causes embryonic lethality, and introduction of the human SMN2 gene partially rescues embryonic lethality in mice with symptoms resembling human SMA. 22, 23
There has been no report of a homozygous SMN1 deletion in an infant who was suspected to have SIDS. One report described an infant who almost suffered sudden death at the age of 7 weeks due to diaphragmatic paralysis and was later diagnosed to have type I SMA. 24 Diaphragmatic paralysis and resulting respiratory distress was an initial manifestation of SMA in reports of six infants. 25, 26, 27 However, the status of SMN1 in these infants has not been determined. We speculate that the cause of some clinically unexplained infant deaths may be SMA. Strictly speaking, the diagnosis of SIDS is inappropriate if a homozygous SMN1 mutation confirms the diagnosis of SMA. More studies are necessary to draw a definitive conclusion about the frequency of undiagnosed SMA being misdiagnosed as SIDS.
Arthrogryposis represents a group of heterogeneous syndromes, all of which have in common joint contractures that are present at birth. 28 The major clinical form, arthrogryposis multiplex congenita (AMC), is caused by decreased fetal movements in utero due to a large number of conditions, including neuropathies, myopathies, connective tissue diseases, and conditions that limit the space within the uterus. 29 Burglen et al 30 showed that 6 of 12 patients with “neurogenic AMC” had homozygous deletions of SMN1. Bingham et al 31 showed that two of four patients with arthrogryposis, not otherwise specified, had homozygous deletions of SMN1. By contrast, Rudnik-Schöneborn et al 32 showed that two of two “neurogenic arthrogryposis” patients did not have homozygous SMN1 deletions. We detected a homozygous deletion of SMN1 in only one of seven cases referred for “arthrogryposis” (one negative case was AMC) and tested by the SMA diagnostic test. However, comparing these data is problematic because of the heterogeneous etiologies for arthrogryposis and the lack of uniform inclusion criteria in these studies.
We report five unrelated individuals who have two alleles that differ in length by a single base due to a single nucleotide thymidine insertion in a pre-existing polythymidine tract (8T) in SMN1 intron 6. Two were symptomatic (one with type II SMA and the other with type III SMA) and the other three were asymptomatic. Interestingly, all five of the samples contained the expected “normal 8T allele” SMN1 peak in addition to the one-base-larger 9T SMN1 peak, and each peak corresponded to one copy of SMN1 by gene dosage analysis. Despite careful scrutiny, we were unable to identify any individuals with only the 9T SMN1 peak without the normal 8T SMN1 peak. The biological significance of this single nucleotide insertion is unknown. It may merely be a polymorphism with no clinical significance.
In conclusion, comprehensive SMA testing, including SMA deletion analysis, SMN1 dosage analysis and linkage analysis, offers the most complete evaluation of SMA patients and their families at this time.
Acknowledgments
We thank Patricia E. McAndrew and Thomas W. Prior for generously providing us with the plasmids containing the SMN and CFTR internal standard inserts; and Cynthia Turino, Juliane Mills, Treasa Smith, Hui-Jia Dong, Ke-Lian Chen, Konrad Gueverra, and Anthony Taylor for performing DNA extractions and clinical SMA genetic testing.
Address reprint requests to Robert B. Wilson, M.D., Ph.D., Department of Pathology and Laboratory Medicine, University of Pennsylvania Medical Center, Room 509A, Stellar-Chance Laboratories, 422 Curie Blvd., Philadelphia, PA 19104. E-mail: wilsonr@mail.med.upenn.edu.
Footnotes
Shuji Ogino’s present address is Department of Pathology, Brigham and Women’s Hospital, 75 Francis Street, Boston, MA 02115.
References
- 1.Munsat TL, Davies KE: International SMA consortium meeting (June 26–28, 1992, Bonn, Germany). Neuromuscul Disord 1992, 2:423-428 [DOI] [PubMed] [Google Scholar]
- 2.Zerres K, Wirth B, Rudnik-Schoeneborn S: Spinal muscular atrophy: clinical and genetic correlation. Neuromuscul Disord 1997, 7:202-207 [DOI] [PubMed] [Google Scholar]
- 3.Clermont O, Burlet P, Burglen L, Lefebvre S, Pascal F, McPherson J, Wasmuth JJ, Cohen D, Le Paslier D, Weissenbach J, Lathrop M, Munnich A, Melki J: Use of genetic and physical mapping to locate the spinal muscular atrophy locus between two new highly polymorphic DNA markers. Am J Hum Genet 1994, 54:687-694 [PMC free article] [PubMed] [Google Scholar]
- 4.Soares VM, Brzustowicz LM, Kleyn PW, Knowles JA, Palmer DA, Asokan S, Penchaszadeh GK, Munsat TL, Gilliam TC: Refinement of the spinal muscular atrophy locus to the interval between D5S435 and MAP1B. Genomics 1993, 15:365-371 [DOI] [PubMed] [Google Scholar]
- 5.Melki J, Lefebvre S, Burglen L, Burlet P, Clermont O, Millasseau P, Reboullet S, Benichou B, Zeviani M, Le Paslier D, Cohen D, Weissenbach J, Munnich A: De novo and inherited deletions of the 5q13 region in spinal muscular atrophies. Science 1994, 264:1474-1477 [DOI] [PubMed] [Google Scholar]
- 6.Lefebvre S, Burglen L, Reboullet S, Clermont O, Burlet P, Viollet L, Benichou B, Cruaud C, Millasseau P, Zeviani M, Le Paslier D, Frezal J, Cohen D, Weissenbach J, Munnich A, Melki J: Identification and characterization of a spinal muscular atrophy-determining gene. Cell 1995, 80:155-165 [DOI] [PubMed] [Google Scholar]
- 7.Burglen L, Lefebvre S, Clermont O, Burlet P, Viollet L, Cruaud C, Munnich A, Melki J: Structure and organization of the human survival motor neurone (SMN) gene. Genomics 1996, 32:479-482 [DOI] [PubMed] [Google Scholar]
- 8.Pellizzoni L, Kataoka N, Charroux B, Dreyfuss G: A novel function for SMN, the spinal muscular atrophy disease gene product, in pre-mRNA splicing. Cell 1998, 95:615-624 [DOI] [PubMed] [Google Scholar]
- 9.Monani UR, Lorson CL, Parsons DW, Prior TW, Androphy EJ, Burghes AH, McPherson JD: A single nucleotide difference that alters splicing patterns distinguishes the SMA gene SMN1 from the copy gene SMN2. Hum Mol Genet 1999, 8:1177-1183 [DOI] [PubMed] [Google Scholar]
- 10.Jong YJ, Chang JG, Lin SP, Yang TY, Wang JC, Chang CP, Lee CC, Li H, Hsieh-Li HM, Tsai CH: Analysis of the mRNA transcripts of the survival motor neuron (SMN) gene in the tissue of an SMA fetus and the peripheral blood mononuclear cells of normals, carriers, and SMA patients. J Neurol Sci 2000, 173:147-153 [DOI] [PubMed] [Google Scholar]
- 11.Wirth B: An update of the mutation spectrum of the survival motor neuron gene (SMN1) in autosomal recessive spinal muscular atrophy (SMA). Hum Mutat 2000, 15:228-237 [DOI] [PubMed] [Google Scholar]
- 12.van der Steege G, Grootscholten P, van der Vlies P, Draaijers TG, Osinga J, Cobben JM, Scheffer H, Buys CHCM: PCR-based DNA test to confirm clinical diagnosis of autosomal recessive spinal muscular atrophy. Lancet 1995, 345:985-986 [PubMed] [Google Scholar]
- 13.McAndrew PE, Parsons DW, Simard LR, Rochette C, Ray PN, Mendell JR, Prior TW, Burghes AH: Identification of proximal spinal muscular atrophy carriers and patients by analysis of SMNT and SMNC gene copy number. Am J Hum Genet 1997, 60:1411-1422 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chen KL, Wang YL, Rennert H, Joshi I, Mills JK, Leonard DG, Wilson RB: Duplications and de novo deletions of the SMNt gene demonstrated by fluorescence-based carrier testing for spinal muscular atrophy. Am J Med Genet 1999, 85:463-469 [PubMed] [Google Scholar]
- 15.Wang CH, Kleyn PW, Vitale E, Ross BM, Lien L, Xu J, Carter TA, Brzustowicz LM, Obici S, Selig S, Pavone L, Parano E, Penchaszadeh GK, Munsat T, Kunkel LM, Gilliam TC: Refinement of the spinal muscular atrophy locus by genetic and physical mapping. Am J Hum Genet 1995, 56:202-209 [PMC free article] [PubMed] [Google Scholar]
- 16.Ogino S, Leonard DGB, Rennert H, Gao S, Wilson RB: Heteroduplex formation in SMN gene dosage analysis. J Mol Diagn 2001, 3:150-157 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lorson CL, Androphy EJ: An exonic enhancer is required for inclusion of an essential exon in the SMA-determining gene SMN. Hum Mol Genet 2000, 9:259-265 [DOI] [PubMed] [Google Scholar]
- 18.Hahnen E, Schonling J, Rudnik-Schoneborn S, Zerres K, Wirth B: Hybrid survival motor neuron genes in patients with autosomal recessive spinal muscular atrophy: new insights into molecular mechanisms responsible for the disease. Am J Hum Genet 1996, 59:1057-1065 [PMC free article] [PubMed] [Google Scholar]
- 19.DiDonato CJ, Ingraham SE, Mendell JR, Prior TW, Lenard S, Moxley III RT, Florence J, Burghes AH: Deletion and conversion in spinal muscular atrophy patients: is there a relationship to severity? Ann Neurol 1997, 41:230–237 [DOI] [PubMed]
- 20.Burghes AH: When is a deletion not a deletion? When it is converted. Am J Hum Genet 1997, 61:9-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gambardella A, Mazzei R, Toscano A, Annesi G, Pasqua A, Annesi F, Quattrone F, Oliveri RL, Valentino P, Bono F, Aguglia U, Zappia M, Vita G, Quattrone A: Spinal muscular atrophy due to an isolated deletion of exon 8 of the telomeric survival motor neuron gene. Ann Neurol 1998, 44:836-839 [DOI] [PubMed] [Google Scholar]
- 22.Monani UR, Sendtner M, Coovert DD, Parsons DW, Andreassi C, Le TT, Jablonka S, Schrank B, Rossol W, Prior TW, Morris GE, Burghes AH: The human centromeric survival motor neuron gene (SMN2) rescues embryonic lethality in Smn(−/−) mice and results in a mouse with spinal muscular atrophy. Hum Mol Genet 2000, 9:333-339 [DOI] [PubMed] [Google Scholar]
- 23.Hsieh-Li HM, Chang JG, Jong YJ, Wu MH, Wang NM, Tsai CH, Li H: A mouse model for spinal muscular atrophy. Nat Genet 2000, 24:66-70 [DOI] [PubMed] [Google Scholar]
- 24.Poets C, Heyer R, von der Hardt H, Walter GF: Acute respiratory insufficiency as the initial clinical manifestation of spinal muscular atrophy. Monatsschr Kinderheilkd 1990, 138:157-159 [PubMed] [Google Scholar]
- 25.Sivan Y, Galvis A: Early diaphragmatic paralysis: in infants with genetic disorders. Clin Pediatr 1990, 29:169-171 [DOI] [PubMed] [Google Scholar]
- 26.Mellins RB, Hays AP, Gold AP, Berdon WE, Bowdler JD: Respiratory distress as the initial manifestation of Werdnig-Hoffmann disease. Pediatrics 1974, 53:33-40 [PubMed] [Google Scholar]
- 27.McWilliam RC, Gardner-Medwin D, Doyle D, Stephenson JB: Diaphragmatic paralysis due to spinal muscular atrophy: an unrecognised cause of respiratory failure in infancy? Arch Dis Child 1985, 60:145-149 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Goldberg MJ: Syndrome of orthopaedic importance. Lovell and Winter’s Pediatric Orthopaedics. Edited by RT Morrissy, SL Weinstein. Philadelphia, Lippincott-Raven Publishers, 1996, pp 262–271
- 29.Gordon N: Arthrogryposis multiplex congenita. Brain Dev 1998, 20:507-511 [DOI] [PubMed] [Google Scholar]
- 30.Burglen L, Amiel J, Viollet L, Lefebvre S, Burlet P, Clermont O, Raclin V, Landrieu P, Verloes A, Munnich A, Melki J: Survival motor neuron gene deletion in the arthrogryposis multiplex congenita-spinal muscular atrophy association. J Clin Invest 1996, 98:1130-1132 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Bingham PM, Shen N, Rennert H, Rorke LB, Black AW, Marin-Padilla MM, Nordgren RE: Arthrogryposis due to infantile neuronal degeneration associated with deletion of the SMNT gene. Neurology 1997, 49:848-851 [DOI] [PubMed] [Google Scholar]
- 32.Rudnik-Schoneborn S, Forkert R, Hahnen E, Wirth B, Zerres K: Clinical spectrum and diagnostic criteria of infantile spinal muscular atrophy: further delineation on the basis of SMN gene deletion findings. Neuropediatrics 1996, 27:8-15 [DOI] [PubMed] [Google Scholar]