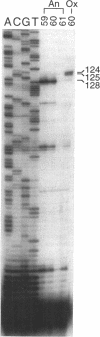
Abstract
A newly identified anaerobically expressed locus, aeg-46.5, which is located at min 46.5 on Escherichia coli linkage map, was cloned and analyzed. The phenotype of this gene was studied by using a lacZ operon fusion. aeg-46.5 is induced anaerobically in the presence of nitrate in wild-type and narL cells. It is repressed by the narL gene product, as it showed derepressed anaerobic expression in narL mutant cells. We postulate that aeg-46.5 is subject to multiple regulatory systems, activation as a result of anaerobiosis, narL-independent nitrate-dependent activation, and narL-mediated repression. The regulatory region of aeg-46.5 was identified. A 304-bp DNA sequence which includes the regulatory elements was obtained, and the 5' end of aeg-46.5 mRNA was identified. It was verified that the anaerobic regulation of aeg-46.5 expression is controlled on the transcriptional level. Computer analysis predicted possible control sites for the NarL and FNR proteins. The proposed NarL site was found in a perfect-symmetry element. The aeg-46.5 regulatory elements are adjacent to, but divergent from, those of the eco gene.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bremer E., Silhavy T. J., Weinstock G. M. Transposable lambda placMu bacteriophages for creating lacZ operon fusions and kanamycin resistance insertions in Escherichia coli. J Bacteriol. 1985 Jun;162(3):1092–1099. doi: 10.1128/jb.162.3.1092-1099.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bremer E., Silhavy T. J., Weinstock G. M. Transposition of lambda placMu is mediated by the A protein altered at its carboxy-terminal end. Gene. 1988 Nov 15;71(1):177–186. doi: 10.1016/0378-1119(88)90089-3. [DOI] [PubMed] [Google Scholar]
- Choe M., Reznikoff W. S. Anaerobically expressed Escherichia coli genes identified by operon fusion techniques. J Bacteriol. 1991 Oct;173(19):6139–6146. doi: 10.1128/jb.173.19.6139-6146.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cotter P. A., Gunsalus R. P. Oxygen, nitrate, and molybdenum regulation of dmsABC gene expression in Escherichia coli. J Bacteriol. 1989 Jul;171(7):3817–3823. doi: 10.1128/jb.171.7.3817-3823.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Erpel T., Hwang P., Craik C. S., Fletterick R. J., McGrath M. E. Physical map location of the new Escherichia coli gene eco, encoding the serine protease inhibitor ecotin. J Bacteriol. 1992 Mar;174(5):1704–1704. doi: 10.1128/jb.174.5.1704.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene. 1984 Jun;28(3):351–359. doi: 10.1016/0378-1119(84)90153-7. [DOI] [PubMed] [Google Scholar]
- Iuchi S., Lin E. C. The narL gene product activates the nitrate reductase operon and represses the fumarate reductase and trimethylamine N-oxide reductase operons in Escherichia coli. Proc Natl Acad Sci U S A. 1987 Jun;84(11):3901–3905. doi: 10.1073/pnas.84.11.3901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jamieson D. J., Higgins C. F. Two genetically distinct pathways for transcriptional regulation of anaerobic gene expression in Salmonella typhimurium. J Bacteriol. 1986 Oct;168(1):389–397. doi: 10.1128/jb.168.1.389-397.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones H. M., Gunsalus R. P. Regulation of Escherichia coli fumarate reductase (frdABCD) operon expression by respiratory electron acceptors and the fnr gene product. J Bacteriol. 1987 Jul;169(7):3340–3349. doi: 10.1128/jb.169.7.3340-3349.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalman L. V., Gunsalus R. P. Identification of a second gene involved in global regulation of fumarate reductase and other nitrate-controlled genes for anaerobic respiration in Escherichia coli. J Bacteriol. 1989 Jul;171(7):3810–3816. doi: 10.1128/jb.171.7.3810-3816.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohara Y., Akiyama K., Isono K. The physical map of the whole E. coli chromosome: application of a new strategy for rapid analysis and sorting of a large genomic library. Cell. 1987 Jul 31;50(3):495–508. doi: 10.1016/0092-8674(87)90503-4. [DOI] [PubMed] [Google Scholar]
- McGrath M. E., Hines W. M., Sakanari J. A., Fletterick R. J., Craik C. S. The sequence and reactive site of ecotin. A general inhibitor of pancreatic serine proteases from Escherichia coli. J Biol Chem. 1991 Apr 5;266(10):6620–6625. [PubMed] [Google Scholar]
- Miller C. G., Miller J. L., Bagga D. A. Cloning and nucleotide sequence of the anaerobically regulated pepT gene of Salmonella typhimurium. J Bacteriol. 1991 Jun;173(11):3554–3558. doi: 10.1128/jb.173.11.3554-3558.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawers G., Böck A. Anaerobic regulation of pyruvate formate-lyase from Escherichia coli K-12. J Bacteriol. 1988 Nov;170(11):5330–5336. doi: 10.1128/jb.170.11.5330-5336.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schulz V. P., Reznikoff W. S. In vitro secondary structure analysis of mRNA from lacZ translation initiation mutants. J Mol Biol. 1990 Jan 20;211(2):427–445. doi: 10.1016/0022-2836(90)90363-Q. [DOI] [PubMed] [Google Scholar]
- Singer M., Baker T. A., Schnitzler G., Deischel S. M., Goel M., Dove W., Jaacks K. J., Grossman A. D., Erickson J. W., Gross C. A. A collection of strains containing genetically linked alternating antibiotic resistance elements for genetic mapping of Escherichia coli. Microbiol Rev. 1989 Mar;53(1):1–24. doi: 10.1128/mr.53.1.1-24.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spiro S., Guest J. R. FNR and its role in oxygen-regulated gene expression in Escherichia coli. FEMS Microbiol Rev. 1990 Aug;6(4):399–428. doi: 10.1111/j.1574-6968.1990.tb04109.x. [DOI] [PubMed] [Google Scholar]
- Stewart V. Nitrate respiration in relation to facultative metabolism in enterobacteria. Microbiol Rev. 1988 Jun;52(2):190–232. doi: 10.1128/mr.52.2.190-232.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stewart V. Requirement of Fnr and NarL functions for nitrate reductase expression in Escherichia coli K-12. J Bacteriol. 1982 Sep;151(3):1320–1325. doi: 10.1128/jb.151.3.1320-1325.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]