Abstract
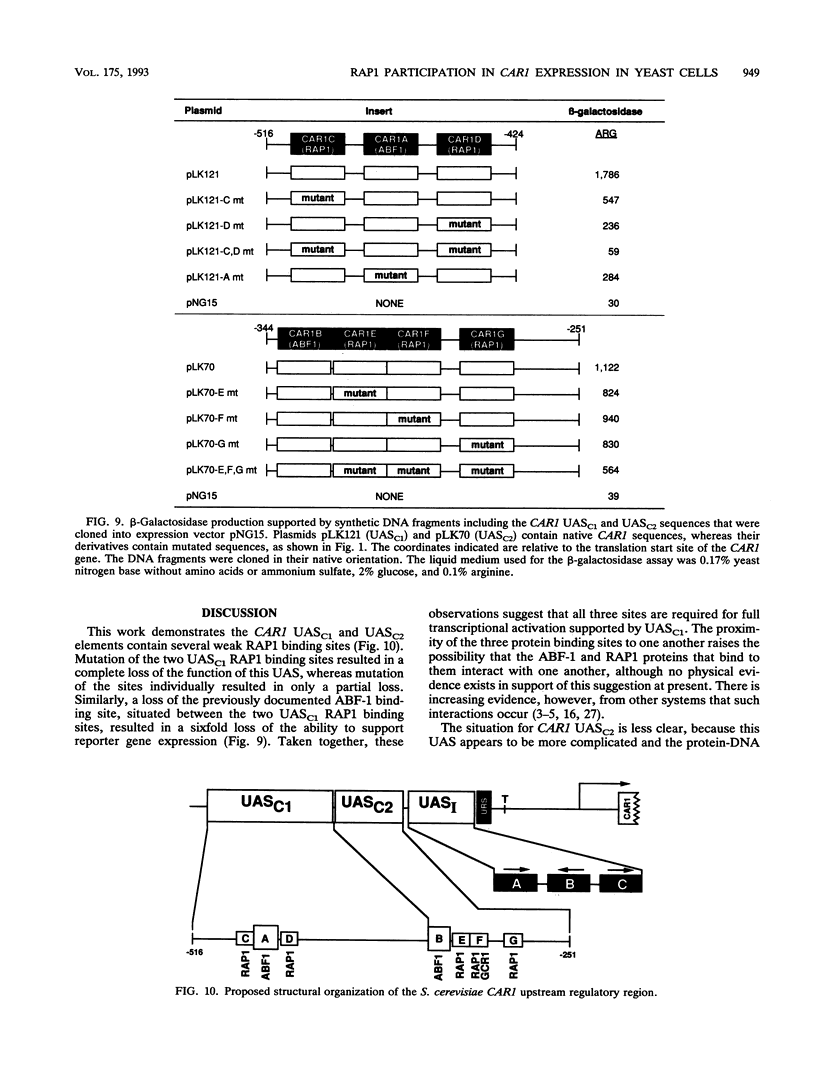
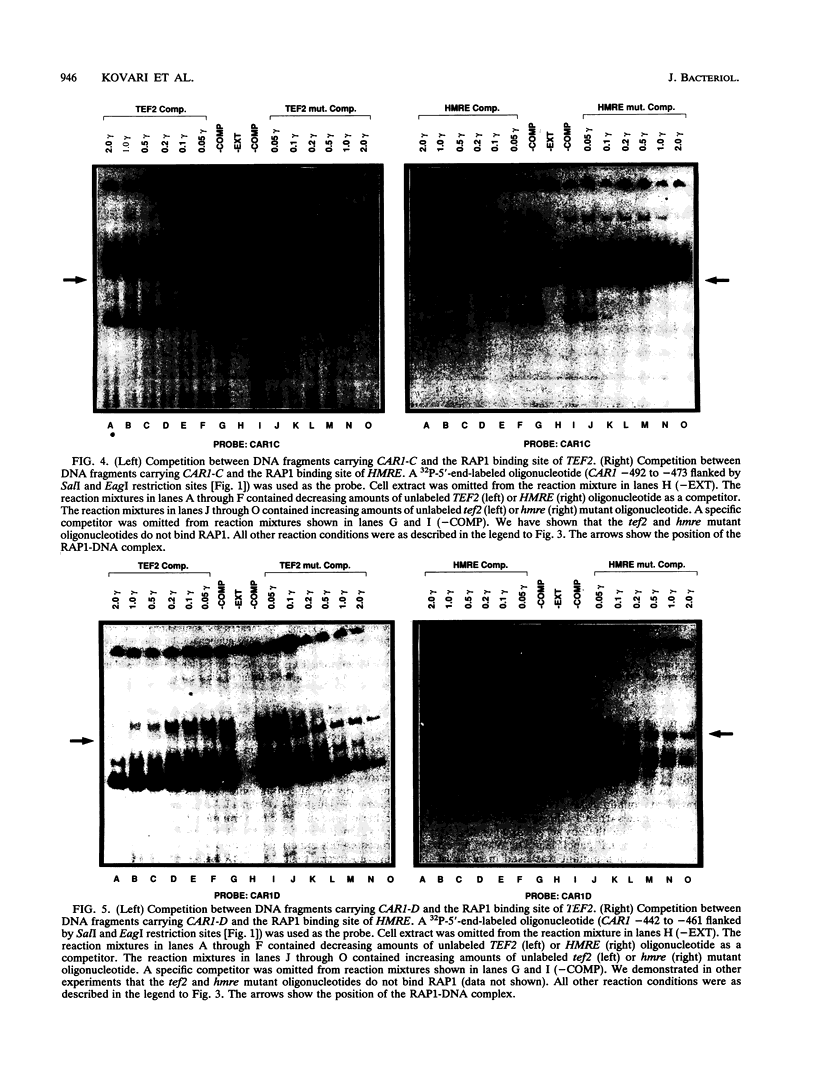
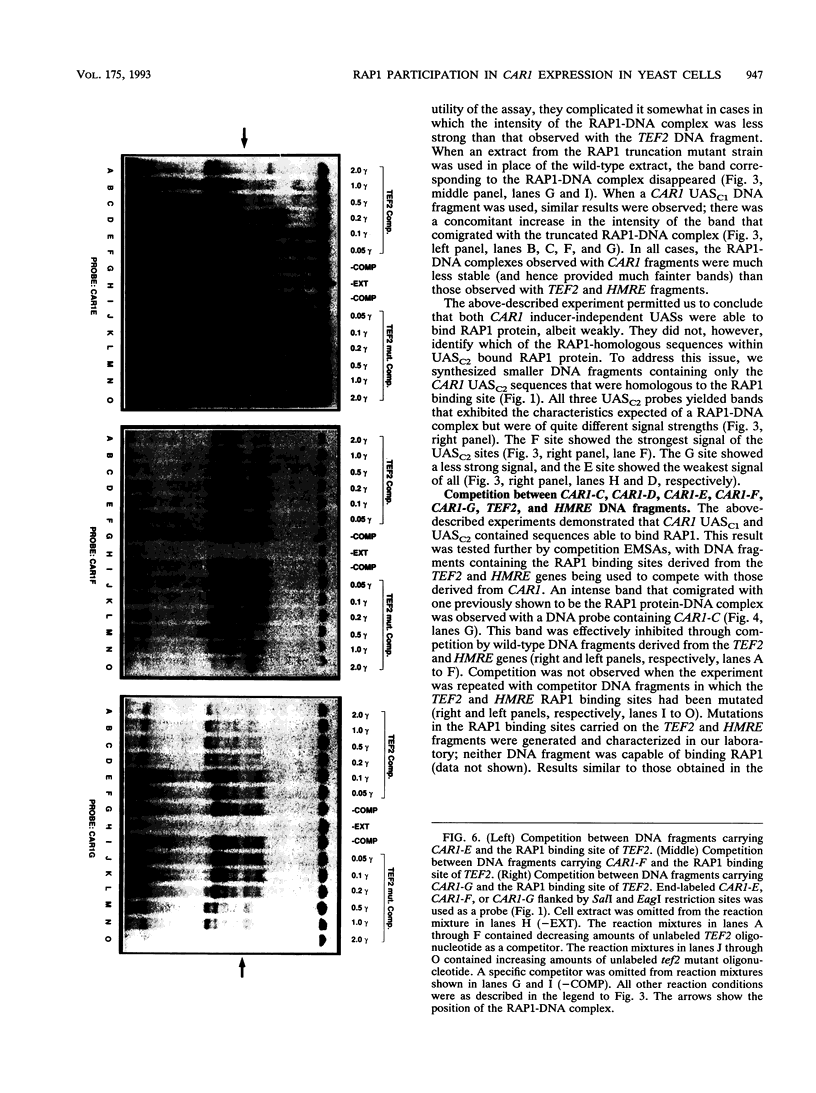
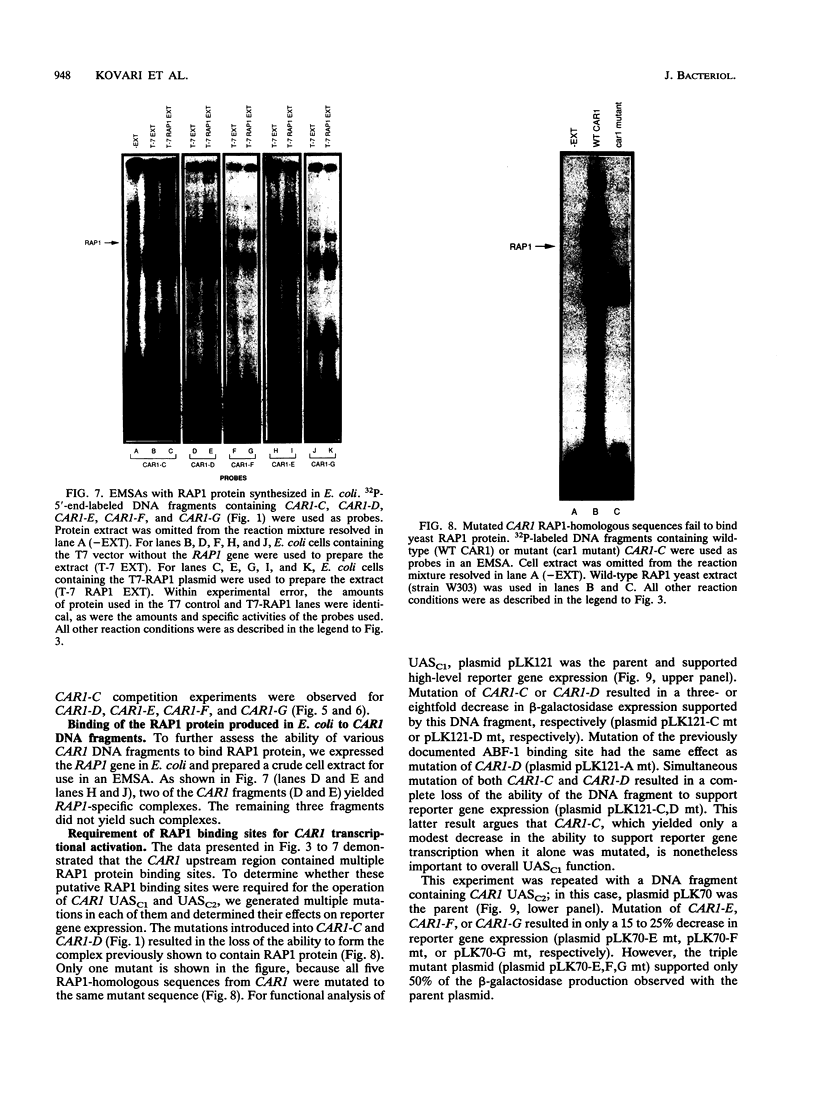
Regulated expression of the inducible arginase (CAR1) gene of Saccharomyces cerevisiae has been shown to require three upstream activation sequences (UASs) and an upstream repression sequence, URS1. Two of the UAS elements, UASC1 and UASC2, operate in an inducer-independent manner, while the third, UASI, is inducer dependent. UASC1 and UASC2 were previously shown to contain ABF-1 binding sites that were required for normal transcription. In this work, we demonstrate that UASC1 and UASC2 also contain two and three sites, respectively, that are able to bind RAP1 protein. RAP1 binding to these sites, however, is significantly weaker than that to sites in TEF2 and HMRE. The effects of mutating the sites individually or in combination suggest that at least three of them, two in UASC1 and one in UASC2, probably participate in CAR1 expression.
Full text
PDF



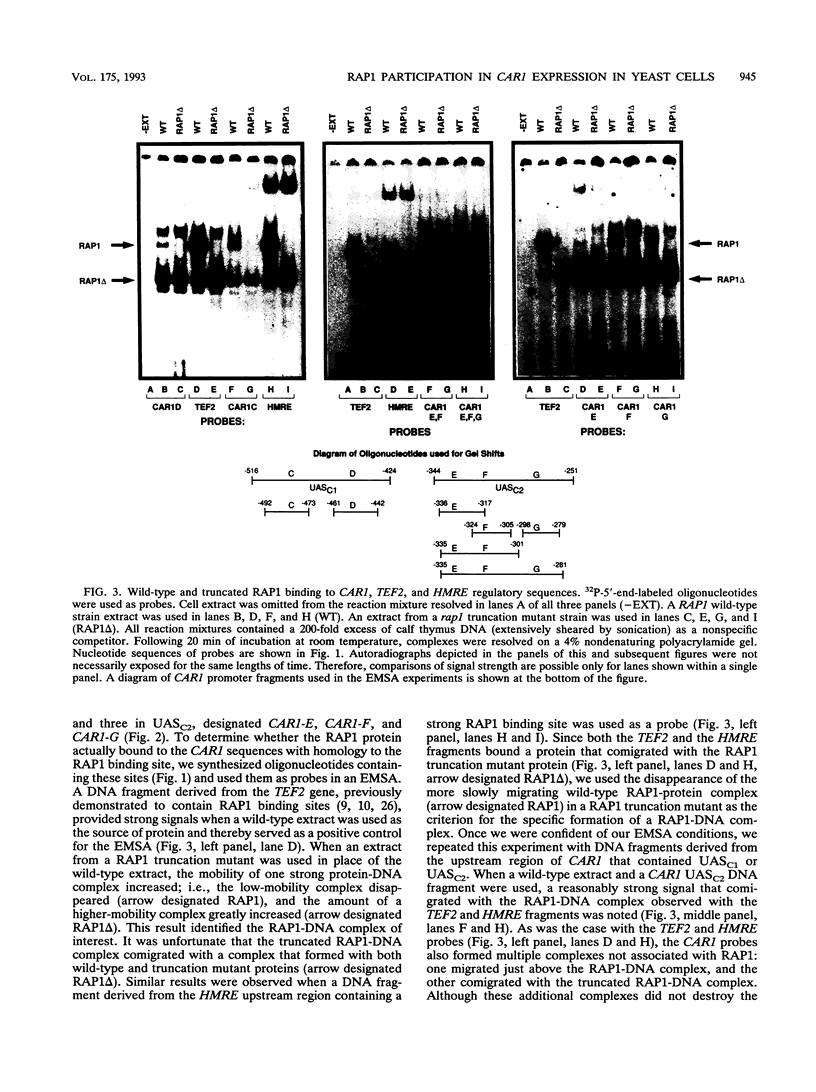



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baker H. V. GCR1 of Saccharomyces cerevisiae encodes a DNA binding protein whose binding is abolished by mutations in the CTTCC sequence motif. Proc Natl Acad Sci U S A. 1991 Nov 1;88(21):9443–9447. doi: 10.1073/pnas.88.21.9443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bossinger J., Cooper T. G. Molecular events associated with induction of arginase in Saccharomyces cerevisiae. J Bacteriol. 1977 Jul;131(1):163–173. doi: 10.1128/jb.131.1.163-173.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Brindle P. K., Holland J. P., Willett C. E., Innis M. A., Holland M. J. Multiple factors bind the upstream activation sites of the yeast enolase genes ENO1 and ENO2: ABFI protein, like repressor activator protein RAP1, binds cis-acting sequences which modulate repression or activation of transcription. Mol Cell Biol. 1990 Sep;10(9):4872–4885. doi: 10.1128/mcb.10.9.4872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buchman A. R., Kimmerly W. J., Rine J., Kornberg R. D. Two DNA-binding factors recognize specific sequences at silencers, upstream activating sequences, autonomously replicating sequences, and telomeres in Saccharomyces cerevisiae. Mol Cell Biol. 1988 Jan;8(1):210–225. doi: 10.1128/mcb.8.1.210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Compagno C., Ranzi B. M., Martegani E. The promoter of Saccharomyces cerevisiae FBA1 gene contains a single positive upstream regulatory element. FEBS Lett. 1991 Nov 18;293(1-2):97–100. doi: 10.1016/0014-5793(91)81160-a. [DOI] [PubMed] [Google Scholar]
- Cooper T. G., Kovari L., Sumrada R. A., Park H. D., Luche R. M., Kovari I. Nitrogen catabolite repression of arginase (CAR1) expression in Saccharomyces cerevisiae is derived from regulated inducer exclusion. J Bacteriol. 1992 Jan;174(1):48–55. doi: 10.1128/jb.174.1.48-55.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henry Y. A., Chambers A., Tsang J. S., Kingsman A. J., Kingsman S. M. Characterisation of the DNA binding domain of the yeast RAP1 protein. Nucleic Acids Res. 1990 May 11;18(9):2617–2623. doi: 10.1093/nar/18.9.2617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huet J., Sentenac A. TUF, the yeast DNA-binding factor specific for UASrpg upstream activating sequences: identification of the protein and its DNA-binding domain. Proc Natl Acad Sci U S A. 1987 Jun;84(11):3648–3652. doi: 10.1073/pnas.84.11.3648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kovari L. Z., Cooper T. G. Participation of ABF-1 protein in expression of the Saccharomyces cerevisiae CAR1 gene. J Bacteriol. 1991 Oct;173(20):6332–6338. doi: 10.1128/jb.173.20.6332-6338.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kovari L., Sumrada R., Kovari I., Cooper T. G. Multiple positive and negative cis-acting elements mediate induced arginase (CAR1) gene expression in Saccharomyces cerevisiae. Mol Cell Biol. 1990 Oct;10(10):5087–5097. doi: 10.1128/mcb.10.10.5087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luche R. M., Sumrada R., Cooper T. G. A cis-acting element present in multiple genes serves as a repressor protein binding site for the yeast CAR1 gene. Mol Cell Biol. 1990 Aug;10(8):3884–3895. doi: 10.1128/mcb.10.8.3884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MIDDELHOVEN W. J. THE PATHWAY OF ARGININE BREAKDOWN IN SACCHAROMYCES CEREVISIAE. Biochim Biophys Acta. 1964 Dec 9;93:650–652. doi: 10.1016/0304-4165(64)90349-6. [DOI] [PubMed] [Google Scholar]
- Mager W. H., Planta R. J. Multifunctional DNA-binding proteins mediate concerted transcription activation of yeast ribosomal protein genes. Biochim Biophys Acta. 1990 Aug 27;1050(1-3):351–355. doi: 10.1016/0167-4781(90)90193-6. [DOI] [PubMed] [Google Scholar]
- Messenguy F., Dubois E., Boonchird C. Determination of the DNA-binding sequences of ARGR proteins to arginine anabolic and catabolic promoters. Mol Cell Biol. 1991 May;11(5):2852–2863. doi: 10.1128/mcb.11.5.2852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Middelhoven W. J. Enzyme repression in the arginine pathway of Saccharomyces cerevisiae. Antonie Van Leeuwenhoek. 1969;35(2):215–226. doi: 10.1007/BF02219132. [DOI] [PubMed] [Google Scholar]
- Middelhoven W. J. Induction and repression of arginase and ornithine transaminase in baker's yeast. Antonie Van Leeuwenhoek. 1970;36(1):1–19. doi: 10.1007/BF02069003. [DOI] [PubMed] [Google Scholar]
- Middelhoven W. J. The derepression of arginase and of ornithine transaminase in nitrogen-starved baker's yeast. Biochim Biophys Acta. 1968 Mar 11;156(2):440–443. doi: 10.1016/0304-4165(68)90284-5. [DOI] [PubMed] [Google Scholar]
- Park H. D., Luche R. M., Cooper T. G. The yeast UME6 gene product is required for transcriptional repression mediated by the CAR1 URS1 repressor binding site. Nucleic Acids Res. 1992 Apr 25;20(8):1909–1915. doi: 10.1093/nar/20.8.1909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qui H. F., Dubois E., Messenguy F. Dissection of the bifunctional ARGRII protein involved in the regulation of arginine anabolic and catabolic pathways. Mol Cell Biol. 1991 Apr;11(4):2169–2179. doi: 10.1128/mcb.11.4.2169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shore D., Nasmyth K. Purification and cloning of a DNA binding protein from yeast that binds to both silencer and activator elements. Cell. 1987 Dec 4;51(5):721–732. doi: 10.1016/0092-8674(87)90095-x. [DOI] [PubMed] [Google Scholar]
- Shore D., Stillman D. J., Brand A. H., Nasmyth K. A. Identification of silencer binding proteins from yeast: possible roles in SIR control and DNA replication. EMBO J. 1987 Feb;6(2):461–467. doi: 10.1002/j.1460-2075.1987.tb04776.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Studier F. W., Rosenberg A. H., Dunn J. J., Dubendorff J. W. Use of T7 RNA polymerase to direct expression of cloned genes. Methods Enzymol. 1990;185:60–89. doi: 10.1016/0076-6879(90)85008-c. [DOI] [PubMed] [Google Scholar]
- Sumrada R. A., Cooper T. G. Isolation of the CAR1 gene from Saccharomyces cerevisiae and analysis of its expression. Mol Cell Biol. 1982 Dec;2(12):1514–1523. doi: 10.1128/mcb.2.12.1514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumrada R. A., Cooper T. G. Point mutation generates constitutive expression of an inducible eukaryotic gene. Proc Natl Acad Sci U S A. 1985 Feb;82(3):643–647. doi: 10.1073/pnas.82.3.643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumrada R. A., Cooper T. G. Ubiquitous upstream repression sequences control activation of the inducible arginase gene in yeast. Proc Natl Acad Sci U S A. 1987 Jun;84(12):3997–4001. doi: 10.1073/pnas.84.12.3997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitney P. A., Magasanik B. The induction of arginase in Saccharomyces cerevisiae. J Biol Chem. 1973 Sep 10;248(17):6197–6202. [PubMed] [Google Scholar]