Abstract
Mutational bias is a potentially important agent of evolution, but it is difficult to disentangle the effects of mutation from those of natural selection. Mutation-accumulation experiments, in which mutations are allowed to accumulate at very small population size, thus minimizing the efficiency of natural selection, are the best way to separate the effects of mutation from those of selection. Body size varies greatly among species of nematode in the family rhabditidae; mutational biases are both a potential cause and a consequence of that variation. We report data on the cumulative effects of mutations that affect body size in three species of rhabditid nematode that vary fivefold in adult size. Results are very consistent with previous studies of mutations underlying fitness in the same strains: two strains of Caenorhabditis briggsae decline in body size about twice as fast as two strains of C. elegans, with a concomitant higher point estimate of the genomic mutation rate; the confamilial Oscheius myriophila is intermediate. There is an overall mutational bias, such that mutations reduce size on average, but the bias appears consistent between species. The genetic correlation between mutations that affect size and those underlying fitness is large and positive, on average.
THE importance of mutation to the evolutionary process is universally appreciated by biologists, both in terms of the deleterious effects on fitness (Morgan 1903; Fisher 1930; Haldane 1937; Sturtevant 1937) and as the ultimate source of potentially adaptive genetic variation. It has been recognized for a long time that there is substantial variation in the mutational process at a variety of taxonomic levels, even among genotypes within species (Sturtevant 1937 and references therein; Woodruff et al. 1984; Fry 2004b; Baer et al. 2005; Ávila et al. 2006; Haag-Liautard et al. 2007). The factors responsible for that variation are poorly understood, but there are two classes of potential explanations. First, the mutation rate may be primarily a by-product of intrinsic or extrinsic environmental factors, e.g., temperature, metabolic rate, UV exposure, etc. (Martin and Palumbi 1993; Hebert et al. 2002; Gillooly et al. 2005). Alternatively, the mutation rate may be an evolutionarily optimized property, with either the optimum or the deviation from the optimum varying among taxa (Kimura 1967; Leigh 1973; Kondrashov 1995; Dawson 1998). Elucidating the taxonomic distribution of variation in mutational properties may provide important insights into several disparate areas of evolutionary biology, among them the causes of adaptive radiation (Bjedov et al. 2003; Sikorski and Nevo 2005) and cladogenesis (Shpak 2005), the rate of molecular evolution (Martin and Palumbi 1993; Gillooly et al. 2005), the nature of selection on modifier loci (Kondrashov 1995), the evolution of genetic architecture underlying the phenotype (Jones et al. 2003), and the evolution of mating system and sexual reproduction (Kondrashov 1988, 1995; Keightley and Otto 2006).
Of particular interest to quantitative geneticists is the relationship between the average phenotypic effect of a new mutation and the starting phenotype. If mutational effects are biased, the evolutionary process will be biased from the start (Jones et al. 2003) and long-term evolutionary trends may have more to do with mutation and drift than with natural selection (Lande 1975). For example, if new mutations are more likely to decrease size than increase it, all else being equal, an evolutionary decrease in size is more probable. Conversely, if mutational effects are not biased, change in any direction is equally likely and, at least in principle, any phenotype will be eventually achievable. Obviously, mutational effects must ultimately be constrained (e.g., size must be positive and finite) and thus biased at the boundary of the possible phenotypic range, but the range of allowable mutational space is generally not known.
Disentangling the relative contributions of the phenotypic effects of mutation per se and natural selection to an apparent mutational bias is not straightforward, because all alleles potentially have pleiotropic effects on fitness. For a fanciful but illustrative example, consider the vertebrate head. Adult individuals with two heads are occasionally found in nature; adults with no head never are. Presumably this bias is not due to the greater frequency of mutations for two than zero heads, but rather because mutations that result in no head are invariably lethal early in development.
The confounding of mutational vs. selective bias can never be fully overcome, but it can be minimized by allowing mutations to accumulate under conditions in which natural selection is minimized. The method of mutation accumulation (MA) minimizes the effects of natural selection by allowing replicate lines of a highly homozygous genotype to evolve at very small population size; mutations with effects on fitness s < 1/4Ne will accumulate at approximately the neutral rate (Kimura 1962).
To date, most of the relevant data are for traits that are closely associated with fitness and are expected to be under strong directional selection (reviewed in Drake et al. 1998; Keightley and Eyre-Walker 1999; Lynch et al. 1999). There is a large body of theoretical (e.g., Lande 1975; Turelli 1984; Jones et al. 2003; Waxman and Peck 2003) and empirical work concerning the mutational properties of traits that are expected to be under stabilizing selection (especially bristle number in Drosophila melanogaster, e.g., Clayton and Robertson 1955; Fry et al. 1995; Nuzhdin et al. 1995; Mackay and Lyman 1998; García-Dorado et al. 2000), but relatively few studies that allow a straightforward post hoc comparison of the same trait among taxa, especially metazoans (see Houle et al. 1996). To our knowledge no study has been explicitly designed to elucidate the variation in mutational properties among taxa in a trait not expected to be highly correlated with fitness.
Body size in nematodes in the family Rhabditidae provides an ideal opportunity to investigate the issues considered above. There is considerable variation within the family in several components of body size, including absolute adult size and percentage of growth following maturation (Flemming et al. 2000). Two studies with MA lines of the N2 strain of Caenorhabditis elegans showed that there is a mutational bias, with spontaneous mutations tending to reduce body size (Azevedo et al. 2002; Estes et al. 2005). Here we report a comparative study of the cumulative effects of spontaneous mutations on body size in three species of nematode in the family Rhabditidae that vary fivefold in maximum adult body volume.
MATERIALS AND METHODS
Systematics and natural history of nematode strains:
Justification for choice of species and strains is given in Baer et al. (2005). We used two species in the genus Caenorhabditis, C. elegans and C. briggsae, and the confamilial species Oscheius myriophila. All are androdioecious hermaphrodites; androdioecy appears to have evolved independently in these three species (Kiontke et al. 2004) Hermaphrodites can outcross to males only (Wood 1988), which are rare in laboratory cultures of all three species (∼0.1% in most strains of C. elegans). Generation time of all three species at 20° is ∼3.5 days, and fecundity is similar in all species. Each species is represented by two strains (iso-hermaphrodite lines): N2 and PB306 in C. elegans, HK104 and PB800 in C. briggsae, and EM435 and DF5020 in O. myriophila. C. briggsae and C. elegans are believed to have diverged at least 50 MYA (Denver et al. 2003), with Caenorhabditis and Oscheius having diverged well before then. Collection information on all strains is available from the Caenorhabditis Genetics Center.
Mutation accumulation:
MA protocols employed in this study have been outlined in detail elsewhere (Vassilieva and Lynch 1999; Baer et al. 2005). The principle is simple: many replicate lines of a highly inbred stock population are allowed to evolve in the relative absence of natural selection, thereby allowing deleterious mutations to accumulate. Descendant populations are then compared to the ancestral control stock. If the average effect of new mutations is nonzero, the mean phenotype will change over time. Since different lines accumulate different mutations, the variance among lines will increase over time, even if the average mutational effect is zero.
For each of the four Caenorhabditis strains we assayed 68 (of the initial 100) MA lines that had accumulated mutations for 200 generations and 30 ancestral control lines. Fewer MA lines were available for the O. myriophila strains due to loss of lines during freezing (DF5020, n = 47; EM435, n = 43). MA and control lines were randomly assigned to two blocks of equal size; each MA line was present in only one block (i.e., line is nested within block). At the beginning of a block, 34 randomly chosen MA lines from each strain (half of the remaining O. myriophila lines) were thawed. A sample of each control population was thawed and 15 worms were chosen to begin replicate lines and allowed to reproduce. Three replicates were started from a single worm and maintained by single-worm transfers for two generations (P1 and P2). Each plate was assigned a random number and was handled only in random numerical order after the first generation. If a worm failed to reproduce during the P1 generation, we started the plate again. A single gravid adult (∼96-hr) P3 worm was collected from each P2 parent. The gravid P3 adult was then allowed to lay eggs on a fresh plate for ∼2 hr; 72 hr after egg laying, 10 adult worms were collected into microcentrifuge tubes containing a fixative (4% glutaraldehyde buffered with PBS). If a worm did not reproduce during the 2-hr period, another gravid adult was selected from the P2 plate and the process was repeated. From each replicate, 5–10 worms were randomly picked out of the fixative, suspended in PBS buffer, and photographed at 50× magnification. Replacement of worms that failed to reproduce at either the P1 or the egg-laying stage potentially imposes selection. Because size is positively correlated with fitness (see below), differences between MA and control groups would have been underestimated, more so in C. briggsae than in the other two species.
Worm measurements:
Our measurement protocol follows that of Azevedo et al. (2002). Adult worms were photographed using a Leica MZ75 dissecting microscope. Images were captured using a Leica DFC280 camera connected to a computer running the Leica IM50 software (Leica Microsystems Imaging Solutions). Images were imported into the public domain ImageJ software (http://rsb.info.nih.gov/ij/), and individuals were measured by manually adjusting the threshold of the image and automatically tracing animal outlines using the “Analyze Particles” option within the “Analyze” menu. Area (A) and perimeter (P) were calculated for each individual and used to estimate body volume (S) under the assumption that the worm is cylindrical using the equation  (Azevedo et al. 2002; note that a typographical error in the original publication omitted the exponent in the numerator).
(Azevedo et al. 2002; note that a typographical error in the original publication omitted the exponent in the numerator).
DATA ANALYSIS
Differences among groups in the change in mean phenotype:
The change in mean phenotype due to the accumulation of new mutations  = U
= U t, where U is the genomic mutation rate, 2a is the homozygous effect of a mutation, and t is the number of generations of mutation accumulation (Lynch and Walsh 1998, p. 341). The average effect,
t, where U is the genomic mutation rate, 2a is the homozygous effect of a mutation, and t is the number of generations of mutation accumulation (Lynch and Walsh 1998, p. 341). The average effect,  , is typically expressed as a fraction of the starting mean. To allow meaningful comparisons among groups, data were natural log-transformed prior to analysis so that equivalent proportional changes in groups with different control means (
, is typically expressed as a fraction of the starting mean. To allow meaningful comparisons among groups, data were natural log-transformed prior to analysis so that equivalent proportional changes in groups with different control means ( ) have equivalent slopes (Rm), where Rm =
) have equivalent slopes (Rm), where Rm =  . Residuals of log-transformed replicate means were slightly left skewed but not significantly different from normal (Shapiro–Wilks' W = 0.998, P < W = 0.35); obvious outliers were removed by eye. We first tested for a change in mean body volume in each strain individually with the linear model log(volume) = Generation + Block + Line(Generation × Block) + Error as implemented in SAS v. 9.1 PROC MIXED with Generation (gen) (control, gen 0; MA, gen 200) coded as a class variable (supplemental Table 4 at http://www.genetics.org/supplemental/). Generation was modeled as a fixed effect; block and line were considered random effects. Degrees of freedom were determined by the Satterthwaite approximation for unequal sample size. Six tests require an individual Bonferroni-corrected significance level of α = 0.05/6 = 0.0083 to maintain an experimentwide type I error rate of 5%. The slope of the regression of log(volume) on Generation (Rm) and its standard error were calculated using the same model in PROC MIXED with the SOLUTION option and Generation was coded as a continuous variable.
. Residuals of log-transformed replicate means were slightly left skewed but not significantly different from normal (Shapiro–Wilks' W = 0.998, P < W = 0.35); obvious outliers were removed by eye. We first tested for a change in mean body volume in each strain individually with the linear model log(volume) = Generation + Block + Line(Generation × Block) + Error as implemented in SAS v. 9.1 PROC MIXED with Generation (gen) (control, gen 0; MA, gen 200) coded as a class variable (supplemental Table 4 at http://www.genetics.org/supplemental/). Generation was modeled as a fixed effect; block and line were considered random effects. Degrees of freedom were determined by the Satterthwaite approximation for unequal sample size. Six tests require an individual Bonferroni-corrected significance level of α = 0.05/6 = 0.0083 to maintain an experimentwide type I error rate of 5%. The slope of the regression of log(volume) on Generation (Rm) and its standard error were calculated using the same model in PROC MIXED with the SOLUTION option and Generation was coded as a continuous variable.
We next fit the full model log(volume) = Block + Generation + Species + Generation × Species + Strain(Species) + Generation × Strain(Species) + Line(Block(Gen(Strain(Species))) + Error. Generation, Species, and their interaction were modeled as fixed effects; other terms were considered random. Species and the interaction of species with generation were considered fixed because prior results suggested particular hypotheses about relative rates of change (Rm) in the different species, e.g., that C. briggsae should change faster than C. elegans (Baer et al. 2005, 2006). Fixed effects were tested for significance using PROC MIXED with degrees of freedom determined by the Satterthwaite method. The specific effect of interest is the species-by-generation interaction, i.e., whether Rm differs between species (supplemental Table 5 at http://www.genetics.org/supplemental/)? Comparisons between pairs of species were performed similarly (supplemental Table 6 at http://www.genetics.org/supplemental/). Three between-species comparisons require a Bonferroni-corrected P < 0.05/(1 + 3) = 0.0125 (the 1 in the sum in the denominator accounts for the test of the full model). Differences between strains within each species were tested for using likelihood-ratio tests with the (random) species-by-generation interaction term included and excluded from the model. Three between-strain comparisons require a Bonferroni-corrected P < 0.05/(1 + 3 + 3) = 0.007 to maintain an experimentwide 5% probability of type I error.
The most intuitive measure of the cumulative effects of new mutations is the change in the (untransformed) mean expressed as a proportion of the control mean, Δ = Rm/
= Rm/ . We estimated Δ
. We estimated Δ by means of a bootstrap procedure. Details of the procedure are presented in Baer et al. (2005). Briefly, lines (control and MA) were sampled with replacement from each assay block. The pseudomean values for MA and control lines were calculated using SAS v. 9.1 PROC MEANS and a pseudovalue of Δ
by means of a bootstrap procedure. Details of the procedure are presented in Baer et al. (2005). Briefly, lines (control and MA) were sampled with replacement from each assay block. The pseudomean values for MA and control lines were calculated using SAS v. 9.1 PROC MEANS and a pseudovalue of Δ was determined for each block and averaged over blocks for a final estimate. This procedure was repeated 1000 times; the upper and lower 2.5% of the distribution constitute ∼95% confidence limits (Efron and Tibshirani 1993). Groups with nonoverlapping confidence limits (CL) are considered significantly different at the 5% level. This protocol accounts for variation both within and between blocks.
was determined for each block and averaged over blocks for a final estimate. This procedure was repeated 1000 times; the upper and lower 2.5% of the distribution constitute ∼95% confidence limits (Efron and Tibshirani 1993). Groups with nonoverlapping confidence limits (CL) are considered significantly different at the 5% level. This protocol accounts for variation both within and between blocks.
Differences among groups in the mutational variance:
The per-generation input of genetic variation from new mutation, VM, is one-half the among-line component of variance divided by the number of generations of mutation accumulation (Lynch and Walsh 1998, p. 330). This calculation is predicated on the assumption that the among-line component of variance in the ancestral control is zero. Comparisons of mutational variances among groups are complicated by scaling effects (see Houle et al. 1996; Fry and Heinsohn 2002; Baer et al. 2006) that are not necessarily obviated by log-transformation. Mutational variances are commonly scaled either by the environmental variance [mutational heritability  = VM/VE, where VE is the environmental (error) variance] or by the control mean [mutational coefficient of variation, CVM = [
= VM/VE, where VE is the environmental (error) variance] or by the control mean [mutational coefficient of variation, CVM = [ ]/z0 (Houle et al. 1996)]. Both of these measures of mutational variability have potentially serious limitations when used in a comparative MA context. Mutational heritability depends on VE, so differences among groups in environmental variance, for whatever reason, can potentially provide a misleading picture of the variation actually due to new mutations (Houle 1992; Houle et al. 1996). Mutational CVs do not depend on VE and do account for scaling effects. However, the CVM does not account for differences among groups in the change in the mean over time (i.e., differences in Rm), so comparisons of CVM may become misleading over time if groups differ in Rm. We have previously found that different ways of scaling mutational variances can result in qualitatively different inferences about relative variability of different groups (Baer et al. 2005, 2006; see also Fry and Heinsohn 2002). We report several measures of mutational variability: VM of raw and natural log-transformed data,
]/z0 (Houle et al. 1996)]. Both of these measures of mutational variability have potentially serious limitations when used in a comparative MA context. Mutational heritability depends on VE, so differences among groups in environmental variance, for whatever reason, can potentially provide a misleading picture of the variation actually due to new mutations (Houle 1992; Houle et al. 1996). Mutational CVs do not depend on VE and do account for scaling effects. However, the CVM does not account for differences among groups in the change in the mean over time (i.e., differences in Rm), so comparisons of CVM may become misleading over time if groups differ in Rm. We have previously found that different ways of scaling mutational variances can result in qualitatively different inferences about relative variability of different groups (Baer et al. 2005, 2006; see also Fry and Heinsohn 2002). We report several measures of mutational variability: VM of raw and natural log-transformed data,  calculated from raw and log-transformed data, and CVM scaled by the untransformed control mean (CVM,Control) and by the untransformed MA mean (CVM,MA).
calculated from raw and log-transformed data, and CVM scaled by the untransformed control mean (CVM,Control) and by the untransformed MA mean (CVM,MA).
Variance components for each strain/generation combination were calculated from the linear model y = Block + Line + Error using restricted maximum likelihood (REML) as implemented in SAS v. 9.1 PROC MIXED (for VM) or PROC VARCOMP ( and CVM). The EM435 strain of O. myriophila was omitted from the analysis because of the nonzero among-line variance of the controls. Standard errors of VM were estimated by REML using the COVTEST option in PROC MIXED; standard errors of
and CVM). The EM435 strain of O. myriophila was omitted from the analysis because of the nonzero among-line variance of the controls. Standard errors of VM were estimated by REML using the COVTEST option in PROC MIXED; standard errors of  and CVM were estimated by a bootstrap protocol analogous to that for Δ
and CVM were estimated by a bootstrap protocol analogous to that for Δ . Lines were resampled with replacement with block structure maintained; standard deviations of the pseudodistribution are approximate standard errors when data are normally distributed (Efron and Tibshirani 1993). Plus/minus two standard deviations corresponded very closely to empirical 95% confidence limits in all cases. We used VARCOMP rather than MIXED for the bootstrap analysis because of much faster run times. Means used in calculations of CVM were calculated from the same pseudosample as the variance components.
. Lines were resampled with replacement with block structure maintained; standard deviations of the pseudodistribution are approximate standard errors when data are normally distributed (Efron and Tibshirani 1993). Plus/minus two standard deviations corresponded very closely to empirical 95% confidence limits in all cases. We used VARCOMP rather than MIXED for the bootstrap analysis because of much faster run times. Means used in calculations of CVM were calculated from the same pseudosample as the variance components.
Mutational covariance between body size and fitness:
Among-group components of (co)variance were determined for body size (generation 200, this study) and fitness (generation 200, Baer et al. 2005) from the linear model y = Trait + Block + Line + Error, where the fixed effect Trait represents log(volume) and log(1 + fitness). “Fitness” is lifetime fecundity, weighted by the probability of survivorship. The among-line component of covariance between traits is the genetic covariance, which in this case is the mutational covariance. Genetic correlations rG and their standard errors were calculated from the among-line components of (co)variance. The analysis was implemented in SAS v. 9.1 PROC MIXED with the unstructured covariance (“TYPE=UNR”) option. If mutational effects on fitness and size are perfectly correlated, the expected genetic correlation is 1. The null hypotheses rG = 1 were tested by constraining the among-line correlation to 1 (0.999 in practice) with a “PARMS” statement and comparing the likelihood to that of the unconstrained model (Fry 2004a, pp. 19–23). We also report the correlation of line means (r ).
).
Mutation rate and mutational effects:
The mutation rate per diploid genome per generation (U) and average homozygous mutational effect ( ) were estimated by the Bateman–Mukai (BM) method (Lynch and Walsh 1998, pp. 341–343). The BM method estimates two parameters, the (downwardly biased) mutation rate UMIN = (Rm)2/VM and the (upwardly biased) average effect
) were estimated by the Bateman–Mukai (BM) method (Lynch and Walsh 1998, pp. 341–343). The BM method estimates two parameters, the (downwardly biased) mutation rate UMIN = (Rm)2/VM and the (upwardly biased) average effect  = VM/Rm, and requires the restrictive assumption of equal mutational effects. We used Rm and VM calculated from log-transformed data, which is appropriate if mutational effects are multiplicative across loci. Estimates of
= VM/Rm, and requires the restrictive assumption of equal mutational effects. We used Rm and VM calculated from log-transformed data, which is appropriate if mutational effects are multiplicative across loci. Estimates of  are not expressed as proportions of the starting mean because they are calculated on log-transformed data, but differences across taxa are comparable. Standard errors of UMIN and
are not expressed as proportions of the starting mean because they are calculated on log-transformed data, but differences across taxa are comparable. Standard errors of UMIN and  were calculated by the Delta method (Vassilieva and Lynch 1999).
were calculated by the Delta method (Vassilieva and Lynch 1999).
We attempted to calculate U and the distribution of mutational effects by maximum likelihood (ML), as implemented by the mlGenomeU v. 2.08 software of Keightley (http://homepages.ed.ac.uk/eang33/mlgenomeu/mlginstructions.html). The method assumes that mutational effects are gamma distributed and accommodates four parameters: U,  , the gamma shape parameter β, and the fraction of mutations with positive effects P. Models in which β and P were allowed to vary simultaneously failed to converge in reasonable time. Rather than consider only a subset of plausible parameter space, we do not report the results of the ML analysis. It has recently been brought to our attention that ML analysis of MA data in which positive effects are allowed is greatly facilitated by the presence of multiple generations in the data set (F. Shaw, personal communication; also see Keightley and Bataillon 2000).
, the gamma shape parameter β, and the fraction of mutations with positive effects P. Models in which β and P were allowed to vary simultaneously failed to converge in reasonable time. Rather than consider only a subset of plausible parameter space, we do not report the results of the ML analysis. It has recently been brought to our attention that ML analysis of MA data in which positive effects are allowed is greatly facilitated by the presence of multiple generations in the data set (F. Shaw, personal communication; also see Keightley and Bataillon 2000).
RESULTS
Change in the mean:
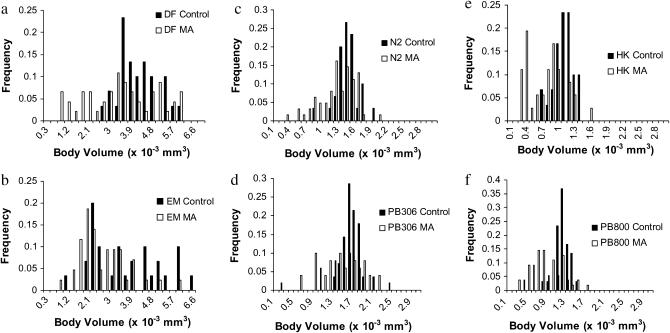
Body volume at maturity decreased over 200 generations of MA in all six strains (Rm < 0, P < 0.0083 in all cases; Figure 1, Table 1, supplemental Table 4 at http://www.genetics.org/supplemental/). ANOVA revealed significant main effects of species and generation and a significant interaction between the two (supplemental Table 5 at http://www.genetics.org/supplemental/). Averaged over strains, C. briggsae declined twice as fast as C. elegans (Rm,Cbr < Rm,Cel, P < 0.001, supplemental Table 6 at http://www.genetics.org/supplemental/), with O. myriophila intermediate. In no case did different strains of the same species differ (data not shown). After 72 hr growth at 20°, the O. myriophila were about three times as large as the two Caenorhabditis species, less than the fivefold difference in maximum (96-hr) body volume reported by Flemming et al. (2000).
Figure 1.—
Distributions of mean body volumes in control (solid bars) and mutation-accumulation (open bars) lines for the two strains of Oscheius myriophila [DF (a) and EM (b)], two strains of Caenorhabditis elegans [N2 (c) and PB306 (d)], and C. briggsae [HK (e) and PB800 (f)].
TABLE 1.
Summary statistics for mean body volume
| Strain (species) | DF (O. myr) | EM (O. myr) | N2 (C. el) | PB306 (C. el) | HK (C. br) | PB800 (C. br) |
|---|---|---|---|---|---|---|
 (×103) (×103) |
4.060 (0.449) | 3.723 (0.344) | 1.374 (0.0.089) | 1.604 (0.0.054) | 1.060 (0.0.051) | 1.116 (0.0.058) |
 (×103) (×103) |
3.348 (0.489) | 2.686 (0.260) | 1.171 (0.0.093) | 1.369 (0.0.073) | 0.725 (0.0.067) | 0.843 (0.070) |
log( ) ) |
−5.571 (0.117) | −5.756 (0.114) | −6.623 (0.079) | −6.468 (0.026) | −6.895 (0.059) | −6.778 (0.057) |
log( ) ) |
−5.894 (0.142) | −6.061 (0.096) | −6.837 (0.088) | −6.679 (0.061) | −7.392 (0.098) | −7.194 (0.077) |
| Rm (×103) | −1.62 (0.47)ab | −1.53 (0.53)ab | −1.07 (0.26)a | −1.11 (0.33)a | −2.49 (0.47)b | −2.10 (0.30)b |
Δ (×103) (×103) |
−0.891 | −1.358 | −0.746 | −0.706 | −1.714 | −1.409 |
| (−1.437, −0.294) | (−1.990, −0.635) | (−1.066, −0.388) | (−1.104, −0.268) | (−2.220, −1.157) | (−1.802, −0.988) |
Values are means, standard errors are in parentheses. The 95% CL of Δz is shown below the mean.  , mean body volume (×102 mm3) at generation zero (control);
, mean body volume (×102 mm3) at generation zero (control);  , mean body volume after 200 generations of MA; Rm, regression slope of log(volume) on generation; Δz, per-generation percentage of change in mean (untransformed) body size. O. myr, O. myriophila; C. el, C. elegans; C. br, C. briggsae.
, mean body volume after 200 generations of MA; Rm, regression slope of log(volume) on generation; Δz, per-generation percentage of change in mean (untransformed) body size. O. myr, O. myriophila; C. el, C. elegans; C. br, C. briggsae.
Rm differs significantly between species with different superscripts.
Change in the variance:
As we (Baer et al. 2006) and others (Houle et al. 1996; Fry and Heinsohn 2002) have previously observed, genetic variance, mutational or otherwise, depends crucially on scaling. To illustrate, consider the two strains of C. elegans, N2 and PB306. When the genetic variance is scaled relative to the environmental variance (mutational heritability,  ), PB306 is twice as variable as N2, but when the genetic variance is scaled by either the control or the MA mean, the variance in PB306 is only ∼20% greater than in N2. The difference between the two measures is due to greater within-line (environmental) variance in PB306. We report the variance scaled in several ways (Table 2), each of which has been widely used in studies of mutational variation. We believe the most meaningful estimate of genetic variability is the coefficient of variation scaled by the MA mean (CVM,
), PB306 is twice as variable as N2, but when the genetic variance is scaled by either the control or the MA mean, the variance in PB306 is only ∼20% greater than in N2. The difference between the two measures is due to greater within-line (environmental) variance in PB306. We report the variance scaled in several ways (Table 2), each of which has been widely used in studies of mutational variation. We believe the most meaningful estimate of genetic variability is the coefficient of variation scaled by the MA mean (CVM,  ); see Crow (1958) for discussion of the interpretation of the variance scaled in this way. It is worth noting that the “mutational CV” is generally reported as the genetic variance scaled by the control mean in studies in which there is a control and necessarily as the genetic variance scaled by the MA mean in studies in which there is no control. The distinction is important in a comparative context if the mean changes at different rates in different groups and the variance scales with the mean.
); see Crow (1958) for discussion of the interpretation of the variance scaled in this way. It is worth noting that the “mutational CV” is generally reported as the genetic variance scaled by the control mean in studies in which there is a control and necessarily as the genetic variance scaled by the MA mean in studies in which there is no control. The distinction is important in a comparative context if the mean changes at different rates in different groups and the variance scales with the mean.
TABLE 2.
Summary statistics for mutational (co)variance
| Strain (species) | DF (O. myr) | EM (O. myr) | N2 (C. el) | PB306 (C. el) | HK (C. br) | PB800 (C. br) |
|---|---|---|---|---|---|---|
Vb ( ) ) |
12.69 (35.52) | 1552 (680.1) | 0.766 (2.69) | 3.734 (5.965) | 16.82 (10.74) | 3.544 (3.503) |
VM ( ) ) |
38.06 (11.06) | — | 1.839 (0.439) | 3.806 (1.103) | 1.107 (0.988) | 1.721 (0.450) |
VM (log ) × 103 ) × 103
|
0.613 (0.180) | — | 0.234 (0.060) | 0.375 (0.096) | 0.389 (0.163) | 0.294 (0.088) |
 ( ( ) × 103 ) × 103
|
2.907 (1.182) | — | 3.477 (1.151) | 7.601 (3.269) | 2.048 (2.961) | 2.547 (0.833) |
 (log (log ) × 103 ) × 103
|
3.161 (2.014) | — | 3.223 (1.444) | 9.593 (7.079) | 2.775 (3.030) | 2.872 (1.051) |
CVM, 
|
1.524 (0.233) | — | 0.988 (0.124) | 1.219 (0.181) | 0.994 (0.550) | 1.126 (0.156) |
CVM, 
|
1.855 (0.294) | — | 1.162 (0.159) | 1.420 (0.208) | 1.515 (0.795) | 1.568 (0.203) |
| rG | 0.43 (0.21) | 0.83 (0.35) | 0.70 (0.20) | 0.93 (0.07) | 0.22 (0.42) | 0.64 (0.16) |
r
|
0.30 | 0.37 | 0.35 | 0.51 | 0.18 | 0.43 |
Values are strain means; standard errors are in parentheses. Vb ( ), among-line variance at generation 0 (control); VM, mutational variance;
), among-line variance at generation 0 (control); VM, mutational variance;  , mutational heritability; CVM,
, mutational heritability; CVM,  , mutational coefficient of variation, scaled by the control (gen 0) mean; CVM,
, mutational coefficient of variation, scaled by the control (gen 0) mean; CVM,  , mutational CV scaled by the MA (gen 200) mean; rG, genetic correlation between log(volume) and log(
, mutational CV scaled by the MA (gen 200) mean; rG, genetic correlation between log(volume) and log( ); r
); r , correlation of line means between log(volume) and log(
, correlation of line means between log(volume) and log( ). O. myr, O. myriophila; C. el, C. elegans; C. br, C. briggsae.
). O. myr, O. myriophila; C. el, C. elegans; C. br, C. briggsae.
Focusing on the MA-scaled CV (CVM,  ), the increase in genetic variance is on the order of 1–2%/generation, very consistent with previous studies (Houle et al. 1996; Drake et al. 1998; Azevedo et al. 2002). The per-generation increase in genetic variance does not differ significantly between strains or species.
), the increase in genetic variance is on the order of 1–2%/generation, very consistent with previous studies (Houle et al. 1996; Drake et al. 1998; Azevedo et al. 2002). The per-generation increase in genetic variance does not differ significantly between strains or species.
We have omitted estimates of genetic variance for the EM435 strain because the among-line variance in the control stock of that strain was significantly greater than zero (Table 2). The consistent presence of among-line variance in the EM435 control stock suggests that six generations of inbreeding did not render that stock completely homozygous. Inadvertent contamination of the ancestral EM435 cannot be ruled out, but since worms have exceedingly short ranges when not in liquid or on agar media, contamination in worm experiments is almost always due either to mislabeling of plates or tubes, in which case a stock will be homozygous for a different genotype than expected, or to poor handling during high-throughput cryopreservation, at which time worms are in liquid. Ancestral stocks were handled one at a time during cryopreservation, so residual segregating genetic variation, perhaps due to a new mutation that occurred during the six generations of inbreeding, seems more likely than contamination of the EM435 control stock.
Rate and average effects of new mutations:
UMIN estimated by BM is “small,” on the order of 0.2–2% per generation (Table 3), with average homozygous effects on the order of 0.15–0.3. UMIN does not differ significantly between species, but point estimates of UMIN are larger for C. briggsae than for C. elegans, consistent with the significantly greater rate of change in the mean in C. briggsae. Overall, the BM estimates are quite consistent with the values obtained by Azevedo et al. (2002) for the N2 strain of C. elegans (Table 3). The point estimates of UMIN for body volume are somewhat smaller than those for fitness in the same strains (Baer et al. 2005), which suggests that body size provides a smaller (but not much smaller) mutational target than does fitness, as expected.
TABLE 3.
Estimates of mutational parameters
| Strain (species) | UMIN (×102) |  |
|---|---|---|
| DF (O. myr) | 0.45 (0.54) | −0.37 |
| EM (O. myr) | — | — |
| N2 (C. el) | 0.44 (0.44) | −0.23 (0.15) |
| N2–KC (C. el) | 0.06 (0.31) | NA |
| N2–VL (C. el) | 0.36 (0.40) | NA |
| PB306 (C. el) | 0.32 (0.39) | −0.34 (0.14) |
| HK (C. br) | 1.59 (1.37) | −0.16 (0.07) |
| PB800 (C. br) | 1.50 (0.96) | −0.14 (0.05) |
Estimates of UMIN from additional N2 data sets (Azevedo et al. 2002) are included for comparison; note that haploid values of UMIN are reported in the original publication. UMIN, Bateman–Mukai estimate of diploid U (standard error in parentheses);  , Bateman–Mukai estimate of average homozygous effect (not scaled to the control mean); NA, not applicable. O. myr, O. myriophila; C. el, C. elegans; C. br, C. briggsae.
, Bateman–Mukai estimate of average homozygous effect (not scaled to the control mean); NA, not applicable. O. myr, O. myriophila; C. el, C. elegans; C. br, C. briggsae.
Mutational correlation of body size with fitness:
Averaged over all six strains, the genetic correlation between mutations for body volume and mutations that affect fitness ( ; Baer et al. 2005) is large and positive, on the order of 0.6, although there is considerable variation between strains in the point estimates (Table 2). Genetic correlations did not differ significantly from 1 except in DF5020. The correlation of line means is considerably smaller, on the order of 0.35 averaged over strains.
; Baer et al. 2005) is large and positive, on the order of 0.6, although there is considerable variation between strains in the point estimates (Table 2). Genetic correlations did not differ significantly from 1 except in DF5020. The correlation of line means is considerably smaller, on the order of 0.35 averaged over strains.
DISCUSSION
This study was motivated by two considerations. First, as part of an ongoing effort to characterize natural variation in the rate and properties of spontaneous mutation, the objective was to determine the extent to which the conclusions about mutational properties derived from estimates of fitness generalize to a character with a different mutational target and (presumably) mode of selection. We previously determined that the two strains of C. briggsae decline in fitness approximately twice as fast as the two strains of C. elegans, with the DF5020 strain of O. myriophila declining at about the same rate as C. elegans (Baer et al. 2005). The EM435 strain of O. myriophila is anomalous because, although the MA lines exhibit an increase in variance in fitness over 200 generations, mean fitness after 200 generations is not significantly different from that at time 0.
Overall, the results from this study are very consistent with our previous findings concerning mutations underlying fitness. On average, the two strains of C. briggsae decline in mean body volume twice as fast as the two C. elegans, with a concomitantly higher estimated U; the two O. myriophila are intermediate between the two Caenorhabditis (Tables 1 and 3). Conversely, the rate of increase of genetic variance was quite similar both within and among species (Table 2).
The second objective of this study was to characterize the relationship between starting phenotype and the direction and magnitude of mutational effects, with the hope of shedding light on the nature of evolutionary biases. Adult O. myriophila are larger than either of the Caenorhabditis species. There are three possible explanations: natural selection, genetic drift, and mutation. Given that the two genera diverged on the order of 50 MYA (Denver et al. 2003) and given the plentiful input of genetic variation for body size, genetic drift is an obvious possibility. Similarly, given the abundant genetic variation, diversifying selection on body size could clearly have led to the observed difference between these species in body size. Two less orthodox possibilities are (1) there are underlying mutational biases (ultimately inseparable from selective biases) such that mutations that increase body size are more common in O. myriophila and/or that mutations that decrease body size are more common in Caenorhabditis or, alternatively, (2) it may be that the nature of the developmental architecture of the worm allows a worm to be so large or small only before any further increase/decrease in size becomes prohibitively deleterious. If the species in question are pushing the limits of the functionally allowable distribution of body size, the result would be a mutational bias such that a greater fraction of allowed mutations decrease size in the large species and increase size in the small species.
The results of this study provide strong evidence for an overall mutational bias: on average, new mutations decrease body size in all six strains. The generally large, positive genetic correlation between size and fitness suggests that, not surprisingly, mutations that decrease size are deleterious. There is no trend for larger species to have smaller average mutational effects (Table 3). It is at least conceivable that the smaller size of C. briggsae relative to C. elegans is in part due to a higher mutation rate, leading to the occasional fixation of slightly deleterious alleles that reduce size, although that possibility does not extend to the relationship between Caenorhabditis and Oscheius.
Given the importance of body size to the biology of metazoans, it is surprising how few estimates of mutational parameters for body size are available [Keightley and Hill 1992, Caballero et al. 1995 (mouse); Toline and Lynch 1994, Lynch et al. 1998 (Daphnia pulex); Santiago et al. 1992, Wayne and Mackay 1998 (Drosophila melanogaster); Azevedo et al. 2002, Estes et al. 2005 (N2 strain of C. elegans)]. Estimates of mutational correlations between body size and fitness are available only for D. melanogaster and C. elegans. Wayne and Mackay (1998) found no significant correlation between MA line means for body size and competitive fitness in D. melanogaster. In C. elegans, Azevedo et al. (2002) reported correlations of line means on the order of 0.2–0.3 between body volume and productivity (r = 0.32 for the VL stock of N2, compared to our estimate of r
= 0.32 for the VL stock of N2, compared to our estimate of r = 0.35 for the same strain). Estes et al. (2005) reported a genetic correlation between body width and intrinsic rate of increase of rG = 0.58 for the VL stock of N2. Correlations of line means will underestimate the true genetic correlation between traits when the traits are measured on different individuals because the sampling covariance between traits is zero (Table 2); thus, our results appear very consistent with previous studies. Azevedo et al. (2002) also performed artificial selection on new mutations for body volume. They observed direct responses to selection for both increased and decreased volume, but they observed no significant correlated response in productivity, which they interpreted as indicative of lack of genetic correlation between size and productivity. However, artificial selection will preferentially fix mutations without deleterious pleiotropic effects on fitness. Moreover, compensatory mutations that increase fitness of mutationally degraded lines are not trivially rare in the N2 strain of C. elegans (Estes and Lynch 2003). Given the consistent mutational decline in both body size and fitness and the observed positive genetic correlation between size and fitness in this study and that of Estes et al. (2005), we believe that the response to selection was largely due to mutant alleles with atypically nonnegative effects on fitness.
= 0.35 for the same strain). Estes et al. (2005) reported a genetic correlation between body width and intrinsic rate of increase of rG = 0.58 for the VL stock of N2. Correlations of line means will underestimate the true genetic correlation between traits when the traits are measured on different individuals because the sampling covariance between traits is zero (Table 2); thus, our results appear very consistent with previous studies. Azevedo et al. (2002) also performed artificial selection on new mutations for body volume. They observed direct responses to selection for both increased and decreased volume, but they observed no significant correlated response in productivity, which they interpreted as indicative of lack of genetic correlation between size and productivity. However, artificial selection will preferentially fix mutations without deleterious pleiotropic effects on fitness. Moreover, compensatory mutations that increase fitness of mutationally degraded lines are not trivially rare in the N2 strain of C. elegans (Estes and Lynch 2003). Given the consistent mutational decline in both body size and fitness and the observed positive genetic correlation between size and fitness in this study and that of Estes et al. (2005), we believe that the response to selection was largely due to mutant alleles with atypically nonnegative effects on fitness.
Prior studies with these strains showed that C. briggsae declines in fitness due to the cumulative effects of spontaneous mutations at about twice the rate as C. elegans, with the DF5020 strain of O. myriophila similar to C. elegans (Baer et al. 2005, 2006). This study shows that that result extends to body size, leading to the conclusion that variation in the mutational properties of these strains is not peculiar to a particular trait but instead appears to be a general property. The strength and direction of the mutational bias–mutations make worms smaller, on average—are similar among genotypes and species. The possibility that the great variation in body size among Rhabditid species may result from fundamentally different mutational architecture (sensu Jones et al. 2003) is not supported.
Acknowledgments
This experiment was begun when C.F.B. was a postdoc in the lab of Mike Lynch at Indiana University. We thank the Indiana University worm crew for 200 generations of hard work, especially AHawk and Cat Steding. We also thank R. Azevedo, A. Custer, D. Denver, S. Estes, D. Houle, P. Keightley, M. Lynch, F. Shaw, M. Wayne, and the reviewers for helpful advice and/or comments. Worms were provided by the Caenorhabditis Stock Center at the University of Minnesota. This work was supported by National Institutes of Health/National Research Service Award (NIH/NRSA) Postdoctoral Fellowship 1 F32 GM20887-01, NIH grant RO1-GM072639, and start-up funds from the University of Florida to C.F.B. and by NIH grant RO1-GM36827 to M. Lynch.
References
- Ávila, V., D. Chavarrías, E. Sánchez, A. Manrique, C. López-Fanjul et al., 2006. Increase of spontaneous mutation rate in a long-term experiment with Drosophila melanogaster. Genetics 173: 267–277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Azevedo, R. B., P. D. Keightley, C. Lauren-Maatta, L. L. Vassilieva, M. Lynch et al., 2002. Spontaneous mutational variation for body size in Caenorhabditis elegans. Genetics 162: 755–765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baer, C. F., F. Shaw, C. Steding, M. Baurngartner, A. Hawkins et al., 2005. Comparative evolutionary genetics of spontaneous mutations affecting fitness in rhabditid nematodes. Proc. Natl. Acad. Sci. USA 102: 5785–5790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baer, C. F., N. Phillips, D. Ostrow, A. Avalos, D. Blanton et al., 2006. Cumulative effects of spontaneous mutations for fitness in Caenorhabditis: role of genotype, environment and stress. Genetics 174: 1387–1395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bjedov, I., O. Tenaillon, B. Gerard, V. Souza, E. Denamur et al., 2003. Stress-induced mutagenesis in bacteria. Science 300: 1404–1409. [DOI] [PubMed] [Google Scholar]
- Caballero, A., P. D. Keightley and W. G. Hill, 1995. Accumulation of mutations affecting body-weight in inbred mouse lines. Genet. Res. 65: 145–149. [DOI] [PubMed] [Google Scholar]
- Clayton, G., and A. Robertson, 1955. Mutation and quantitative variation. Am. Nat. 89: 151–158. [Google Scholar]
- Crow, J. F., 1958. Some possibilities for measuring selection intensities in man. Hum. Biol. 30: 1–13. [PubMed] [Google Scholar]
- Dawson, K. J., 1998. Evolutionarily stable mutation rates. J. Theor. Biol. 194: 143–157. [DOI] [PubMed] [Google Scholar]
- Denver, D. R., K. Morris and W. K. Thomas, 2003. Phylogenetics in Caenorhabditis elegans: an analysis of divergence and outcrossing. Mol. Biol. Evol. 20: 393–400. [DOI] [PubMed] [Google Scholar]
- Drake, J. W., B. Charlesworth, D. Charlesworth and J. F. Crow, 1998. Rates of spontaneous mutation. Genetics 148: 1667–1686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Efron, B., and R. J. Tibshirani, 1993. An Introduction to the Bootstrap. Chapman & Hall, New York.
- Estes, S., and M. Lynch, 2003. Rapid fitness recovery in mutationally degraded lines of Caenorhabditis elegans. Evolution 57: 1022–1030. [DOI] [PubMed] [Google Scholar]
- Estes, S., B. C. Ajie, M. Lynch and P. C. Phillips, 2005. Spontaneous mutational correlations for life-history, morphological and behavioral characters in Caenorhabditis elegans. Genetics 170: 645–653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher, R. A., 1930. The Genetical Theory of Natural Selection. Clarendon Press, Oxford.
- Flemming, A. J., Z.-Z. Shen, A. Cunha, S. W. Emmons and A. M. Leroi, 2000. Somatic polyploidization and cellular proliferation drive body size evolution in nematodes. Proc. Natl. Acad. Sci. USA 97: 5285–5290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fry, J. D., 2004. a Estimation of genetic variances and covariances by REML using PROC MIXED, pp. 11–34 in Genetic Analysis of Complex Traits, edited by A. M. Saxton. SAS Publishing, Cary, NC.
- Fry, J. D., 2004. b On the rate and linearity of viability declines in Drosophila mutation-accumulation experiments: genomic mutation rates and synergistic epistasis revisited. Genetics 166: 797–806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fry, J. D., and S. L. Heinsohn, 2002. Environment dependence of mutational parameters for viability in Drosophila melanogaster. Genetics 161: 1155–1167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fry, J. D., K. A. Deronde and T. F. C. Mackay, 1995. Polygenic mutation in Drosophila melanogaster: genetic analysis of selection lines. Genetics 139: 1293–1307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- García-Dorado, A., J. Fernandez and C. Lopez-Fanjul, 2000. Temporal uniformity of the spontaneous mutational variance of quantitative traits in Drosophila melanogaster. Genet. Res. 75: 47–51. [DOI] [PubMed] [Google Scholar]
- Gillooly, J. F., A. P. Allen, G. B. West and J. H. Brown, 2005. The rate of DNA evolution: effects of body size and temperature on the molecular clock. Proc. Natl. Acad. Sci. USA 102: 140–145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haag-Liautard, C., M. Dorris, X. Maside, S. Macaskill, D. L. Halligan et al., 2007. Direct estimation of per nucleotide and genomic deleterious mutation rates in Drosophila. Nature 445: 82–85. [DOI] [PubMed] [Google Scholar]
- Haldane, J. B. S., 1937. The effect of variation on fitness. Am. Nat. 71: 337–349. [Google Scholar]
- Hebert, P. D. N., E. A. Remigio, J. K. Colbourne, D. J. Taylor and C. C. Wilson, 2002. Accelerated molecular evolution in halophilic crustaceans. Evolution 56: 909–926. [DOI] [PubMed] [Google Scholar]
- Houle, D., 1992. Comparing evolvability and variability of quantitative traits. Genetics 130: 195–204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Houle, D., B. Morikawa and M. Lynch, 1996. Comparing mutational variabilities. Genetics 143: 1467–1483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones, A. G., S. J. Arnold and R. Bürger, 2003. Stability of the G-matrix in a population experiencing pleiotropic mutation, stabilizing selection, and genetic drift. Evolution 57: 1747–1760. [DOI] [PubMed] [Google Scholar]
- Keightley, P. D., and T. M. Bataillon, 2000. Multigeneration maximum-likelihood analysis applied to mutation-accumulation experiments in Caenorhabditis elegans. Genetics 154: 1193–1201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keightley, P. D., and A. Eyre-Walker, 1999. Terumi Mukai and the riddle of deleterious mutation rates. Genetics 153: 515–523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keightley, P. D., and W. G. Hill, 1992. Quantitative genetic-variation in body size of mice from new mutations. Genetics 131: 693–700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keightley, P. D., and S. P. Otto, 2006. Interference among deleterious mutations favours sex and recombination in finite populations. Nature 443: 89–92. [DOI] [PubMed] [Google Scholar]
- Kimura, M., 1962. On probability of fixation of mutant genes in a population. Genetics 47: 713–719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimura, M., 1967. On the evolutionary adjustment of spontaneous mutation rates. Genet. Res. 9: 23–34. [Google Scholar]
- Kiontke, K., N. P. Gavin, Y. Raynes, C. Roehrig, F. Piano et al., 2004. Caenorhabditis phylogeny predicts convergence of hermaphroditism and extensive intron loss. Proc. Natl. Acad. Sci. USA 101: 9003–9008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kondrashov, A. S., 1988. Deleterious mutations and the evolution of sexual reproduction. Nature 336: 435–440. [DOI] [PubMed] [Google Scholar]
- Kondrashov, A. S., 1995. Modifiers of mutation-selection balance: general approach and the evolution of mutation rates. Genet. Res. 66: 53–69. [Google Scholar]
- Lande, R., 1975. The maintenance of genetic variability by mutation in a polygenic character with linked loci. Genet. Res. 26: 221–235. [DOI] [PubMed] [Google Scholar]
- Leigh, E. G., 1973. The evolution of mutation rates. Genetics 73(Suppl.): 1–18. [PubMed] [Google Scholar]
- Lynch, M., and J. B. Walsh, 1998. Genetics and Analysis of Quantitative Traits. Sinauer Press, Sunderland, MA.
- Lynch, M., J. Blanchard, D. Houle, T. Kibota, S. Schultz et al., 1999. Perspective: spontaneous deleterious mutation. Evolution 53: 645–663. [DOI] [PubMed] [Google Scholar]
- Lynch, M., L. Latta, J. Hicks and M. Giorgianni, 1998. Mutation, selection, and the maintenance of life-history variation in a natural population. Evolution 52: 727–733. [DOI] [PubMed] [Google Scholar]
- Mackay, T. F. C., and R. F. Lyman, 1998. Polygenic mutation in Drosophila melanogaster: genotype x environment interaction for spontaneous mutations affecting bristle number. Genetica 102: 199–215. [PubMed] [Google Scholar]
- Martin, A. P., and S. R. Palumbi, 1993. Body size, metabolic-rate, generation time, and the molecular clock. Proc. Natl. Acad. Sci. USA 90: 4087–4091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morgan, T. H., 1903. Evolution and Adaptation. MacMillan, New York.
- Nuzhdin, S. V., J. D. Fry and T. F. C. Mackay, 1995. Polygenic mutation in Drosophila melanogaster: the causal relationship of bristle number to fitness. Genetics 139: 861–872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santiago, E., J. Albornoz, A. Domínguez, M. A. Toro and C. López-Fanjul, 1992. The distribution of spontaneous mutations on quantitative traits and fitness in Drosophila melanogaster. Genetics 132: 771–781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shpak, M., 2005. The role of deleterious mutations in allopatric speciation. Evolution 59: 1389–1399. [PubMed] [Google Scholar]
- Sikorski, J., and E. Nevo, 2005. Adaptation and incipient sympatric speciation of Bacillus simplex under microclimatic contrast at “Evolution Canyons” I and II, Israel. Proc. Natl. Acad. Sci. USA 102: 15924–15929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sturtevant, A. H., 1937. Essays on evolution. I. On the effect of selection on mutation rate. Q. Rev. Biol. 12: 464–467. [Google Scholar]
- Toline, C. A., and M. Lynch, 1994. Mutational divergence of life-history traits in an obligate parthenogen. Genome 37: 33–35. [DOI] [PubMed] [Google Scholar]
- Turelli, M., 1984. Heritable genetic variation via mutation-selection balance: Lerch's zeta meets the abdominal bristle. Theor. Popul. Biol. 25: 138–193. [DOI] [PubMed] [Google Scholar]
- Vassilieva, L., and M. Lynch, 1999. The fitness effects of spontaneous mutation in Caenorhabditis elegans. Genetics 151: 119–129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waxman, D., and J. R. Peck, 2003. The anomalous effects of biased mutation. Genetics 164: 1615–1626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wayne, M. L., and T. F. C. Mackay, 1998. Quantitative genetics of ovariole number in Drosophila melanogaster. II. Mutational variation and genotype-environment interaction Genetics 148: 201–210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wood, W. B. (Editor), 1988. The Nematode Caenorhabditis elegans. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
- Woodruff, R. C., J. N. Thompson, M. A. Seeger and W. E. Spivey, 1984. Variation in spontaneous mutation and repair in natural population lines of Drosophila melanogaster. Heredity 53: 223–234. [Google Scholar]